4e2cb46c65b3324d68bf03d090982437.ppt
- Количество слайдов: 83

Kinetics and Thermodynamics of Amyloid Fibril Formation Ron Wetzel University of Tennessee

Energetics of Amyloid Fibril Formation Fibril assembly equilibria and G fibril elongation - Aβ(1 -40) amyloid fibrils (Alzheimer’s disease) - polyglutamine amyloid (Huntington’s disease) Kinetics of nucleated growth polymerization and G of nucleus formation - polyglutamine amyloid
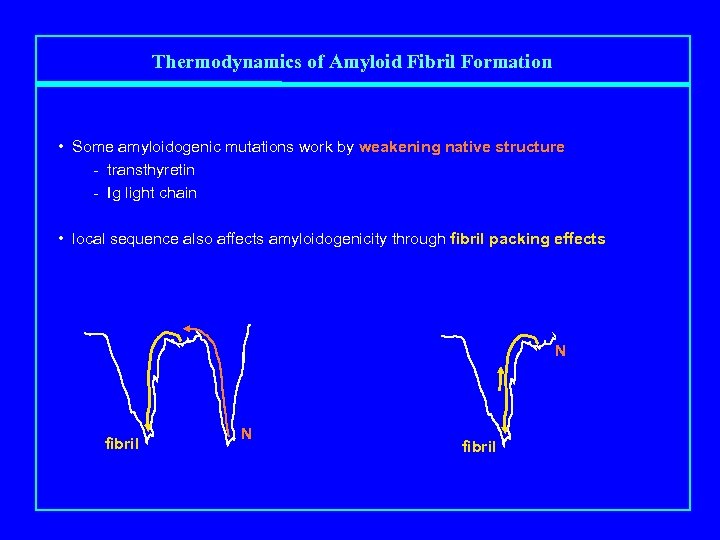
Thermodynamics of Amyloid Fibril Formation • Some amyloidogenic mutations work by weakening native structure - transthyretin - Ig light chain • local sequence also affects amyloidogenicity through fibril packing effects N fibril
![Aβ Amyloid Fibril Formation 35 30 [Monomer], μM 25 20 20 15 15 10 Aβ Amyloid Fibril Formation 35 30 [Monomer], μM 25 20 20 15 15 10](https://present5.com/presentation/4e2cb46c65b3324d68bf03d090982437/image-4.jpg)
Aβ Amyloid Fibril Formation 35 30 [Monomer], μM 25 20 20 15 15 10 10 5 5 0 0 Th. T Fluorescence (au) 30 25 0 lag phase 2 4 Time (days) 6 8 10 Cr
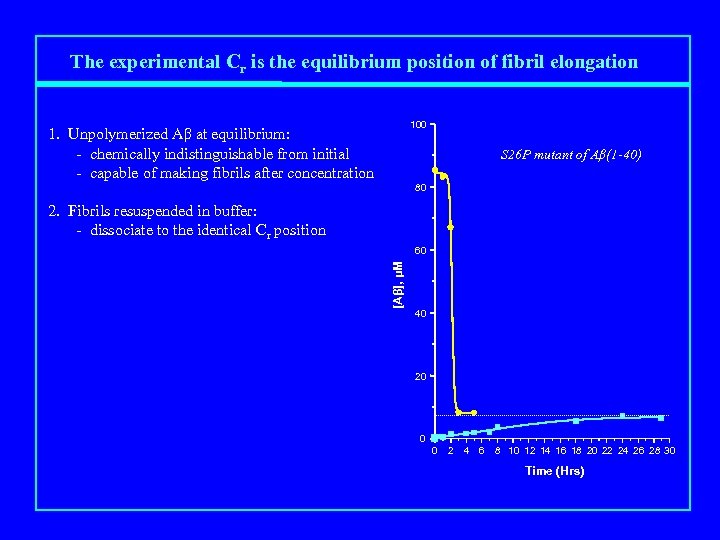
The experimental Cr is the equilibrium position of fibril elongation 100 1. Unpolymerized Aβ at equilibrium: - chemically indistinguishable from initial - capable of making fibrils after concentration S 26 P mutant of Aβ(1 -40) 80 2. Fibrils resuspended in buffer: - dissociate to the identical Cr position [Aβ], μM 60 40 20 0 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 Time (Hrs)
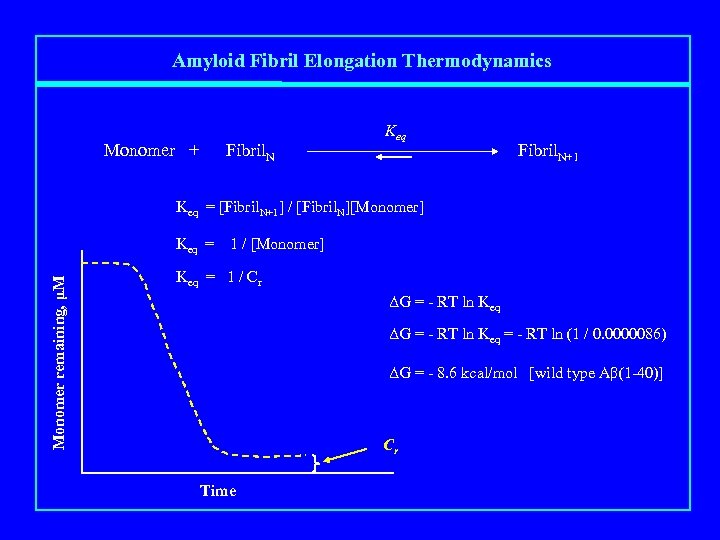
Amyloid Fibril Elongation Thermodynamics Monomer + Fibril. N Keq Fibril. N+1 Keq = [Fibril. N+1] / [Fibril. N][Monomer] Monomer remaining, μM Keq = 1 / [Monomer] Keq = 1 / Cr ΔG = - RT ln Keq = - RT ln (1 / 0. 0000086) ΔG = - 8. 6 kcal/mol [wild type Aβ(1 -40)] Cr Time
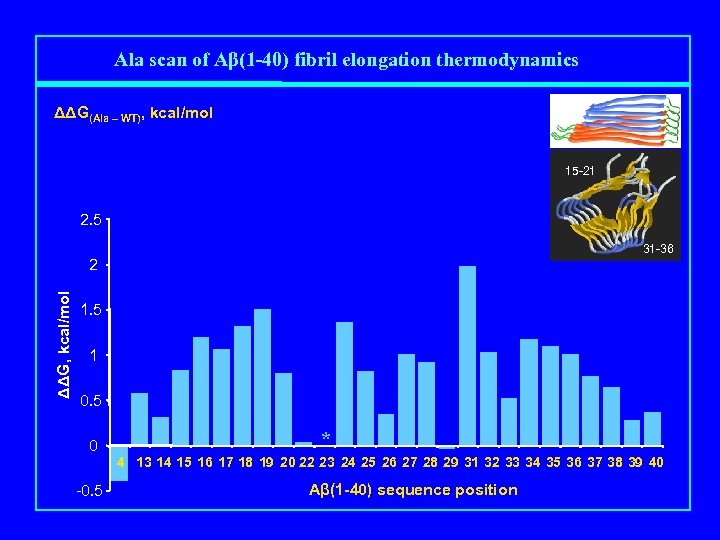
Ala scan of Aβ(1 -40) fibril elongation thermodynamics ΔΔG(Ala – WT), kcal/mol 15 -21 2. 5 31 -36 ΔΔG, kcal/mol 2 1. 5 1 0. 5 0 * 4 13 14 15 16 17 18 19 20 22 23 24 25 26 27 28 29 31 32 33 34 35 36 37 38 39 40 -0. 5 Aβ(1 -40) sequence position
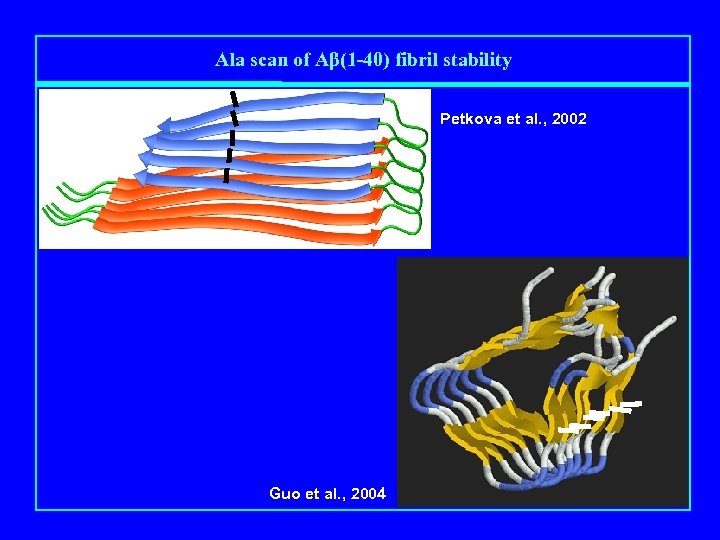
Ala scan of Aβ(1 -40) fibril stability Petkova et al. , 2002 Guo et al. , 2004
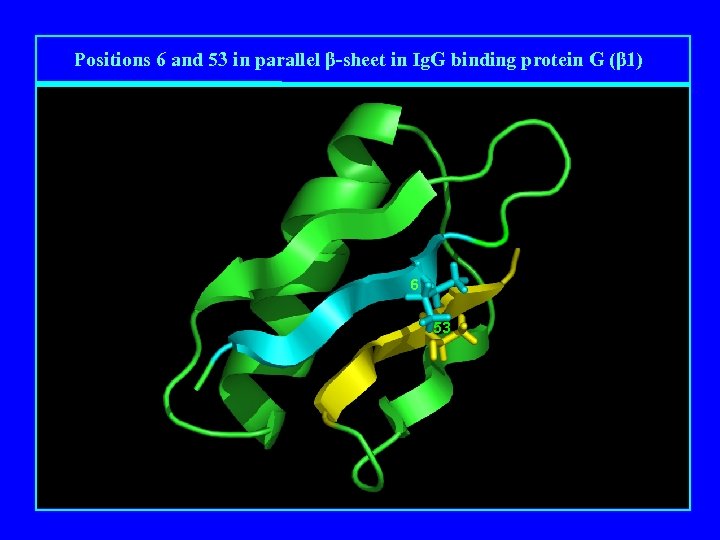
Positions 6 and 53 in parallel β-sheet in Ig. G binding protein G (β 1) 6 53
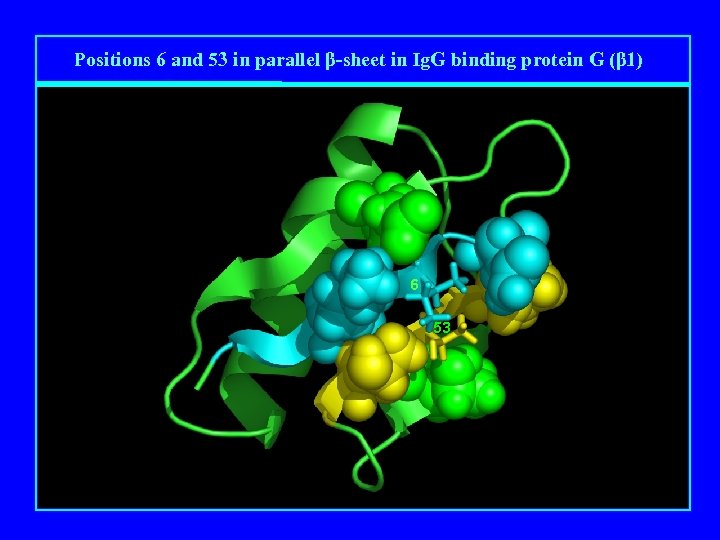
Positions 6 and 53 in parallel β-sheet in Ig. G binding protein G (β 1) 6 53
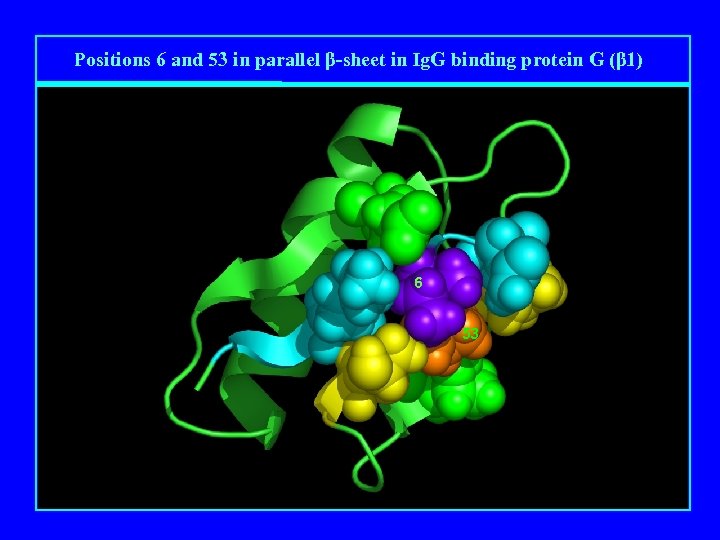
Positions 6 and 53 in parallel β-sheet in Ig. G binding protein G (β 1) 6 53
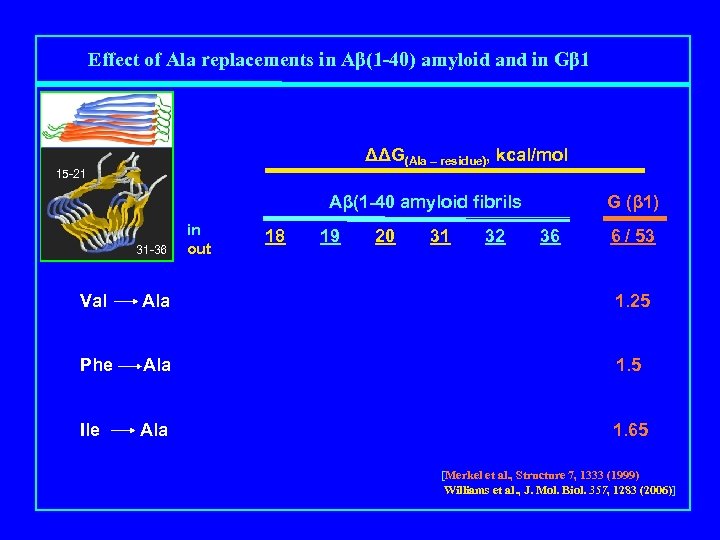

