6f33116be3a77e8ba30c7f7f06ef2420.ppt
- Количество слайдов: 19

Knowledge Management for Disease Coding (KMDC): Background & Introduction Timothy Hays, Ph. D. Project Manager, Knowledge Management for Disease Coding Office of Extramural Research (OER) Office of Portfolio Analysis and Strategic Initiatives (OPASI) 1

Overview of Today’s Presentation Ø Why is NIH Pursuing Knowledge Management for Disease Coding (KMDC)? Ø How Does KMDC Work Conceptually Ø What Insights Were Gained? Ø Next Steps 2

Why is NIH Pursuing Knowledge Management for Disease Coding? 3

External Drivers Ø The public and the Congress have a right to know how NIH money is spent Ø Efforts to find out how money is spent often begin with questions about the amount of money devoted to specific diseases or research topics Ø Currently, all 27 institutes and centers apply different definitions, methods and business rules when coding diseases & topics and determining the dollar 4

Why is NIH Pursuing KMDC? Two recent National Academy of Science reports that recommended that NIH should improve data on funding by disease: Ø ". . . the Committee concludes that the current lack of an information management method and infrastructure to collect, analyze, and report investment data in a timely fashion must be 5 addressed…”

How Does KM for Disease Coding Work Conceptually? 6
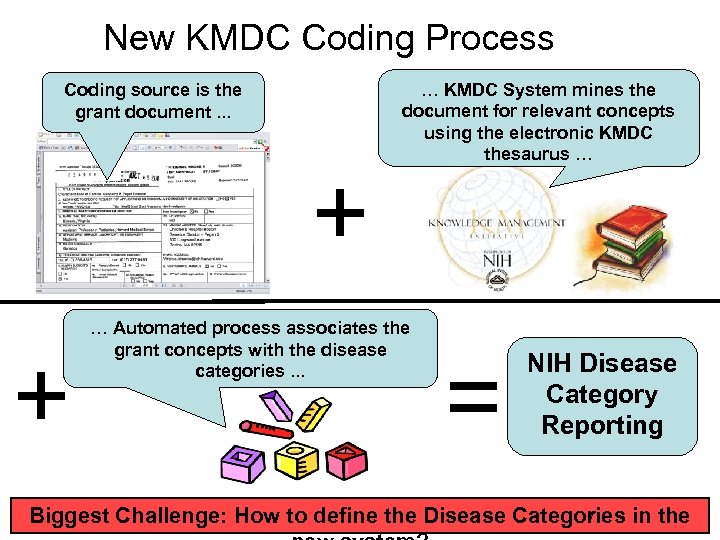
New KMDC Coding Process Coding source is the grant document. . . + + … KMDC System mines the document for relevant concepts using the electronic KMDC thesaurus … … Automated process associates the grant concepts with the disease categories. . . = NIH Disease Category Reporting 7 Biggest Challenge: How to define the Disease Categories in the
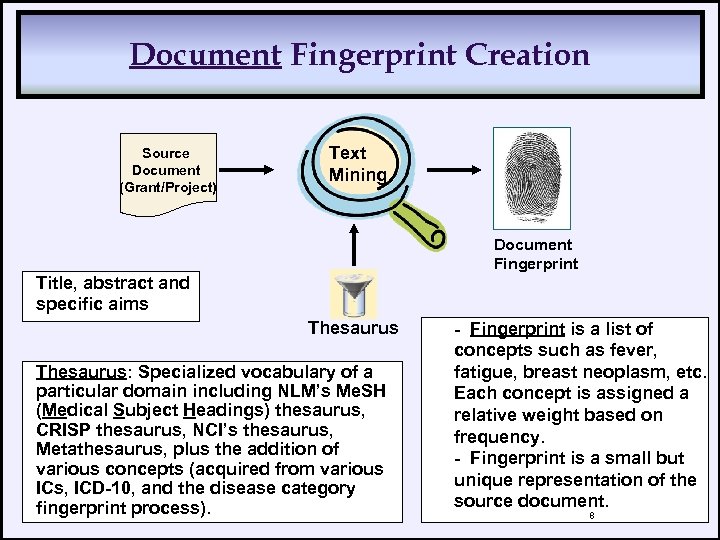
Document Fingerprint Creation Source Document (Grant/Project) Text Mining Document Fingerprint Title, abstract and specific aims Thesaurus: Specialized vocabulary of a particular domain including NLM’s Me. SH (Medical Subject Headings) thesaurus, CRISP thesaurus, NCI’s thesaurus, Metathesaurus, plus the addition of various concepts (acquired from various ICs, ICD-10, and the disease category fingerprint process). - Fingerprint is a list of concepts such as fever, fatigue, breast neoplasm, etc. Each concept is assigned a relative weight based on frequency. - Fingerprint is a small but unique representation of the source document. 8

