359b6335d3cb6d9dc647a17bfff2f06e.ppt
- Количество слайдов: 29
 Integration of data to uncover evolutionary trends and infer protein function: The tale of Rcs 1 M. Madan Babu MRC Laboratory of Molecular Biology Cambridge
Integration of data to uncover evolutionary trends and infer protein function: The tale of Rcs 1 M. Madan Babu MRC Laboratory of Molecular Biology Cambridge
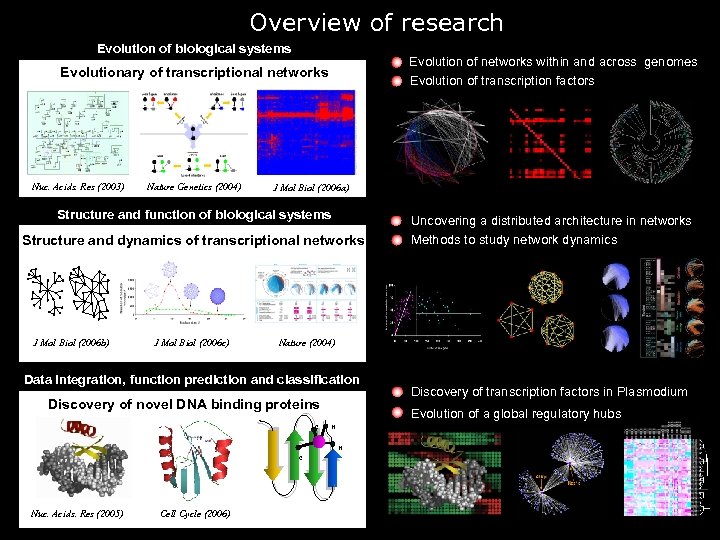 Overview of research Evolution of biological systems Evolution of networks within and across genomes Evolution of transcription factors Evolutionary of transcriptional networks Nuc. Acids. Res (2003) Nature Genetics (2004) J Mol Biol (2006 a) Structure and function of biological systems Structure and dynamics of transcriptional networks J Mol Biol (2006 b) J Mol Biol (2006 c) Nature (2004) Data integration, function prediction and classification Discovery of novel DNA binding proteins C Cell Cycle (2006) Discovery of transcription factors in Plasmodium Evolution of a global regulatory hubs H H C Nuc. Acids. Res (2005) Uncovering a distributed architecture in networks Methods to study network dynamics
Overview of research Evolution of biological systems Evolution of networks within and across genomes Evolution of transcription factors Evolutionary of transcriptional networks Nuc. Acids. Res (2003) Nature Genetics (2004) J Mol Biol (2006 a) Structure and function of biological systems Structure and dynamics of transcriptional networks J Mol Biol (2006 b) J Mol Biol (2006 c) Nature (2004) Data integration, function prediction and classification Discovery of novel DNA binding proteins C Cell Cycle (2006) Discovery of transcription factors in Plasmodium Evolution of a global regulatory hubs H H C Nuc. Acids. Res (2005) Uncovering a distributed architecture in networks Methods to study network dynamics
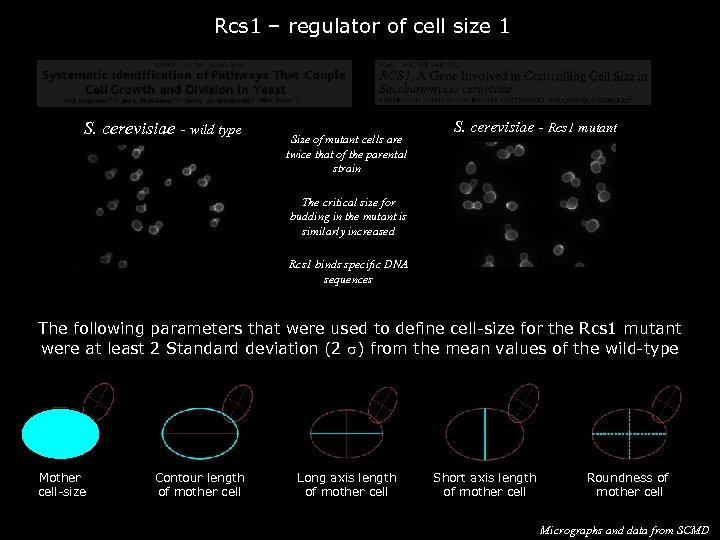 Rcs 1 – regulator of cell size 1 S. cerevisiae - wild type Size of mutant cells are twice that of the parental strain S. cerevisiae - Rcs 1 mutant The critical size for budding in the mutant is similarly increased Rcs 1 binds specific DNA sequences The following parameters that were used to define cell-size for the Rcs 1 mutant were at least 2 Standard deviation (2 s) from the mean values of the wild-type Mother cell-size 874 760 Contour length of mother cell 108 100 Long axis length of mother cell 36 33 Short axis length of mother cell 30 27 Roundness of mother cell 1. 29 1. 20 Micrographs and data from SCMD
Rcs 1 – regulator of cell size 1 S. cerevisiae - wild type Size of mutant cells are twice that of the parental strain S. cerevisiae - Rcs 1 mutant The critical size for budding in the mutant is similarly increased Rcs 1 binds specific DNA sequences The following parameters that were used to define cell-size for the Rcs 1 mutant were at least 2 Standard deviation (2 s) from the mean values of the wild-type Mother cell-size 874 760 Contour length of mother cell 108 100 Long axis length of mother cell 36 33 Short axis length of mother cell 30 27 Roundness of mother cell 1. 29 1. 20 Micrographs and data from SCMD
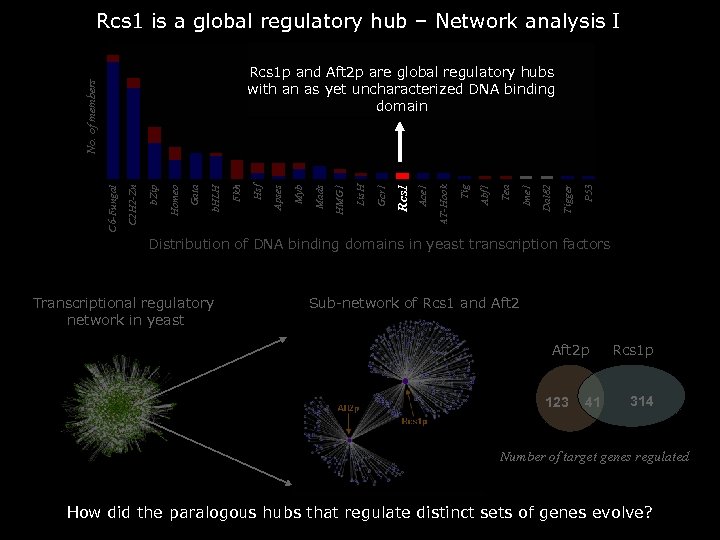 Rcs 1 is a global regulatory hub – Network analysis I P 53 Tigger Dal 82 Ime 1 Tea Abf 1 Tig AT-Hook Ace 1 Rcs 1 Gcr 1 Lis. H HMG 1 Mads Myb Apses Hsf Fkh b. HLH Gata Homeo b. Zip C 2 H 2 -Zn C 6 -Fungal No. of members Rcs 1 p and Aft 2 p are global regulatory hubs with an as yet uncharacterized DNA binding domain Distribution of DNA binding domains in yeast transcription factors Transcriptional regulatory network in yeast Sub-network of Rcs 1 and Aft 2 p 123 41 Rcs 1 p 314 Number of target genes regulated How did the paralogous hubs that regulate distinct sets of genes evolve?
