7bbd6c5699162ba4de531f6876b63c08.ppt
- Количество слайдов: 46
 INSERM TAGC Diversity of UTR Regions INSERM TAGC, Marseille Alternate Transcript Diversity
INSERM TAGC Diversity of UTR Regions INSERM TAGC, Marseille Alternate Transcript Diversity
 INSERM TAGC Part I: EST-based analysis of polyadenylation and UTRs
INSERM TAGC Part I: EST-based analysis of polyadenylation and UTRs
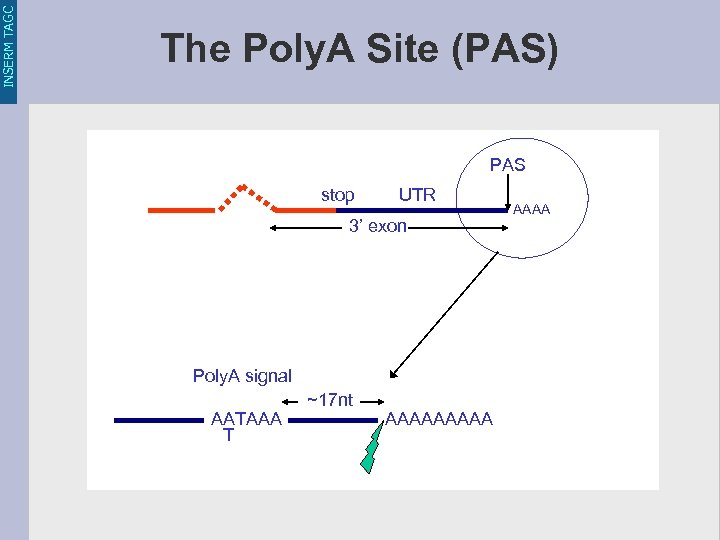 INSERM TAGC The Poly. A Site (PAS) PAS stop UTR 3’ exon Poly. A signal AATAAA T ~17 nt AAAAA
INSERM TAGC The Poly. A Site (PAS) PAS stop UTR 3’ exon Poly. A signal AATAAA T ~17 nt AAAAA
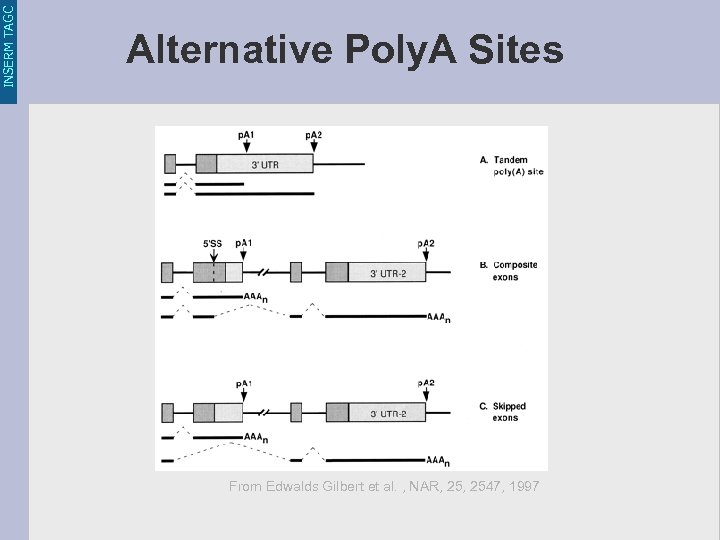 INSERM TAGC Alternative Poly. A Sites From Edwalds Gilbert et al. , NAR, 2547, 1997
INSERM TAGC Alternative Poly. A Sites From Edwalds Gilbert et al. , NAR, 2547, 1997
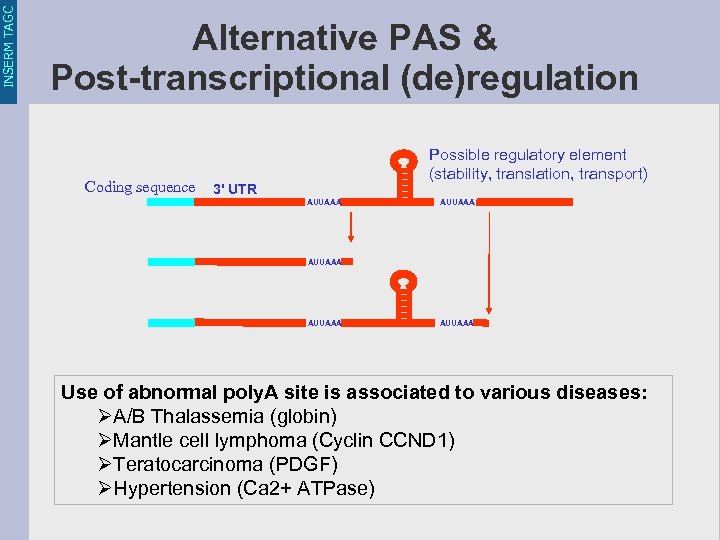 INSERM TAGC Alternative PAS & Post-transcriptional (de)regulation Coding sequence Possible regulatory element (stability, translation, transport) 3' UTR AUUAAA AUUAAA Use of abnormal poly. A site is associated to various diseases: ØA/B Thalassemia (globin) ØMantle cell lymphoma (Cyclin CCND 1) ØTeratocarcinoma (PDGF) ØHypertension (Ca 2+ ATPase)
INSERM TAGC Alternative PAS & Post-transcriptional (de)regulation Coding sequence Possible regulatory element (stability, translation, transport) 3' UTR AUUAAA AUUAAA Use of abnormal poly. A site is associated to various diseases: ØA/B Thalassemia (globin) ØMantle cell lymphoma (Cyclin CCND 1) ØTeratocarcinoma (PDGF) ØHypertension (Ca 2+ ATPase)
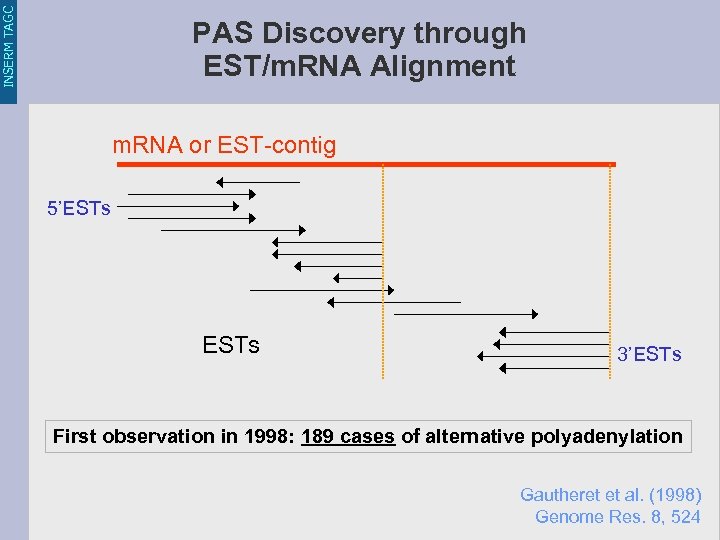 INSERM TAGC PAS Discovery through EST/m. RNA Alignment m. RNA or EST-contig 5’ESTs 3’ESTs First observation in 1998: 189 cases of alternative polyadenylation Gautheret et al. (1998) Genome Res. 8, 524
INSERM TAGC PAS Discovery through EST/m. RNA Alignment m. RNA or EST-contig 5’ESTs 3’ESTs First observation in 1998: 189 cases of alternative polyadenylation Gautheret et al. (1998) Genome Res. 8, 524
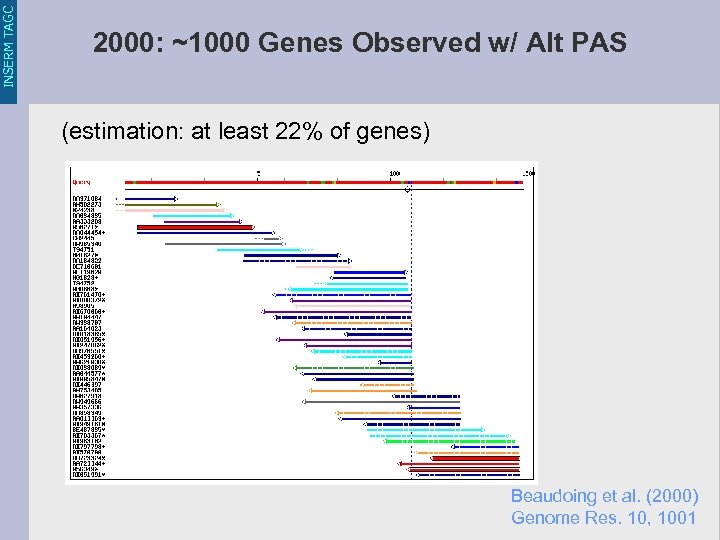 INSERM TAGC 2000: ~1000 Genes Observed w/ Alt PAS (estimation: at least 22% of genes) Beaudoing et al. (2000) Genome Res. 10, 1001
INSERM TAGC 2000: ~1000 Genes Observed w/ Alt PAS (estimation: at least 22% of genes) Beaudoing et al. (2000) Genome Res. 10, 1001
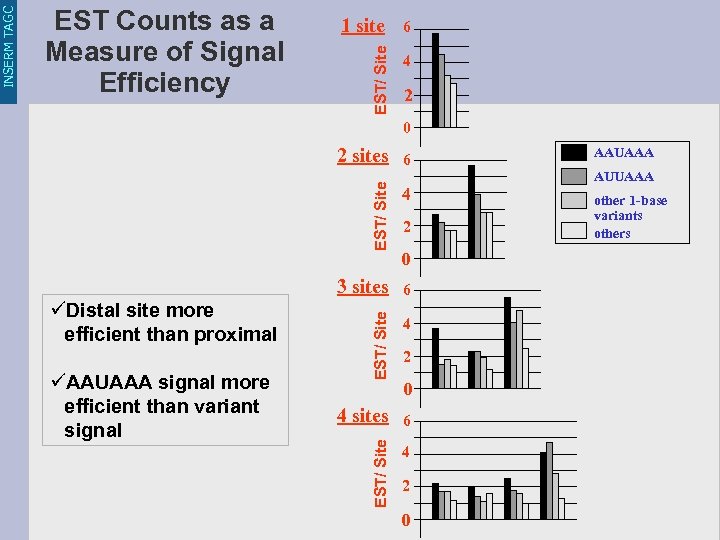 1 site 6 EST/ Site 4 2 0 6 EST/ Site 2 sites 3 sites üAAUAAA signal more efficient than variant signal 6 2 0 EST/ Site üDistal site more efficient than proximal 4 4 4 sites 6 EST/ Site INSERM TAGC EST Counts as a Measure of Signal Efficiency 4 2 0 AAUAAA AUUAAA other 1 -base variants others