Effect of Ala replacements in Aβ(1 -40) amyloid and in Gβ 1 ΔΔG(Ala – residue), kcal/mol 15 -21 Aβ(1 -40 amyloid fibrils Mutation 31 -36 in out 18 19 20 31 32 G (β 1) 36 6 / 53 Val Ala 1. 25 Phe Ala 1. 5 Ile Ala 1. 65 [Merkel et al. , Structure 7, 1333 (1999) Williams et al. , J. Mol. Biol. 357, 1283 (2006)]
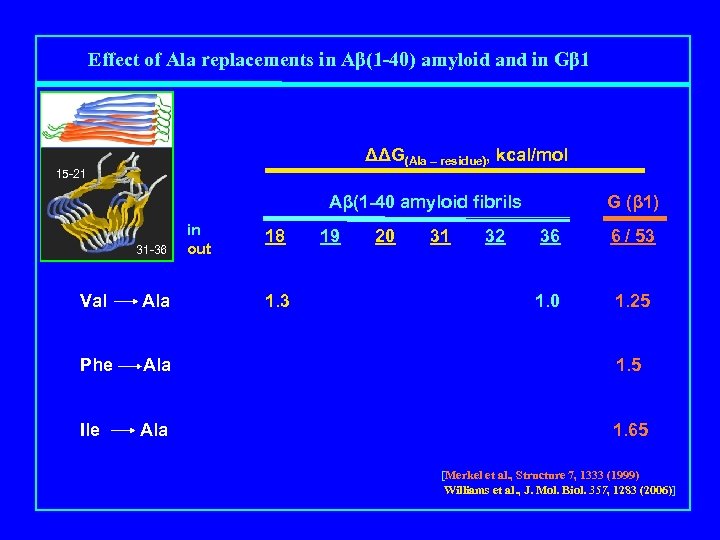

Effect of Ala replacements in Aβ(1 -40) amyloid and in Gβ 1 ΔΔG(Ala – residue), kcal/mol 15 -21 Aβ(1 -40 amyloid fibrils Mutation 31 -36 in out 18 1. 3 19 20 31 32 G (β 1) 36 6 / 53 1. 0 1. 25 Val Ala Phe Ala 1. 5 Ile Ala 1. 65 [Merkel et al. , Structure 7, 1333 (1999) Williams et al. , J. Mol. Biol. 357, 1283 (2006)]
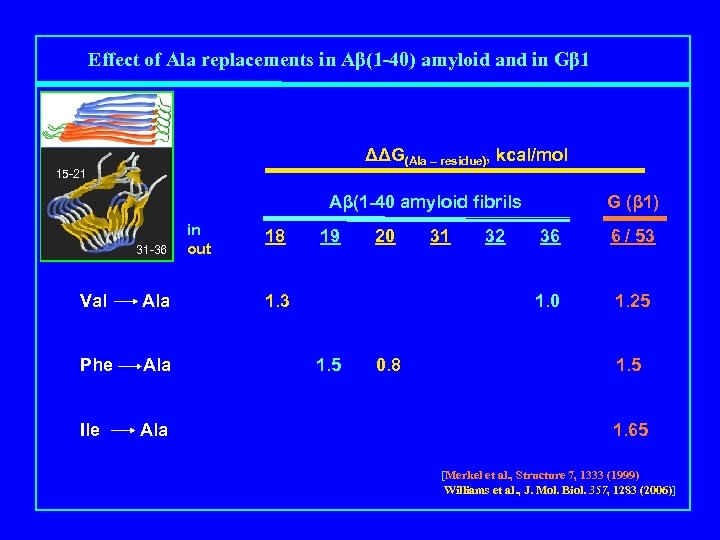

Effect of Ala replacements in Aβ(1 -40) amyloid and in Gβ 1 ΔΔG(Ala – residue), kcal/mol 15 -21 Aβ(1 -40 amyloid fibrils Mutation 31 -36 Val Ala Phe Ala Ile Ala in out 18 19 20 1. 5 0. 8 32 36 6 / 53 1. 0 1. 3 31 G (β 1) 1. 25 1. 65 [Merkel et al. , Structure 7, 1333 (1999) Williams et al. , J. Mol. Biol. 357, 1283 (2006)]
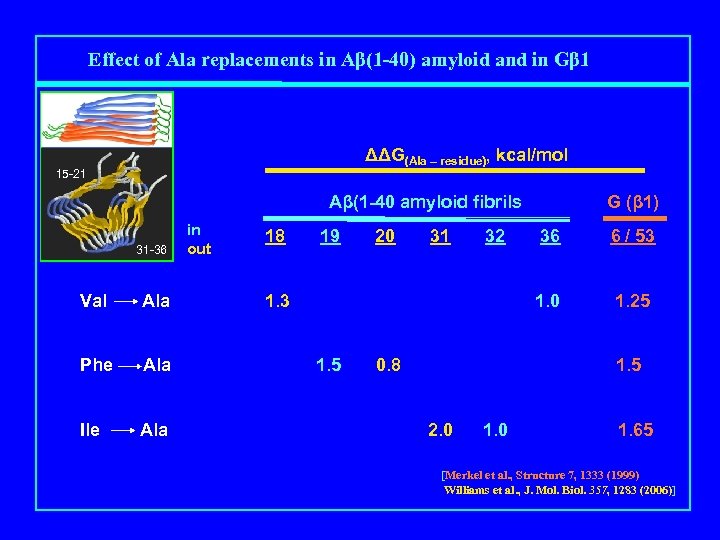

Effect of Ala replacements in Aβ(1 -40) amyloid and in Gβ 1 ΔΔG(Ala – residue), kcal/mol 15 -21 Aβ(1 -40 amyloid fibrils Mutation 31 -36 Val Ala Phe Ala Ile Ala in out 18 19 20 31 32 G (β 1) 1. 5 0. 8 6 / 53 1. 0 1. 3 36 1. 25 1. 5 2. 0 1. 65 [Merkel et al. , Structure 7, 1333 (1999) Williams et al. , J. Mol. Biol. 357, 1283 (2006)]
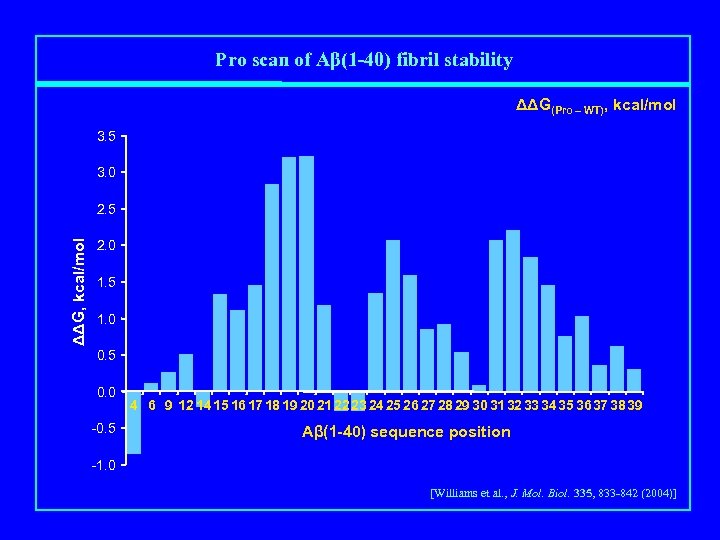
Pro scan of Aβ(1 -40) fibril stability ΔΔG(Pro – WT), kcal/mol 3. 5 3. 0 ΔΔG, kcal/mol 2. 5 2. 0 1. 5 1. 0 0. 5 0. 0 -0. 5 4 6 9 12 14 15 16 1718 19 20 21 22 23 24 25 26 2728 29 30 3132 33 34 35 3637 38 39 Aβ(1 -40) sequence position -1. 0 [Williams et al. , J. Mol. Biol. 335, 833 -842 (2004)]