Rcs 1 is a global regulatory hub – Network analysis I P 53 Tigger Dal 82 Ime 1 Tea Abf 1 Tig AT-Hook Ace 1 Rcs 1 Gcr 1 Lis. H HMG 1 Mads Myb Apses Hsf Fkh b. HLH Gata Homeo b. Zip C 2 H 2 -Zn C 6 -Fungal No. of members Rcs 1 p and Aft 2 p are global regulatory hubs with an as yet uncharacterized DNA binding domain Distribution of DNA binding domains in yeast transcription factors Transcriptional regulatory network in yeast Sub-network of Rcs 1 and Aft 2 p 123 41 Rcs 1 p 314 Number of target genes regulated How did the paralogous hubs that regulate distinct sets of genes evolve?
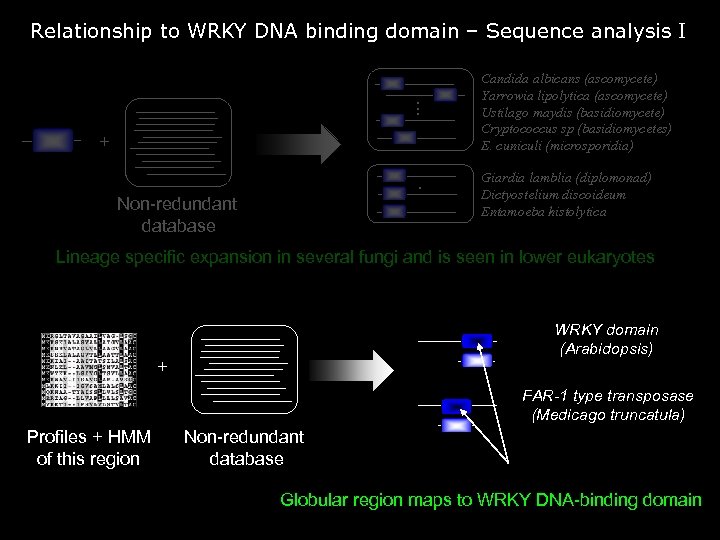 Relationship to WRKY DNA binding domain – Sequence analysis I . . . + . Non-redundant database Candida albicans (ascomycete) Yarrowia lipolytica (ascomycete) Ustilago maydis (basidiomycete) Cryptococcus sp (basidiomycetes) E. cuniculi (microsporidia) Giardia lamblia (diplomonad) Dictyostelium discoideum Entamoeba histolytica Lineage specific expansion in several fungi and is seen in lower eukaryotes WRKY domain (Arabidopsis) + FAR-1 type transposase (Medicago truncatula) Profiles + HMM of this region Non-redundant database Globular region maps to WRKY DNA-binding domain
Relationship to WRKY DNA binding domain – Sequence analysis I . . . + . Non-redundant database Candida albicans (ascomycete) Yarrowia lipolytica (ascomycete) Ustilago maydis (basidiomycete) Cryptococcus sp (basidiomycetes) E. cuniculi (microsporidia) Giardia lamblia (diplomonad) Dictyostelium discoideum Entamoeba histolytica Lineage specific expansion in several fungi and is seen in lower eukaryotes WRKY domain (Arabidopsis) + FAR-1 type transposase (Medicago truncatula) Profiles + HMM of this region Non-redundant database Globular region maps to WRKY DNA-binding domain
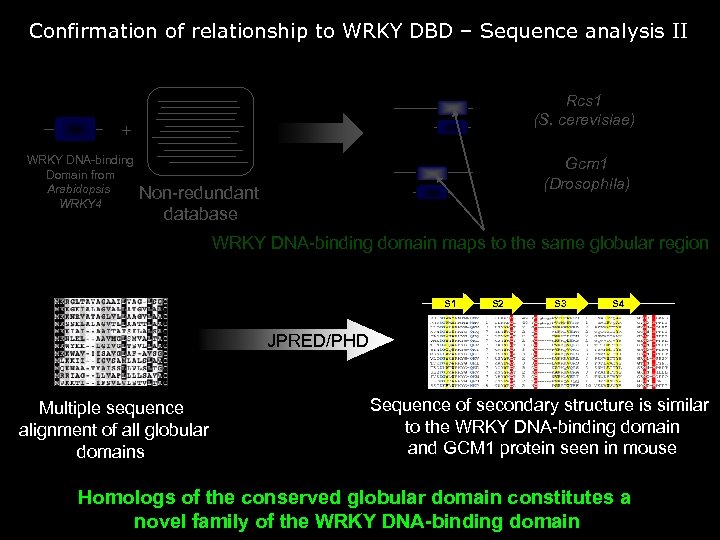 Confirmation of relationship to WRKY DBD – Sequence analysis II Rcs 1 (S. cerevisiae) + WRKY DNA-binding Domain from Arabidopsis WRKY 4 Gcm 1 (Drosophila) Non-redundant database WRKY DNA-binding domain maps to the same globular region S 1 S 2 S 3 S 4 JPRED/PHD Multiple sequence alignment of all globular domains Sequence of secondary structure is similar to the WRKY DNA-binding domain and GCM 1 protein seen in mouse Homologs of the conserved globular domain constitutes a novel family of the WRKY DNA-binding domain
Confirmation of relationship to WRKY DBD – Sequence analysis II Rcs 1 (S. cerevisiae) + WRKY DNA-binding Domain from Arabidopsis WRKY 4 Gcm 1 (Drosophila) Non-redundant database WRKY DNA-binding domain maps to the same globular region S 1 S 2 S 3 S 4 JPRED/PHD Multiple sequence alignment of all globular domains Sequence of secondary structure is similar to the WRKY DNA-binding domain and GCM 1 protein seen in mouse Homologs of the conserved globular domain constitutes a novel family of the WRKY DNA-binding domain
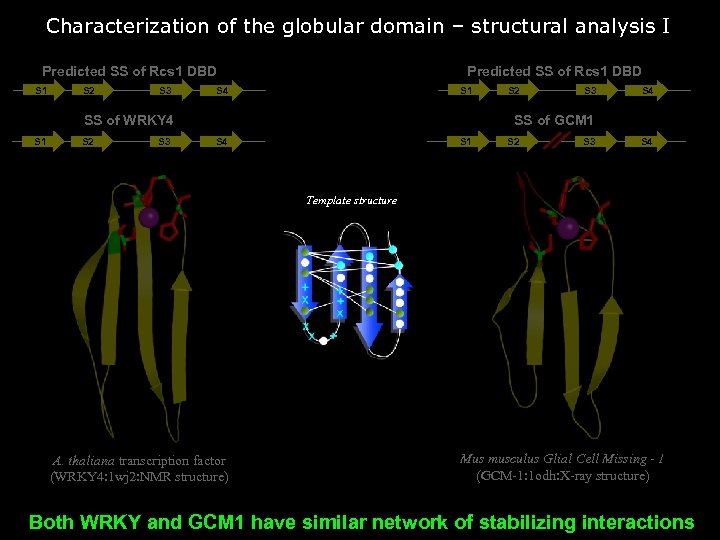 Characterization of the globular domain – structural analysis I Predicted SS of Rcs 1 DBD S 1 S 2 S 3 Predicted SS of Rcs 1 DBD S 4 S 1 SS of WRKY 4 S 1 S 2 S 3 S 4 SS of GCM 1 S 4 S 1 S 2 S 3 S 4 Template structure A. thaliana transcription factor (WRKY 4: 1 wj 2: NMR structure) Mus musculus Glial Cell Missing - 1 (GCM-1: 1 odh: X-ray structure) Both WRKY and GCM 1 have similar network of stabilizing interactions
Characterization of the globular domain – structural analysis I Predicted SS of Rcs 1 DBD S 1 S 2 S 3 Predicted SS of Rcs 1 DBD S 4 S 1 SS of WRKY 4 S 1 S 2 S 3 S 4 SS of GCM 1 S 4 S 1 S 2 S 3 S 4 Template structure A. thaliana transcription factor (WRKY 4: 1 wj 2: NMR structure) Mus musculus Glial Cell Missing - 1 (GCM-1: 1 odh: X-ray structure) Both WRKY and GCM 1 have similar network of stabilizing interactions
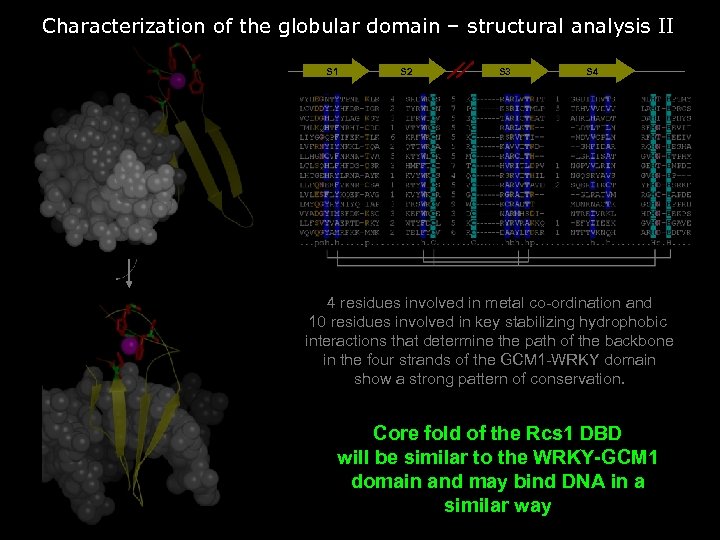 Characterization of the globular domain – structural analysis II S 1 S 2 S 3 S 4 4 residues involved in metal co-ordination and 10 residues involved in key stabilizing hydrophobic interactions that determine the path of the backbone in the four strands of the GCM 1 -WRKY domain show a strong pattern of conservation. Core fold of the Rcs 1 DBD will be similar to the WRKY-GCM 1 domain and may bind DNA in a similar way
