cc07f42da8b0adf2507e64245bdd3392.ppt
- Количество слайдов: 29

Joint meeting of the Molecular Libraries Screening Centers Network (MLSCN) and the Exploratory Centers for Cheminformatics Research (ECCR): Talk I July 17 2006 Geoffrey Fox Computer Science, Informatics, Physics Pervasive Technology Laboratories With apologies for my credentials. I have written a few papers on Biology, Chemistry and Crystallography while at Cambridge, Caltech and Syracuse Indiana University Bloomington IN 47401 Mostly on applications of parallel computing gcf@indiana. edu http: //www. infomall. org 1 http: //www. chembiogrid. org

Start-up and Organization n n n n Local Teams, successful Prototypes and International Collaboration set up in 3 major focus areas • “Tool and Data” Cyberinfrastructure • “Archival Database and Simulation” Cyberinfrastructure • Education Wiki chosen to support project as a shared editable web space Web site http: //www. chembiogrid. org Building Collaboratory involving Pub. Chem – Global Information System accessible anywhere and at any time – enhance Pub. Chem with distributed tools (clustering, simulation, annotation etc. ) and data Initial results discussed at conferences/workshops/papers • Gordon Conferences, ACS, SDSC tutorial First new Cheminformatics courses offered Advisory board set up and met Videoconferencing-based meetings with Peter Murray-Rust and group at Cambridge roughly every 2 -3 weeks Good interactions with NIH DTP, Lilly and Michigan ECCR 2

http: //www. chembiogrid. org 3

CICC Senior Personnel n n n n Geoffrey C. Fox Mu-Hyun (Mookie) Baik Dennis B. Gannon Marlon Pierce Beth A. Plale Gary D. Wiggins David J. Wild Yuqing (Melanie) Wu n n n From Biology, Chemistry, Computer Science, Informatics at IU Bloomington and IUPUI (Indianapolis) n n Peter T. Cherbas Mehmet M. Dalkilic Charles H. Davis A. Keith Dunker Kelsey M. Forsythe Kevin E. Gilbert John C. Huffman Malika Mahoui Daniel J. Mindiola Santiago D. Schnell William Scott Craig A. Stewart David R. Williams 4

CICC Advisory Board n n n n Alan D. Palkowitz (Eli Lilly) Andrew Martin (Kalypsys) David Spellmeyer (IBM) Dimitris K. Agrafiotis (Johnson & Johnson) Horst Hemmerle (Eli Lilly) James M. Caruthers (Purdue University) Jeremy G. Frey (University of Southampton) Joel Saltz (Ohio State University/University of Maryland/Johns Hopkins University) John M. Barnard (Digital Chemistry) John Reynders (Eli Lilly) Peter Murray-Rust (University of Cambridge) Industry and Academia Peter Willett (University of Sheffield) Met October 2005 Thompson Doman (Eli Lilly) will meet this fall Val Gillet (University of Sheffield) 5

Publications Baik says he is especially productive due to Cyberinfrastructure 6

Our Meetings are on the Web 7
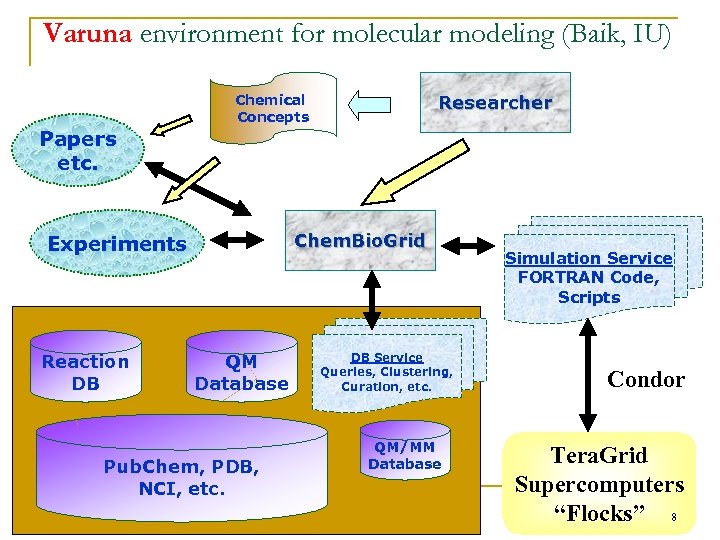
Varuna environment for molecular modeling (Baik, IU) Researcher Chemical Concepts Papers etc. Chem. Bio. Grid Experiments Reaction DB QM Database Pub. Chem, PDB, NCI, etc. DB Service Queries, Clustering, Curation, etc. QM/MM Database Simulation Service FORTRAN Code, Scripts Condor Tera. Grid Supercomputers “Flocks” 8

