9e6bcbca39d2d6db54a2b48041b1bc2d.ppt
- Количество слайдов: 22
 Introduction to DAS / State of the Union Tim Hubbard th@sanger. ac. uk DAS developer workshop 10 th March 2009 Wellcome Trust Genome Campus
Introduction to DAS / State of the Union Tim Hubbard th@sanger. ac. uk DAS developer workshop 10 th March 2009 Wellcome Trust Genome Campus
 Distributed Annotation System or How I Learnt to Stop Worrying and Love Data Federation Credit: Andreas Prlić
Distributed Annotation System or How I Learnt to Stop Worrying and Love Data Federation Credit: Andreas Prlić
 Distributed Annotation System • Origins: – – – xml client/server specification (http: //biodas. org/) Lincoln Stein, Sean Eddy, Robin Dowell and La. Deana Hillier acedb based prototype server Java based prototype client Dowell, R. D. , Jokerst, R. M. , Day, A. , Eddy, S. R. & Stein, L. (2001) Bio. Med. Central Bioinformatics 2. • Genome campus adoption – Initially via Ensembl becoming a DAS client (now also a DAS server) – Code: Dazzle and Proservers; Bio: : DASLite and biojava client libraries – Hosts DAS registry (http: //www. dasregistry. org/)
Distributed Annotation System • Origins: – – – xml client/server specification (http: //biodas. org/) Lincoln Stein, Sean Eddy, Robin Dowell and La. Deana Hillier acedb based prototype server Java based prototype client Dowell, R. D. , Jokerst, R. M. , Day, A. , Eddy, S. R. & Stein, L. (2001) Bio. Med. Central Bioinformatics 2. • Genome campus adoption – Initially via Ensembl becoming a DAS client (now also a DAS server) – Code: Dazzle and Proservers; Bio: : DASLite and biojava client libraries – Hosts DAS registry (http: //www. dasregistry. org/)
 DAS in a nutshell • Standardized set of web services – Reference servers (the sequence) – Annotation servers (features: chr: start-end) – Alignment servers (chr: start-end matches chr: start-end) – Identifier based servers (ref item X rather than coordinate) • Standardization allows clients to connect to different DAS sources without additional programming
DAS in a nutshell • Standardized set of web services – Reference servers (the sequence) – Annotation servers (features: chr: start-end) – Alignment servers (chr: start-end matches chr: start-end) – Identifier based servers (ref item X rather than coordinate) • Standardization allows clients to connect to different DAS sources without additional programming
 Data integration • Complete genomes provide the framework to pull all biological data together such that each piece says something about biology as a whole • Biology is too complex for any organisation to have a monopoly of ideas or data • The more organisations provide data or analysis separately, the harder it becomes for anyone to make use of the results
Data integration • Complete genomes provide the framework to pull all biological data together such that each piece says something about biology as a whole • Biology is too complex for any organisation to have a monopoly of ideas or data • The more organisations provide data or analysis separately, the harder it becomes for anyone to make use of the results
 Scientific impact Utility of bioinformatics Too little bioinformatics Too many databases Too diverse interfaces
Scientific impact Utility of bioinformatics Too little bioinformatics Too many databases Too diverse interfaces
 Split data and presentation • Databases responsible for curating data and serving it as primitive datatypes defined by open standards (high cost) • Different front ends or components of front ends compete for users (development of each low cost) c. f. browsers.
Split data and presentation • Databases responsible for curating data and serving it as primitive datatypes defined by open standards (high cost) • Different front ends or components of front ends compete for users (development of each low cost) c. f. browsers.