Characterization of the globular domain – structural analysis II S 1 S 2 S 3 S 4 4 residues involved in metal co-ordination and 10 residues involved in key stabilizing hydrophobic interactions that determine the path of the backbone in the four strands of the GCM 1 -WRKY domain show a strong pattern of conservation. Core fold of the Rcs 1 DBD will be similar to the WRKY-GCM 1 domain and may bind DNA in a similar way
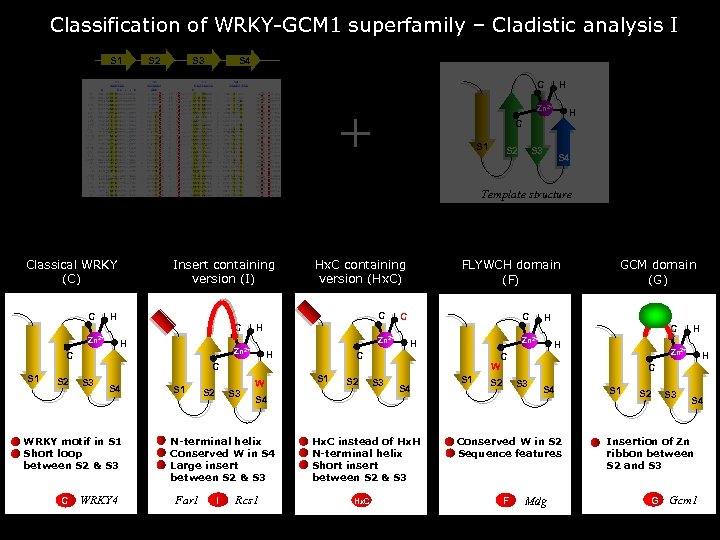 Classification of WRKY-GCM 1 superfamily – Cladistic analysis I S 1 S 2 S 3 S 4 C + H Zn 2+ H C S 1 S 2 S 3 S 4 Template structure Classical WRKY (C) C Insert containing version (I) C H Zn 2+ Hx. C containing version (Hx. C) C H C C H Zn 2+ C FLYWCH domain (F) H H S 2 S 3 S 4 WRKY motif in S 1 Short loop between S 2 & S 3 C WRKY 4 S 1 S 2 S 3 W I S 2 S 3 S 4 S 1 S 2 S 4 N-terminal helix Conserved W in S 4 Large insert between S 2 & S 3 Far 1 S 1 Rcs 1 Hx. C instead of Hx. H N-terminal helix Short insert between S 2 & S 3 Hx. C C H H C S 3 S 4 Conserved W in S 2 Sequence features F H Zn 2+ C W C S 1 H Zn 2+ C GCM domain (G) Mdg S 1 S 2 S 3 S 4 Insertion of Zn ribbon between S 2 and S 3 G Gcm 1
Classification of WRKY-GCM 1 superfamily – Cladistic analysis I S 1 S 2 S 3 S 4 C + H Zn 2+ H C S 1 S 2 S 3 S 4 Template structure Classical WRKY (C) C Insert containing version (I) C H Zn 2+ Hx. C containing version (Hx. C) C H C C H Zn 2+ C FLYWCH domain (F) H H S 2 S 3 S 4 WRKY motif in S 1 Short loop between S 2 & S 3 C WRKY 4 S 1 S 2 S 3 W I S 2 S 3 S 4 S 1 S 2 S 4 N-terminal helix Conserved W in S 4 Large insert between S 2 & S 3 Far 1 S 1 Rcs 1 Hx. C instead of Hx. H N-terminal helix Short insert between S 2 & S 3 Hx. C C H H C S 3 S 4 Conserved W in S 2 Sequence features F H Zn 2+ C W C S 1 H Zn 2+ C GCM domain (G) Mdg S 1 S 2 S 3 S 4 Insertion of Zn ribbon between S 2 and S 3 G Gcm 1
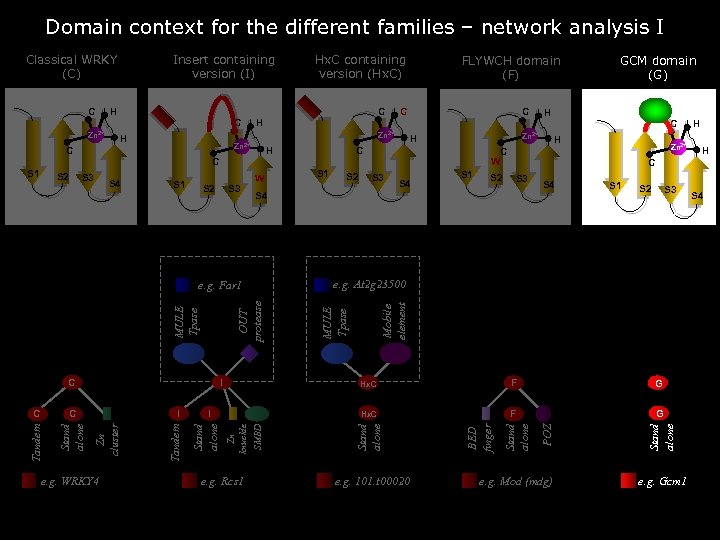 Domain context for the different families – network analysis I C H C H Zn 2+ C FLYWCH domain (F) H H S 2 S 3 W OUT protease MULE Tpase Zn knuckle SMBD Stand alone e. g. WRKY 4 S 1 S 2 S 3 S 4 S 1 S 2 S 3 e. g. At 2 g 23500 e. g. Rcs 1 F G Hx. C I Stand alone Tandem I Tandem C S 3 Hx. C I Zn cluster C S 2 Zn 2+ F G e. g. 101. t 00020 H H C S 4 e. g. Far 1 C S 1 BED finger S 1 H Mobile element S 4 Stand alone S 3 MULE Tpase S 2 C C W C S 1 H Zn 2+ C GCM domain (G) e. g. Mod (mdg) Stand alone Zn 2+ Hx. C containing version (Hx. C) POZ C Insert containing version (I) Stand alone Classical WRKY (C) e. g. Gcm 1 S 4
Domain context for the different families – network analysis I C H C H Zn 2+ C FLYWCH domain (F) H H S 2 S 3 W OUT protease MULE Tpase Zn knuckle SMBD Stand alone e. g. WRKY 4 S 1 S 2 S 3 S 4 S 1 S 2 S 3 e. g. At 2 g 23500 e. g. Rcs 1 F G Hx. C I Stand alone Tandem I Tandem C S 3 Hx. C I Zn cluster C S 2 Zn 2+ F G e. g. 101. t 00020 H H C S 4 e. g. Far 1 C S 1 BED finger S 1 H Mobile element S 4 Stand alone S 3 MULE Tpase S 2 C C W C S 1 H Zn 2+ C GCM domain (G) e. g. Mod (mdg) Stand alone Zn 2+ Hx. C containing version (Hx. C) POZ C Insert containing version (I) Stand alone Classical WRKY (C) e. g. Gcm 1 S 4
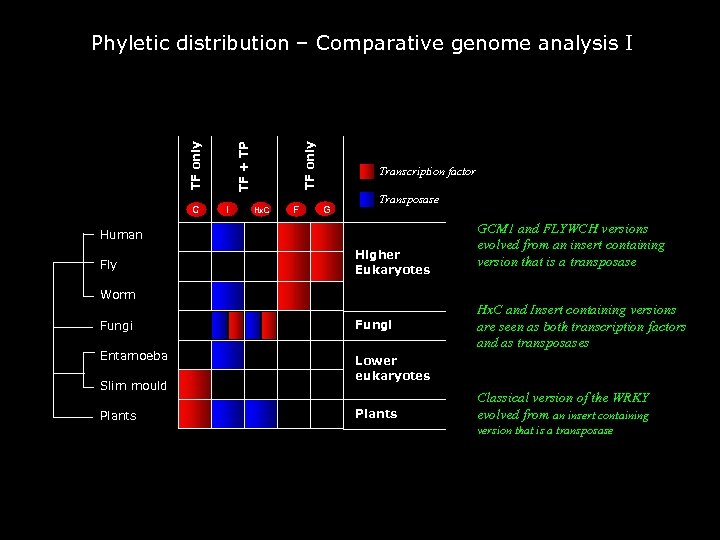 C I TF only TF + TP Phyletic distribution – Comparative genome analysis I Hx. C F Transcription factor G Transposase Human Fly Higher Eukaryotes Worm Fungi Entamoeba GCM 1 and FLYWCH versions evolved from an insert containing version that is a transposase Hx. C and Insert containing versions are seen as both transcription factors and as transposases Lower eukaryotes Slim mould Plants Classical version of the WRKY evolved from an insert containing version that is a transposase
C I TF only TF + TP Phyletic distribution – Comparative genome analysis I Hx. C F Transcription factor G Transposase Human Fly Higher Eukaryotes Worm Fungi Entamoeba GCM 1 and FLYWCH versions evolved from an insert containing version that is a transposase Hx. C and Insert containing versions are seen as both transcription factors and as transposases Lower eukaryotes Slim mould Plants Classical version of the WRKY evolved from an insert containing version that is a transposase