Cyberinfrastructure and Grids n n These support e. Science or distributed Computers, Databases, Instruments, Sensors and People Grids use large scale managed Web services – the current major technology building on modern Industry enterprise and Internet systems • W 3 C, OASIS, OGF or Open Grid Forum (Fox VP for e. Science) develops standards insuring distributed resources interoperate Cheminformatics benefits from 2 styles of Grids • Tera. Grid typifies Grid support of large scale computation of parallel simulations • Bioinformatics (BIRN, ca. BIG, My. Grid …), Earth Science and Astronomy Grids illustrate integration of real-time and archival data(bases) and computation Well designed Grids run faster than older approaches 9

Cheminformatics Grids Need n n n Broad System standards such as WSDL, SOAP, WSRM, JSDL, BPEL Domain specific data structures • CML Cheminformatics • GML Earth Science • Cell. ML, SBML Biology • VOQL Astronomy Use of specific Grid/Web service technologies such as • Web services directly for tools • Web service proxies for large simulation codes – ANYTHING can be made a Web service efficiently if execution/network access time ≥ 20 ms • Portals/Portlets for user interfaces • Workflow for composition n Access to data and compute resources 10
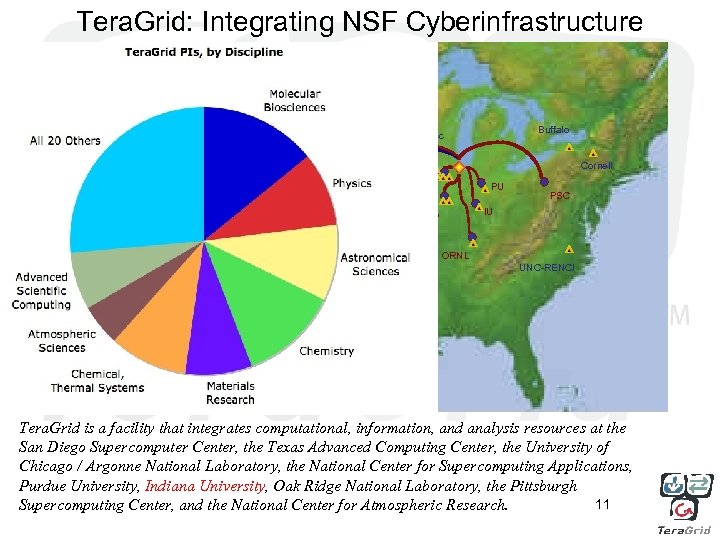
Tera. Grid: Integrating NSF Cyberinfrastructure Buffalo Wisc UC/ANL Utah Cornell Iowa PU NCAR IU NCSA Caltech PSC ORNL USC-ISI UNC-RENCI SDSC TACC Tera. Grid is a facility that integrates computational, information, and analysis resources at the San Diego Supercomputer Center, the Texas Advanced Computing Center, the University of Chicago / Argonne National Laboratory, the National Center for Supercomputing Applications, Purdue University, Indiana University, Oak Ridge National Laboratory, the Pittsburgh 11 Supercomputing Center, and the National Center for Atmospheric Research.
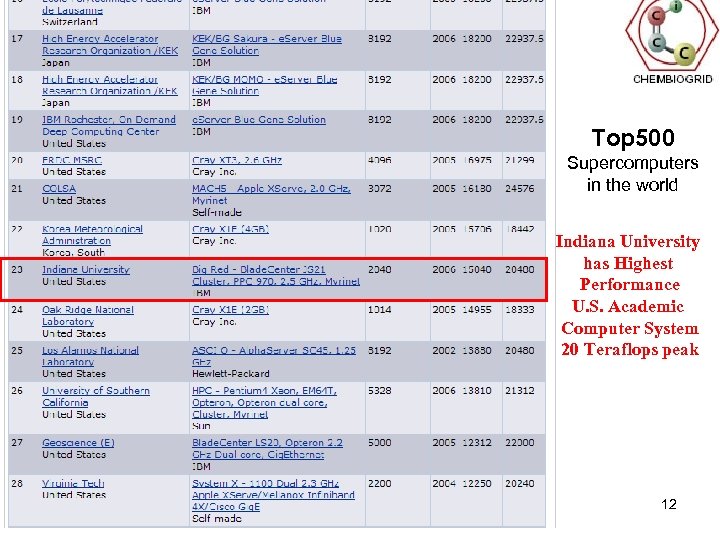
Top 500 Supercomputers in the world Indiana University has Highest Performance U. S. Academic Computer System 20 Teraflops peak 12