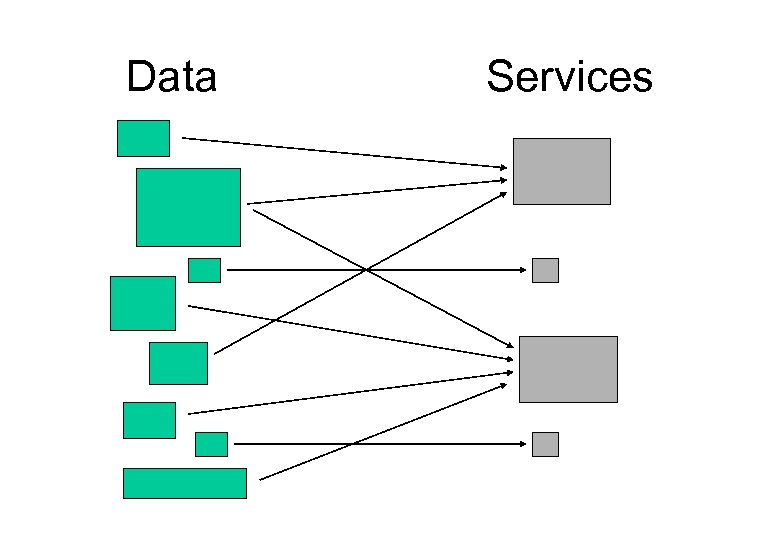 Data Services
Data Services
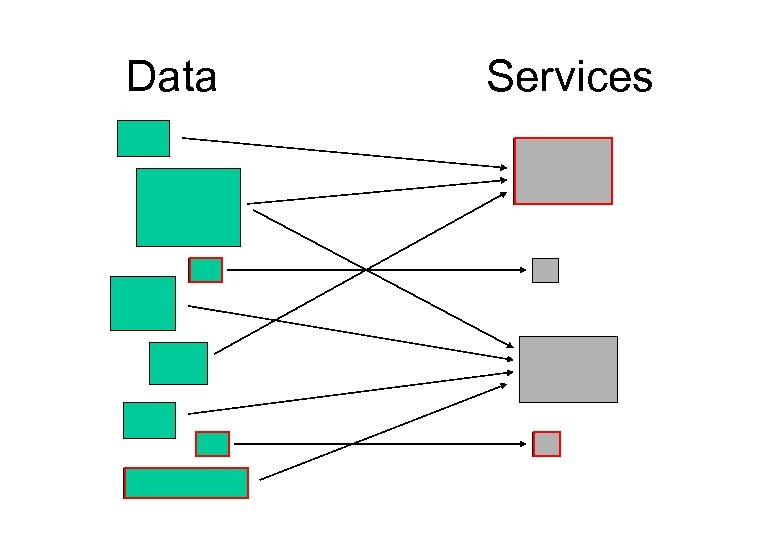 Data Services
Data Services
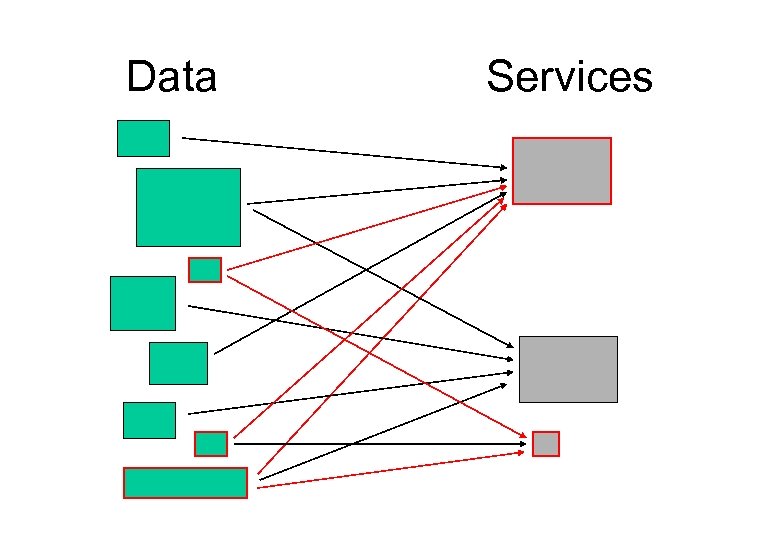 Data Services
Data Services
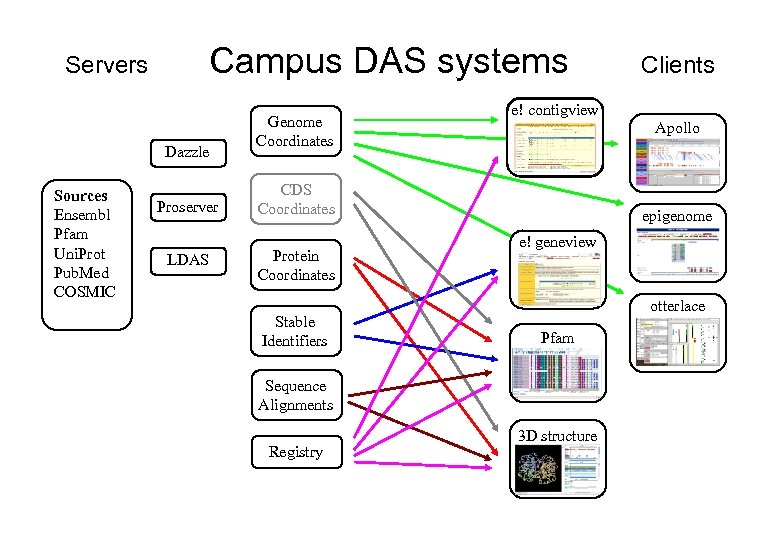 Campus DAS systems Servers Dazzle Sources Ensembl Pfam Uni. Prot Pub. Med COSMIC Proserver LDAS Genome Coordinates e! contigview Apollo CDS Coordinates Protein Coordinates Stable Identifiers epigenome e! geneview otterlace Pfam Sequence Alignments Registry Clients 3 D structure
Campus DAS systems Servers Dazzle Sources Ensembl Pfam Uni. Prot Pub. Med COSMIC Proserver LDAS Genome Coordinates e! contigview Apollo CDS Coordinates Protein Coordinates Stable Identifiers epigenome e! geneview otterlace Pfam Sequence Alignments Registry Clients 3 D structure
 Rise of Federation Technologies • DAS for features • Bio. Mart for data mining • Bio. Mart server is a DAS server • New international genome data projects – routinely using the F word – frequently the D and B words too – e. g. International Cancer Genome Consortium
Rise of Federation Technologies • DAS for features • Bio. Mart for data mining • Bio. Mart server is a DAS server • New international genome data projects – routinely using the F word – frequently the D and B words too – e. g. International Cancer Genome Consortium