 -explain that there has been multiple transitions from transposase to TFs in the fungal genomes -explain how this could have happened by showing the snapshot of the breakup of selfish elements into two distinct products -explain that the transposase can itself regulate the gene expression of itself
-explain that there has been multiple transitions from transposase to TFs in the fungal genomes -explain how this could have happened by showing the snapshot of the breakup of selfish elements into two distinct products -explain that the transposase can itself regulate the gene expression of itself
 Outline of the presentation Rcs 1 and aft 2 have a distinct version of the WRKY type DNA binding domain Sensitive sequence search reveals that Oryza sativa (monocot) Arabidopsis thaliana (dicot) Medicago truncatula (dicot) Nicotiana tabacum (dicot)
Outline of the presentation Rcs 1 and aft 2 have a distinct version of the WRKY type DNA binding domain Sensitive sequence search reveals that Oryza sativa (monocot) Arabidopsis thaliana (dicot) Medicago truncatula (dicot) Nicotiana tabacum (dicot)
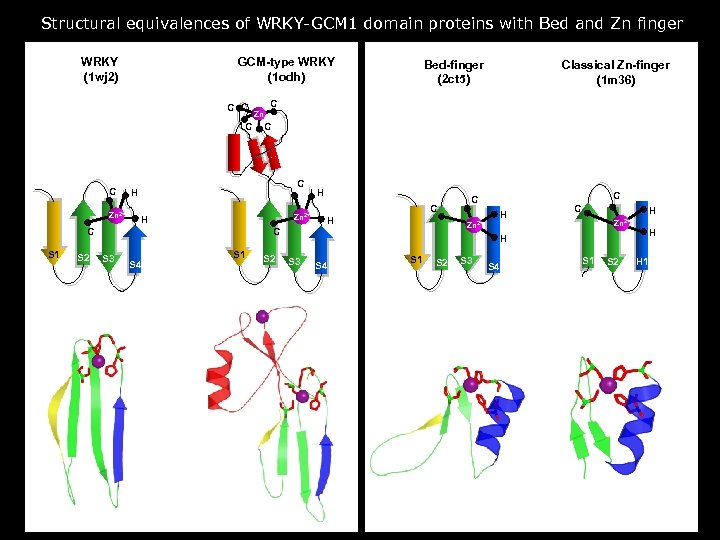 Structural equivalences of WRKY-GCM 1 domain proteins with Bed and Zn finger WRKY (1 wj 2) GCM-type WRKY (1 odh) Zn C Zn 2+ S 2 C C H Zn 2+ H C S 1 H C H S 4 S 1 S 2 C C Zn 2+ C S 3 Classical Zn-finger (1 m 36) C C C Bed-finger (2 ct 5) H C H Zn 2+ H H S 3 S 4 S 1 S 2 H 1
Structural equivalences of WRKY-GCM 1 domain proteins with Bed and Zn finger WRKY (1 wj 2) GCM-type WRKY (1 odh) Zn C Zn 2+ S 2 C C H Zn 2+ H C S 1 H C H S 4 S 1 S 2 C C Zn 2+ C S 3 Classical Zn-finger (1 m 36) C C C Bed-finger (2 ct 5) H C H Zn 2+ H H S 3 S 4 S 1 S 2 H 1
 Why Rcs 1? While systematically analyzing the genes which gave rise to abnormal cell size, We and the other noted that mutants of Rcs 1 give abnormal cell shape. It was known to be an important transcription factor involved in cell size regulation – explain showing graphs and images Independently, during the analysis of the TNET in yeast We looked at the hubs and the DNA binding domains That were present in them. Interestingly, there were two Hubs that did not have any known DNA binding domain Identified in them, but the region which mediates DNA was known – explain showing the family relationship Of the hubs -only two members, and both are hubs -how and when did they evolve? Standard search procedures using Pfam and other databases did not provide any clue about the domain. So we set out to characterize the DNA binding region from Rcs 1 p and its paralog Aft 2 p using sensitive sequence search and other computational methods. -show output from Pfam hits
Why Rcs 1? While systematically analyzing the genes which gave rise to abnormal cell size, We and the other noted that mutants of Rcs 1 give abnormal cell shape. It was known to be an important transcription factor involved in cell size regulation – explain showing graphs and images Independently, during the analysis of the TNET in yeast We looked at the hubs and the DNA binding domains That were present in them. Interestingly, there were two Hubs that did not have any known DNA binding domain Identified in them, but the region which mediates DNA was known – explain showing the family relationship Of the hubs -only two members, and both are hubs -how and when did they evolve? Standard search procedures using Pfam and other databases did not provide any clue about the domain. So we set out to characterize the DNA binding region from Rcs 1 p and its paralog Aft 2 p using sensitive sequence search and other computational methods. -show output from Pfam hits
 WRKY DNA binding domain – Structure analysis I Structural aspects of the DNA binding domain Explain the residues involved in metal chelating -DNA contacting surface -Inserts in the loops -Stabilizing contacts involved
WRKY DNA binding domain – Structure analysis I Structural aspects of the DNA binding domain Explain the residues involved in metal chelating -DNA contacting surface -Inserts in the loops -Stabilizing contacts involved
 WRKY DNA binding domain – Structure analysis II Structure comparisons identify several other Known transcription factors including the GCM protein in eukaryotes -Explain the insert of a zinc ribbon in the loop In fact sequence comparison without the insert can pick these WRKY proteins