1 site 6 EST/ Site 4 2 0 6 EST/ Site 2 sites 3 sites üAAUAAA signal more efficient than variant signal 6 2 0 EST/ Site üDistal site more efficient than proximal 4 4 4 sites 6 EST/ Site INSERM TAGC EST Counts as a Measure of Signal Efficiency 4 2 0 AAUAAA AUUAAA other 1 -base variants others
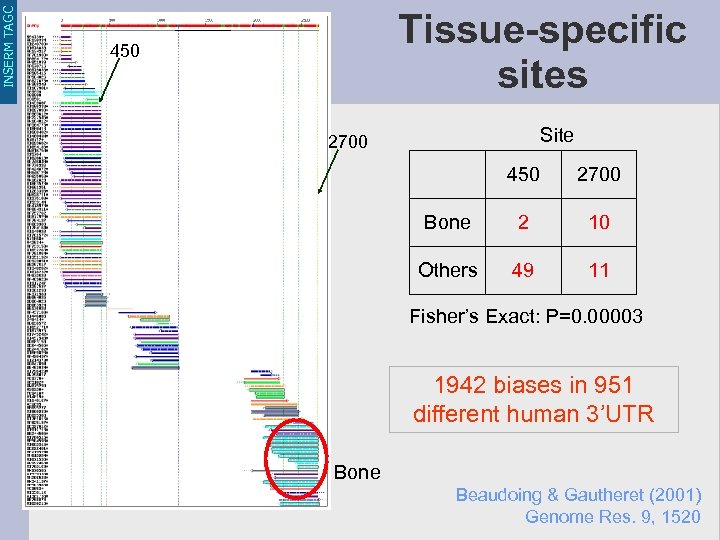 INSERM TAGC Tissue-specific sites 450 Site 2700 450 2700 Bone 2 10 Others 49 11 Fisher’s Exact: P=0. 00003 1942 biases in 951 different human 3’UTR Bone Beaudoing & Gautheret (2001) Genome Res. 9, 1520
INSERM TAGC Tissue-specific sites 450 Site 2700 450 2700 Bone 2 10 Others 49 11 Fisher’s Exact: P=0. 00003 1942 biases in 951 different human 3’UTR Bone Beaudoing & Gautheret (2001) Genome Res. 9, 1520
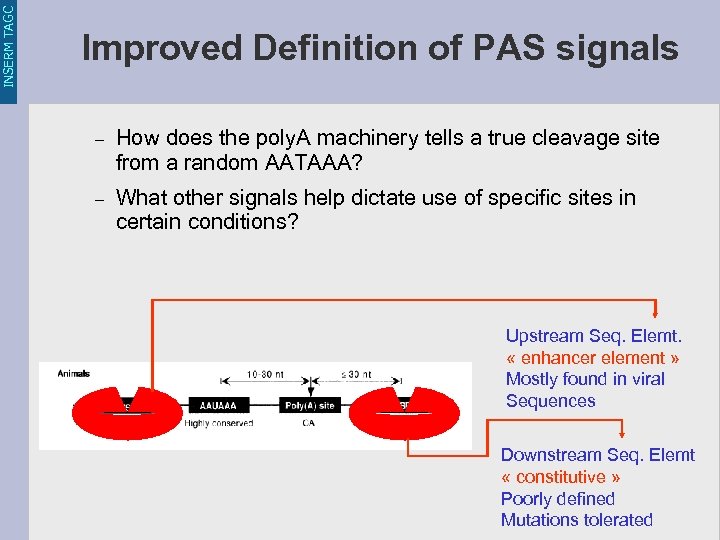 INSERM TAGC Improved Definition of PAS signals – How does the poly. A machinery tells a true cleavage site from a random AATAAA? – What other signals help dictate use of specific sites in certain conditions? Upstream Seq. Elemt. « enhancer element » Mostly found in viral Sequences Downstream Seq. Elemt « constitutive » Poorly defined Mutations tolerated
INSERM TAGC Improved Definition of PAS signals – How does the poly. A machinery tells a true cleavage site from a random AATAAA? – What other signals help dictate use of specific sites in certain conditions? Upstream Seq. Elemt. « enhancer element » Mostly found in viral Sequences Downstream Seq. Elemt « constitutive » Poorly defined Mutations tolerated
 INSERM TAGC Analysis of PAS-flanking regions Genomic
INSERM TAGC Analysis of PAS-flanking regions Genomic
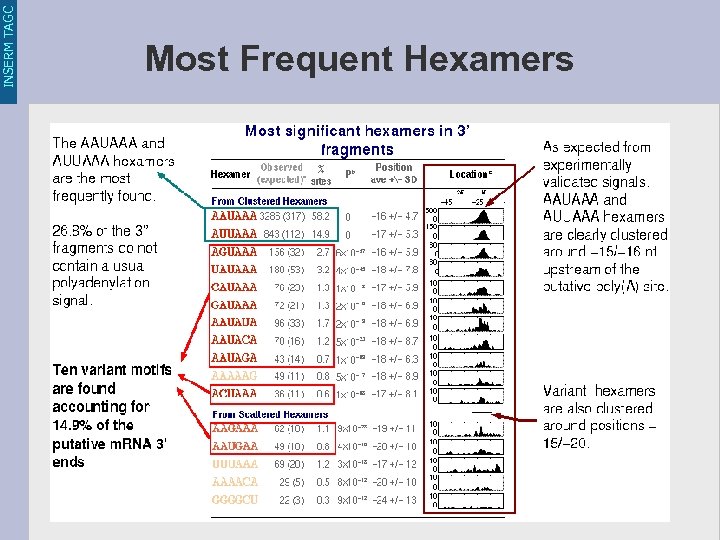 INSERM TAGC Most Frequent Hexamers
INSERM TAGC Most Frequent Hexamers
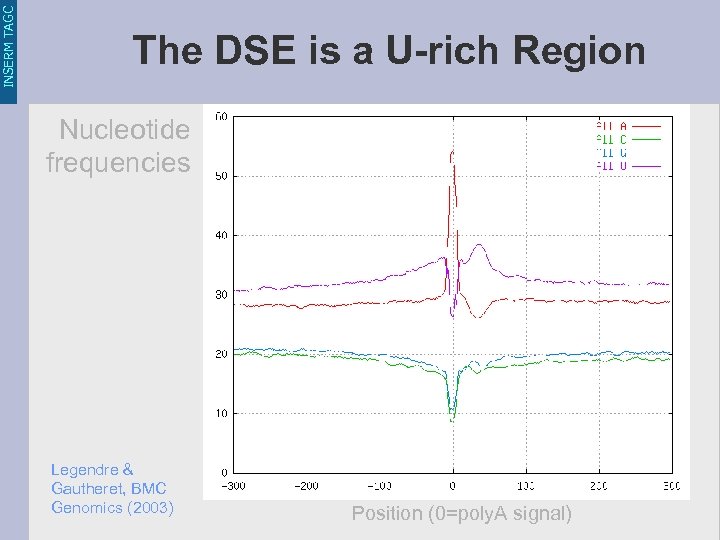 INSERM TAGC The DSE is a U-rich Region Nucleotide frequencies Legendre & Gautheret, BMC Genomics (2003) Position (0=poly. A signal)
INSERM TAGC The DSE is a U-rich Region Nucleotide frequencies Legendre & Gautheret, BMC Genomics (2003) Position (0=poly. A signal)
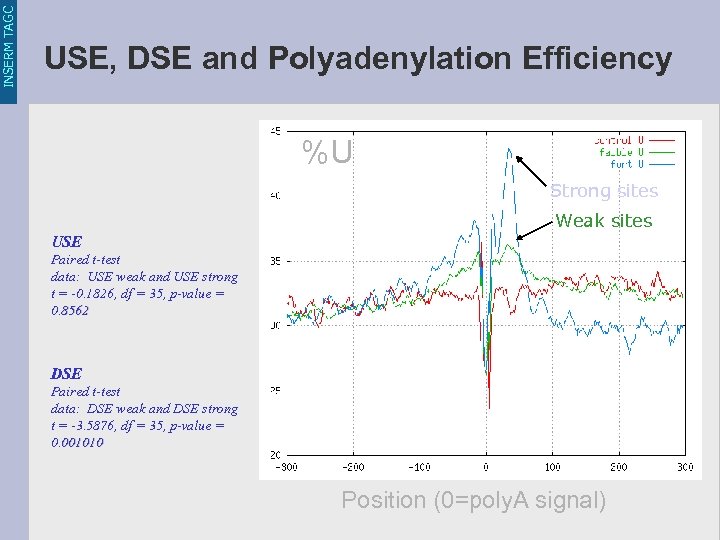 INSERM TAGC USE, DSE and Polyadenylation Efficiency %U Strong sites Weak sites USE Paired t-test data: USE weak and USE strong t = -0. 1826, df = 35, p-value = 0. 8562 DSE Paired t-test data: DSE weak and DSE strong t = -3. 5876, df = 35, p-value = 0. 001010 Position (0=poly. A signal)
INSERM TAGC USE, DSE and Polyadenylation Efficiency %U Strong sites Weak sites USE Paired t-test data: USE weak and USE strong t = -0. 1826, df = 35, p-value = 0. 8562 DSE Paired t-test data: DSE weak and DSE strong t = -3. 5876, df = 35, p-value = 0. 001010 Position (0=poly. A signal)
 INSERM TAGC EST-based PAS Map ● 3’ ESTs mapped directly to Genome ● 2005 67, 000 poly. A sites identified – – ● 28000 sites within 2 K of an Ensembl gene 29000 sites not within 10 K of an Ensembl gene 57% of genes have 2 or more poly. A sites – (was 22% in 2000!)
