feeef08274202ed07688f776257594bb.ppt
- Количество слайдов: 29

Identification of Protein-Protein Interactions by the Yeast 2 -Hybrid System Alliance for Cell Signaling Myriad Genetics Inc.
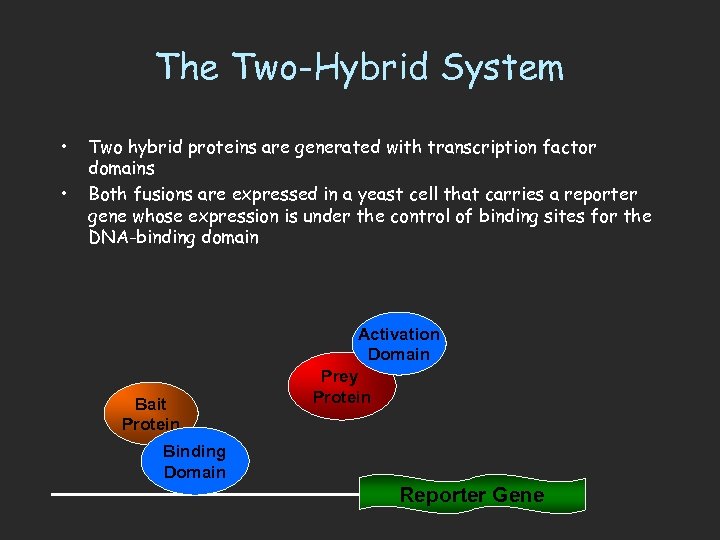
The Two-Hybrid System • • Two hybrid proteins are generated with transcription factor domains Both fusions are expressed in a yeast cell that carries a reporter gene whose expression is under the control of binding sites for the DNA-binding domain Bait Protein Activation Domain Prey Protein Binding Domain Reporter Gene
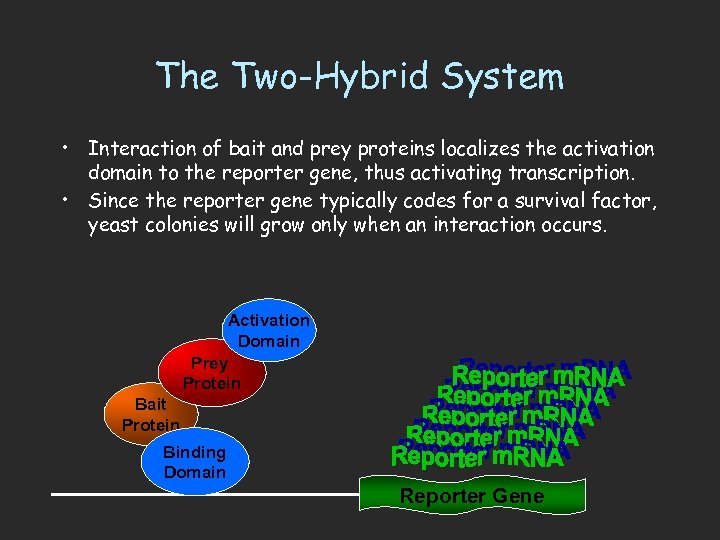
The Two-Hybrid System • Interaction of bait and prey proteins localizes the activation domain to the reporter gene, thus activating transcription. • Since the reporter gene typically codes for a survival factor, yeast colonies will grow only when an interaction occurs. Activation Domain Prey Protein Bait Protein Binding Domain Reporter Gene

Myriad’s Pro. Net Two-Hybrid Process • Industrial-scale application of the yeast two-hybrid system • Roboticized bait creation process • Custom activation domain libraries • Efficient mating strategy • Automated for quality and throughput – Positive sample tracking – Robots for media preparation, liquid handling, colony picking – Statistical quality control
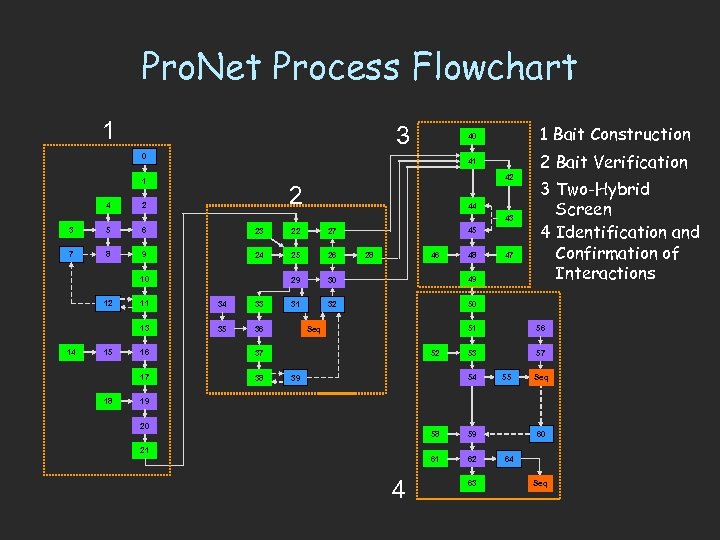
Pro. Net Process Flowchart 1 3 1 Bait Construction 40 0 41 1 42 2 4 2 3 5 6 23 22 27 7 8 9 24 25 26 29 30 49 31 32 2 Bait Verification 3 Two-Hybrid Screen 4 Identification and Confirmation of Interactions 50 44 43 10 12 15 34 33 13 14 11 35 36 38 46 28 48 47 51 Seq 37 17 18 16 45 52 56 53 57 54 39 55 Seq 19 20 58 61 21 4 62 60 59 63 64 Seq