Products and Demonstrations www. chembiogrid. org Note mixture of In-house Out of House Commercial Academic 13
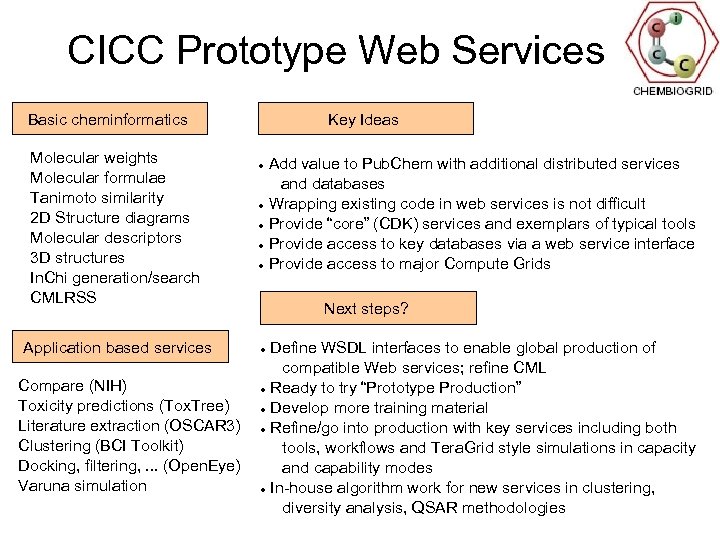
CICC Prototype Web Services Basic cheminformatics Molecular weights Molecular formulae Tanimoto similarity 2 D Structure diagrams Molecular descriptors 3 D structures In. Chi generation/search CMLRSS Application based services Compare (NIH) Toxicity predictions (Tox. Tree) Literature extraction (OSCAR 3) Clustering (BCI Toolkit) Docking, filtering, . . . (Open. Eye) Varuna simulation Key Ideas Add value to Pub. Chem with additional distributed services and databases Wrapping existing code in web services is not difficult Provide “core” (CDK) services and exemplars of typical tools Provide access to key databases via a web service interface Provide access to major Compute Grids Next steps? Define WSDL interfaces to enable global production of compatible Web services; refine CML Ready to try “Prototype Production” Develop more training material Refine/go into production with key services including both tools, workflows and Tera. Grid style simulations in capacity and capability modes In-house algorithm work for new services in clustering, diversity analysis, QSAR methodologies
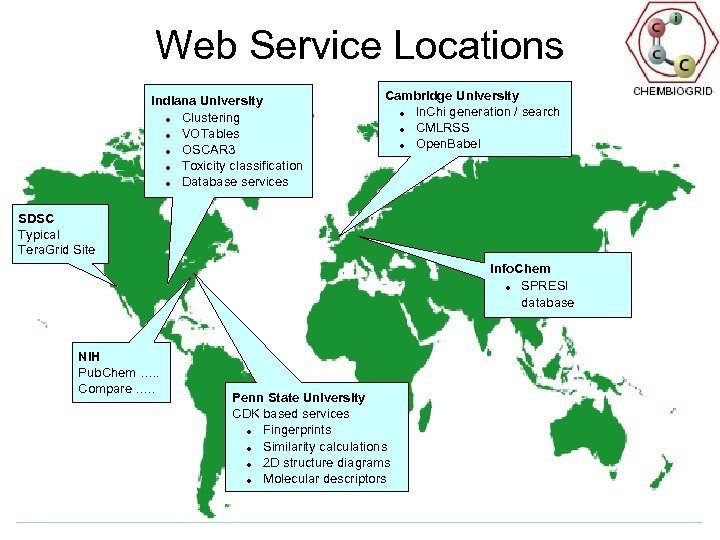
Web Service Locations Indiana University Clustering VOTables OSCAR 3 Toxicity classification Database services Cambridge University In. Chi generation / search CMLRSS Open. Babel SDSC Typical Tera. Grid Site Info. Chem SPRESI database NIH Pub. Chem …. . Compare …. . Penn State University CDK based services Fingerprints Similarity calculations 2 D structure diagrams Molecular descriptors