INSERM TAGC EST-based PAS Map ● 3’ ESTs mapped directly to Genome ● 2005 67, 000 poly. A sites identified – – ● 28000 sites within 2 K of an Ensembl gene 29000 sites not within 10 K of an Ensembl gene 57% of genes have 2 or more poly. A sites – (was 22% in 2000!)
 INSERM TAGC The ATD Project ü Integrate Splice+Poly. A+Init variants ü Quality control ü Tissue-specific Isoforms ü Regulatory motifs ü Isoform specific oligos ü RT-PCR validation of selected isoforms The ATD project is funded by the European Commission within its FP 6 Programme, under thematic area "Life sciences, genomics and biotechnology for health", contract number LHSG-CT-2003 -503329
INSERM TAGC The ATD Project ü Integrate Splice+Poly. A+Init variants ü Quality control ü Tissue-specific Isoforms ü Regulatory motifs ü Isoform specific oligos ü RT-PCR validation of selected isoforms The ATD project is funded by the European Commission within its FP 6 Programme, under thematic area "Life sciences, genomics and biotechnology for health", contract number LHSG-CT-2003 -503329
 INSERM TAGC Revisiting UTR Length
INSERM TAGC Revisiting UTR Length
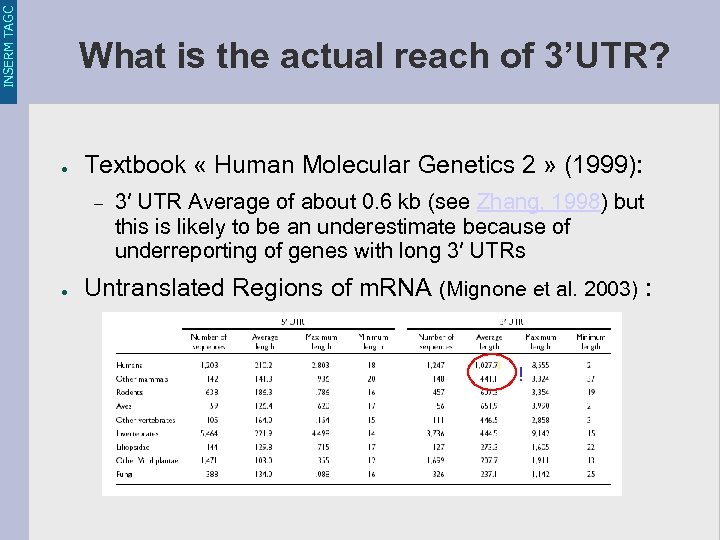 INSERM TAGC What is the actual reach of 3’UTR? ● Textbook « Human Molecular Genetics 2 » (1999): – ● 3′ UTR Average of about 0. 6 kb (see Zhang, 1998) but this is likely to be an underestimate because of underreporting of genes with long 3′ UTRs Untranslated Regions of m. RNA (Mignone et al. 2003) : !
INSERM TAGC What is the actual reach of 3’UTR? ● Textbook « Human Molecular Genetics 2 » (1999): – ● 3′ UTR Average of about 0. 6 kb (see Zhang, 1998) but this is likely to be an underestimate because of underreporting of genes with long 3′ UTRs Untranslated Regions of m. RNA (Mignone et al. 2003) : !
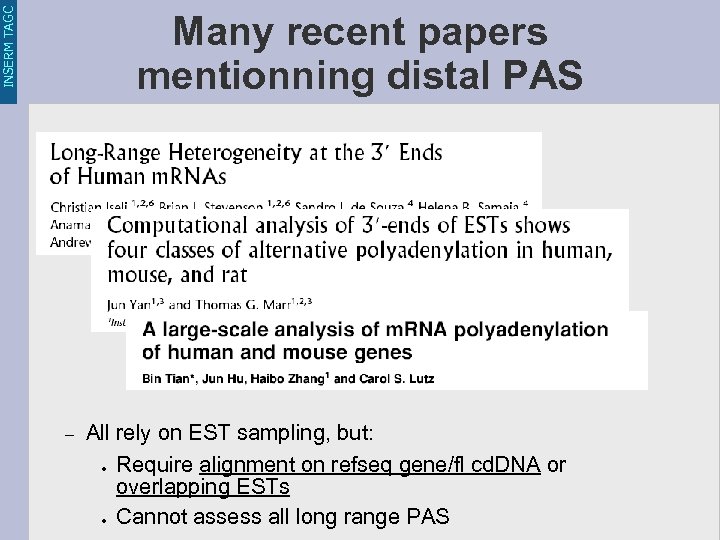 INSERM TAGC Many recent papers mentionning distal PAS – All rely on EST sampling, but: ● Require alignment on refseq gene/fl cd. DNA or overlapping ESTs ● Cannot assess all long range PAS
INSERM TAGC Many recent papers mentionning distal PAS – All rely on EST sampling, but: ● Require alignment on refseq gene/fl cd. DNA or overlapping ESTs ● Cannot assess all long range PAS
 INSERM TAGC How can you make sure a PAS pertains to the nearest 5’ gene ? ● In the absence of overlapping ESTs: danger! – – ● There could be another short gene in the interval PAS could be just noise (remember: 29000 PAS are >10 kb from any Ensembl gene) => We need a gauge to evaluate PAS reality
INSERM TAGC How can you make sure a PAS pertains to the nearest 5’ gene ? ● In the absence of overlapping ESTs: danger! – – ● There could be another short gene in the interval PAS could be just noise (remember: 29000 PAS are >10 kb from any Ensembl gene) => We need a gauge to evaluate PAS reality
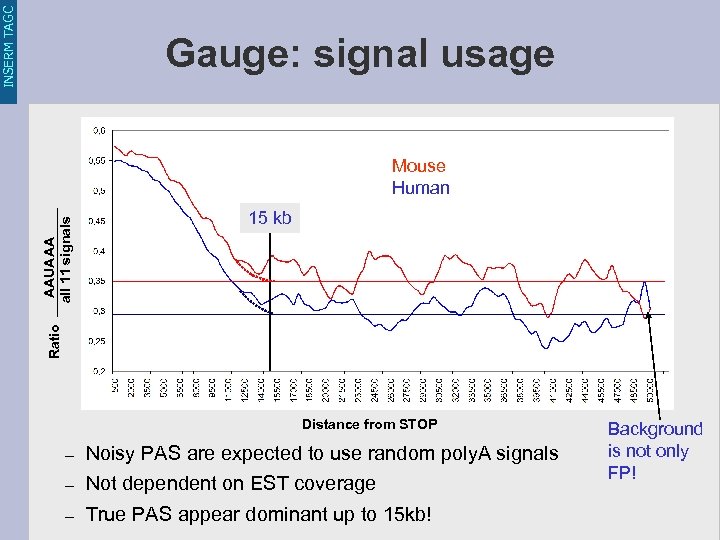 INSERM TAGC Gauge: signal usage 15 kb Ratio AAUAAA all 11 signals Mouse Human Distance from STOP – Noisy PAS are expected to use random poly. A signals – Not dependent on EST coverage – True PAS appear dominant up to 15 kb! Background is not only FP!
INSERM TAGC Gauge: signal usage 15 kb Ratio AAUAAA all 11 signals Mouse Human Distance from STOP – Noisy PAS are expected to use random poly. A signals – Not dependent on EST coverage – True PAS appear dominant up to 15 kb! Background is not only FP!