How Does Proline Destabilize β-Sheet? • Backbone Effects - no N-H proton: lost H-bond - loss of planarity in extended chain • Side Chain Packing Effects - Pro “side chain” is compact loop that does not extend far out of plane
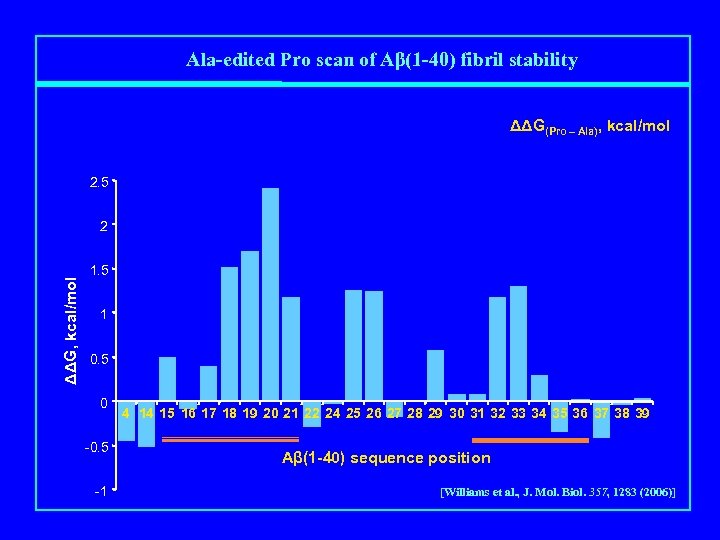
Ala-edited Pro scan of Aβ(1 -40) fibril stability ΔΔG(Pro – Ala), kcal/mol 2. 5 ΔΔG, kcal/mol 2 1. 5 1 0. 5 0 -0. 5 -1 4 14 15 16 17 18 19 20 21 22 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 Aβ(1 -40) sequence position [Williams et al. , J. Mol. Biol. 357, 1283 (2006)]
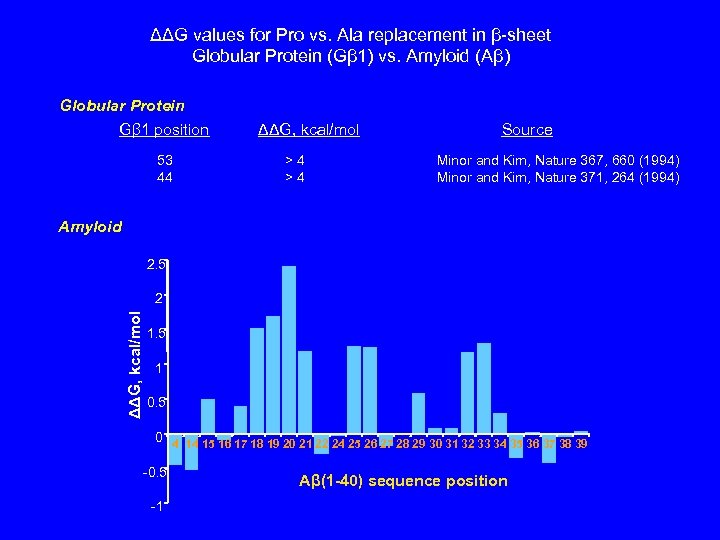
ΔΔG values for Pro vs. Ala replacement in β-sheet Globular Protein (Gβ 1) vs. Amyloid (Aβ) Globular Protein Gβ 1 position 53 44 ΔΔG, kcal/mol >4 >4 Source Minor and Kim, Nature 367, 660 (1994) Minor and Kim, Nature 371, 264 (1994) Amyloid 2. 5 ΔΔG, kcal/mol 2 1. 5 1 0. 5 0 4 14 15 16 17 18 19 20 21 22 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 -0. 5 -1 Aβ(1 -40) sequence position
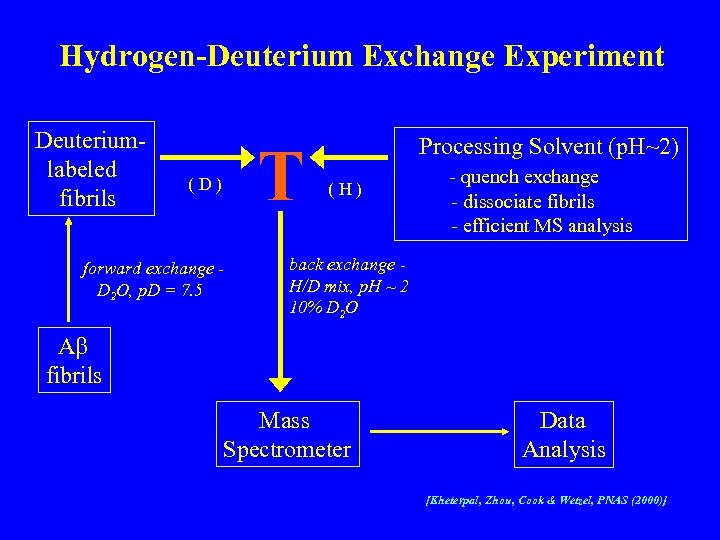
Hydrogen-Deuterium Exchange Experiment Deuteriumlabeled fibrils T (D) forward exchange D 2 O, p. D = 7. 5 Processing Solvent (p. H~2) (H) - quench exchange - dissociate fibrils - efficient MS analysis back exchange H/D mix, p. H ~ 2 10% D 2 O Ab fibrils Mass Spectrometer Data Analysis [Kheterpal, Zhou, Cook & Wetzel, PNAS (2000)]
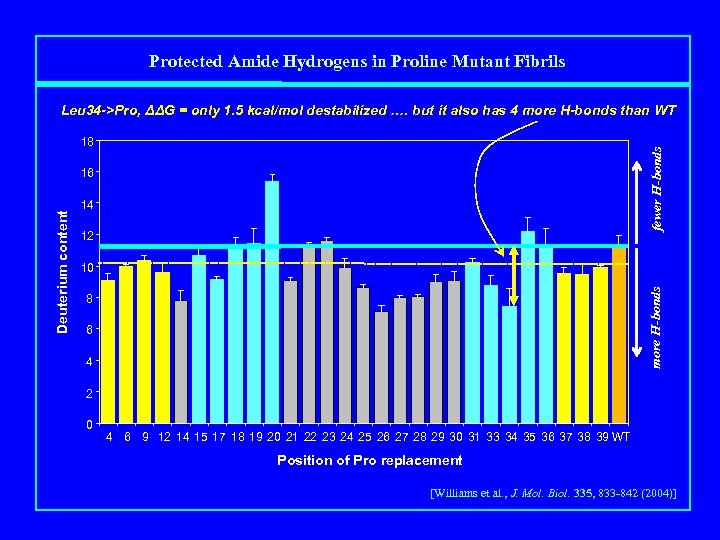
Protected Amide Hydrogens in Proline Mutant Fibrils Leu 34 ->Pro, ΔΔG = only 1. 5 kcal/mol destabilized …. but it also has 4 more H-bonds than WT fewer H-bonds 18 14 12 10 more H-bonds Deuterium content 16 8 6 4 2 0 4 6 9 12 14 15 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 33 34 35 36 37 38 39 WT Position of Pro replacement [Williams et al. , J. Mol. Biol. 335, 833 -842 (2004)]
Thermodynamics of Amyloid Fibril Formation Results: - Aβ(1 -40) fibril growth tends to a reversible equilibrium position with a Keq and ΔG - ΔΔGs from Ala mutations agree with data from parallel β-sheet in globular protein … propagated structural changes suggest a fundamental difference from globular proteins - some ΔΔG effects attributable to energy changes within the monomer ensemble N fibril
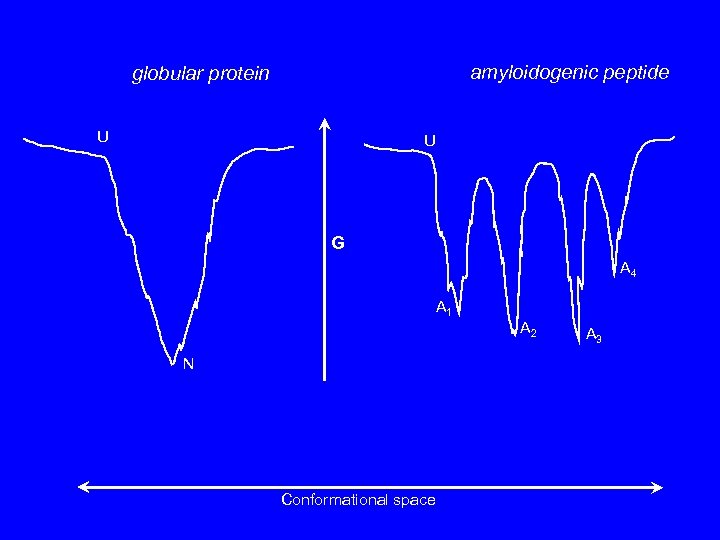
amyloidogenic peptide globular protein U U G A 4 A 1 A 2 N Conformational space A 3
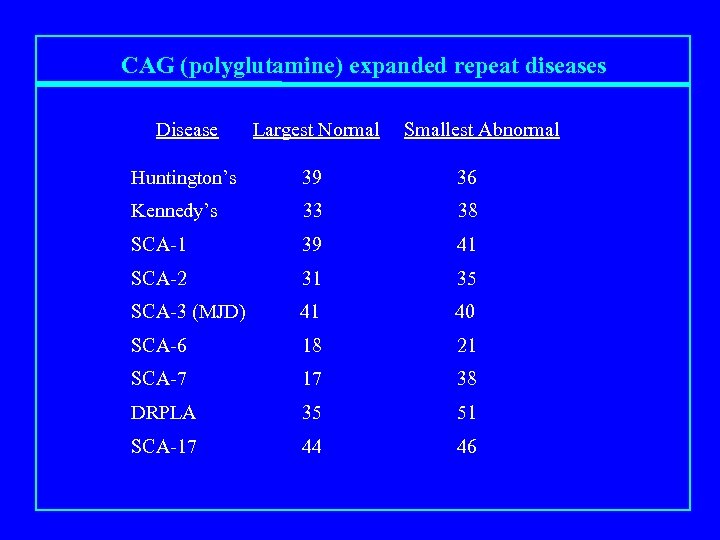
CAG (polyglutamine) expanded repeat diseases Disease Largest Normal Smallest Abnormal Huntington’s 39 36 Kennedy’s 33 38 SCA-1 39 41 SCA-2 31 35 SCA-3 (MJD) 41 40 SCA-6 18 21 SCA-7 17 38 DRPLA 35 51 SCA-17 44 46
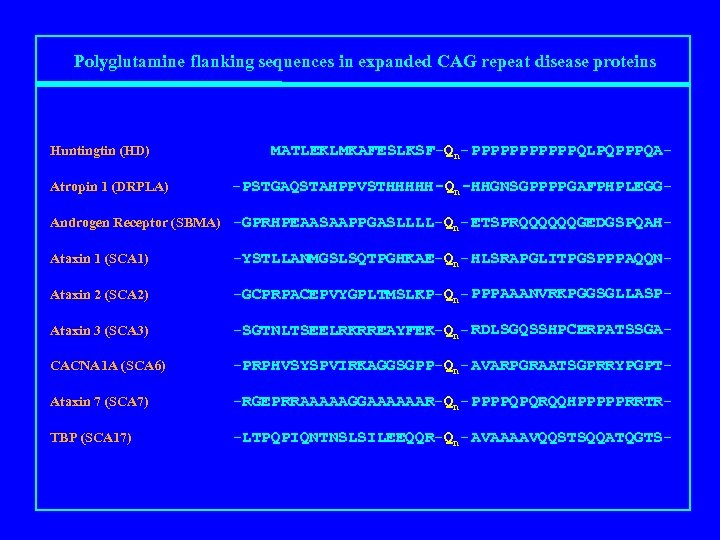
Polyglutamine flanking sequences in expanded CAG repeat disease proteins Huntingtin (HD) Atropin 1 (DRPLA) MATLEKLMKAFESLKSF-Qn- PPPPPPQLPQPPPQA-PSTGAQSTAHPPVSTHHHHH-Qn-HHGNSGPPPPGAFPHPLEGG- Androgen Receptor (SBMA) -GPRHPEAASAAPPGASLLLL-Qn- ETSPRQQQQQQGEDGSPQAHAtaxin 1 (SCA 1) -YSTLLANMGSLSQTPGHKAE-Qn- HLSRAPGLITPGSPPPAQQN- Ataxin 2 (SCA 2) -GCPRPACEPVYGPLTMSLKP-Qn- PPPAAANVRKPGGSGLLASP- Ataxin 3 (SCA 3) -SGTNLTSEELRKRREAYFEK-Qn- RDLSGQSSHPCERPATSSGA- CACNA 1 A (SCA 6) -PRPHVSYSPVIRKAGGSGPP-Qn- AVARPGRAATSGPRRYPGPT- Ataxin 7 (SCA 7) -RGEPRRAAAAAGGAAAAAAR-Qn- PPPPQPQRQQHPPPPPRRTR- TBP (SCA 17) -LTPQPIQNTNSLSILEEQQR-Qn- AVAAAAVQQSTSQQATQGTS-
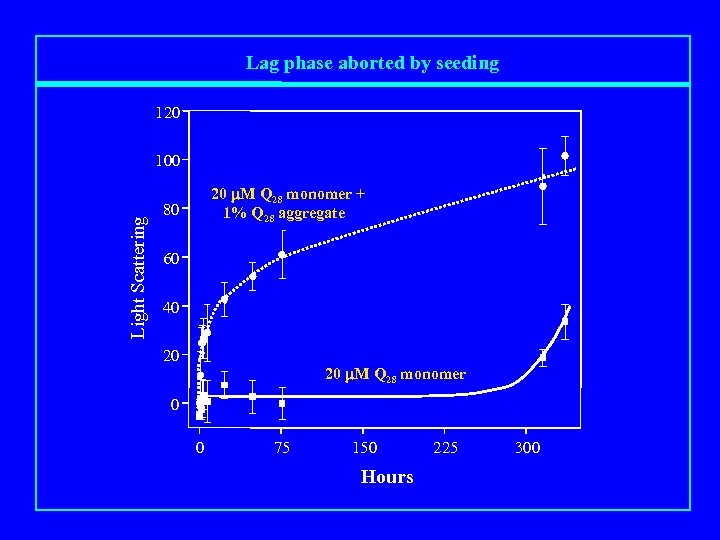
Lag phase aborted by seeding 120 Light Scattering 100 20 m. M Q 28 monomer + 1% Q 28 aggregate 80 60 40 20 20 m. M Q 28 monomer 0 0 75 150 Hours 225 300
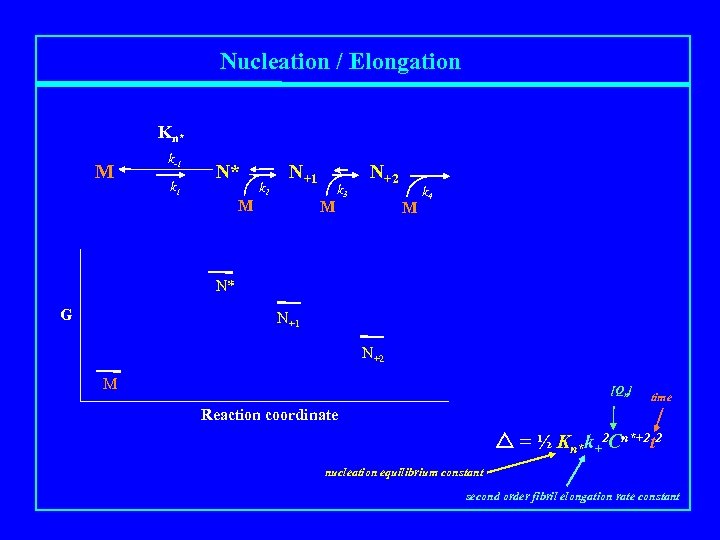
Nucleation / Elongation Kn* M k-1 k 1 N* M k 2 N+1 M k 3 N+2 M k 4 N* G N+1 N+2 M [Qn] time Reaction coordinate = ½ Kn*k+2 Cn*+2 t 2 nucleation equilibrium constant second order fibril elongation rate constant
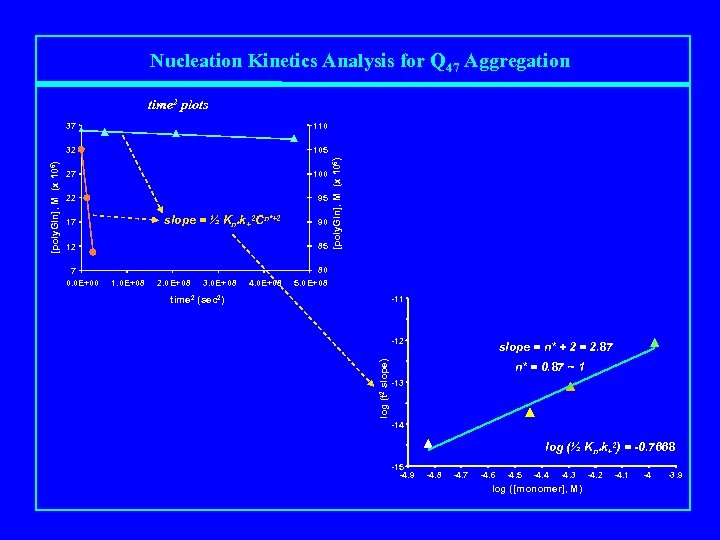
Nucleation Kinetics Analysis for Q 47 Aggregation time 2 plots 105 27 100 22 95 slope = ½ Kn*k+2 Cn*+2 85 12 7 0. 0 E+00 90 1. 0 E+08 2. 0 E+08 3. 0 E+08 4. 0 E+08 80 5. 0 E+08 -11 time 2 (sec 2) -12 log (t 2 slope) 17 [poly. Gln], M (x 106) 110 32 [poly. Gln], M (x 106) 37 slope = n* + 2 = 2. 87 n* = 0. 87 ~ 1 -13 -14 log (½ Kn*k+2) = -0. 7668 -15 -4. 9 -4. 8 -4. 7 -4. 6 -4. 5 -4. 4 -4. 3 log ([monomer], M) -4. 2 -4. 1 -4 -3. 9
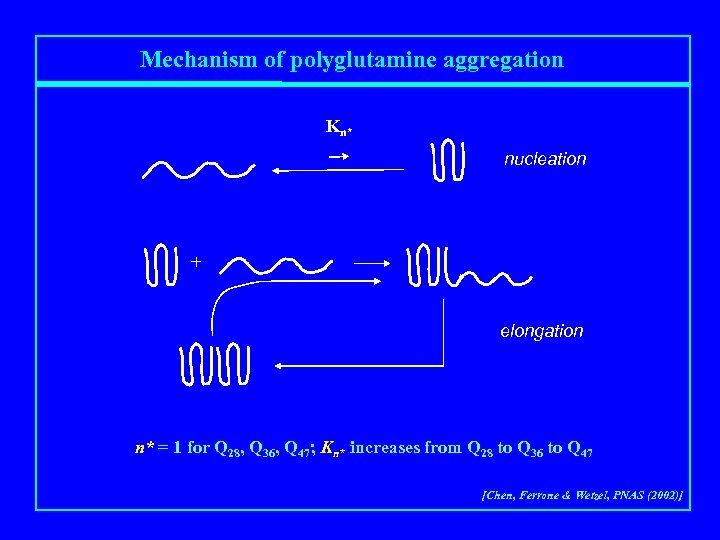
Mechanism of polyglutamine aggregation Kn* nucleation + elongation n* = 1 for Q 28, Q 36, Q 47; Kn* increases from Q 28 to Q 36 to Q 47 [Chen, Ferrone & Wetzel, PNAS (2002)]
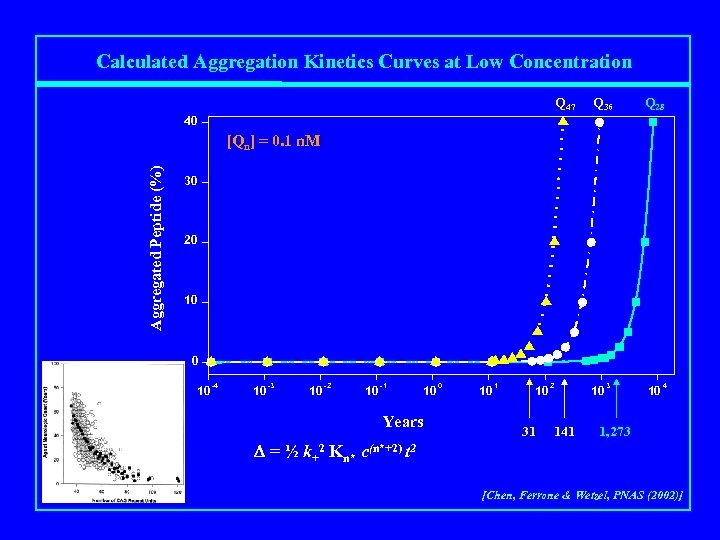
Calculated Aggregation Kinetics Curves at Low Concentration Q 47 Q 36 Q 28 40 Aggregated Peptide (%) [Qn] = 0. 1 n. M 30 20 10 -4 10 -3 10 -2 10 -1 Years 10 0 10 1 10 31 2 141 10 3 10 4 1, 273 D = ½ k+2 Kn* c(n*+2) t 2 [Chen, Ferrone & Wetzel, PNAS (2002)]
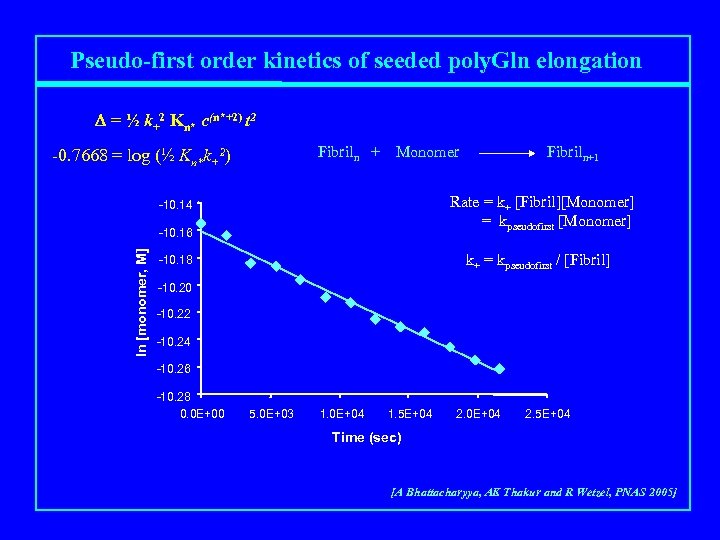
Pseudo-first order kinetics of seeded poly. Gln elongation D = ½ k+2 Kn* c(n*+2) t 2 Fibriln + -0. 7668 = log (½ Kn*k+2) Monomer Rate = k+ [Fibril][Monomer] = kpseudofirst [Monomer] -10. 14 -10. 16 ln [monomer, M] Fibriln+1 k+ = kpseudofirst / [Fibril] -10. 18 -10. 20 -10. 22 -10. 24 -10. 26 -10. 28 0. 0 E+00 5. 0 E+03 1. 0 E+04 1. 5 E+04 2. 0 E+04 2. 5 E+04 Time (sec) [A Bhattacharyya, AK Thakur and R Wetzel, PNAS 2005]
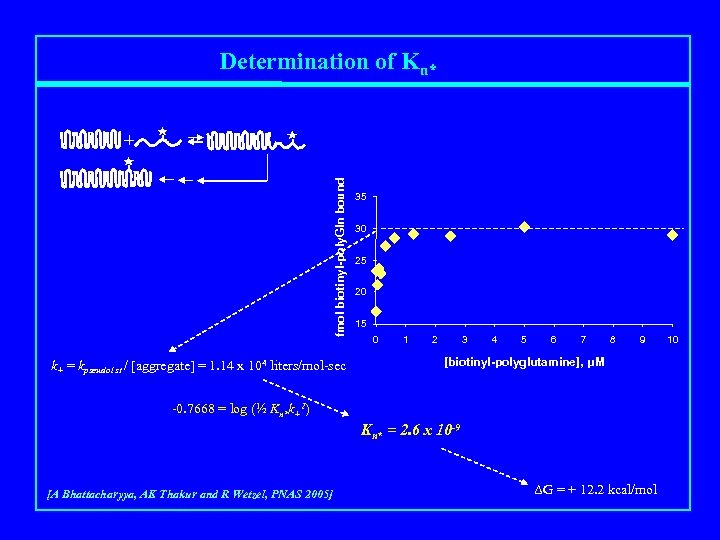
Determination of Kn* fmol biotinyl-poly. Gln bound + k+ = kpseudo 1 st / [aggregate] = 1. 14 x 104 liters/mol-sec 35 30 25 20 15 0 1 2 3 4 5 6 7 8 9 [biotinyl-polyglutamine], μM -0. 7668 = log (½ Kn*k+2) Kn* = 2. 6 x 10 -9 [A Bhattacharyya, AK Thakur and R Wetzel, PNAS 2005] ΔG = + 12. 2 kcal/mol 10
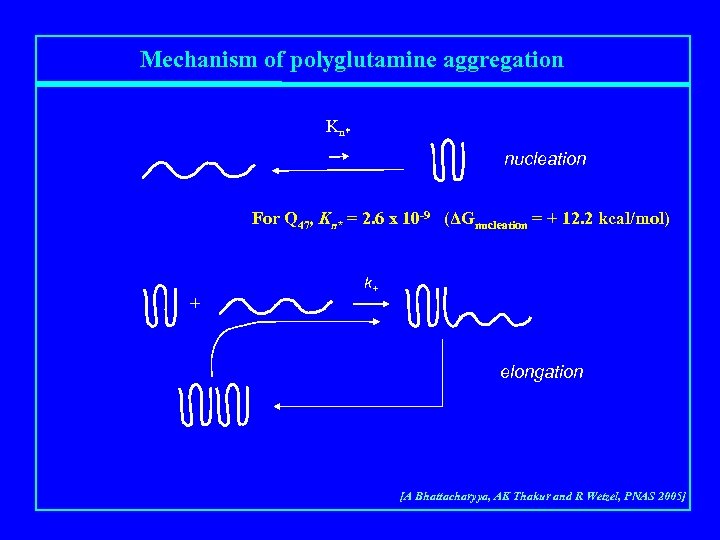
Mechanism of polyglutamine aggregation Kn* nucleation For Q 47, Kn* = 2. 6 x 10 -9 (ΔGnucleation = + 12. 2 kcal/mol) + k+ elongation [A Bhattacharyya, AK Thakur and R Wetzel, PNAS 2005]