WRKY DNA binding domain – Structure analysis II Structure comparisons identify several other Known transcription factors including the GCM protein in eukaryotes -Explain the insert of a zinc ribbon in the loop In fact sequence comparison without the insert can pick these WRKY proteins
 Classification of WRKY domains – Cladistic analysis I Multiple starting points identified all homologs in the different species This allowed us to classify the sequences into different families Each with a specific feature suggesting common evolutionary relationship Based on shared and derived features of the domains - List the 5 families and point to features involved using a structure template
Classification of WRKY domains – Cladistic analysis I Multiple starting points identified all homologs in the different species This allowed us to classify the sequences into different families Each with a specific feature suggesting common evolutionary relationship Based on shared and derived features of the domains - List the 5 families and point to features involved using a structure template
 Phylogenetic distribution and domain architecture for the different families - I Phyletic profiles of the different domains points to the possibility that these transcription factors could have evolved from transposases With at least two distinct recruitment into transcription factors. -In plants in one case -In the base of the fungal genomes in the other case
Phylogenetic distribution and domain architecture for the different families - I Phyletic profiles of the different domains points to the possibility that these transcription factors could have evolved from transposases With at least two distinct recruitment into transcription factors. -In plants in one case -In the base of the fungal genomes in the other case
 Phylogenetic distribution and domain architecture for the different families - II
Phylogenetic distribution and domain architecture for the different families - II
 Comparative genomics using the fungal genomes provides the clue for the evolution of these TFs -explain that there has been multiple transitions from transposase to TFs in the fungal genomes -explain how this could have happened by showing the snapshot of the breakup of selfish elements into two distinct products -explain that the transposase can itself regulate the gene expression of itself
Comparative genomics using the fungal genomes provides the clue for the evolution of these TFs -explain that there has been multiple transitions from transposase to TFs in the fungal genomes -explain how this could have happened by showing the snapshot of the breakup of selfish elements into two distinct products -explain that the transposase can itself regulate the gene expression of itself
 Comparative genomics using the fungal genomes provides the clue for the evolution of these TFs -extensive recruitment of the transposase in the different fungal lineages -multiple jumps within the fungal lineage -very recent duplication event in the order Saccharomycetales suggest hubs could Evolve rapidly -Candida rbf 1 and other TFs independently duplicated and evolved as global regulators
Comparative genomics using the fungal genomes provides the clue for the evolution of these TFs -extensive recruitment of the transposase in the different fungal lineages -multiple jumps within the fungal lineage -very recent duplication event in the order Saccharomycetales suggest hubs could Evolve rapidly -Candida rbf 1 and other TFs independently duplicated and evolved as global regulators
 Analysis of the gene expression data in plants Since it happened in fungal genomes, we ask how does this behave in the plants. -show the gene expression patterns for the different subfamilies. We see two trends one where divergence has primarily occurred in the expression changes rather than in the protein sequence, and the other in which proteins with the same expression pattern have different binding site residues. -spatio-temporal changes in gene expression -It is experimentally well known that the FLYWCH and the GCM proteins are developmentally important regulatory proteins. So in three lineages there has been recruitment of the transposase into becoming a developmentally important global regulator.