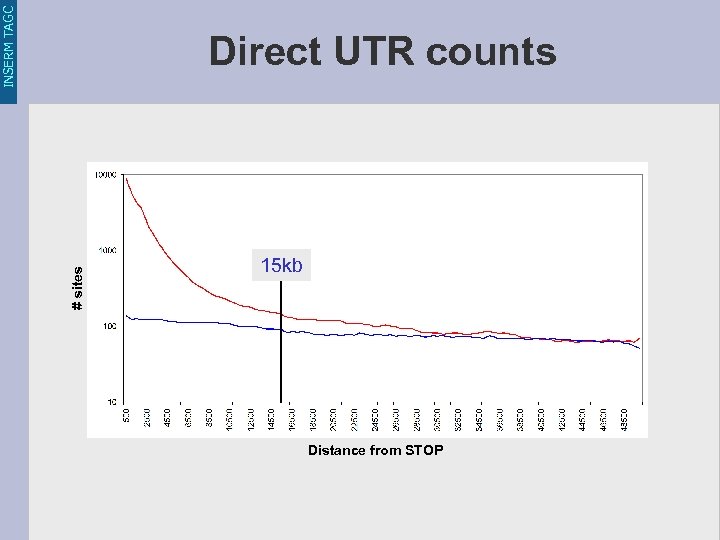 INSERM TAGC # sites Direct UTR counts 15 kb Distance from STOP
INSERM TAGC # sites Direct UTR counts 15 kb Distance from STOP
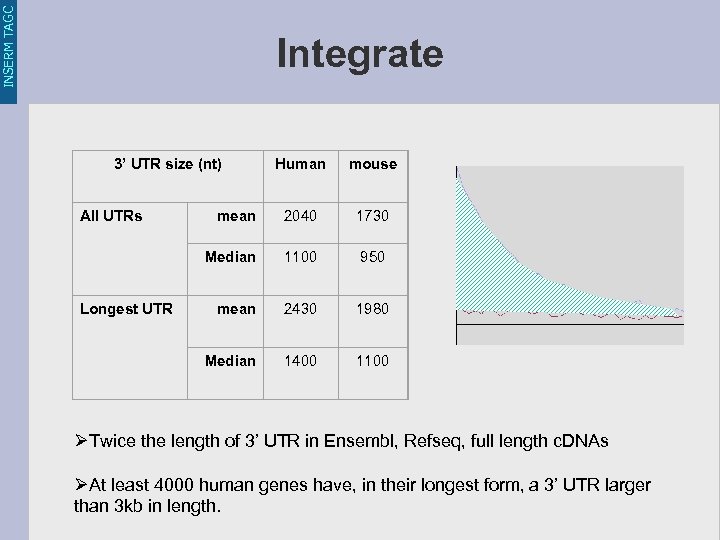 INSERM TAGC Integrate 3’ UTR size (nt) mean 2040 1730 1100 950 mean 2430 1980 Median Longest UTR mouse Median All UTRs Human 1400 1100 ØTwice the length of 3’ UTR in Ensembl, Refseq, full length c. DNAs ØAt least 4000 human genes have, in their longest form, a 3’ UTR larger than 3 kb in length.
INSERM TAGC Integrate 3’ UTR size (nt) mean 2040 1730 1100 950 mean 2430 1980 Median Longest UTR mouse Median All UTRs Human 1400 1100 ØTwice the length of 3’ UTR in Ensembl, Refseq, full length c. DNAs ØAt least 4000 human genes have, in their longest form, a 3’ UTR larger than 3 kb in length.
 INSERM TAGC Intergenic poly. A ● ● ● About 50% of predicted poly. A sites fall in intergenic regions (>15 kb from Stop) Consistent with recent tiling microarray data (Rosetta etc. ) We estimate that 75% of our intergenic poly. A sites are true
INSERM TAGC Intergenic poly. A ● ● ● About 50% of predicted poly. A sites fall in intergenic regions (>15 kb from Stop) Consistent with recent tiling microarray data (Rosetta etc. ) We estimate that 75% of our intergenic poly. A sites are true
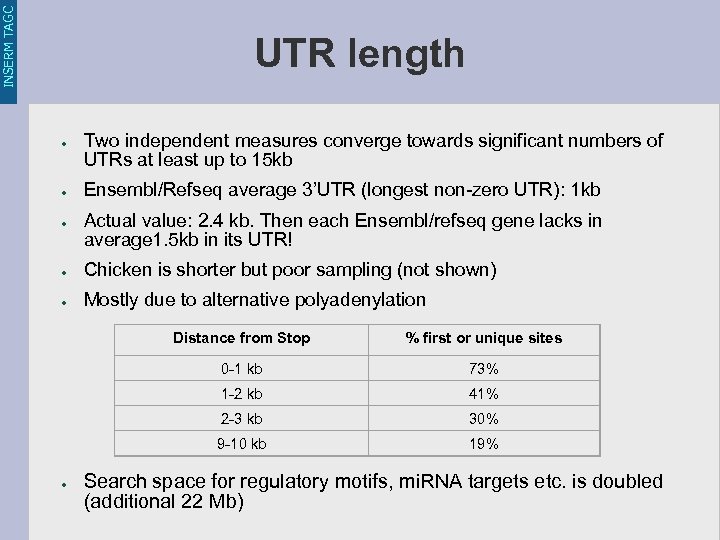 INSERM TAGC UTR length ● ● ● Two independent measures converge towards significant numbers of UTRs at least up to 15 kb Ensembl/Refseq average 3’UTR (longest non-zero UTR): 1 kb Actual value: 2. 4 kb. Then each Ensembl/refseq gene lacks in average 1. 5 kb in its UTR! ● Chicken is shorter but poor sampling (not shown) ● Mostly due to alternative polyadenylation Distance from Stop 0 -1 kb 73% 1 -2 kb 41% 2 -3 kb 30% 9 -10 kb ● % first or unique sites 19% Search space for regulatory motifs, mi. RNA targets etc. is doubled (additional 22 Mb)
INSERM TAGC UTR length ● ● ● Two independent measures converge towards significant numbers of UTRs at least up to 15 kb Ensembl/Refseq average 3’UTR (longest non-zero UTR): 1 kb Actual value: 2. 4 kb. Then each Ensembl/refseq gene lacks in average 1. 5 kb in its UTR! ● Chicken is shorter but poor sampling (not shown) ● Mostly due to alternative polyadenylation Distance from Stop 0 -1 kb 73% 1 -2 kb 41% 2 -3 kb 30% 9 -10 kb ● % first or unique sites 19% Search space for regulatory motifs, mi. RNA targets etc. is doubled (additional 22 Mb)
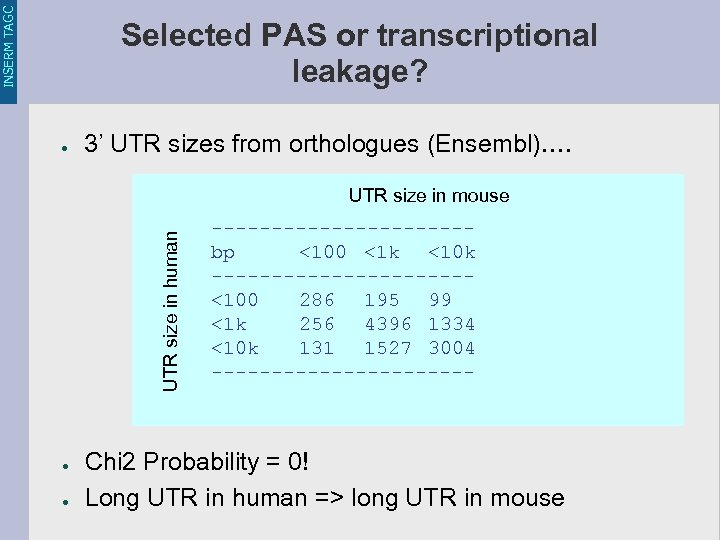 INSERM TAGC Selected PAS or transcriptional leakage? ● 3’ UTR sizes from orthologues (Ensembl)…. UTR size in human UTR size in mouse ● ● ----------- bp <100 <1 k <10 k -----------<100 286 195 99 <1 k 256 4396 1334 <10 k 131 1527 3004 ----------- Chi 2 Probability = 0! Long UTR in human => long UTR in mouse
INSERM TAGC Selected PAS or transcriptional leakage? ● 3’ UTR sizes from orthologues (Ensembl)…. UTR size in human UTR size in mouse ● ● ----------- bp <100 <1 k <10 k -----------<100 286 195 99 <1 k 256 4396 1334 <10 k 131 1527 3004 ----------- Chi 2 Probability = 0! Long UTR in human => long UTR in mouse
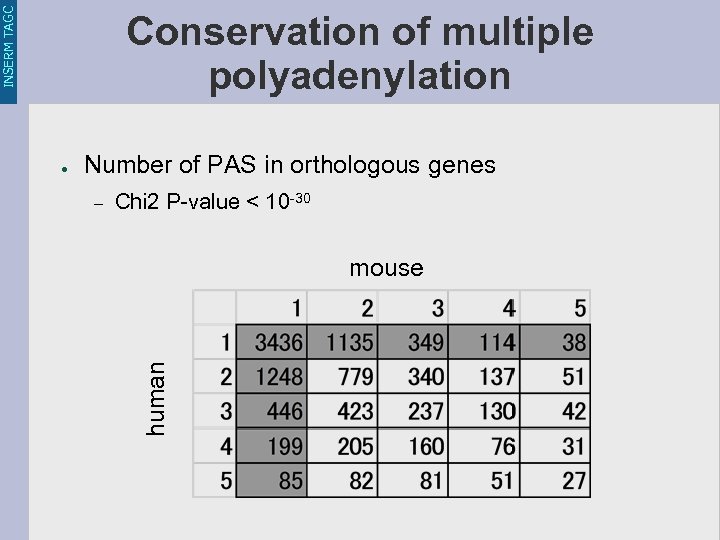 INSERM TAGC Conservation of multiple polyadenylation Number of PAS in orthologous genes – Chi 2 P-value < 10 -30 mouse human ●
INSERM TAGC Conservation of multiple polyadenylation Number of PAS in orthologous genes – Chi 2 P-value < 10 -30 mouse human ●
 INSERM TAGC Conservation & function ● ● Alternative polyadenylation appears to be regulated Increased importance of UTR extension as target for postranscriptional regulation
INSERM TAGC Conservation & function ● ● Alternative polyadenylation appears to be regulated Increased importance of UTR extension as target for postranscriptional regulation