Usage of Open Source Projects A number of open source projects are used in our infrastructure CDK provides the underlying cheminformatics toolkit R provides the back-end modeling capabilities OSCAR is used for literature mining Tox. Tree is used to provide toxicity classification Open data and standards as promoted by the Blue Obelisk project

Contributions to Open Source Projects We also contribute functionality to these projects Molecular descriptor development to the CDK Modifications of various CDK functionality to make them suitable for web service usage Infrastructure for accessing R from the CDK Packages to use the CDK from within R Quality control, testing and documentation Steinbeck, C. et al. ; Curr. Pharm. Des. , 2006, 12(17), 2110 -2120 Guha, R. ; CDK News, 2005, 2(1), 7 -13
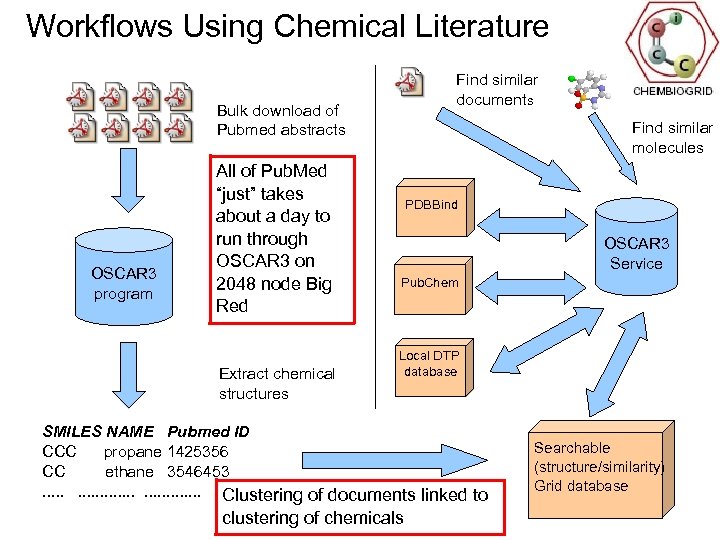
Workflows Using Chemical Literature Find similar documents Bulk download of Pubmed abstracts OSCAR 3 program All of Pub. Med “just” takes about a day to run through OSCAR 3 on 2048 node Big Red Extract chemical structures Find similar molecules PDBBind OSCAR 3 Service Pub. Chem Local DTP database SMILES NAME Pubmed ID CCC propane 1425356 CC ethane 3546453. . . . Clustering of documents linked to clustering of chemicals Searchable (structure/similarity) Grid database
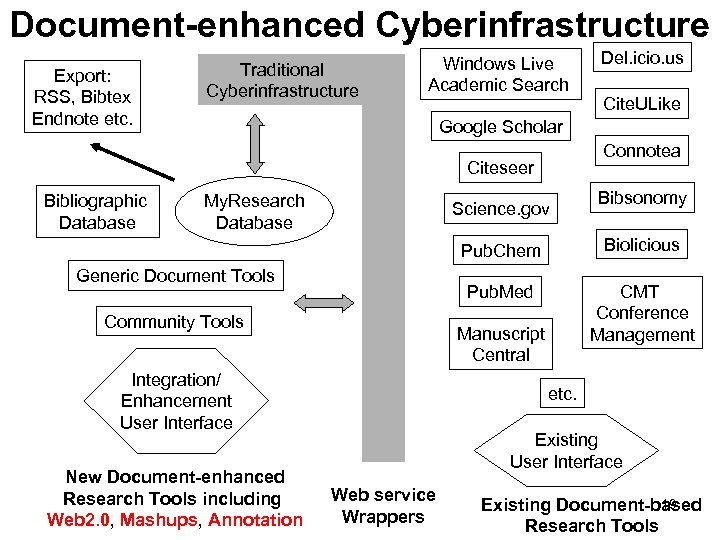
Document-enhanced Cyberinfrastructure Export: RSS, Bibtex Endnote etc. Traditional Cyberinfrastructure Windows Live Academic Search Cite. ULike Google Scholar Connotea Citeseer Bibliographic Database Del. icio. us My. Research Database Science. gov Bibsonomy Pub. Chem Pub. Med Generic Document Tools Community Tools CMT Conference Management Manuscript Central Integration/ Enhancement User Interface New Document-enhanced Research Tools including Web 2. 0, Mashups, Annotation Biolicious etc. Existing User Interface Web service Wrappers 19 Existing Document-based Research Tools