Acknowledgments Aβ Team Poly. Gln Team Angela Williams Shankari Shivaprasad Anusri Bhattacharyya Ashwani Thakur Brian O’Nuallain Indu Kheterpal Eric Portelius Songming Chen Frank Ferrone (Drexel Univ. ) Trevor Creamer (Univ. Kentucky) Veronique Hermann (Univ. Kentucky)

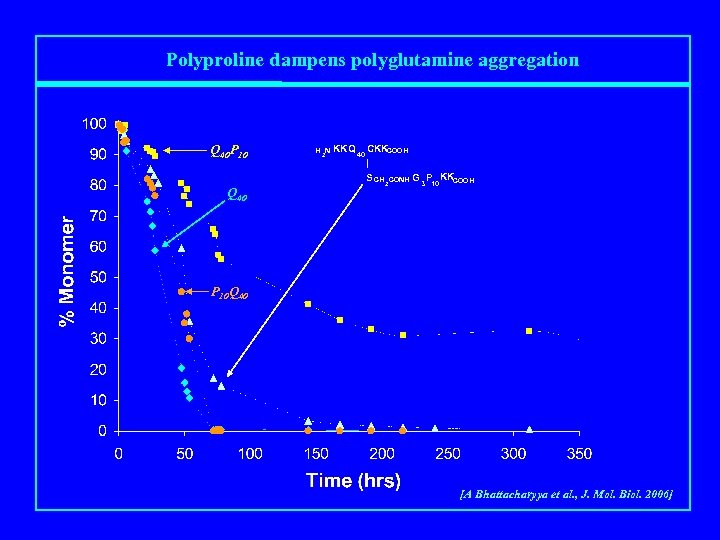
Polyproline dampens polyglutamine aggregation Q 40 P 10 Q 40 H 2 N KKQ 40 CKKCOOH | S CH 2 CONH G 3 P KKCOOH 10 P 10 Q 40 [A Bhattacharyya et al. , J. Mol. Biol. 2006]
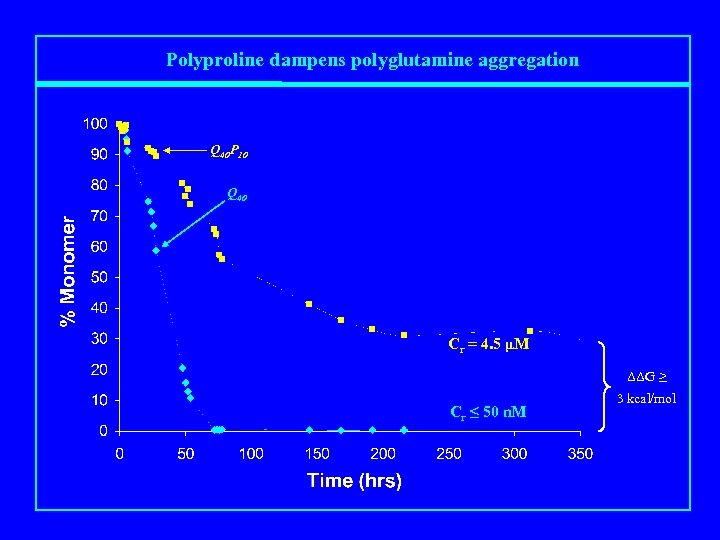
Polyproline dampens polyglutamine aggregation Q 40 P 10 Q 40 Cr = 4. 5 μM ΔΔG ≥ Cr ≤ 50 n. M 3 kcal/mol
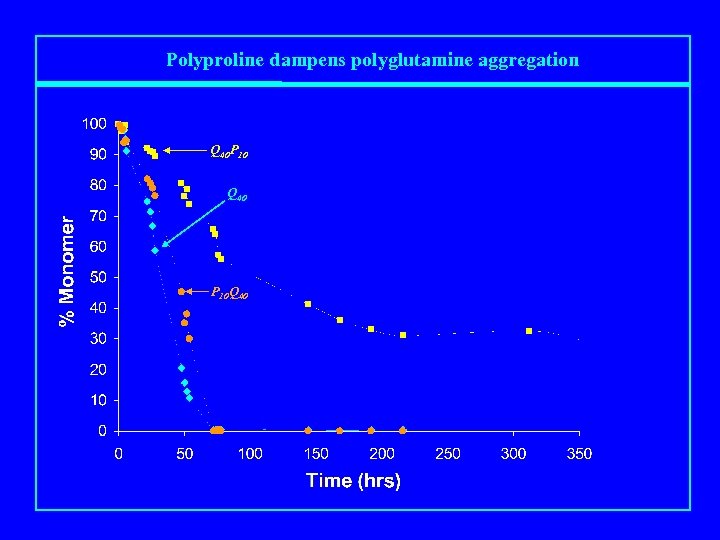
Polyproline dampens polyglutamine aggregation Q 40 P 10 Q 40 P 10 Q 40
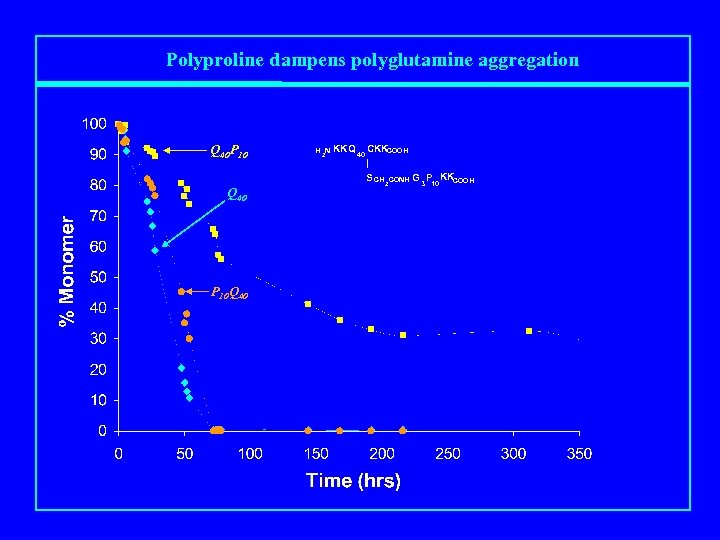
Polyproline dampens polyglutamine aggregation Q 40 P 10 Q 40 P 10 Q 40 H 2 N KKQ 40 CKKCOOH | S CH 2 CONH G 3 P KKCOOH 10
![Is the Plateau a Real Thermodynamic Cr? 12 [Monomer], µM 10 8 Q 40 Is the Plateau a Real Thermodynamic Cr? 12 [Monomer], µM 10 8 Q 40](https://present5.com/presentation/4e2cb46c65b3324d68bf03d090982437/image-40.jpg)
Is the Plateau a Real Thermodynamic Cr? 12 [Monomer], µM 10 8 Q 40 6 Q 40 P 10 4 2 Q 40 P 10 Q 40 0 0 100 200 300 400 Time (hrs) [A Bhattacharyya et al. , J. Mol. Biol. 2006]
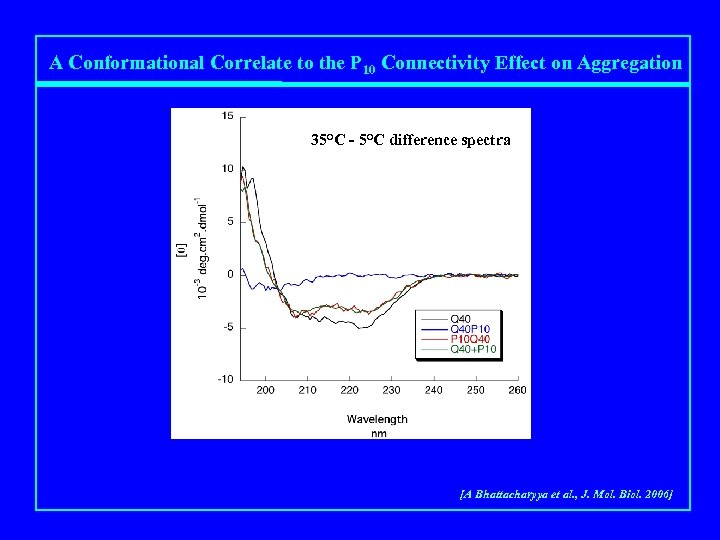
A Conformational Correlate to the P 10 Connectivity Effect on Aggregation 35°C - 5°C difference spectra [A Bhattacharyya et al. , J. Mol. Biol. 2006]
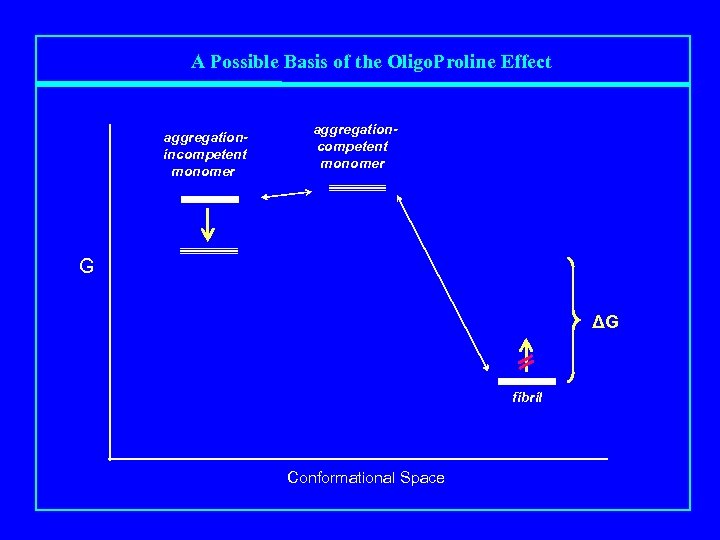
A Possible Basis of the Oligo. Proline Effect aggregationincompetent monomer aggregationcompetent monomer G ΔG fibril Conformational Space