 INSERM TAGC Alternative PAS Conservation across Species Identifying Regulated Alternative PAS (ongoing work)
INSERM TAGC Alternative PAS Conservation across Species Identifying Regulated Alternative PAS (ongoing work)
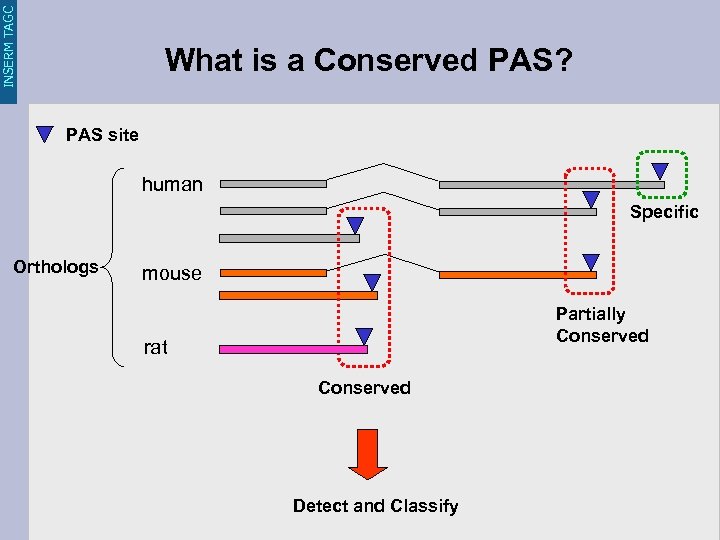 INSERM TAGC What is a Conserved PAS? PAS site human Specific Orthologs mouse Partially Conserved rat Conserved Detect and Classify
INSERM TAGC What is a Conserved PAS? PAS site human Specific Orthologs mouse Partially Conserved rat Conserved Detect and Classify
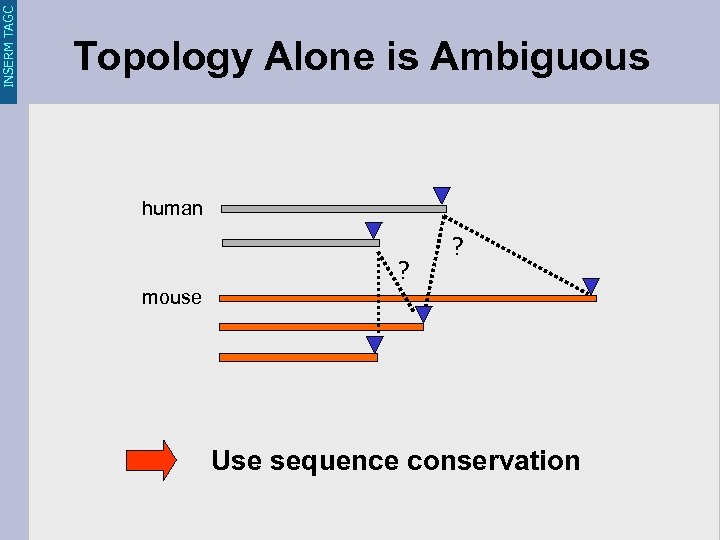 INSERM TAGC Topology Alone is Ambiguous human ? ? mouse Use sequence conservation
INSERM TAGC Topology Alone is Ambiguous human ? ? mouse Use sequence conservation
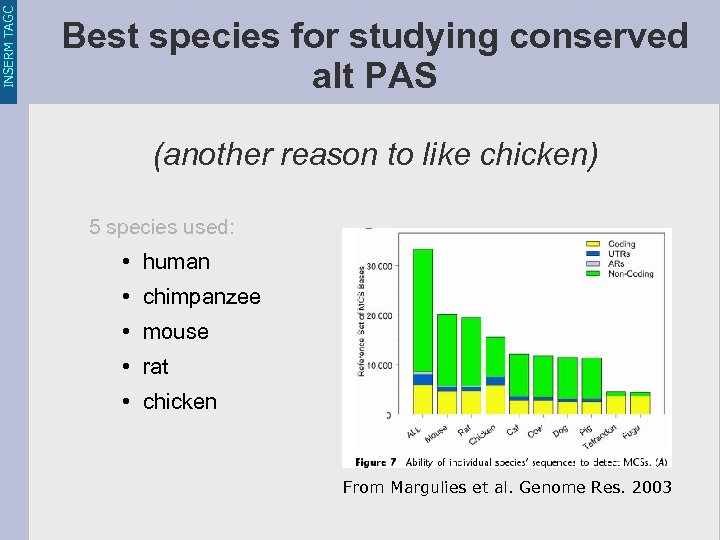 INSERM TAGC Best species for studying conserved alt PAS (another reason to like chicken) 5 species used: • human • chimpanzee • mouse • rat • chicken From Margulies et al. Genome Res. 2003
INSERM TAGC Best species for studying conserved alt PAS (another reason to like chicken) 5 species used: • human • chimpanzee • mouse • rat • chicken From Margulies et al. Genome Res. 2003
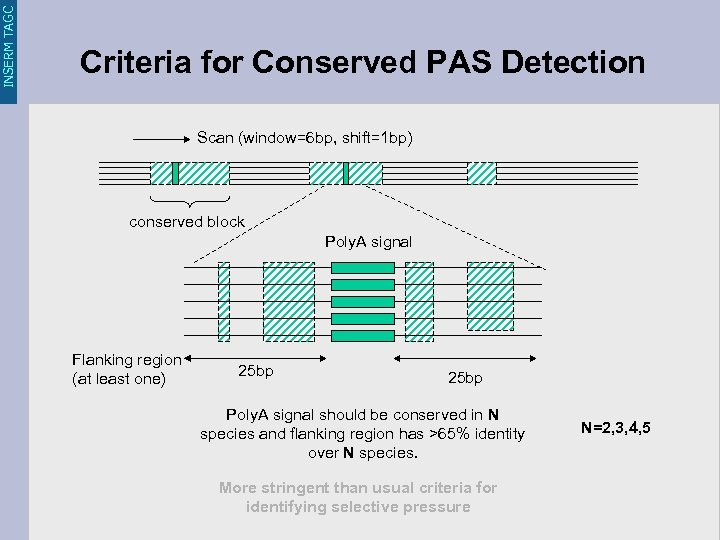 INSERM TAGC Criteria for Conserved PAS Detection Scan (window=6 bp, shift=1 bp) conserved block Poly. A signal Flanking region (at least one) 25 bp Poly. A signal should be conserved in N species and flanking region has >65% identity over N species. More stringent than usual criteria for identifying selective pressure N=2, 3, 4, 5
INSERM TAGC Criteria for Conserved PAS Detection Scan (window=6 bp, shift=1 bp) conserved block Poly. A signal Flanking region (at least one) 25 bp Poly. A signal should be conserved in N species and flanking region has >65% identity over N species. More stringent than usual criteria for identifying selective pressure N=2, 3, 4, 5
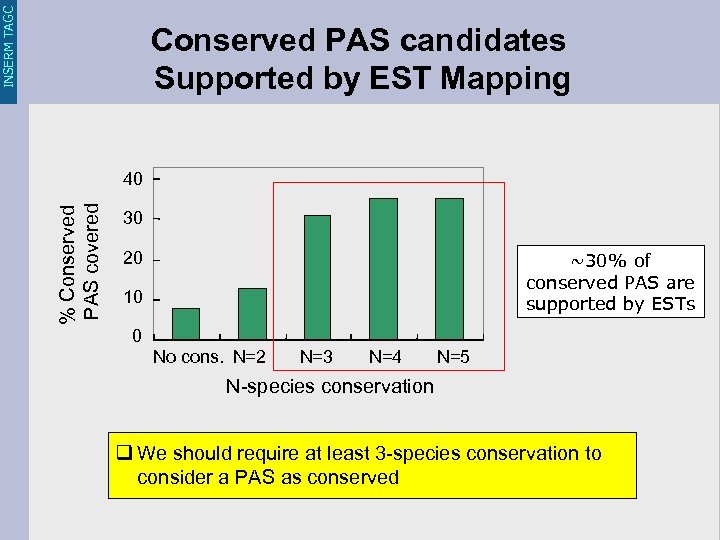 INSERM TAGC Conserved PAS candidates Supported by EST Mapping % Conserved PAS covered 40 30 20 ~30% of conserved PAS are supported by ESTs 10 0 No cons. N=2 N=3 N=4 N=5 N-species conservation q We should require at least 3 -species conservation to consider a PAS as conserved
INSERM TAGC Conserved PAS candidates Supported by EST Mapping % Conserved PAS covered 40 30 20 ~30% of conserved PAS are supported by ESTs 10 0 No cons. N=2 N=3 N=4 N=5 N-species conservation q We should require at least 3 -species conservation to consider a PAS as conserved
 INSERM TAGC A Significant Fraction of Genes has Conserved PAS ● Over 22000 annotated human genes: – – ● 10% have multiple putative CONPAS – ● >20% have a putative CONPAS (at least 3 -species) 7% have a putative CONPAS supported by ESTs Compares to 10 -15% conserved alt-splice variants Suggests selective pressure for many poly. A site sequences in animal genomes
INSERM TAGC A Significant Fraction of Genes has Conserved PAS ● Over 22000 annotated human genes: – – ● 10% have multiple putative CONPAS – ● >20% have a putative CONPAS (at least 3 -species) 7% have a putative CONPAS supported by ESTs Compares to 10 -15% conserved alt-splice variants Suggests selective pressure for many poly. A site sequences in animal genomes
 INSERM TAGC Why should PAS be embedded in conserved sequences? ● Regulatory protein binding site? ● Regulatory RNA structure? ● Antisense or mi. RNA targets? Probably not In some cases Probably most cases
INSERM TAGC Why should PAS be embedded in conserved sequences? ● Regulatory protein binding site? ● Regulatory RNA structure? ● Antisense or mi. RNA targets? Probably not In some cases Probably most cases
 INSERM TAGC Part II Erpin News
INSERM TAGC Part II Erpin News