Disease Category Fingerprint Creation in the New System In the new system a defined Disease Category Definition is called a Fingerprint. Ø A Fingerprint is a list of concepts from thesaurus. Ø Concepts are selected by NIH Scientific Experts to define that disease category. Ø Concepts can be weighted to fine-tune the system. Ø The Disease Category fingerprints are matched to the grant concepts to produce disease reporting. 9
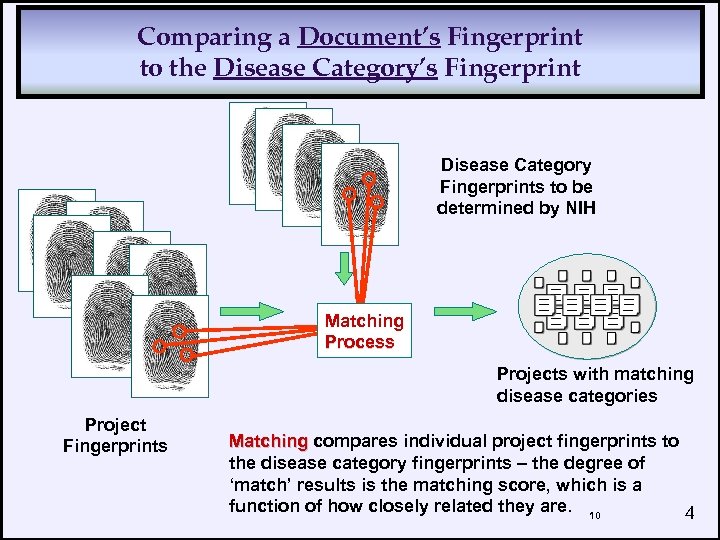
Comparing a Document’s Fingerprint to the Disease Category’s Fingerprint Disease Category Fingerprints to be determined by NIH Matching Process Projects with matching disease categories Project Fingerprints Matching compares individual project fingerprints to the disease category fingerprints – the degree of ‘match’ results is the matching score, which is a function of how closely related they are. 10 4

What Insights Were Gained… …a work in progress 11

What Insights Were Gained? Ø Clear direction and support from the NIH Director and Senior Leadership has been crucial Ø Sufficient resources Ø Build cross-agency teams to address key issues Ø Be open to feedback: customer service focus Ø Careful attention to build a process that capitalizes on what NIH experts do best & minimize “time burden” Ø Allow time for business process change Ø Keep the train moving… 12

Next Steps Ø Define taxonomy/thesaurus Ø Define disease/reporting category definitions through use of thesaurus Ø Develop customer-friendly interface to access and operate the tool Ø Use clear communication Ø Be transparent 13
Thank You 14

Thank You 15
16

NATIONAL LIBRARY OF MEDICINE (NLM) SBIR-STTR The NLM supports research and development projects in biomedical informatics and bioinformatics. NLM defines biomedical informatics as the intersection of basic informational and computing sciences with a wide range of application domains in biomedicine and public health. Bioinformatics is the intersection of basic informational and computing sciences with the biological sciences. For additional information about areas of interest to the NLM, please visit our home page at http: //www. nlm. nih. gov/ep. For additional information on research topics, contact: Hua-Chuan Sim, M. D. Program Officer Division of Extramural Programs National Library of Medicine (301) 496 -4253, Fax: (301) 402 -2952 Email: simh@mail. nih. gov For administrative and business management questions, contact: Mr. Dwight Mowery Grants Management Officer Extramural Programs Division National Library of Medicine (301) 496 -4221, Fax: (301) 402 -0421 17

NATIONAL LIBRARY OF MEDICINE (NLM) SBIR-STTR Biomedical Informatics There are broad needs for informatics concepts, tools and systems to manage the information of health care delivery, reduce medical errors, provide decision support for clinicians, extract outcome and public health information from large datasets, and predict health events. To support such projects, NLM is interested in: A. Mechanisms to capture and integrate new information into existing knowledge bases, including integration of heterogeneous data sets. B. Approaches for retrieving, extracting and analyzing data from large healthrelated and heterogeneous databases, such as patient data, population health data, or image databases. C. Systems, devices, or programs that facilitate utilization of electronic medical record systems in clinical practice, for such functions as chart entry, decision support, reduction of errors. D. Systems, devices, or programs that facilitate utilization of electronic record systems and tools in public health, for such functions as syndromic surveillance and trend analysis. E. Projects relevant to the informatics of disaster management, including management and delivery of information in disaster settings. 18

NATIONAL LIBRARY OF MEDICINE (NLM) SBIR-STTR Bioinformatics High through-put scientific research has greatly increased the volume of research data and has magnified the problem of information management and interpretation. To help manage such data, NLM is interested in: A. Software algorithms and database query methods capable of capturing scientific data from published knowledge sources or multiple related factual databases. B. Tools for data management and analysis for genetic linkage mapping, physical mapping, DNA sequencing, and proteomics. C. Tools and systems for bringing "bench to bedside, " applying research data to clinical problems. D. Algorithms capable of predicting structure and/or function in model biological systems. E. Algorithms capable of enhanced computational modeling of biological, biomedical and behavioral sciences data at multiple scales of research, ranging from molecular to population. 19
6f33116be3a77e8ba30c7f7f06ef2420.ppt