Transportability of the P 10 Effect Peptide Aβ(1 -40)-P 10 C r, μM 0. 9 μM 21. 5 μM [A Bhattacharyya et al. , J. Mol. Biol. 2006]
![Side Chain Packing by Disulfide Formation HS HS SH HS [O] HS SH SH Side Chain Packing by Disulfide Formation HS HS SH HS [O] HS SH SH](https://present5.com/presentation/4e2cb46c65b3324d68bf03d090982437/image-44.jpg)
Side Chain Packing by Disulfide Formation HS HS SH HS [O] HS SH SH HS S [S. Shivaprasad and R. Wetzel, Biochem. 43, 15310 (2004)] S
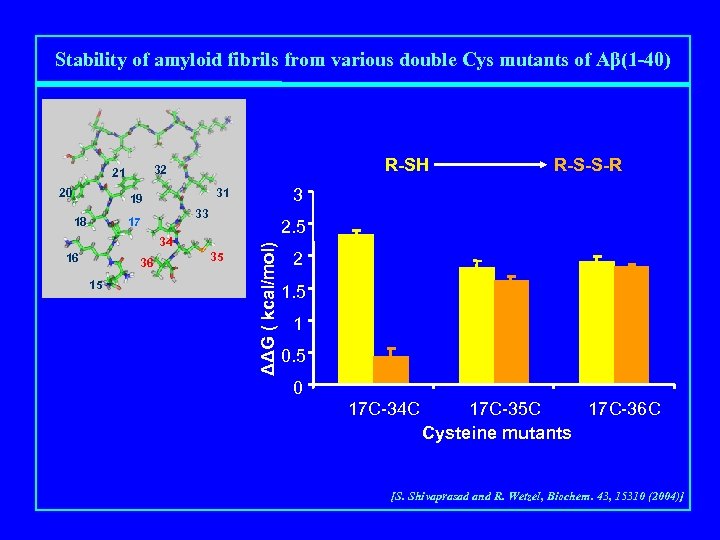
Stability of amyloid fibrils from various double Cys mutants of Aβ(1 -40) 33 17 18 2. 5 34 16 36 15 R-S-S-R 3 31 19 35 ΔΔG ( kcal/mol) 20 R-SH 32 21 2 1. 5 1 0. 5 0 17 C-34 C 17 C-35 C 17 C-36 C Cysteine mutants [S. Shivaprasad and R. Wetzel, Biochem. 43, 15310 (2004)]
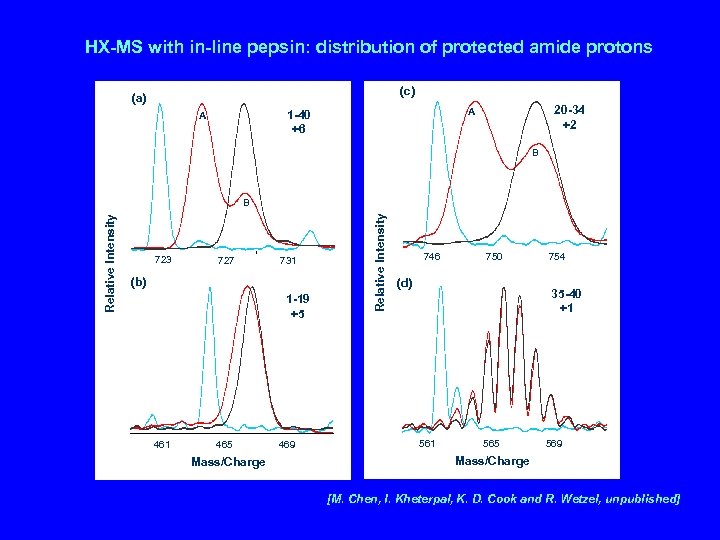
HX-MS with in-line pepsin: distribution of protected amide protons (c) (a) 20 -34 +2 A 1 -40 +6 A B 723 727 731 (b) 1 -19 +5 461 465 Mass/Charge 469 Relative Intensity B 746 750 (d) 754 35 -40 +1 565 569 Mass/Charge [M. Chen, I. Kheterpal, K. D. Cook and R. Wetzel, unpublished]
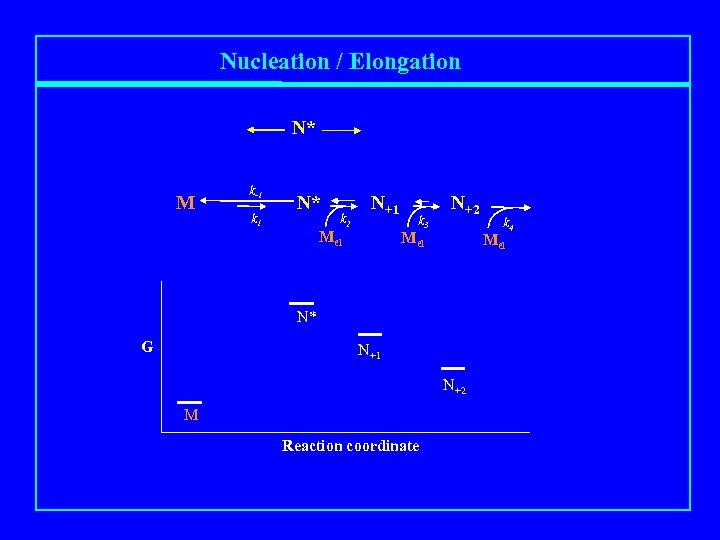
Nucleation / Elongation N* M k-1 k 1 N* k 2 N+1 Mel k 3 N+2 Mel N* G N+1 N+2 M Reaction coordinate k 4
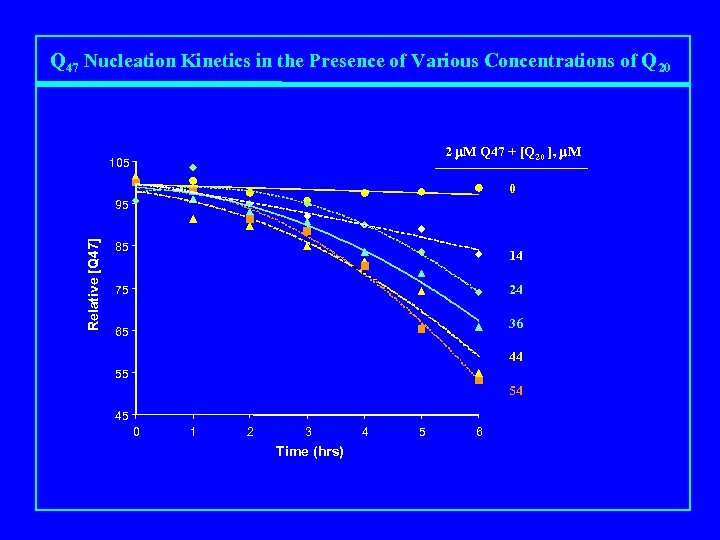
Q 47 Nucleation Kinetics in the Presence of Various Concentrations of Q 20 2 m. M Q 47 + [Q 20 ], m. M 105 0 Relative [Q 47] 95 85 14 24 75 36 65 44 55 54 45 0 1 2 3 Time (hrs) 4 5 6
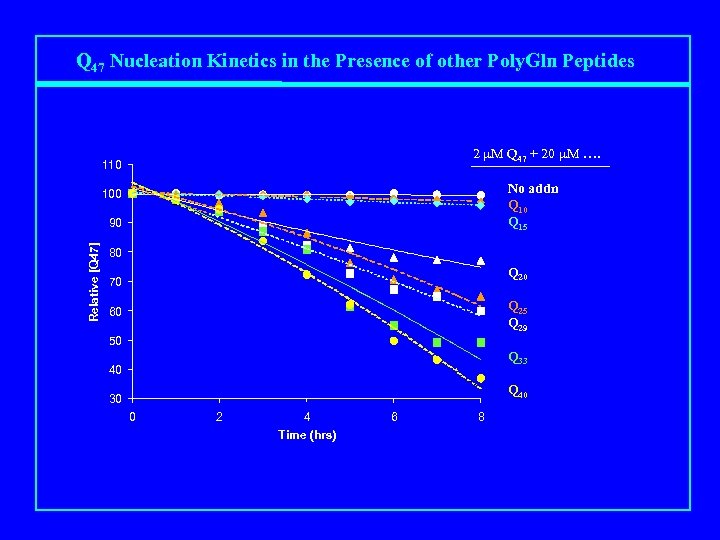
Q 47 Nucleation Kinetics in the Presence of other Poly. Gln Peptides 2 m. M Q 47 + 20 m. M …. 110 No addn Q 10 Q 15 100 Relative [Q 47] 90 80 Q 20 70 Q 25 Q 29 60 50 Q 33 40 Q 40 30 0 2 4 Time (hrs) 6 8
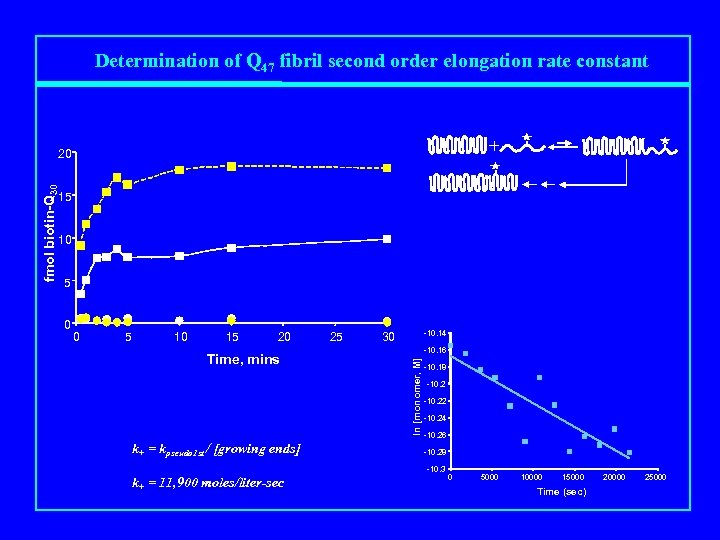
Determination of Q 47 fibril second order elongation rate constant + fmol biotin-Q 30 20 15 10 15 20 Time, mins k+ = kpseudo 1 st / [growing ends] 25 -10. 14 30 -10. 16 ln [monomer, M] 0 -10. 18 -10. 22 -10. 24 -10. 26 -10. 28 -10. 3 k+ = 11, 900 moles/liter-sec 0 5000 10000 15000 Time (sec) 20000 25000