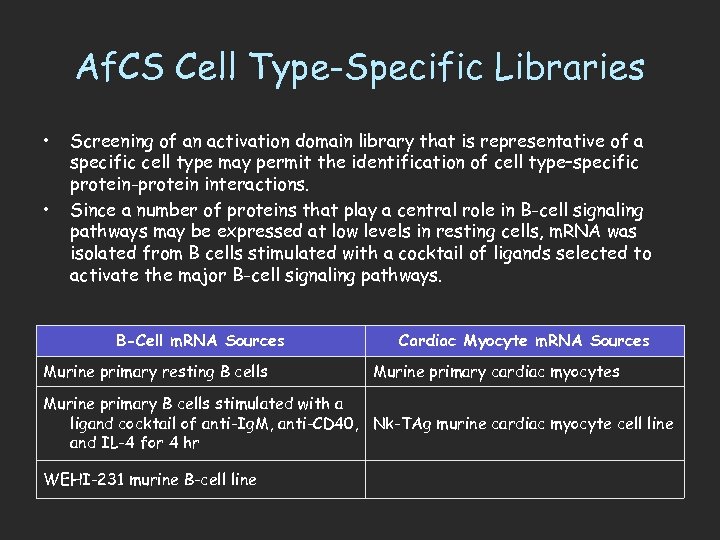
Af. CS Cell Type-Specific Libraries • • Screening of an activation domain library that is representative of a specific cell type may permit the identification of cell type–specific protein-protein interactions. Since a number of proteins that play a central role in B-cell signaling pathways may be expressed at low levels in resting cells, m. RNA was isolated from B cells stimulated with a cocktail of ligands selected to activate the major B-cell signaling pathways. B-Cell m. RNA Sources Murine primary resting B cells Cardiac Myocyte m. RNA Sources Murine primary cardiac myocytes Murine primary B cells stimulated with a ligand cocktail of anti-Ig. M, anti-CD 40, Nk-TAg murine cardiac myocyte cell line and IL-4 for 4 hr WEHI-231 murine B-cell line

Bait Selection • A number of criteria were taken into consideration in the selection of bait protein candidates: – Proteins known to be expressed in B cells and/or cardiac myocytes. – Proteins known or suspected to be involved in signaling pathways in B cells and/or cardiac myocytes. – Proteins believed important in GPCR or BCR signaling leading to PIP 3 generation. These were selected first in an attempt to define interactions within a focused network of signaling molecules. • Preference was also given to proteins that had a higher likelihood of success in the yeast two-hybrid assay – proteins with well-defined modular domains exhibiting secondary structure suggestive of involvement in protein-protein interactions.