Analysis of the gene expression data in plants Since it happened in fungal genomes, we ask how does this behave in the plants. -show the gene expression patterns for the different subfamilies. We see two trends one where divergence has primarily occurred in the expression changes rather than in the protein sequence, and the other in which proteins with the same expression pattern have different binding site residues. -spatio-temporal changes in gene expression -It is experimentally well known that the FLYWCH and the GCM proteins are developmentally important regulatory proteins. So in three lineages there has been recruitment of the transposase into becoming a developmentally important global regulator.
 Analysis of the gene expression data in plants There are interesting traces of gene expression pattern when we see for the different WRKY containing proteins. TPases are expressed in the root and in the pollen enhancing the possibility of rapidly expanding themselves during evolution.
Analysis of the gene expression data in plants There are interesting traces of gene expression pattern when we see for the different WRKY containing proteins. TPases are expressed in the root and in the pollen enhancing the possibility of rapidly expanding themselves during evolution.
 Acknowledgements Aravind group L Aravind S Balaji Lakshminarayan Iyer
Acknowledgements Aravind group L Aravind S Balaji Lakshminarayan Iyer
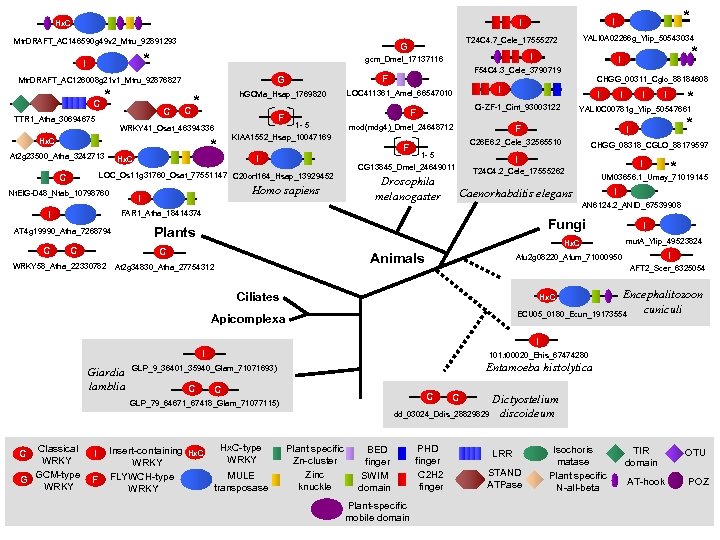 Mtr. DRAFT_AC 146590 g 49 v 2_Mtru_92891293 * I C * TTR 1_Atha_30694675 C h. GCMa_Hsap_1769820 * C Hx. C Nt. EIG-D 48_Ntab_10798760 I Homo sapiens I I Ci-ZF-1_Cint_93003122 mod(mdg 4)_Dmel_24648712 F 1 - 5 CG 13845_Dmel_24649011 Drosophila melanogaster I I Caenorhabditis elegans AN 6124. 2_ANID_67539908 Hx. C Animals At 2 g 34830_Atha_27754312 * I UM 03656. 1_Umay_71019145 Fungi C WRKY 58_Atha_22330782 CHGG_08318_CGLO_88179597 T 24 C 4. 2_Cele_17555262 Plants C * * I I FAR 1_Atha_18414374 AT 4 g 19990_Atha_7268794 I I YALI 0 C 00781 g_Ylip_50547661 C 26 E 6. 2_Cele_32565510 F I LOC_Os 11 g 31760_Osat_77551147 C 20 orf 164_Hsap_13929452 C C 1 - 5 KIAA 1552_Hsap_10047169 * Hx. C CHGG_00311_Cglo_88184608 I LOC 411361_Amel_66547010 F F * I F 54 C 4. 3_Cele_3790719 F G WRKY 41_Osat_46394336 At 2 g 23500_Atha_3242713 I gcm_Dmel_17137116 Mtr. DRAFT_AC 126008 g 21 v 1_Mtru_92876827 YALI 0 A 02266 g_Ylip_50543034 T 24 C 4. 7_Cele_17555272 G * I I Hx. C I mut. A_Ylip_49523824 I Afu 2 g 08220_Afum_71000950 AFT 2_Scer_6325054 Encephalitozoon cuniculi ECU 05_0180_Ecun_19173554 Ciliates Hx. C Apicomplexa I I Giardia lamblia 101. t 00020_Ehis_67474280 GLP_9_36401_35940_Glam_71071693) Entamoeba histolytica C Classical WRKY GCM-type G WRKY C I F Insert-containing WRKY FLYWCH-type WRKY Hx. C-type WRKY MULE transposase C Dictyostelium dd_03024_Ddis_28829829 discoideum GLP_79_64671_67418_Glam_71077115) Plant specific Zn-cluster Zinc knuckle BED finger SWIM domain Plant-specific mobile domain PHD finger C 2 H 2 finger LRR STAND ATPase Isochoris matase Plant specific N-all-beta TIR domain AT-hook OTU POZ
Mtr. DRAFT_AC 146590 g 49 v 2_Mtru_92891293 * I C * TTR 1_Atha_30694675 C h. GCMa_Hsap_1769820 * C Hx. C Nt. EIG-D 48_Ntab_10798760 I Homo sapiens I I Ci-ZF-1_Cint_93003122 mod(mdg 4)_Dmel_24648712 F 1 - 5 CG 13845_Dmel_24649011 Drosophila melanogaster I I Caenorhabditis elegans AN 6124. 2_ANID_67539908 Hx. C Animals At 2 g 34830_Atha_27754312 * I UM 03656. 1_Umay_71019145 Fungi C WRKY 58_Atha_22330782 CHGG_08318_CGLO_88179597 T 24 C 4. 2_Cele_17555262 Plants C * * I I FAR 1_Atha_18414374 AT 4 g 19990_Atha_7268794 I I YALI 0 C 00781 g_Ylip_50547661 C 26 E 6. 