Products and Demonstrations II 20
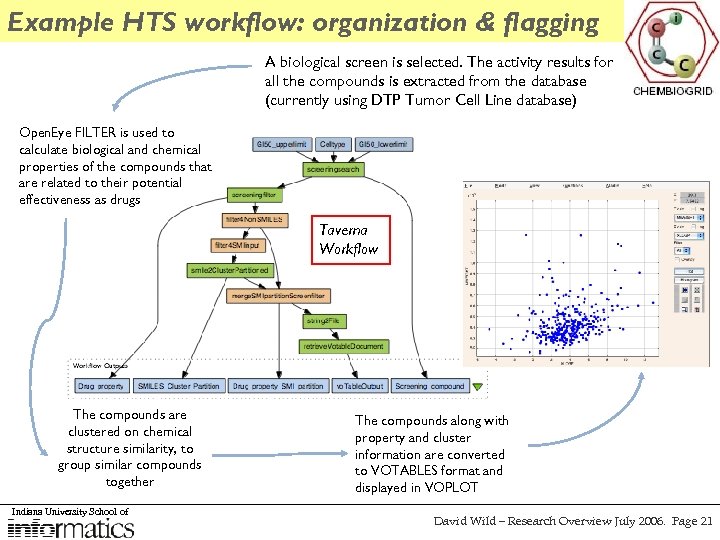
Example HTS workflow: organization & flagging A biological screen is selected. The activity results for all the compounds is extracted from the database (currently using DTP Tumor Cell Line database) Open. Eye FILTER is used to calculate biological and chemical properties of the compounds that are related to their potential effectiveness as drugs Taverna Workflow The compounds are clustered on chemical structure similarity, to group similar compounds together Indiana University School of The compounds along with property and cluster information are converted to VOTABLES format and displayed in VOPLOT David Wild – Research Overview July 2006. Page 21
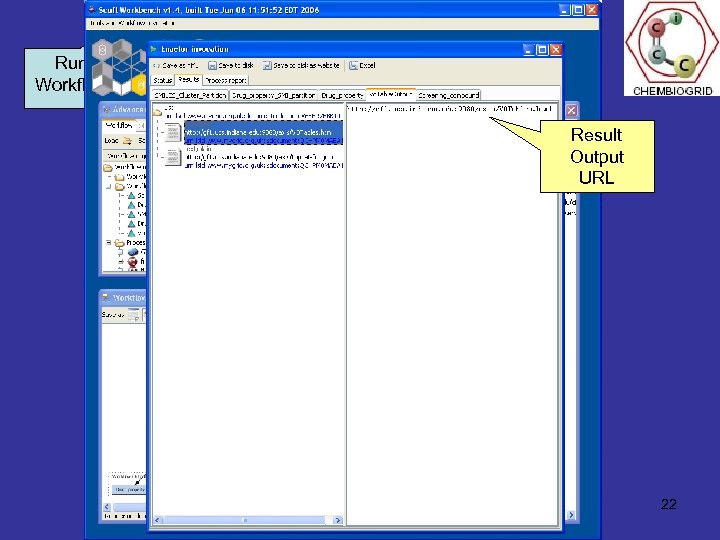
Run Workflow Load Workflow Result Output URL Current Process 22