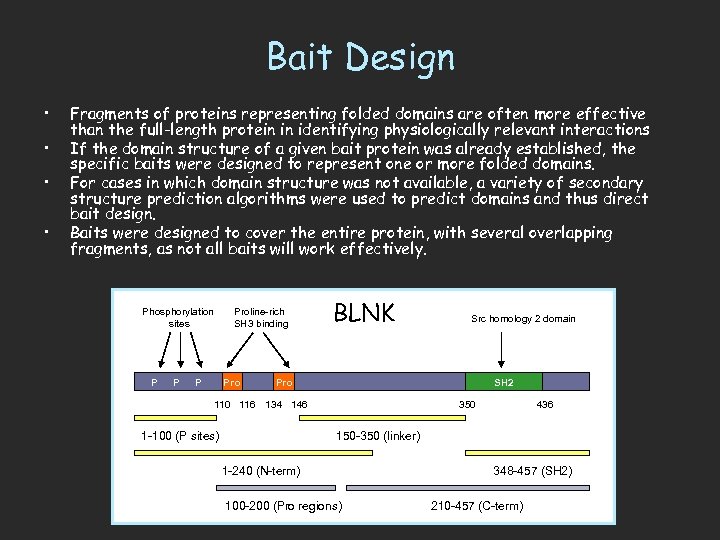
Bait Design • • Fragments of proteins representing folded domains are often more effective than the full-length protein in identifying physiologically relevant interactions If the domain structure of a given bait protein was already established, the specific baits were designed to represent one or more folded domains. For cases in which domain structure was not available, a variety of secondary structure prediction algorithms were used to predict domains and thus direct bait design. Baits were designed to cover the entire protein, with several overlapping fragments, as not all baits will work effectively. Phosphorylation sites P Proline-rich SH 3 binding Pro BLNK