2_Cele_32565510 F I LOC_Os 11 g 31760_Osat_77551147 C 20 orf 164_Hsap_13929452 C C 1 - 5 KIAA 1552_Hsap_10047169 * Hx. C CHGG_00311_Cglo_88184608 I LOC 411361_Amel_66547010 F F * I F 54 C 4. 3_Cele_3790719 F G WRKY 41_Osat_46394336 At 2 g 23500_Atha_3242713 I gcm_Dmel_17137116 Mtr. DRAFT_AC 126008 g 21 v 1_Mtru_92876827 YALI 0 A 02266 g_Ylip_50543034 T 24 C 4. 7_Cele_17555272 G * I I Hx. C I mut. A_Ylip_49523824 I Afu 2 g 08220_Afum_71000950 AFT 2_Scer_6325054 Encephalitozoon cuniculi ECU 05_0180_Ecun_19173554 Ciliates Hx. C Apicomplexa I I Giardia lamblia 101. t 00020_Ehis_67474280 GLP_9_36401_35940_Glam_71071693) Entamoeba histolytica C Classical WRKY GCM-type G WRKY C I F Insert-containing WRKY FLYWCH-type WRKY Hx. C-type WRKY MULE transposase C Dictyostelium dd_03024_Ddis_28829829 discoideum GLP_79_64671_67418_Glam_71077115) Plant specific Zn-cluster Zinc knuckle BED finger SWIM domain Plant-specific mobile domain PHD finger C 2 H 2 finger LRR STAND ATPase Isochoris matase Plant specific N-all-beta TIR domain AT-hook OTU POZ
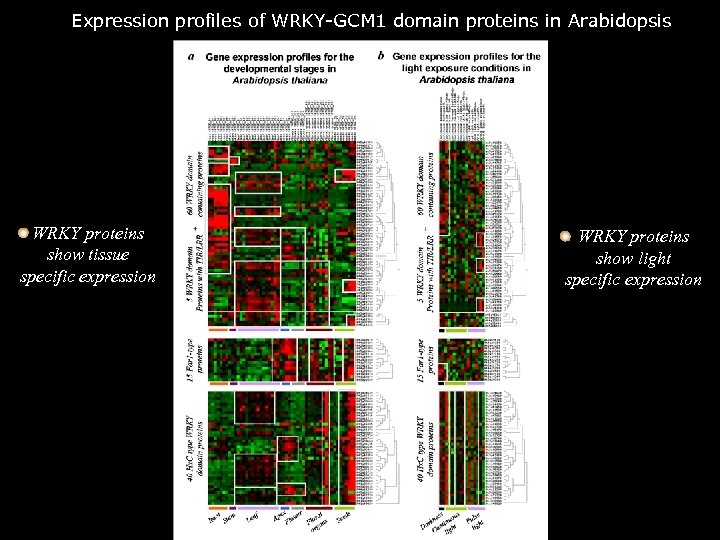 Expression profiles of WRKY-GCM 1 domain proteins in Arabidopsis WRKY proteins show tissue specific expression WRKY proteins show light specific expression
Expression profiles of WRKY-GCM 1 domain proteins in Arabidopsis WRKY proteins show tissue specific expression WRKY proteins show light specific expression
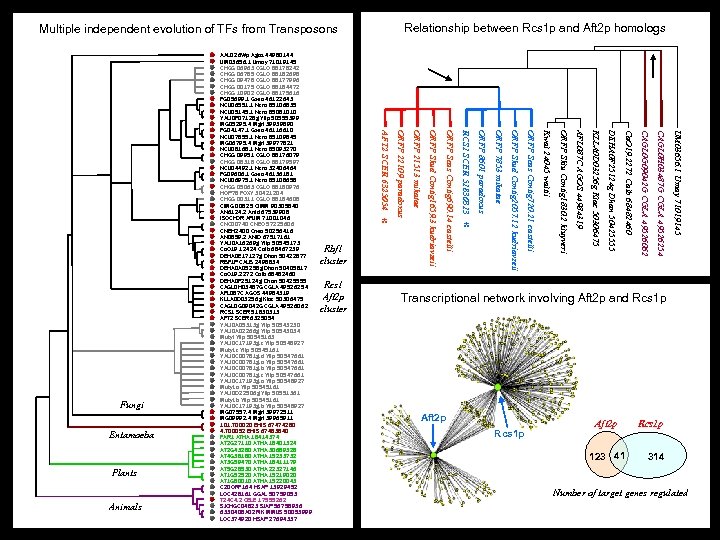 Relationship between Rcs 1 p and Aft 2 p homologs Multiple independent evolution of TFs from Transposons UM 03656. 1 Umay 71019145 CAGL 0 H 03487 G CGLA 49526254 CAGL 0 G 09042 G CGLA 49526062 Ca. O 19. 2272 Calb 68482460 DEHA 0 F 25124 g Dhan 50425555 KLLA 0 D 03256 g Klac 50306475 AFL 087 C AGOS 44984319 ORFP Sklu Contig 1830. 2 kluyveri Kwal 24045 waltii ORFP Scas Contig 720. 21 castelli ORFP Skud Contig 2057. 12 kudriavzeii ORFP 7853 mikatae * ORFP 8601 paradoxus RCS 1 SCER 51830313 ORFP Scas Contig 690. 14 castelli Rcs 1 Aft 2 p cluster ORFP Skud Contig 1659. 3 kudriavzeii Animals Rbf 1 cluster ORFP 21513 mikatae Plants * ORFP 22109 paradoxus Entamoeba AFT 2 SCER 6325054 Fungi AAL 026 Wp Agos 44980144 UM 03656. 1 Umay 71019145 CHGG 06963 CGLO 88178242 CHGG 06785 CGLO 88182698 CHGG 09478 CGLO 88177996 CHGG 00175 CGLO 88184472 CHGG 10902 CGLO 88175616 FG 05699. 1 Gzea 46122643 NCU 06551. 1 Ncra 85106835 NCU 05145. 1 Ncra 85081010 YALI 0 F 07128 g Ylip 50555399 MG 05295. 4 Mgri 39939890 FG 04147. 1 Gzea 46116610 NCU 07855. 1 Ncra 85109845 MG 06795. 4 Mgri 39977821 NCU 08168. 1 Ncra 85093270 CHGG 09951 CGLO 88176079 CHGG 08318 CGLO 88179597 NCU 04492. 1 Ncra 32406464 FG 09606. 1 Gzea 46136181 NCU 06975. 1 Ncra 85108658 CHGG 05063 CGLO 88180976 HOP 78 FOXY 30421204 CHGG 00311 CGLO 88184608 CIMG 00825 CIMM 90305840 AN 6124. 2 Anid 67539908 ISOCHOR AFUM 71001046 CNC 00740 CNEO 57225606 CNBH 2400 Cneo 50256416 AN 0859. 2 ANID 67517161 YALI 0 A 16269 g Ylip 50545173 Ca. O 19 12424 Calb 68467239 DEHA 0 E 17127 g Dhan 50422877 RBF 1 P CALB 2498834 DEHA 0 A 05258 g Dhan 50405817 Ca. O 19. 2272 Calb 68482460 DEHA 0 F 25124 g Dhan 50425555 CAGL 0 H 03487 G CGLA 49526254 AFL 087 C AGOS 44984319 KLLA 0 D 03256 g Klac 50306475 CAGL 0 G 09042 G CGLA 49526062 RCS 1 SCER 51830313 AFT 2 SCER 6325054 YALI 0 A 05313 g Ylip 50543230 YALI 0 A 02266 g Ylip 50543034 Mutyl Ylip 50545163 YALI 0 C 17193 g. c Ylip 50548927 Mutyl. c Ylip 50545161 YALI 0 C 00781 g. d Ylip 50547661 YALI 0 C 00781 g. a Ylip 50547661 YALI 0 C 00781 g. b Ylip 50547661 YALI 0 C 00781 g. c Ylip 50547661 YALI 0 C 17193 g. a Ylip 50548927 Mutyl. a Ylip 50545161 YALI 0 D 22506 g Ylip 50551361 Mutyl. b Ylip 50545161 YALI 0 C 17193 g. b Ylip 50548927 MG 07557. 4 Mgri 39972511 MG 09992. 4 Mgri 39965911 101. T 00020 EHIS 67474280 4. T 00052 EHIS 67483840 FAR 1 ATHA 18414374 AT 2 G 27110 ATHA 18401324 AT 2 G 43280 ATHA 30689328 AT 4 G 38180 ATHA 15233732 AT 3 G 59470 ATHA 18411179 AT 5 G 28530 ATHA 22327146 AT 1 G 52520 ATHA 15219020 AT 1 G 80010 ATHA 15220043 C 20 ORF 164 HSAP 13929452 LOC 428161 GGAL 50759053 T 24 C 4. 2 CELE 17555262 SJCHGC 04823 SJAP 56758936 6330408 A 02 RIK MMUS 50053999 LOC 374920 HSAP 27694337 Transcriptional network involving Aft 2 p and Rcs 1 p Aft 2 p 123 41 Rcs 1 p 314 Number of target genes regulated