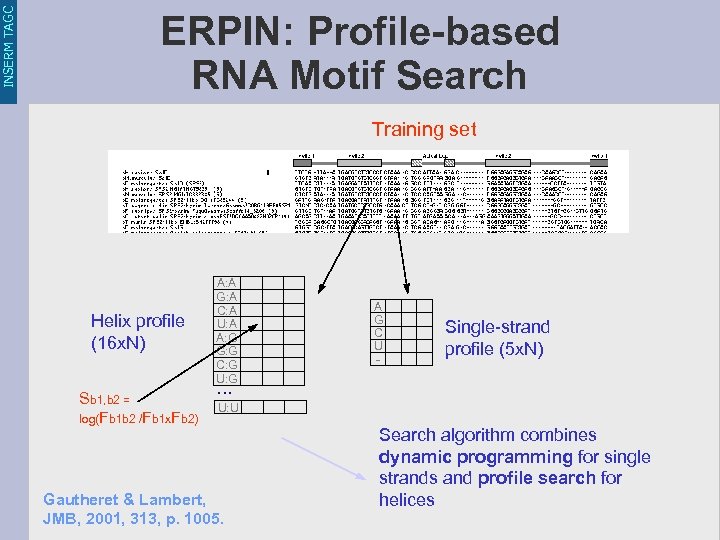 INSERM TAGC ERPIN: Profile-based RNA Motif Search Training set Helix profile (16 x. N) Sb 1, b 2 = log(Fb 1 b 2 /Fb 1 x. Fb 2) A: A G: A C: A U: A A: G G: G C: G U: G A G C U - Single-strand profile (5 x. N) . . . U: U Gautheret & Lambert, JMB, 2001, 313, p. 1005. Search algorithm combines dynamic programming for single strands and profile search for helices
INSERM TAGC ERPIN: Profile-based RNA Motif Search Training set Helix profile (16 x. N) Sb 1, b 2 = log(Fb 1 b 2 /Fb 1 x. Fb 2) A: A G: A C: A U: A A: G G: G C: G U: G A G C U - Single-strand profile (5 x. N) . . . U: U Gautheret & Lambert, JMB, 2001, 313, p. 1005. Search algorithm combines dynamic programming for single strands and profile search for helices
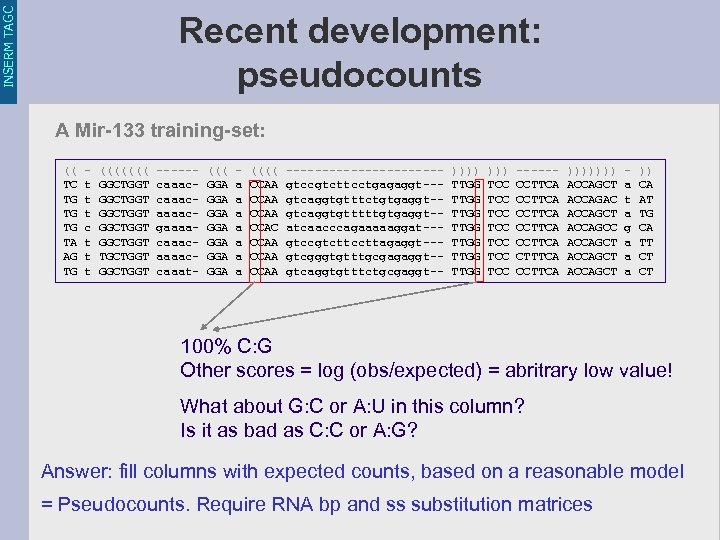 INSERM TAGC Recent development: pseudocounts A Mir-133 training-set: (( - ((((((( ------ (((( ----------- ))))))) - )) TC t GGCTGGT caaac- GGA a CCAA gtccgtcttcctgagaggt--- TTGG TCC CCTTCA ACCAGCT a CA TG t GGCTGGT caaac- GGA a CCAA gtcaggtgtttctgtgaggt-- TTGG TCC CCTTCA ACCAGAC t AT TG t GGCTGGT aaaac- GGA a CCAA gtcaggtgtttttgtgaggt-- TTGG TCC CCTTCA ACCAGCT a TG TG c GGCTGGT gaaaa- GGA a CCAC atcaacccagaaaaaggat--- TTGG TCC CCTTCA ACCAGCC g CA TA t GGCTGGT caaac- GGA a CCAA gtccgtcttccttagaggt--- TTGG TCC CCTTCA ACCAGCT a TT AG t TGCTGGT aaaac- GGA a CCAA gtcgggtgtttgcgagaggt-- TTGG TCC CTTTCA ACCAGCT a CT TG t GGCTGGT caaat- GGA a CCAA gtcaggtgtttctgcgaggt-- TTGG TCC CCTTCA ACCAGCT a CT 100% C: G Other scores = log (obs/expected) = abritrary low value! What about G: C or A: U in this column? Is it as bad as C: C or A: G? Answer: fill columns with expected counts, based on a reasonable model = Pseudocounts. Require RNA bp and ss substitution matrices
INSERM TAGC Recent development: pseudocounts A Mir-133 training-set: (( - ((((((( ------ (((( ----------- ))))))) - )) TC t GGCTGGT caaac- GGA a CCAA gtccgtcttcctgagaggt--- TTGG TCC CCTTCA ACCAGCT a CA TG t GGCTGGT caaac- GGA a CCAA gtcaggtgtttctgtgaggt-- TTGG TCC CCTTCA ACCAGAC t AT TG t GGCTGGT aaaac- GGA a CCAA gtcaggtgtttttgtgaggt-- TTGG TCC CCTTCA ACCAGCT a TG TG c GGCTGGT gaaaa- GGA a CCAC atcaacccagaaaaaggat--- TTGG TCC CCTTCA ACCAGCC g CA TA t GGCTGGT caaac- GGA a CCAA gtccgtcttccttagaggt--- TTGG TCC CCTTCA ACCAGCT a TT AG t TGCTGGT aaaac- GGA a CCAA gtcgggtgtttgcgagaggt-- TTGG TCC CTTTCA ACCAGCT a CT TG t GGCTGGT caaat- GGA a CCAA gtcaggtgtttctgcgaggt-- TTGG TCC CCTTCA ACCAGCT a CT 100% C: G Other scores = log (obs/expected) = abritrary low value! What about G: C or A: U in this column? Is it as bad as C: C or A: G? Answer: fill columns with expected counts, based on a reasonable model = Pseudocounts. Require RNA bp and ss substitution matrices
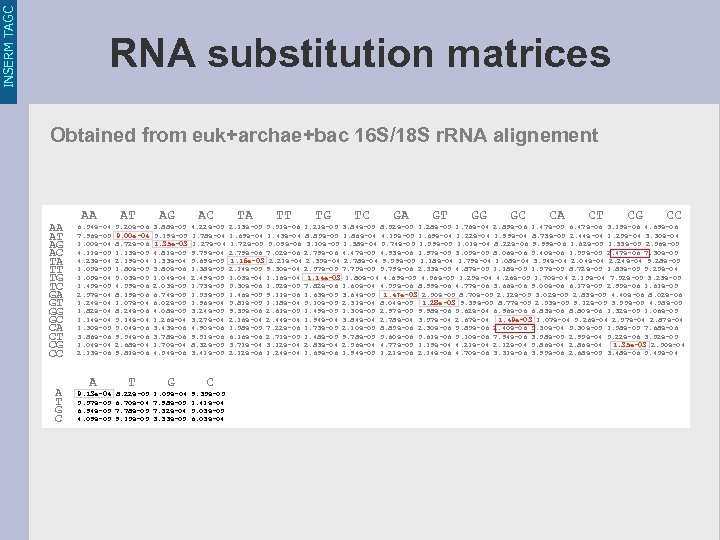 INSERM TAGC RNA substitution matrices Obtained from euk+archae+bac 16 S/18 S r. RNA alignement AA AT AG AC TA TT TG TC GA GT GG GC CA CT CG CC A T G C AA AT AG AC TA TT TG TC GA GT GG GC CA CT CG CC 6. 54 e-04 5. 20 e-06 3. 88 e-05 4. 22 e-05 2. 13 e-05 5. 51 e-06 1. 21 e-05 3. 84 e-05 8. 52 e-05 1. 28 e-05 1. 76 e-04 2. 89 e-06 1. 47 e-05 6. 47 e-06 3. 19 e-06 4. 69 e-06 7. 96 e-05 9. 00 e-04 5. 19 e-05 1. 78 e-04 1. 69 e-04 1. 43 e-04 8. 85 e-05 1. 86 e-04 4. 15 e-05 1. 69 e-04 1. 22 e-04 1. 99 e-04 8. 73 e-05 2. 44 e-04 1. 25 e-04 3. 30 e-04 1. 00 e-04 8. 72 e-06 1. 35 e-03 1. 27 e-04 1. 72 e-05 5. 09 e-06 3. 10 e-05 1. 38 e-04 5. 74 e-05 1. 59 e-05 1. 01 e-04 8. 22 e-06 9. 99 e-06 1. 62 e-05 1. 33 e-05 2. 56 e-05 4. 11 e-05 1. 13 e-05 4. 81 e-05 9. 79 e-04 2. 79 e-06 7. 02 e-06 2. 79 e-06 4. 47 e-05 4. 93 e-06 1. 97 e-05 3. 05 e-05 8. 06 e-06 5. 40 e-06 1. 55 e-05 2. 47 e-06 7. 30 e-05 4. 23 e-04 2. 19 e-04 1. 33 e-04 5. 69 e-05 1. 16 e-03 2. 21 e-04 2. 35 e-04 2. 78 e-04 9. 59 e-05 1. 18 e-04 1. 79 e-04 1. 08 e-04 3. 54 e-04 2. 04 e-04 2. 