Lilly very interested in our new educational programs 23
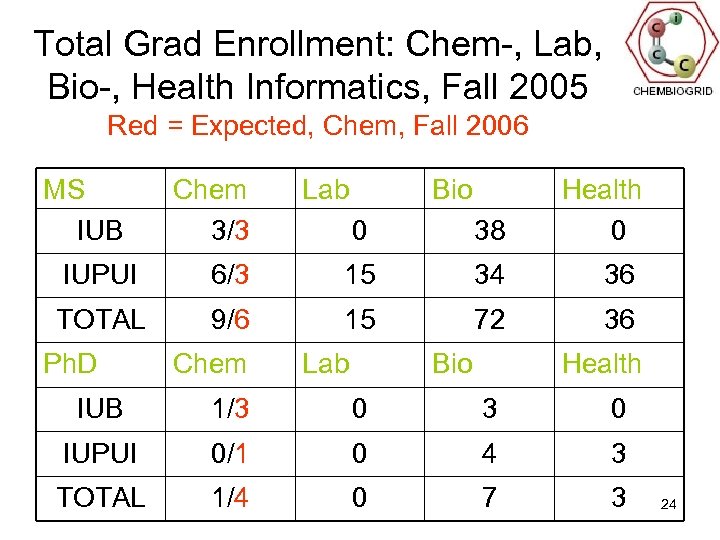
Total Grad Enrollment: Chem-, Lab, Bio-, Health Informatics, Fall 2005 Red = Expected, Chem, Fall 2006 MS IUB Chem 3/3 Lab Bio 0 38 Health 0 IUPUI 6/3 15 34 36 TOTAL 9/6 15 72 36 Ph. D Chem Lab Bio Health IUB 1/3 0 IUPUI 0/1 0 4 3 TOTAL 1/4 0 7 3 24

Formal Cheminformatics Courses • I 571 Chemical Information Technology (3 cr. ) – Distance Ed section had 10 students in Fall 2005, from California to Connecticut • I 572 Computational Chemistry and Molecular Modeling (3 cr. ) • I 573 Programming Techniques for Chemical and Life Science Informatics (3 cr. ) • I 553 Independent Study in Chemical Informatics (3 cr. ) • Above courses required for the new Graduate Certificate Program in Chemical Informatics • Also I 533 (Cheminformatics seminar) 25