Relationship between Rcs 1 p and Aft 2 p homologs Multiple independent evolution of TFs from Transposons UM 03656. 1 Umay 71019145 CAGL 0 H 03487 G CGLA 49526254 CAGL 0 G 09042 G CGLA 49526062 Ca. O 19. 2272 Calb 68482460 DEHA 0 F 25124 g Dhan 50425555 KLLA 0 D 03256 g Klac 50306475 AFL 087 C AGOS 44984319 ORFP Sklu Contig 1830. 2 kluyveri Kwal 24045 waltii ORFP Scas Contig 720. 21 castelli ORFP Skud Contig 2057. 12 kudriavzeii ORFP 7853 mikatae * ORFP 8601 paradoxus RCS 1 SCER 51830313 ORFP Scas Contig 690. 14 castelli Rcs 1 Aft 2 p cluster ORFP Skud Contig 1659. 3 kudriavzeii Animals Rbf 1 cluster ORFP 21513 mikatae Plants * ORFP 22109 paradoxus Entamoeba AFT 2 SCER 6325054 Fungi AAL 026 Wp Agos 44980144 UM 03656. 1 Umay 71019145 CHGG 06963 CGLO 88178242 CHGG 06785 CGLO 88182698 CHGG 09478 CGLO 88177996 CHGG 00175 CGLO 88184472 CHGG 10902 CGLO 88175616 FG 05699. 1 Gzea 46122643 NCU 06551. 1 Ncra 85106835 NCU 05145. 1 Ncra 85081010 YALI 0 F 07128 g Ylip 50555399 MG 05295. 4 Mgri 39939890 FG 04147. 1 Gzea 46116610 NCU 07855. 1 Ncra 85109845 MG 06795. 4 Mgri 39977821 NCU 08168. 1 Ncra 85093270 CHGG 09951 CGLO 88176079 CHGG 08318 CGLO 88179597 NCU 04492. 1 Ncra 32406464 FG 09606. 1 Gzea 46136181 NCU 06975. 1 Ncra 85108658 CHGG 05063 CGLO 88180976 HOP 78 FOXY 30421204 CHGG 00311 CGLO 88184608 CIMG 00825 CIMM 90305840 AN 6124. 2 Anid 67539908 ISOCHOR AFUM 71001046 CNC 00740 CNEO 57225606 CNBH 2400 Cneo 50256416 AN 0859. 2 ANID 67517161 YALI 0 A 16269 g Ylip 50545173 Ca. O 19 12424 Calb 68467239 DEHA 0 E 17127 g Dhan 50422877 RBF 1 P CALB 2498834 DEHA 0 A 05258 g Dhan 50405817 Ca. O 19. 2272 Calb 68482460 DEHA 0 F 25124 g Dhan 50425555 CAGL 0 H 03487 G CGLA 49526254 AFL 087 C AGOS 44984319 KLLA 0 D 03256 g Klac 50306475 CAGL 0 G 09042 G CGLA 49526062 RCS 1 SCER 51830313 AFT 2 SCER 6325054 YALI 0 A 05313 g Ylip 50543230 YALI 0 A 02266 g Ylip 50543034 Mutyl Ylip 50545163 YALI 0 C 17193 g. c Ylip 50548927 Mutyl. c Ylip 50545161 YALI 0 C 00781 g. d Ylip 50547661 YALI 0 C 00781 g. a Ylip 50547661 YALI 0 C 00781 g. b Ylip 50547661 YALI 0 C 00781 g. c Ylip 50547661 YALI 0 C 17193 g. a Ylip 50548927 Mutyl. a Ylip 50545161 YALI 0 D 22506 g Ylip 50551361 Mutyl. b Ylip 50545161 YALI 0 C 17193 g. b Ylip 50548927 MG 07557. 4 Mgri 39972511 MG 09992. 4 Mgri 39965911 101. T 00020 EHIS 67474280 4. T 00052 EHIS 67483840 FAR 1 ATHA 18414374 AT 2 G 27110 ATHA 18401324 AT 2 G 43280 ATHA 30689328 AT 4 G 38180 ATHA 15233732 AT 3 G 59470 ATHA 18411179 AT 5 G 28530 ATHA 22327146 AT 1 G 52520 ATHA 15219020 AT 1 G 80010 ATHA 15220043 C 20 ORF 164 HSAP 13929452 LOC 428161 GGAL 50759053 T 24 C 4. 2 CELE 17555262 SJCHGC 04823 SJAP 56758936 6330408 A 02 RIK MMUS 50053999 LOC 374920 HSAP 27694337 Transcriptional network involving Aft 2 p and Rcs 1 p Aft 2 p 123 41 Rcs 1 p 314 Number of target genes regulated
 Conclusion Integration of different types of experimental data allowed us to Identify the DNA binding domain in Rcs 1 Sequence Structure Expression Interaction
Conclusion Integration of different types of experimental data allowed us to Identify the DNA binding domain in Rcs 1 Sequence Structure Expression Interaction