 DAS infrastructure status • Lots of progress – – Servers: Dazzle, Proserver, My. Das, Bio: : Daslite Clients: Ensembl, Vega, Dasty, SPICE, Pfam, Jalview, Pepper, IGB >500 sources in DAS registry (http: //www. dasregistry. org/) Broadly adopted by large scale projects: Ensembl, biosapiens, efamily, ZFmodels, e. Protein, ENCODE annotation – Extensions in 1. 53 E: stylesheets, semantic zooming, ontology support, timestamps, interactions – Planned 1. 6: incorporating some features of DAS 2 specification – Better adoption of DAS in US • Opportunities – – – Searching, writeback Source ranking, credit, social networking Inter-client communications protocol Async delivery/caching; servers built on servers/workflows Alternative entry points from servers? Next left/right? Date of addition?
DAS infrastructure status • Lots of progress – – Servers: Dazzle, Proserver, My. Das, Bio: : Daslite Clients: Ensembl, Vega, Dasty, SPICE, Pfam, Jalview, Pepper, IGB >500 sources in DAS registry (http: //www. dasregistry. org/) Broadly adopted by large scale projects: Ensembl, biosapiens, efamily, ZFmodels, e. Protein, ENCODE annotation – Extensions in 1. 53 E: stylesheets, semantic zooming, ontology support, timestamps, interactions – Planned 1. 6: incorporating some features of DAS 2 specification – Better adoption of DAS in US • Opportunities – – – Searching, writeback Source ranking, credit, social networking Inter-client communications protocol Async delivery/caching; servers built on servers/workflows Alternative entry points from servers? Next left/right? Date of addition?