How is amyloid formation initiated? Polyglutamine studies There are no kinetically relevant intermediates in nucleation of simple poly. Gln peptides Results: - the nucleus for poly. Gln aggregation is an energetically unfavorable monomer - repeat length dependent nucleation efficiency may help account for ages-of-onset - Kn* for a Q 47 peptide is ~ 10 -9 - short poly. Gln peptides in the environment can enhance nucleation efficiency
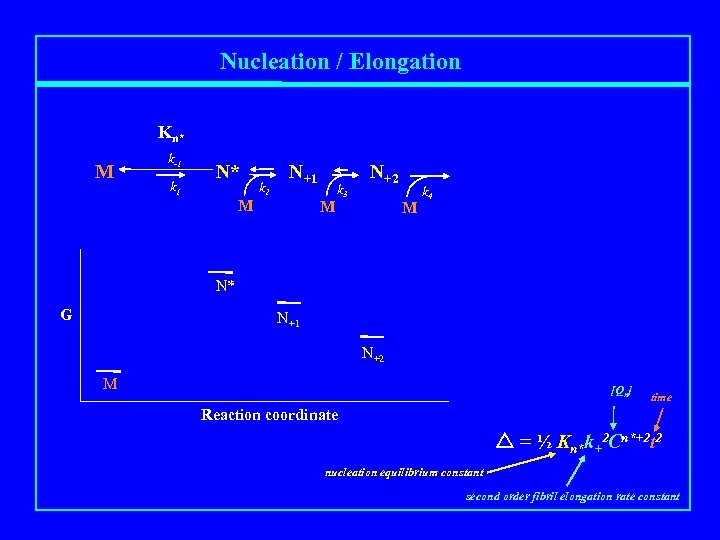
Nucleation / Elongation Kn* M k-1 k 1 N* M k 2 N+1 M k 3 N+2 M k 4 N* G N+1 N+2 M [Qn] time Reaction coordinate = ½ Kn*k+2 Cn*+2 t 2 nucleation equilibrium constant second order fibril elongation rate constant
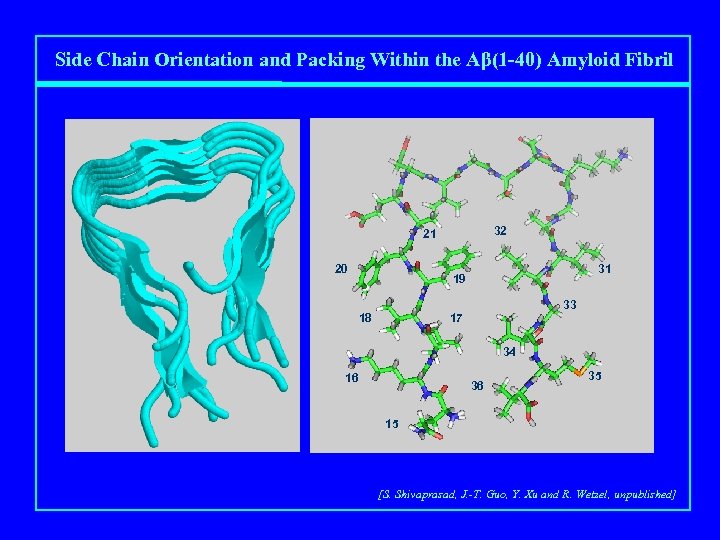
Side Chain Orientation and Packing Within the Aβ(1 -40) Amyloid Fibril 32 21 20 31 19 33 17 18 34 16 36 35 15 [S. Shivaprasad, J. -T. Guo, Y. Xu and R. Wetzel, unpublished]
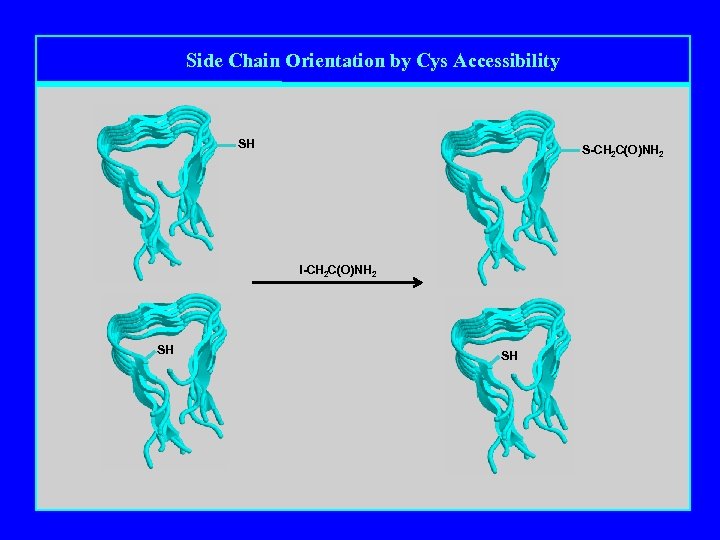
Side Chain Orientation by Cys Accessibility SH S-CH 2 C(O)NH 2 I-CH 2 C(O)NH 2 SH SH
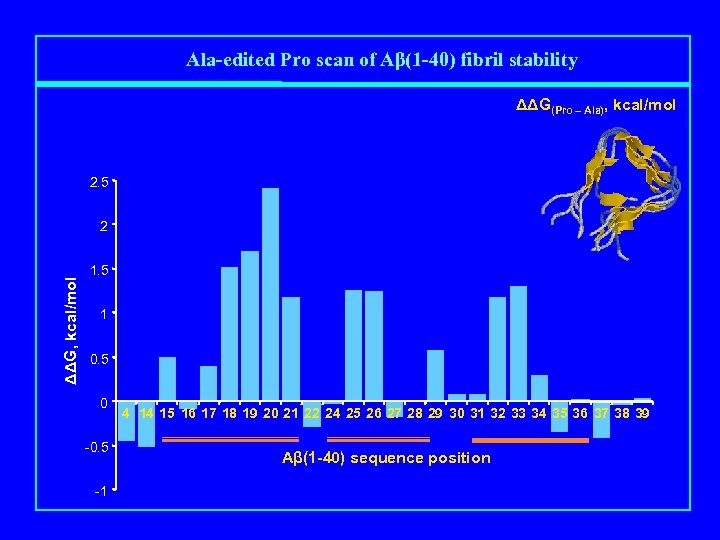
Ala-edited Pro scan of Aβ(1 -40) fibril stability ΔΔG(Pro – Ala), kcal/mol 2. 5 ΔΔG, kcal/mol 2 1. 5 1 0. 5 0 -0. 5 -1 4 14 15 16 17 18 19 20 21 22 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 Aβ(1 -40) sequence position
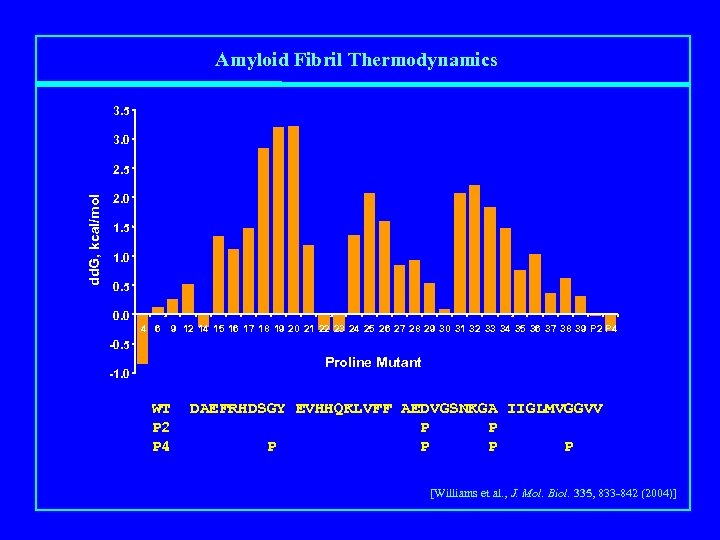
Amyloid Fibril Thermodynamics 3. 5 3. 0 dd. G, kcal/mol 2. 5 2. 0 1. 5 1. 0 0. 5 0. 0 4 6 9 12 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 P 2 P 4 -0. 5 Proline Mutant -1. 0 WT P 2 P 4 DAEFRHDSGY EVHHQKLVFF AEDVGSNKGA IIGLMVGGVV P P P [Williams et al. , J. Mol. Biol. 335, 833 -842 (2004)]
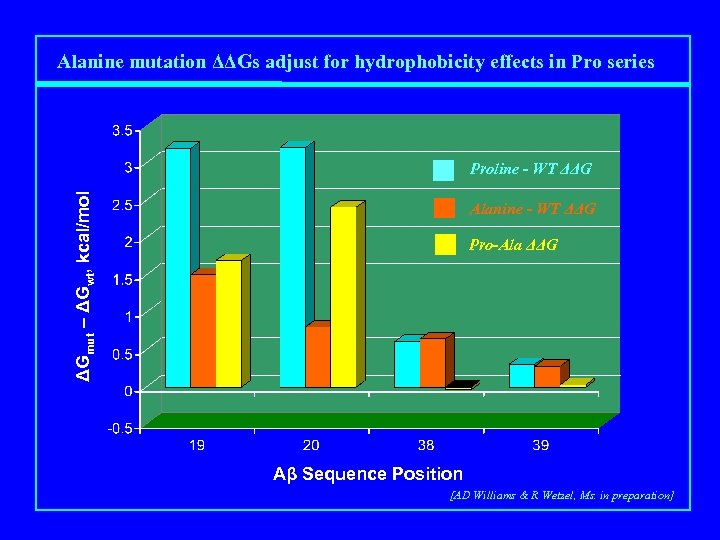
Alanine mutation ΔΔGs adjust for hydrophobicity effects in Pro series ΔGmut – ΔGwt, kcal/mol Proline - WT ΔΔG Alanine - WT ΔΔG Pro-Ala ΔΔG Aβ Sequence Position [AD Williams & R Wetzel, Ms. in preparation]
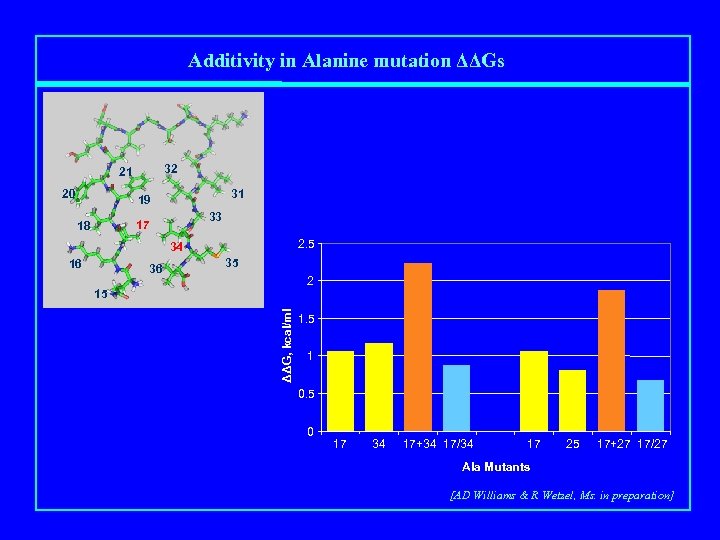
Additivity in Alanine mutation ΔΔGs 32 21 20 31 19 33 17 18 2. 5 34 16 36 35 2 ΔΔG, kcal/ml 15 1. 5 1 0. 5 0 17 34 17+34 17/34 17 25 17+27 17/27 Ala Mutants [AD Williams & R Wetzel, Ms. in preparation]
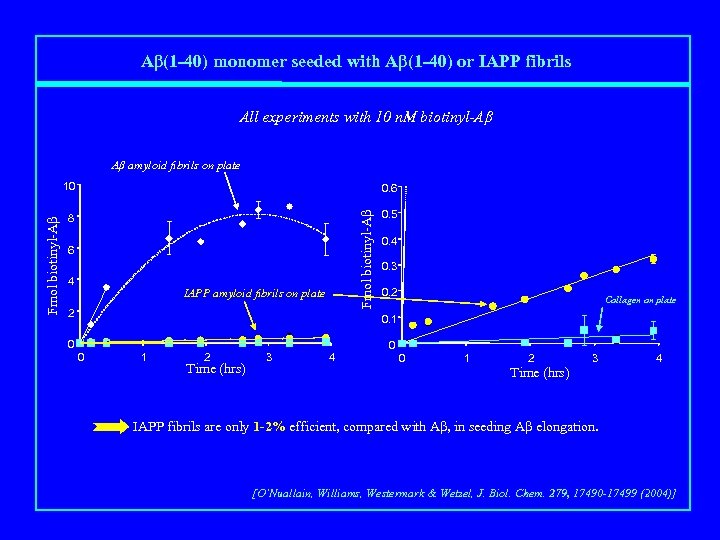
Aβ(1 -40) monomer seeded with Aβ(1 -40) or IAPP fibrils All experiments with 10 n. M biotinyl-Aβ Aβ amyloid fibrils on plate 0. 6 8 0. 5 Fmol biotinyl-Aβ 10 6 4 IAPP amyloid fibrils on plate 2 0. 4 0. 3 0. 2 Collagen on plate 0. 1 0 0 0 1 2 Time (hrs) 3 4 0 1 2 3 4 Time (hrs) IAPP fibrils are only 1 -2% efficient, compared with Aβ, in seeding Aβ elongation. [O’Nuallain, Williams, Westermark & Wetzel, J. Biol. Chem. 279, 17490 -17499 (2004)]
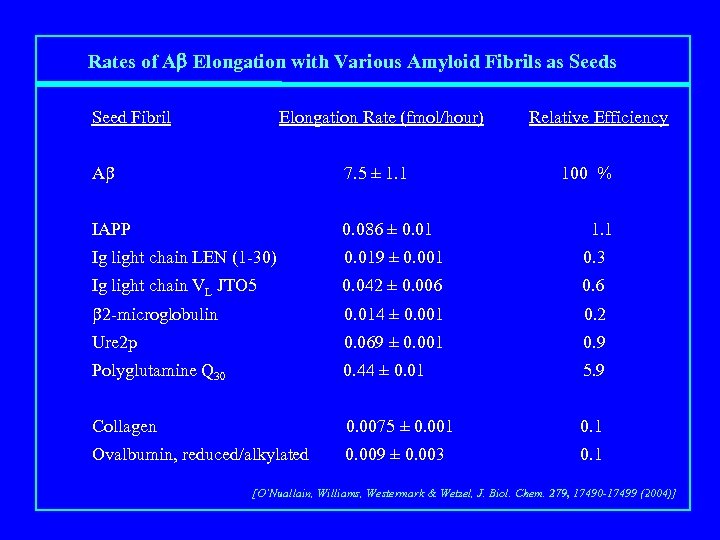
Rates of Ab Elongation with Various Amyloid Fibrils as Seed Fibril Elongation Rate (fmol/hour) Relative Efficiency Ab 7. 5 ± 1. 1 100 % IAPP 0. 086 ± 0. 01 Ig light chain LEN (1 -30) 0. 019 ± 0. 001 0. 3 Ig light chain VL JTO 5 0. 042 ± 0. 006 0. 6 b 2 -microglobulin 0. 014 ± 0. 001 0. 2 Ure 2 p 0. 069 ± 0. 001 0. 9 Polyglutamine Q 30 0. 44 ± 0. 01 5. 9 Collagen 0. 0075 ± 0. 001 0. 1 Ovalbumin, reduced/alkylated 0. 009 ± 0. 003 0. 1 1. 1 [O’Nuallain, Williams, Westermark & Wetzel, J. Biol. Chem. 279, 17490 -17499 (2004)]
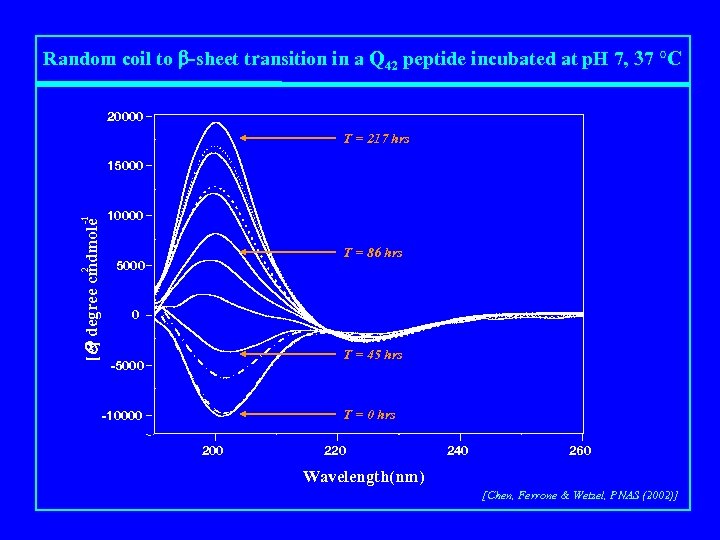
Random coil to b-sheet transition in a Q 42 peptide incubated at p. H 7, 37 °C 20000 T = 217 hrs -1 5000 [Q] degree cmdmole 10000 2 15000 T = 86 hrs 0 T = 45 hrs -5000 T = 0 hrs -10000 220 240 260 Wavelength(nm) [Chen, Ferrone & Wetzel, PNAS (2002)]
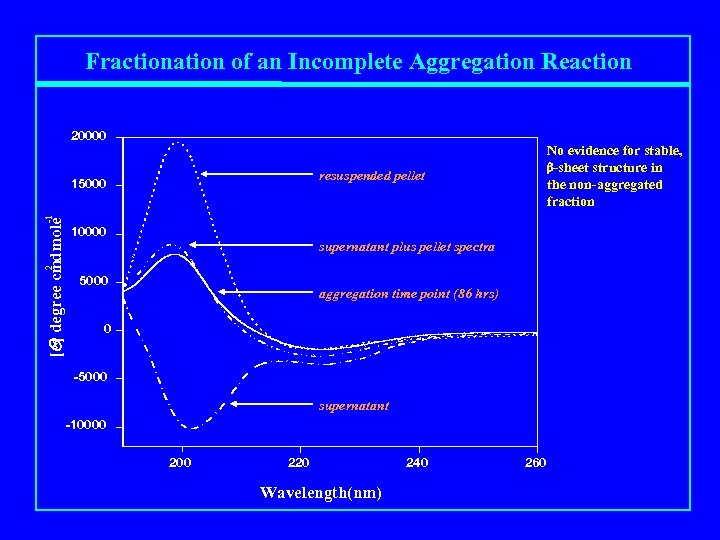
Fractionation of an Incomplete Aggregation Reaction 20000 No evidence for stable, b-sheet structure in the non-aggregated fraction resuspended pellet 10000 supernatant plus pellet spectra 2 [Q] degree cmdmole -1 15000 aggregation time point (86 hrs) 0 -5000 supernatant -10000 220 Wavelength(nm) 240 260
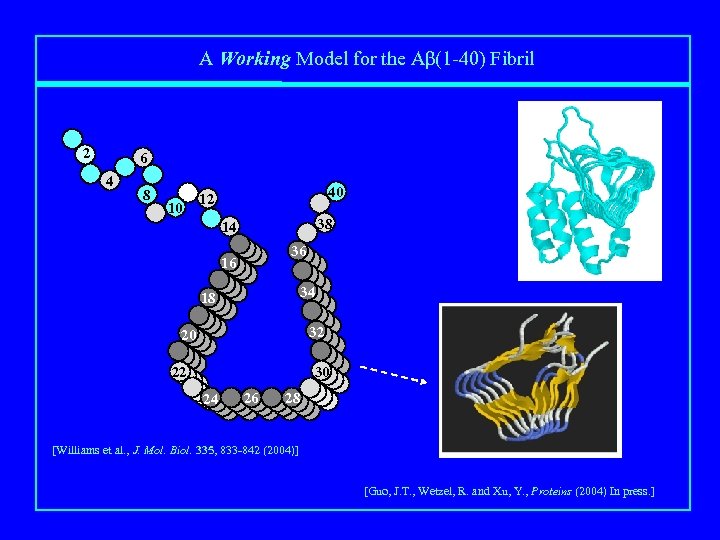