24 e-04 9. 28 e-05 1. 05 e-05 1. 80 e-05 3. 80 e-06 1. 38 e-05 2. 14 e-05 9. 30 e-04 2. 57 e-05 7. 75 e-05 5. 79 e-06 2. 33 e-05 4. 87 e-05 1. 18 e-05 1. 57 e-05 8. 72 e-05 1. 83 e-05 5. 25 e-04 1. 05 e-04 5. 03 e-05 1. 04 e-04 2. 49 e-05 1. 03 e-04 1. 16 e-04 1. 14 e-03 1. 80 e-04 4. 69 e-05 4. 56 e-05 1. 25 e-04 4. 26 e-05 1. 70 e-04 2. 15 e-04 7. 52 e-05 3. 23 e-05 1. 45 e-05 4. 59 e-06 2. 03 e-05 1. 73 e-05 5. 30 e-06 1. 52 e-05 7. 82 e-06 1. 60 e-04 4. 55 e-06 8. 99 e-06 4. 77 e-06 3. 66 e-06 9. 00 e-06 6. 17 e-05 2. 95 e-06 1. 61 e-05 2. 57 e-04 8. 19 e-06 6. 74 e-05 1. 53 e-05 1. 46 e-05 9. 11 e-06 1. 63 e-05 3. 64 e-05 1. 47 e-03 2. 50 e-05 8. 70 e-05 2. 12 e-05 3. 02 e-05 2. 83 e-05 4. 40 e-06 8. 02 e-06 1. 24 e-04 1. 07 e-04 6. 02 e-05 1. 96 e-04 5. 81 e-05 1. 18 e-04 5. 10 e-05 2. 31 e-04 8. 04 e-05 1. 28 e-03 9. 39 e-05 8. 77 e-05 2. 53 e-05 9. 12 e-05 3. 55 e-05 4. 58 e-05 1. 82 e-04 8. 24 e-06 4. 08 e-05 3. 24 e-05 9. 35 e-06 2. 61 e-05 1. 49 e-05 1. 30 e-05 2. 97 e-05 9. 98 e-06 5. 62 e-04 6. 96 e-06 6. 83 e-06 8. 80 e-06 1. 32 e-05 1. 06 e-05 1. 14 e-04 5. 14 e-04 1. 26 e-04 3. 27 e-04 2. 16 e-04 2. 44 e-04 1. 94 e-04 3. 84 e-04 2. 78 e-04 3. 57 e-04 2. 67 e-04 1. 49 e-03 1. 07 e-04 5. 26 e-04 2. 57 e-04 2. 87 e-04 1. 30 e-05 5. 04 e-06 3. 43 e-06 4. 90 e-06 1. 58 e-05 7. 22 e-06 1. 73 e-05 2. 10 e-05 8. 85 e-06 2. 30 e-06 5. 85 e-06 2. 40 e-06 5. 30 e-04 5. 30 e-05 1. 58 e-05 7. 68 e-06 3. 86 e-06 9. 54 e-06 3. 78 e-06 9. 51 e-06 6. 16 e-06 2. 71 e-05 1. 48 e-05 9. 78 e-05 5. 60 e-06 5. 61 e-06 5. 10 e-06 7. 94 e-06 3. 58 e-05 2. 95 e-04 5. 22 e-06 3. 52 e-05 1. 04 e-04 2. 68 e-04 1. 70 e-04 8. 32 e-05 3. 71 e-04 3. 12 e-04 2. 83 e-04 2. 56 e-04 4. 77 e-05 1. 19 e-04 4. 21 e-04 2. 12 e-04 5. 86 e-04 2. 86 e-04 1. 35 e-03 2. 50 e-04 2. 13 e-06 9. 81 e-06 4. 54 e-06 3. 41 e-05 2. 12 e-06 1. 24 e-04 1. 69 e-06 1. 94 e-05 1. 21 e-06 2. 14 e-06 4. 70 e-06 3. 31 e-06 3. 95 e-06 2. 68 e-05 3. 48 e-06 5. 45 e-04 A T G C 9. 13 e-04 8. 22 e-05 1. 05 e-04 9. 35 e-05 5. 57 e-05 6. 70 e-04 7. 98 e-05 1. 41 e-04 6. 94 e-05 7. 78 e-05 7. 32 e-04 5. 03 e-05 4. 09 e-05 9. 15 e-05 3. 33 e-05 6. 03 e-04
INSERM TAGC RNA substitution matrices Obtained from euk+archae+bac 16 S/18 S r. RNA alignement AA AT AG AC TA TT TG TC GA GT GG GC CA CT CG CC A T G C AA AT AG AC TA TT TG TC GA GT GG GC CA CT CG CC 6. 54 e-04 5. 20 e-06 3. 88 e-05 4. 22 e-05 2. 13 e-05 5. 51 e-06 1. 21 e-05 3. 84 e-05 8. 52 e-05 1. 28 e-05 1. 76 e-04 2. 89 e-06 1. 47 e-05 6. 47 e-06 3. 19 e-06 4. 69 e-06 7. 96 e-05 9. 00 e-04 5. 19 e-05 1. 78 e-04 1. 69 e-04 1. 43 e-04 8. 85 e-05 1. 86 e-04 4. 15 e-05 1. 69 e-04 1. 22 e-04 1. 99 e-04 8. 73 e-05 2. 44 e-04 1. 25 e-04 3. 30 e-04 1. 00 e-04 8. 72 e-06 1. 35 e-03 1. 27 e-04 1. 72 e-05 5. 09 e-06 3. 10 e-05 1. 38 e-04 5. 74 e-05 1. 59 e-05 1. 01 e-04 8. 22 e-06 9. 99 e-06 1. 62 e-05 1. 33 e-05 2. 56 e-05 4. 11 e-05 1. 13 e-05 4. 81 e-05 9. 79 e-04 2. 79 e-06 7. 02 e-06 2. 79 e-06 4. 47 e-05 4. 93 e-06 1. 97 e-05 3. 05 e-05 8. 06 e-06 5. 40 e-06 1. 55 e-05 2. 47 e-06 7. 30 e-05 4. 23 e-04 2. 19 e-04 1. 33 e-04 5. 69 e-05 1. 16 e-03 2. 21 e-04 2. 35 e-04 2. 78 e-04 9. 59 e-05 1. 18 e-04 1. 79 e-04 1. 08 e-04 3. 54 e-04 2. 04 e-04 2. 24 e-04 9. 28 e-05 1. 05 e-05 1. 80 e-05 3. 80 e-06 1. 38 e-05 2. 14 e-05 9. 30 e-04 2. 57 e-05 7. 75 e-05 5. 79 e-06 2. 33 e-05 4. 87 e-05 1. 18 e-05 1. 57 e-05 8. 72 e-05 1. 83 e-05 5. 25 e-04 1. 05 e-04 5. 03 e-05 1. 04 e-04 2. 49 e-05 1. 03 e-04 1. 16 e-04 1. 14 e-03 1. 80 e-04 4. 69 e-05 4. 56 e-05 1. 25 e-04 4. 26 e-05 1. 70 e-04 2. 15 e-04 7. 52 e-05 3. 23 e-05 1. 45 e-05 4. 59 e-06 2. 03 e-05 1. 73 e-05 5. 30 e-06 1. 52 e-05 7. 82 e-06 1. 60 e-04 4. 55 e-06 8. 99 e-06 4. 77 e-06 3. 66 e-06 9. 00 e-06 6. 17 e-05 2. 95 e-06 1. 