Src homology 2 domain Pro SH 2 110 116 134 146 1 -100 (P sites) 350 436 150 -350 (linker) 1 -240 (N-term) 100 -200 (Pro regions) 348 -457 (SH 2) 210 -457 (C-term)
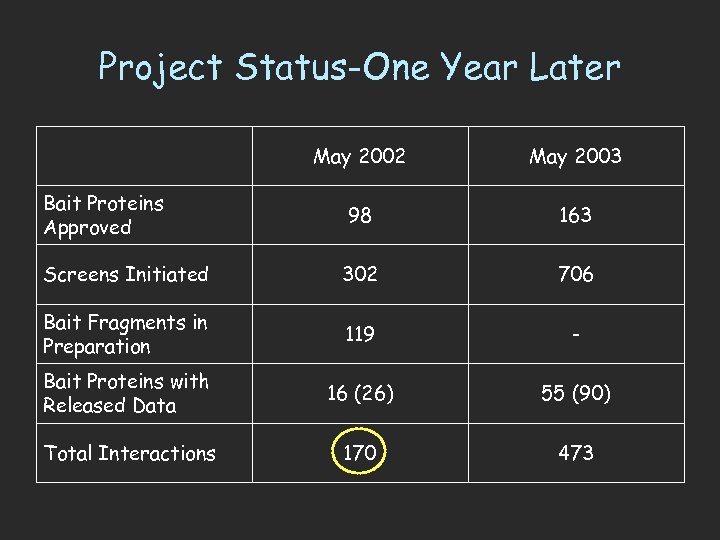
Project Status-One Year Later May 2002 May 2003 98 163 Screens Initiated 302 706 Bait Fragments in Preparation 119 - Bait Proteins with Released Data 16 (26) 55 (90) Total Interactions 170 473 Bait Proteins Approved
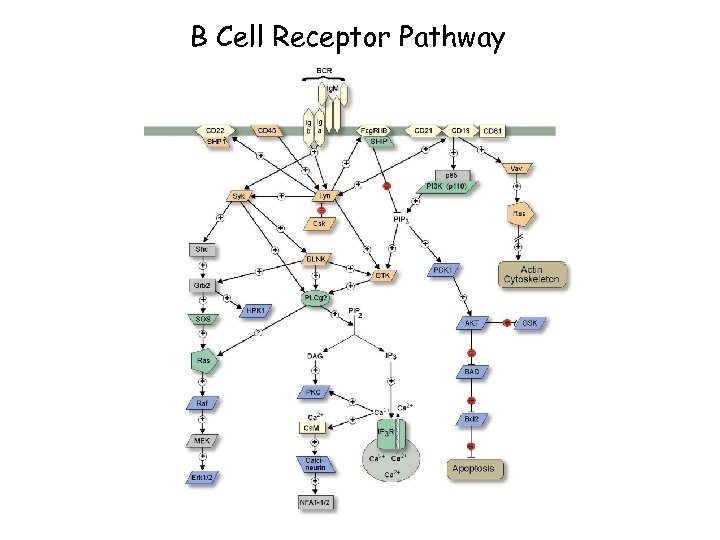
B Cell Receptor Pathway
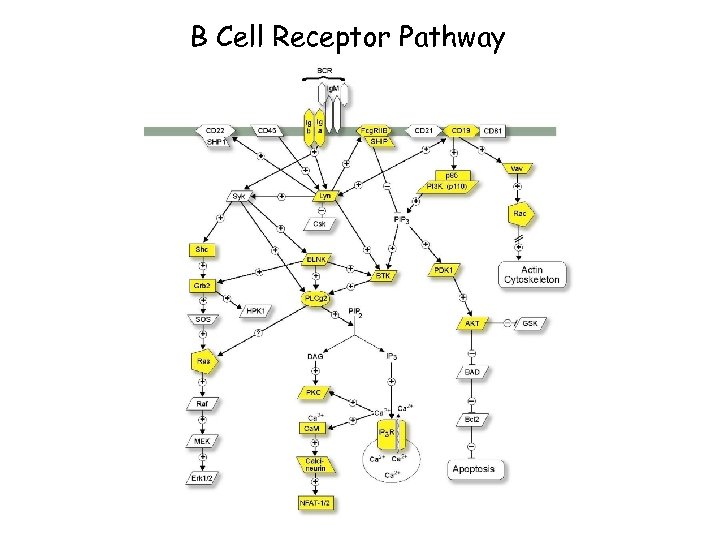
B Cell Receptor Pathway
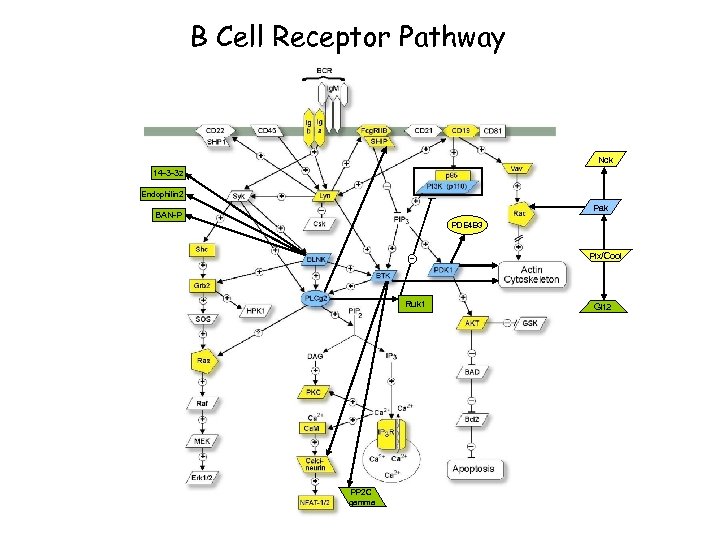
B Cell Receptor Pathway Nck 14 -3 -3 z Endophilin 2 Pak BAN-P PDE 4 B 3 Ruk 1 PP 2 C gamma Pix/Cool Git 2