 2008 the year of… • Open access to publications – PMC, uk. PMC, Zotero, Papers, My. NCBI, Citeulike, Connotea, 2 collab and Hub. Med – All WT funded publications open in 6 months – All NIH funded publications open in 12 months • DAS for publications? – Text is just a new coordinate system • Links to Social Networks? – Google Open. Social • Still waiting…
2008 the year of… • Open access to publications – PMC, uk. PMC, Zotero, Papers, My. NCBI, Citeulike, Connotea, 2 collab and Hub. Med – All WT funded publications open in 6 months – All NIH funded publications open in 12 months • DAS for publications? – Text is just a new coordinate system • Links to Social Networks? – Google Open. Social • Still waiting…
 2009 the year of… • Massive datasets – Track likely to be 50 million solexa transcriptome reads • Need: – Better ways for users to create tracks for large datasets
2009 the year of… • Massive datasets – Track likely to be 50 million solexa transcriptome reads • Need: – Better ways for users to create tracks for large datasets
 Problems of large user data (credits to Jim Kent, UCSC) • Easy to generate 1 GB files with next gen sequencing. – 25 million tag mappings at 40 bytes each – Potential to translate into histograms with 1 floating point number every 12 bases • Slow to load into My. SQL database backend to local DAS server; many users will not want to setup DAS servers • Too large to upload to remote DAS server services (e. g. Ensembl) to create track • Most users only look at 5 -50 sites - less than 1%
Problems of large user data (credits to Jim Kent, UCSC) • Easy to generate 1 GB files with next gen sequencing. – 25 million tag mappings at 40 bytes each – Potential to translate into histograms with 1 floating point number every 12 bases • Slow to load into My. SQL database backend to local DAS server; many users will not want to setup DAS servers • Too large to upload to remote DAS server services (e. g. Ensembl) to create track • Most users only look at 5 -50 sites - less than 1%
 Jim Kent’s idea • User runs program to convert their data into single indexed file (Big. Wig & Big. Bed) • Place on their website • UCSC browser fetches parts of file on demand using http(s) “byte range” queries • Relationship to DAS? – Potential to create DAS server plugin to serve Big. Wig/Big. Bed files as DAS servers
Jim Kent’s idea • User runs program to convert their data into single indexed file (Big. Wig & Big. Bed) • Place on their website • UCSC browser fetches parts of file on demand using http(s) “byte range” queries • Relationship to DAS? – Potential to create DAS server plugin to serve Big. Wig/Big. Bed files as DAS servers
 Acknowledgements Ewan Birney Tony Cox Thomas Down Rob Finn Stefan Graf David Jackson Andreas Kahari Eugene Kulesha Henning Hermjakob Roger Pettett Matt Pocock James Smith Jim Stalker Janet Thornton Jonathan Warren Andy Jenkinson Andreas Prlic Ensembl/Sanger Web team efamily, biosapiens, e. Protein Zebrafish analysis (ZF-models) Anacode/Acedb (otterlace/Zmap)
Acknowledgements Ewan Birney Tony Cox Thomas Down Rob Finn Stefan Graf David Jackson Andreas Kahari Eugene Kulesha Henning Hermjakob Roger Pettett Matt Pocock James Smith Jim Stalker Janet Thornton Jonathan Warren Andy Jenkinson Andreas Prlic Ensembl/Sanger Web team efamily, biosapiens, e. Protein Zebrafish analysis (ZF-models) Anacode/Acedb (otterlace/Zmap)
 2009 the year of… • Massive datasets – Track likely to be 50 million solexa transcriptome reads • Private datasets – EGA requires registration and logins – Even summary data currently not public • Need: – Better ways for users to create tracks for large datasets – Federated access controls for patient data
2009 the year of… • Massive datasets – Track likely to be 50 million solexa transcriptome reads • Private datasets – EGA requires registration and logins – Even summary data currently not public • Need: – Better ways for users to create tracks for large datasets – Federated access controls for patient data
 Todo: tilling array DAS stylesheet magic (Eugene Kulesha)
Todo: tilling array DAS stylesheet magic (Eugene Kulesha)