More detailed Slides not used 26

Tera. Grid Hardware and Software n n n Tera. Grid is coordinated at the University of Chicago and includes 8 partner facilities • NCSA, SDSC, PSC, ORNL, IU, PU, TACC, UC/ANL Tera. Grid hardware totals > 102 teraflops of computing power. • Comprehensive information available from http: //www. teragrid. org/userinfo/hardware/overview. php. • Systems are primarily Linux clusters. Grid software and services (Globus, My. Proxy, etc) provide a uniform means for accessing Tera. Grid resources. • Scheduling, running and monitoring jobs • Monitoring resources • Moving and managing remote files. • Common service APIs simplify the process for building remote tools. 27
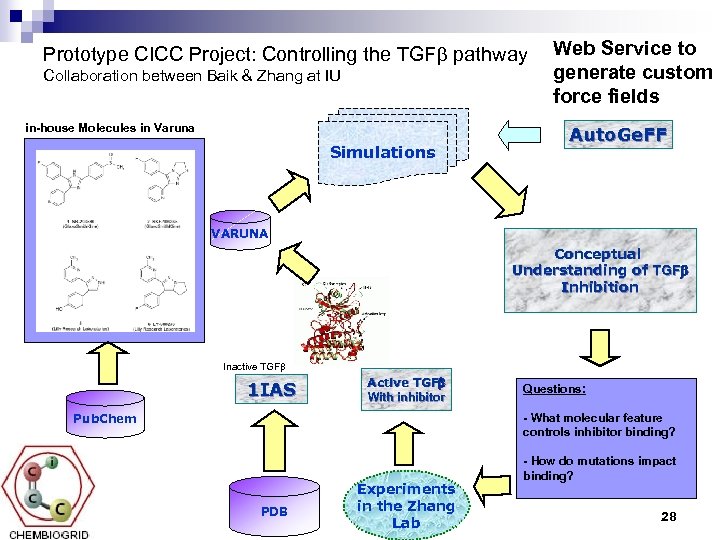
Prototype CICC Project: Controlling the TGFb pathway Collaboration between Baik & Zhang at IU in-house Molecules in Varuna Simulations Web Service to generate custom force fields Auto. Ge. FF VARUNA Conceptual Understanding of TGFb Inhibition Inactive TGFb 1 IAS Active TGFb With inhibitor Pub. Chem Questions: - What molecular feature controls inhibitor binding? PDB Experiments in the Zhang Lab - How do mutations impact binding? 28
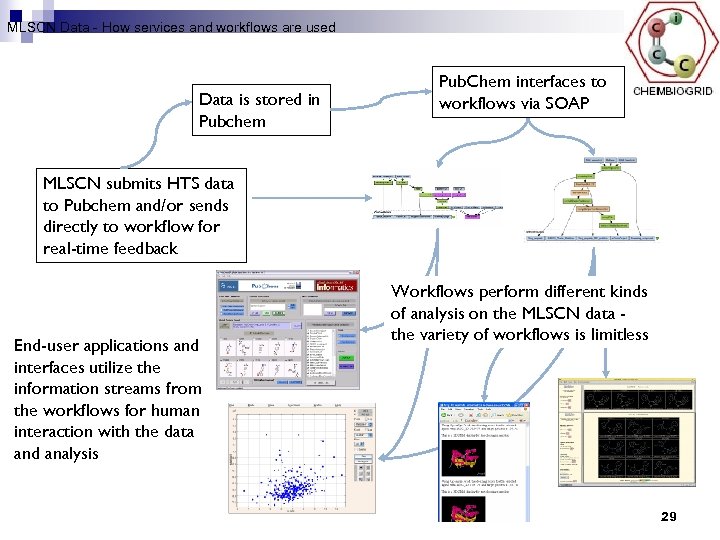
MLSCN Data - How services and workflows are used Data is stored in Pubchem Pub. Chem interfaces to workflows via SOAP MLSCN submits HTS data to Pubchem and/or sends directly to workflow for real-time feedback End-user applications and interfaces utilize the information streams from the workflows for human interaction with the data and analysis Workflows perform different kinds of analysis on the MLSCN data the variety of workflows is limitless 29
cc07f42da8b0adf2507e64245bdd3392.ppt