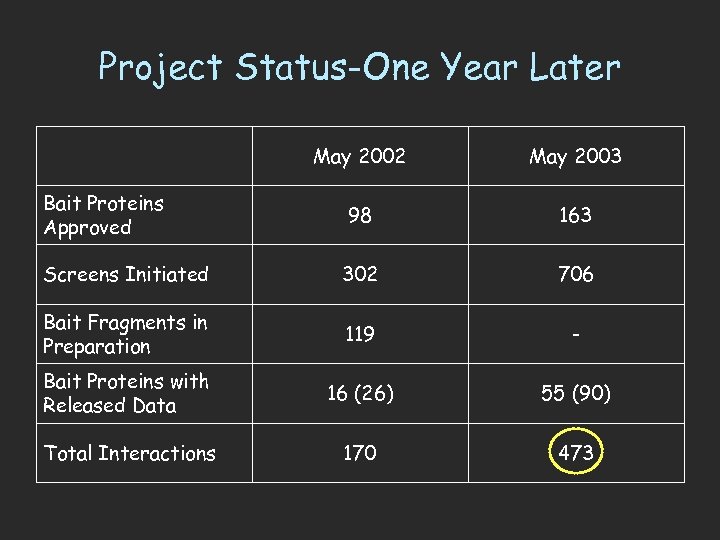
Project Status-One Year Later May 2002 May 2003 98 163 Screens Initiated 302 706 Bait Fragments in Preparation 119 - Bait Proteins with Released Data 16 (26) 55 (90) Total Interactions 170 473 Bait Proteins Approved
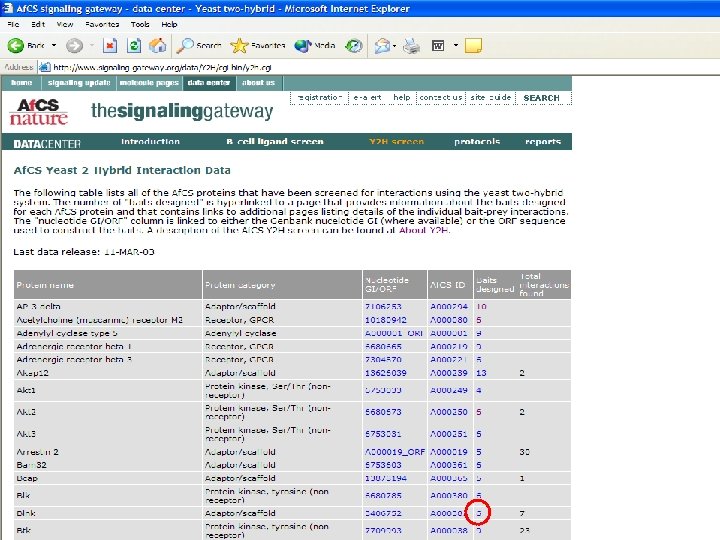
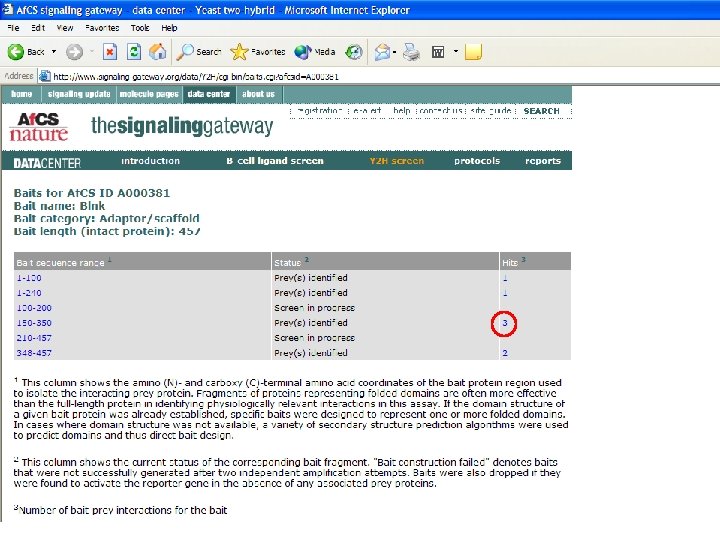

What can we learn from the larger data set? • Last year, 170 interactions derived from just 16 bait proteins – Limited interconnectivity • We now have 473 interactions from 55 bait proteins • How can the average biologist analyze this data set? – With difficulty! • All 473 interactions uploaded to Access database table • Query: which prey IDs are correlated with more than one bait ID?