A Working Model for the Aβ(1 -40) Fibril 2 6 4 8 10 40 12 38 14 36 16 34 18 32 20 30 22 24 26 28 [Williams et al. , J. Mol. Biol. 335, 833 -842 (2004)] [Guo, J. T. , Wetzel, R. and Xu, Y. , Proteins (2004) In press. ]
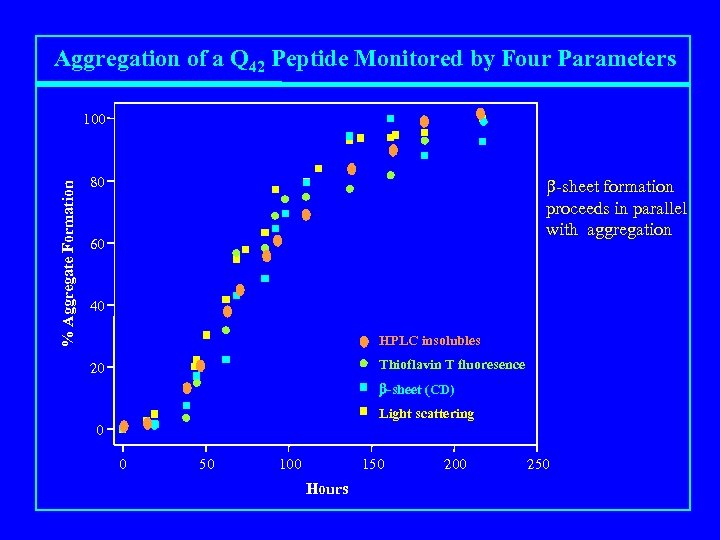
Aggregation of a Q 42 Peptide Monitored by Four Parameters % Aggregate Formation 100 80 b-sheet formation proceeds in parallel with aggregation 60 40 HPLC insolubles Thioflavin T fluoresence 20 b-sheet (CD) Light scattering 0 0 50 100 150 Hours 200 250
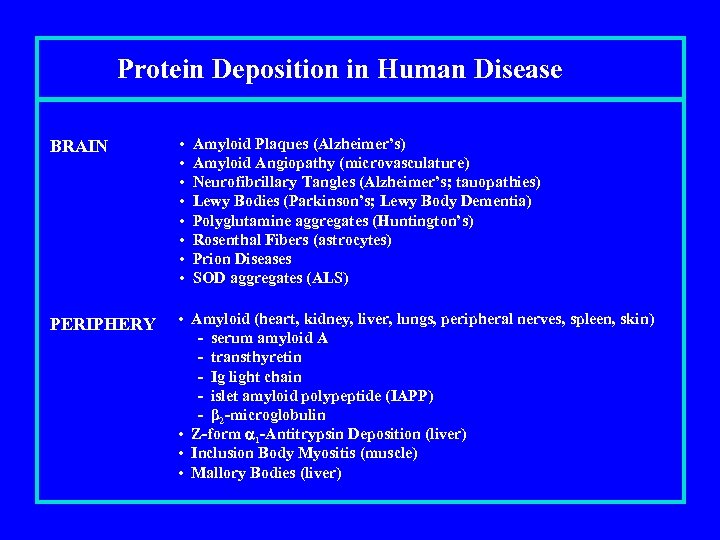
Protein Deposition in Human Disease BRAIN • • PERIPHERY • Amyloid (heart, kidney, liver, lungs, peripheral nerves, spleen, skin) - serum amyloid A - transthyretin - Ig light chain - islet amyloid polypeptide (IAPP) - β 2 -microglobulin • Z-form 1 -Antitrypsin Deposition (liver) • Inclusion Body Myositis (muscle) • Mallory Bodies (liver) Amyloid Plaques (Alzheimer’s) Amyloid Angiopathy (microvasculature) Neurofibrillary Tangles (Alzheimer’s; tauopathies) Lewy Bodies (Parkinson’s; Lewy Body Dementia) Polyglutamine aggregates (Huntington’s) Rosenthal Fibers (astrocytes) Prion Diseases SOD aggregates (ALS)
![Seeded amyloid growth from Aβ(1 -40) 30 Th. T fluorescence or [Aβ] (μM) 25 Seeded amyloid growth from Aβ(1 -40) 30 Th. T fluorescence or [Aβ] (μM) 25](https://present5.com/presentation/4e2cb46c65b3324d68bf03d090982437/image-66.jpg)
Seeded amyloid growth from Aβ(1 -40) 30 Th. T fluorescence or [Aβ] (μM) 25 20 15 10 [Aβ(1 -40)] 5 Cr 0 0 0. 5 1 1. 5 2 Time (h) 2. 5 3 3. 5 4
![Seeded amyloid growth from Aβ(1 -40) 30 Th. T fluorescence or [Aβ] (μM) 25 Seeded amyloid growth from Aβ(1 -40) 30 Th. T fluorescence or [Aβ] (μM) 25](https://present5.com/presentation/4e2cb46c65b3324d68bf03d090982437/image-67.jpg)
Seeded amyloid growth from Aβ(1 -40) 30 Th. T fluorescence or [Aβ] (μM) 25 20 15 10 [Aβ(1 -40)] 5 0 0 0. 5 1 1. 5 2 Time (h) 2. 5 3 3. 5 4
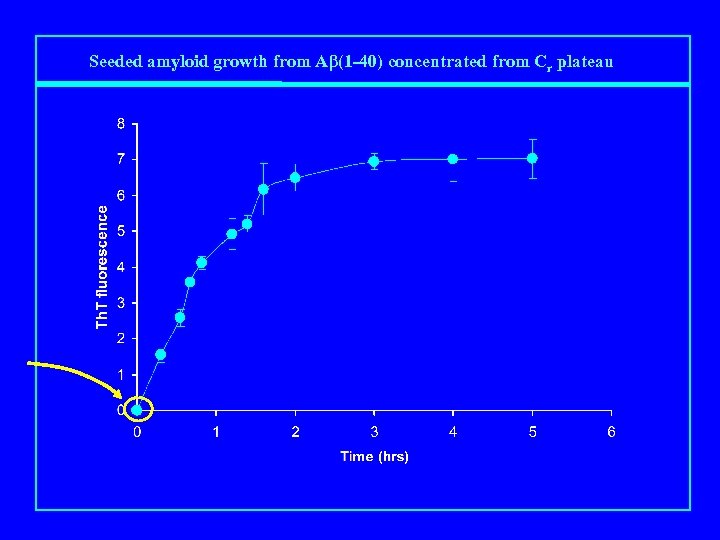
Seeded amyloid growth from Aβ(1 -40) concentrated from Cr plateau
![Seeded amyloid growth from Aβ(1 -40) Th. T fluorescence or [Aβ] (μM) 30 Th. Seeded amyloid growth from Aβ(1 -40) Th. T fluorescence or [Aβ] (μM) 30 Th.](https://present5.com/presentation/4e2cb46c65b3324d68bf03d090982437/image-69.jpg)
Seeded amyloid growth from Aβ(1 -40) Th. T fluorescence or [Aβ] (μM) 30 Th. T 25 20 15 10 [Aβ(1 -40)] 5 0 0 0. 5 1 1. 5 2 Time (h) 2. 5 3 3. 5 4
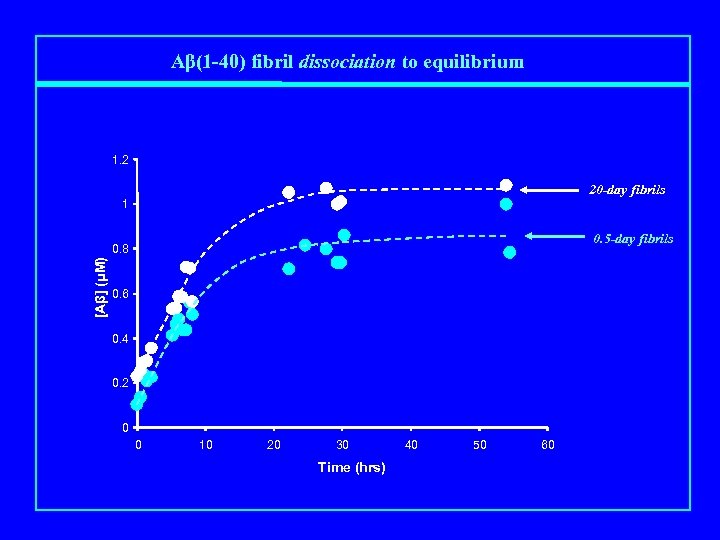
Aβ(1 -40) fibril dissociation to equilibrium 1. 2 20 -day fibrils 1 0. 5 -day fibrils [Aβ] (μM) 0. 8 0. 6 0. 4 0. 2 0 0 10 20 30 Time (hrs) 40 50 60
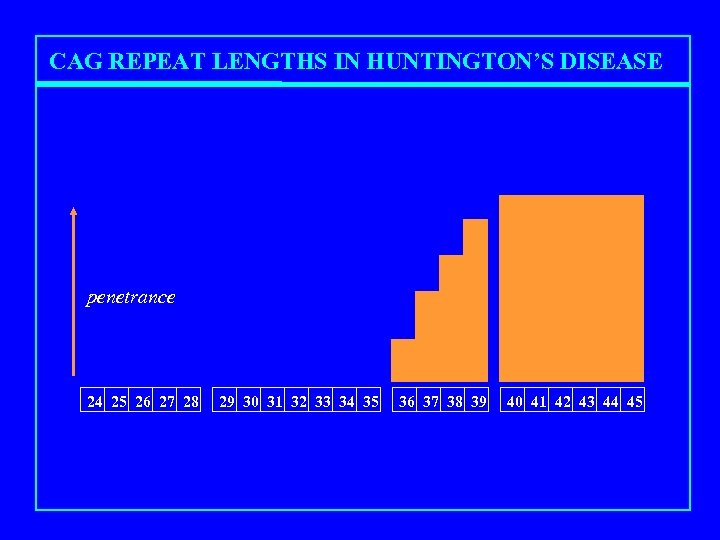
CAG REPEAT LENGTHS IN HUNTINGTON’S DISEASE penetrance 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45
![Repeat Length Dependence of Age of Onset in Huntington’s Disease [Courtesy Marcy Mac. Donald] Repeat Length Dependence of Age of Onset in Huntington’s Disease [Courtesy Marcy Mac. Donald]](https://present5.com/presentation/4e2cb46c65b3324d68bf03d090982437/image-72.jpg)
Repeat Length Dependence of Age of Onset in Huntington’s Disease [Courtesy Marcy Mac. Donald]
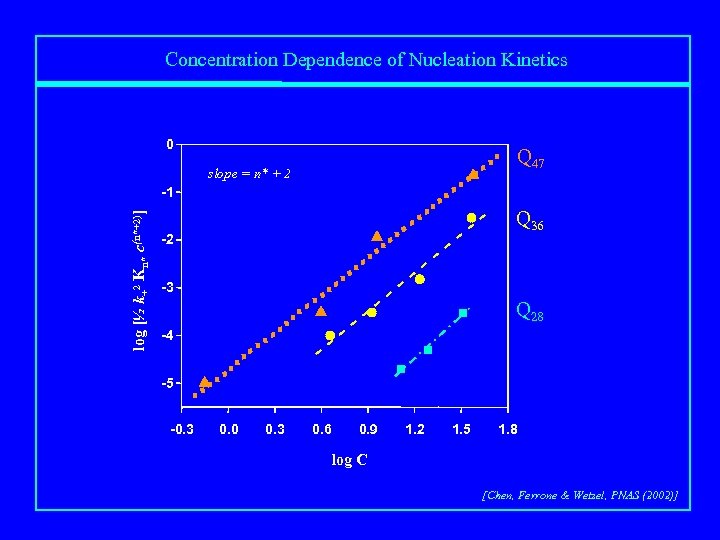
Concentration Dependence of Nucleation Kinetics 0 Q 47 slope = n* + 2 log [½ k+2 Kn* c(n*+2)] -1 Q 36 -2 -3 Q 28 -4 -5 -0. 3 0. 0 0. 3 0. 6 0. 9 1. 2 1. 5 1. 8 log C [Chen, Ferrone & Wetzel, PNAS (2002)]
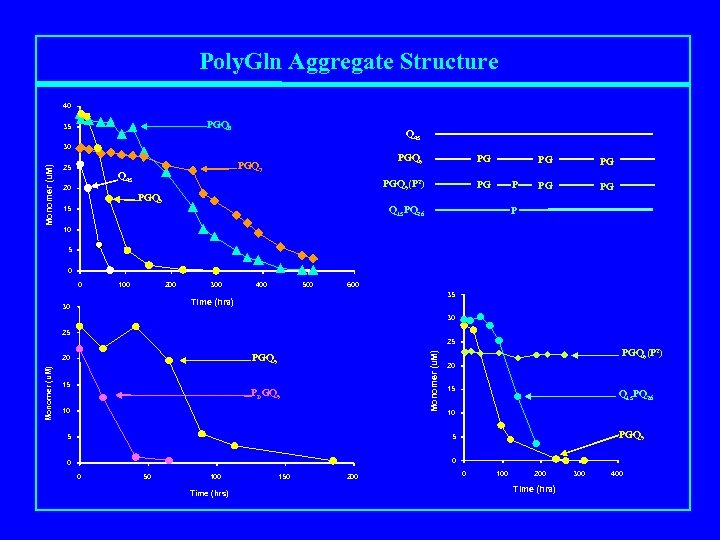
Poly. Gln Aggregate Structure 40 PGQ 8 35 Q 45 Monomer (u. M) 30 25 PGQ 9 15 PG PGQ 9(P 2) Q 45 20 PGQ 9 PGQ 7 PG PG P Q 15 PQ 26 PG PG PG P 10 5 0 0 100 200 300 400 500 600 35 Time (hrs) 30 30 25 PGQ 9 20 Monomer (u. M) 25 15 PDGQ 9 10 PGQ 9(P 2) 20 15 Q 15 PQ 26 10 5 0 PGQ 9 5 0 0 50 100 Time (hrs) 150 200 0 100 200 Time (hrs) 300 400
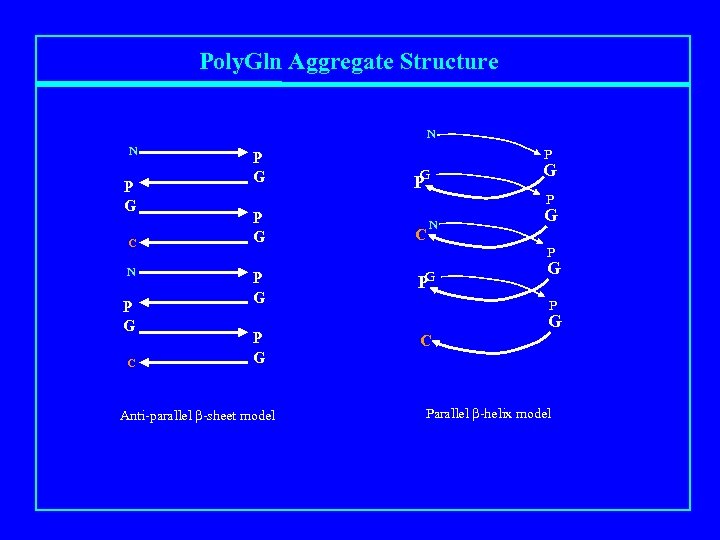