61 e-05 2. 57 e-04 8. 19 e-06 6. 74 e-05 1. 53 e-05 1. 46 e-05 9. 11 e-06 1. 63 e-05 3. 64 e-05 1. 47 e-03 2. 50 e-05 8. 70 e-05 2. 12 e-05 3. 02 e-05 2. 83 e-05 4. 40 e-06 8. 02 e-06 1. 24 e-04 1. 07 e-04 6. 02 e-05 1. 96 e-04 5. 81 e-05 1. 18 e-04 5. 10 e-05 2. 31 e-04 8. 04 e-05 1. 28 e-03 9. 39 e-05 8. 77 e-05 2. 53 e-05 9. 12 e-05 3. 55 e-05 4. 58 e-05 1. 82 e-04 8. 24 e-06 4. 08 e-05 3. 24 e-05 9. 35 e-06 2. 61 e-05 1. 49 e-05 1. 30 e-05 2. 97 e-05 9. 98 e-06 5. 62 e-04 6. 96 e-06 6. 83 e-06 8. 80 e-06 1. 32 e-05 1. 06 e-05 1. 14 e-04 5. 14 e-04 1. 26 e-04 3. 27 e-04 2. 16 e-04 2. 44 e-04 1. 94 e-04 3. 84 e-04 2. 78 e-04 3. 57 e-04 2. 67 e-04 1. 49 e-03 1. 07 e-04 5. 26 e-04 2. 57 e-04 2. 87 e-04 1. 30 e-05 5. 04 e-06 3. 43 e-06 4. 90 e-06 1. 58 e-05 7. 22 e-06 1. 73 e-05 2. 10 e-05 8. 85 e-06 2. 30 e-06 5. 85 e-06 2. 40 e-06 5. 30 e-04 5. 30 e-05 1. 58 e-05 7. 68 e-06 3. 86 e-06 9. 54 e-06 3. 78 e-06 9. 51 e-06 6. 16 e-06 2. 71 e-05 1. 48 e-05 9. 78 e-05 5. 60 e-06 5. 61 e-06 5. 10 e-06 7. 94 e-06 3. 58 e-05 2. 95 e-04 5. 22 e-06 3. 52 e-05 1. 04 e-04 2. 68 e-04 1. 70 e-04 8. 32 e-05 3. 71 e-04 3. 12 e-04 2. 83 e-04 2. 56 e-04 4. 77 e-05 1. 19 e-04 4. 21 e-04 2. 12 e-04 5. 86 e-04 2. 86 e-04 1. 35 e-03 2. 50 e-04 2. 13 e-06 9. 81 e-06 4. 54 e-06 3. 41 e-05 2. 12 e-06 1. 24 e-04 1. 69 e-06 1. 94 e-05 1. 21 e-06 2. 14 e-06 4. 70 e-06 3. 31 e-06 3. 95 e-06 2. 68 e-05 3. 48 e-06 5. 45 e-04 A T G C 9. 13 e-04 8. 22 e-05 1. 05 e-04 9. 35 e-05 5. 57 e-05 6. 70 e-04 7. 98 e-05 1. 41 e-04 6. 94 e-05 7. 78 e-05 7. 32 e-04 5. 03 e-05 4. 09 e-05 9. 15 e-05 3. 33 e-05 6. 03 e-04
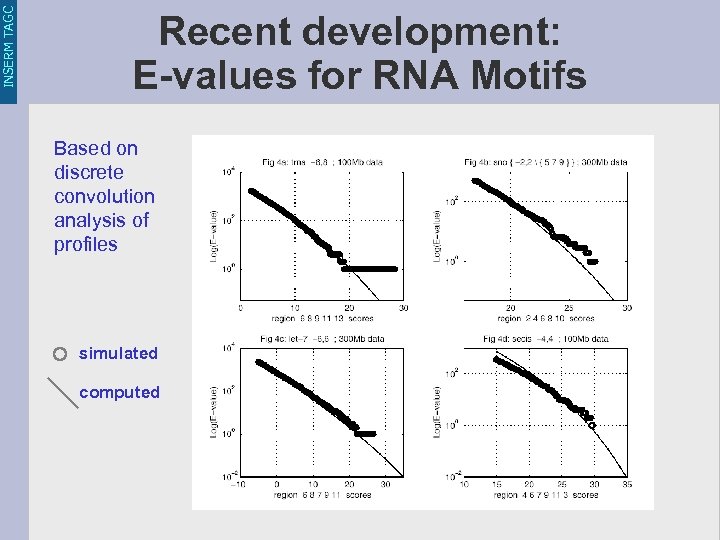 INSERM TAGC Recent development: E-values for RNA Motifs Based on discrete convolution analysis of profiles simulated computed
INSERM TAGC Recent development: E-values for RNA Motifs Based on discrete convolution analysis of profiles simulated computed
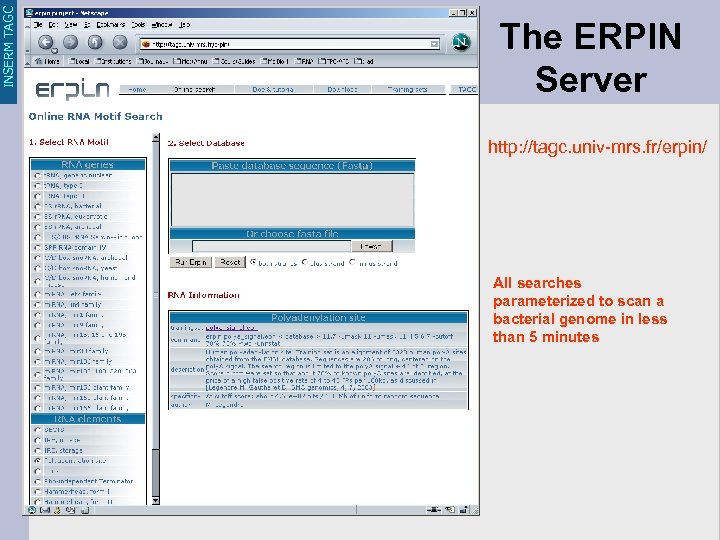 INSERM TAGC The ERPIN Server http: //tagc. univ-mrs. fr/erpin/ All searches parameterized to scan a bacterial genome in less than 5 minutes
INSERM TAGC The ERPIN Server http: //tagc. univ-mrs. fr/erpin/ All searches parameterized to scan a bacterial genome in less than 5 minutes
 INSERM TAGC Micro-RNA Search ● 18 training sets build for 18 mi. RNA families – Using CLUSTALW + Alifold – 10 sequences in average Legendre, Lambert, Gautheret, Bioinformatics 2004
INSERM TAGC Micro-RNA Search ● 18 training sets build for 18 mi. RNA families – Using CLUSTALW + Alifold – 10 sequences in average Legendre, Lambert, Gautheret, Bioinformatics 2004
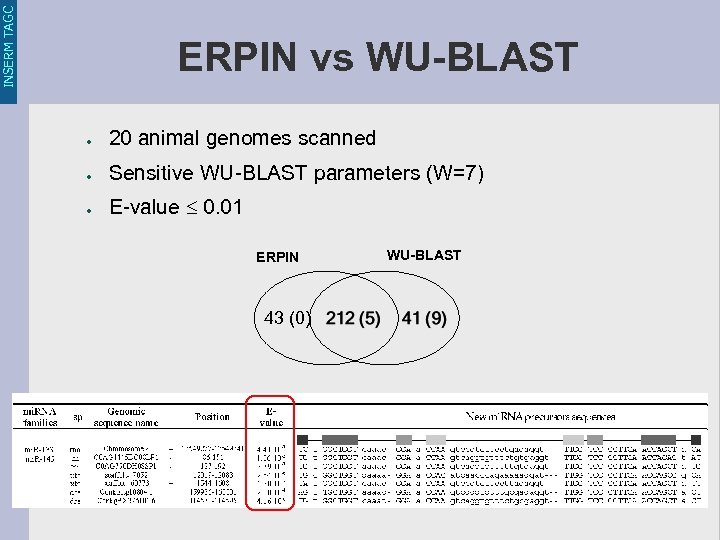 INSERM TAGC ERPIN vs WU-BLAST ● 20 animal genomes scanned ● Sensitive WU-BLAST parameters (W=7) ● E-value 0. 01 ERPIN 43 (0) WU-BLAST
INSERM TAGC ERPIN vs WU-BLAST ● 20 animal genomes scanned ● Sensitive WU-BLAST parameters (W=7) ● E-value 0. 01 ERPIN 43 (0) WU-BLAST
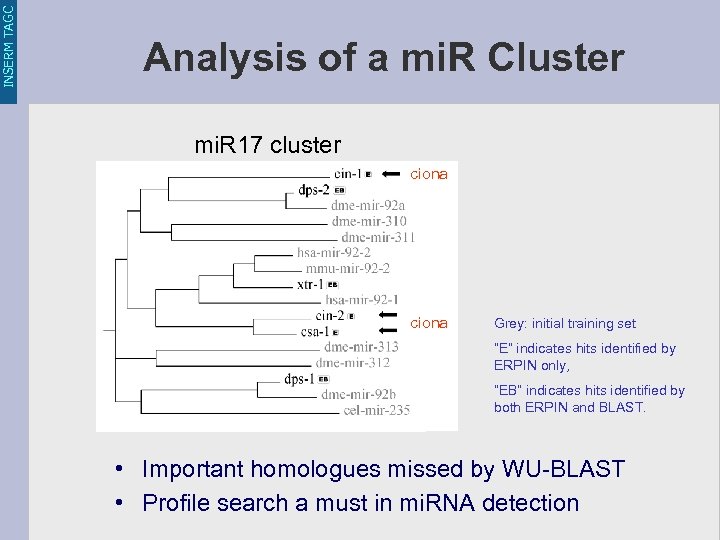 INSERM TAGC Analysis of a mi. R Cluster mi. R 17 cluster ciona Grey: initial training set “E” indicates hits identified by ERPIN only, “EB” indicates hits identified by both ERPIN and BLAST. • Important homologues missed by WU-BLAST • Profile search a must in mi. RNA detection
INSERM TAGC Analysis of a mi. R Cluster mi. R 17 cluster ciona Grey: initial training set “E” indicates hits identified by ERPIN only, “EB” indicates hits identified by both ERPIN and BLAST. • Important homologues missed by WU-BLAST • Profile search a must in mi. RNA detection
 INSERM TAGC lab RNA Bioinformatics • Takeshi Ara • Fabrice Lopez • Matthieu Legendre • William Ritchie • Daniel Gautheret
INSERM TAGC lab RNA Bioinformatics • Takeshi Ara • Fabrice Lopez • Matthieu Legendre • William Ritchie • Daniel Gautheret