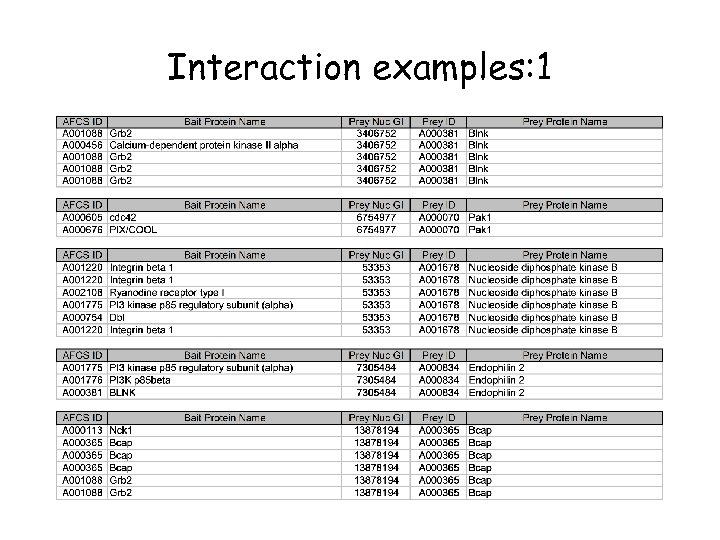
Interaction examples: 1
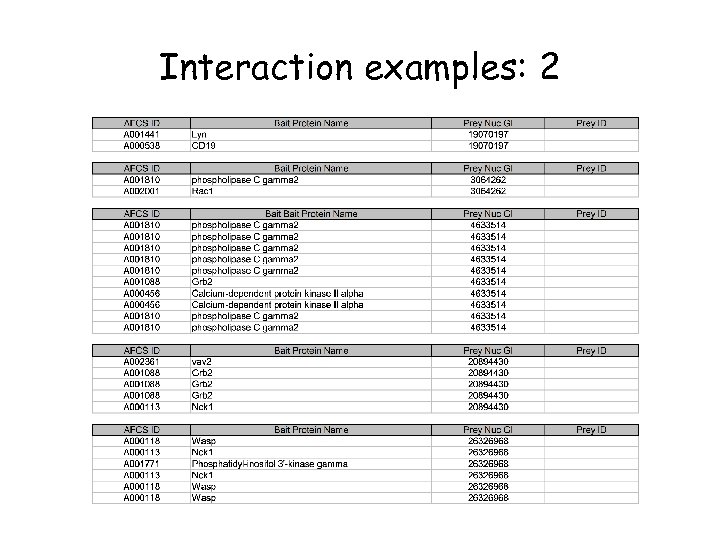
Interaction examples: 2
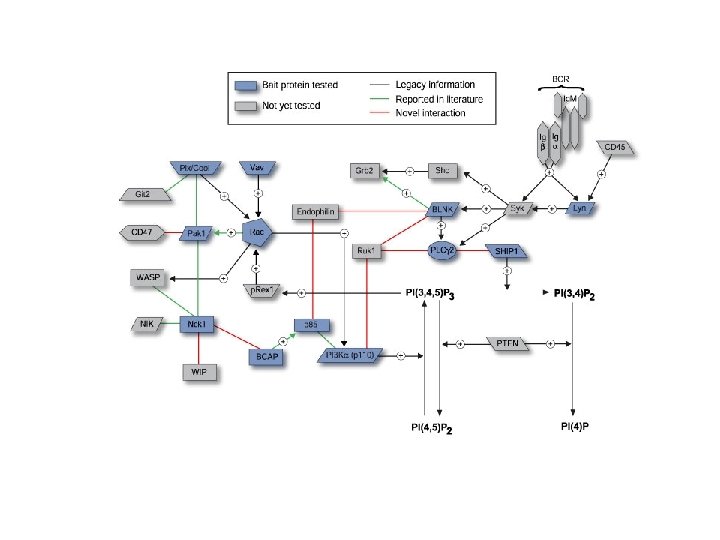
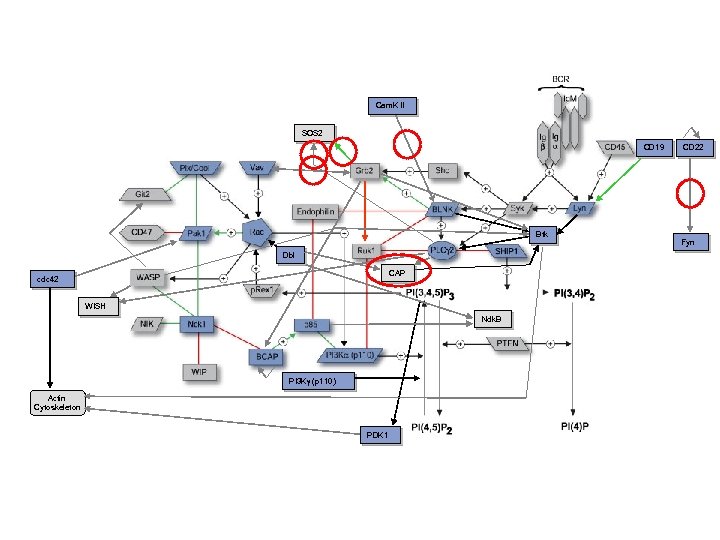
Cam. K II SOS 2 CD 19 Btk Dbl CAP cdc 42 WISH Ndk. B PI 3 Kg (p 110) Actin Cytoskeleton PDK 1 CD 22 Fyn
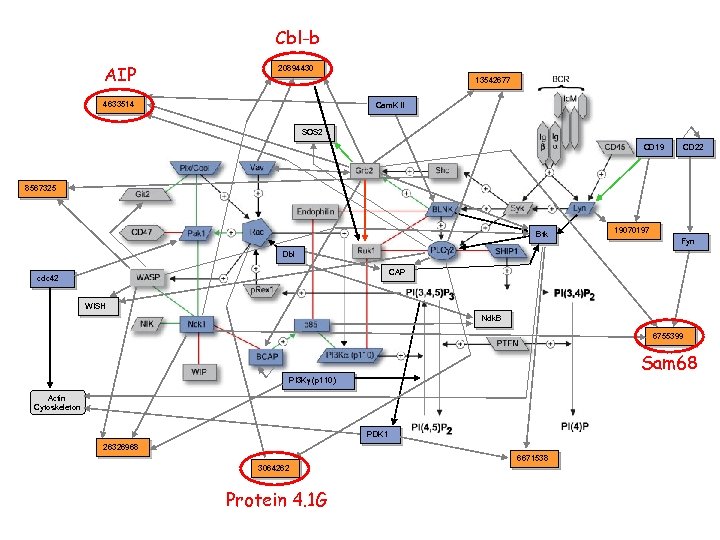
Cbl-b AIP 20894430 13542677 4633514 Cam. K II SOS 2 CD 19 CD 22 8567325 Btk 19070197 Fyn Dbl CAP cdc 42 WISH Ndk. B 6755399 Sam 68 PI 3 Kg (p 110) Actin Cytoskeleton PDK 1 26326968 6671538 3064262 Protein 4. 1 G

Caveats! • Y 2 H data requires validation by secondary assay – Protein complementation – Pull-downs • Didn’t observe source library in data analysis • Didn’t analyze all bait and prey coordinates to map sites of interaction – Cant assume multi-protein complexes since proteins may be competing for same site of interaction – This level of analysis requires more sophisticated computational approach
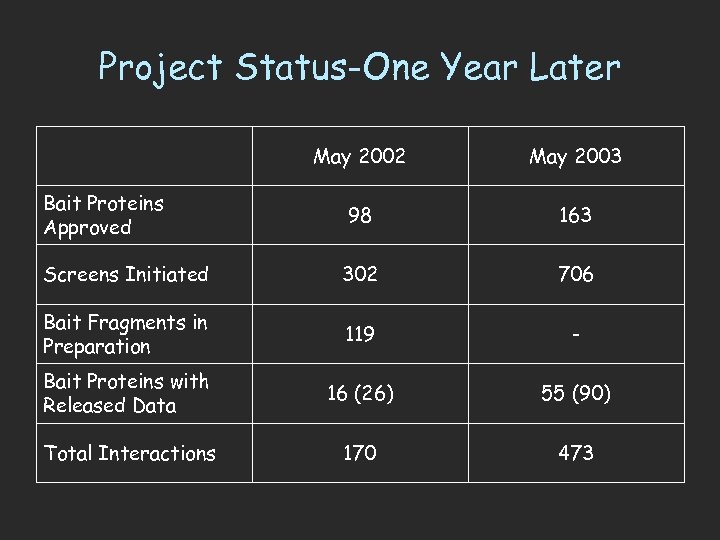
Project Status-One Year Later May 2002 May 2003 98 163 Screens Initiated 302 706 Bait Fragments in Preparation 119 - Bait Proteins with Released Data 16 (26) 55 (90) Total Interactions 170 473 Bait Proteins Approved
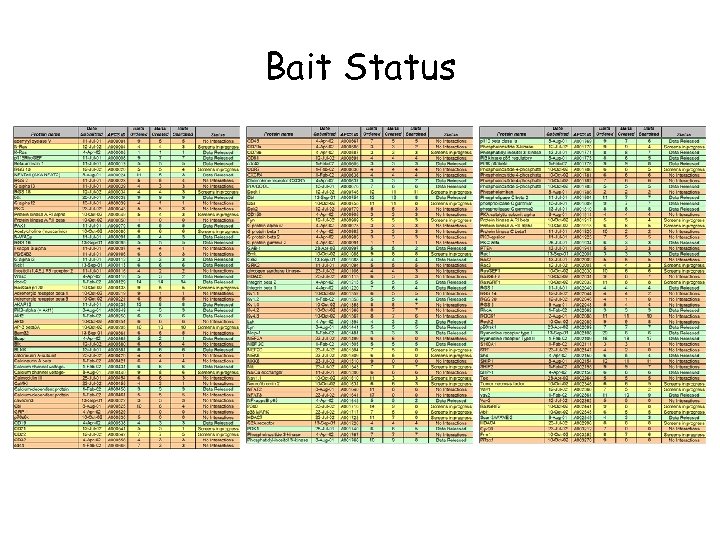
Bait Status
Bait Status

Pattern Among Negative Baits? • Of 72 negative baits, 45 are either membrane proteins (receptors, cell-surface antigens, channels) or are membrane