Poly. Gln Aggregate Structure N N P G C P G P G Anti-parallel b-sheet model P G P C N G P G P G C Parallel b-helix model
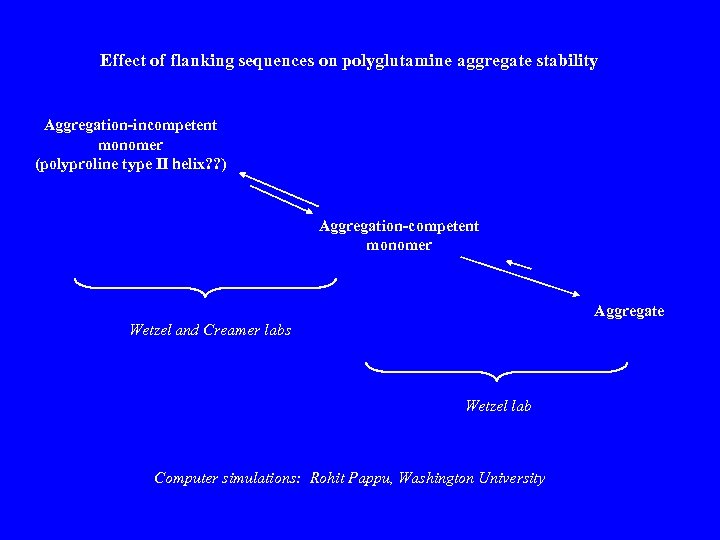
Effect of flanking sequences on polyglutamine aggregate stability Aggregation-incompetent monomer (polyproline type II helix? ? ) Aggregation-competent monomer Aggregate Wetzel and Creamer labs Wetzel lab Computer simulations: Rohit Pappu, Washington University

Summary • as predicted by theory, in vitro amyloid fibrils can achieve an equilibrium with monomer • the position of this equilibrium is proportional to the free energy of fibril formation • measurement of shifted equilibria allows quantitation of mutational effects • amyloid fibrils exhibit a remarkable structural plasticity • in ideal cases, aggregation kinetics can be interpreted mechanistically • the kinetic nucleus for polyglutamine aggregation is an alternatively folded monomer • accumulated sequence changes strongly diminish cross-seeding efficiency

Mutagenesis and Kinetics/Thermodynamics in Globular Protein Structure • studies on “natural” mutants of globular proteins (1970 s) - Gary Ackers (human hemoglobin variants) - Mike Laskowski (ovomucoid variants) • protein engineering approaches to globular protein folding stability (1984 ->) - Ron Wetzel (T 4 lysozyme disulfide bonds) - Brian Matthews (T 4 lysozyme point mutations) - Robert Matthews, Alan Fersht (folding kinetics) • protein folding stability and amyloidogenicity (1993 ->) - Jeff Kelly (transthyretin / TTR amyloidosis) - Ron Wetzel (light chain FV domain / Ig light chain amyloidosis) - Chris Dobson (lysozyme amyloidosis) • amyloid fibril assembly kinetics and thermodynamics…. landscape continuity? - kinetics complicated by protofibrils and by secondary nucleation - can fibril formation reach true equilibrium positions in vitro?
![Aggregation and Packing Interactions [R. Wetzel, Trends Biotech. 12, 193 -198 (1994)] Aggregation and Packing Interactions [R. Wetzel, Trends Biotech. 12, 193 -198 (1994)]](https://present5.com/presentation/4e2cb46c65b3324d68bf03d090982437/image-79.jpg)
Aggregation and Packing Interactions [R. Wetzel, Trends Biotech. 12, 193 -198 (1994)]

ACKNOWLEDGMENTS UGA UT Main Campus UTMCK • Indu Kheterpal • Angela Williams • Shankaramma Shivaprasad • Israel Huff • Tina Richey • Kimberley Salone • Matt Sega • Brian Bledsoe • Valerie Berthelier • Lezlee Dice • Brian O’Nuallain • Anusri Bhattacharyya Mitra • Songming Chen • Wen Yang • Brad Hamilton • Ashwani Thakur • Geetha Thiagarajan • Roopa Kenoth • Merav Geva • Alex Osmand • Erica Johnson Rowe • Erin Newby • Maolian Chen • Erik Portelius • David Kaleta • Shaolian Zhou • Kelsey Cook • Neil Whittemore • Rajesh Mishra • Engin Serpersu • Guangyao Gao • Ying Chen • Peter Zhang • Anna Gardberg • Chris Dealwis • Juntao Guo • Ying Xu Harvard Med • Hilal Lashuel • Peter Lansbury • Prasanna Venkatraman • Fred Goldberg Cal Tech • Jan Ko • Susan Ou • Paul Patterson Uppsala • Per Westermark • Liz Howell • John Dunlap Drexel • Frank Ferrone FUNDING: NIH (NIA, NINDS); Hereditary Disease Foundation

Thermodynamics of Amyloid Fibril Formation • In globular proteins, some amyloidogenic mutations work by weakening native structure - transthyretin (Kelly) - Ig light chain (Wetzel) • local sequence also affects amyloidogenicity through fibril packing effects • simplest systems are where the starting monomer is in coil, …. . - no overlay of a stable native state - reasonable assumption that mutation minimally affects native state G • …. . and where there is an easily and accurately measured Cr • Results: - Aβ(1 -40) fibril growth tends to an easily measured, reversible equilibrium position ΔG = - 8. 6 kcal/mol ΔΔGs from Ala mutations agree with data from parallel β-sheet in globular protein Ala-edited Pro scan reveals sequence segments in rigid structure, …. . … but propagated structural changes in H-bonding complicate interpretation
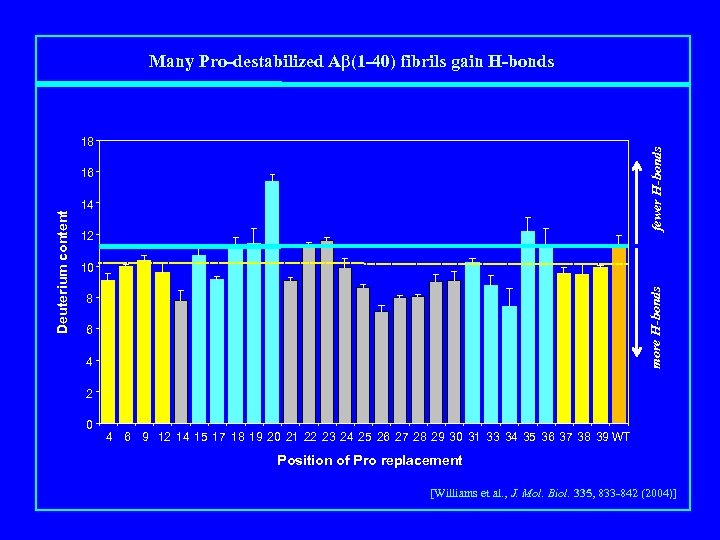
Many Pro-destabilized Aβ(1 -40) fibrils gain H-bonds fewer H-bonds 18 14 12 10 more H-bonds Deuterium content 16 8 6 4 2 0 4 6 9 12 14 15 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 33 34 35 36 37 38 39 WT Position of Pro replacement [Williams et al. , J. Mol. Biol. 335, 833 -842 (2004)]
Normal globular proteins generally have only one stable state
4e2cb46c65b3324d68bf03d090982437.ppt