associated (G proteins, GEFs, RGS etc. ) – May be overcome by generous and creative bait design but these proteins will always give lower “return” in Y 2 H assay • Also several proteins that could be expected to do better (cytosolic protein kinases and some adaptors) – Further baits advisable • Future baits sets should also include some of novel preys identified in primary screen
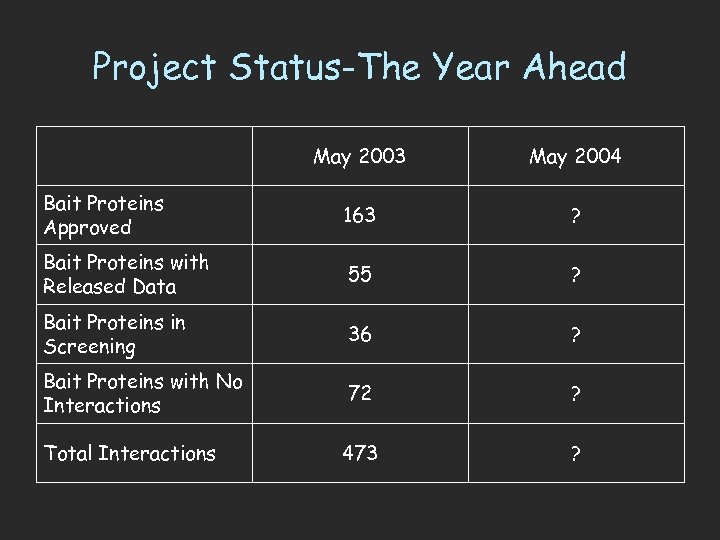
Project Status-The Year Ahead May 2003 May 2004 Bait Proteins Approved 163 ? Bait Proteins with Released Data 55 ? Bait Proteins in Screening 36 ? Bait Proteins with No Interactions 72 ? 473 ? Total Interactions

Acknowledgements Myriad Af. CS Terrece Pearman Data analysis and display Gil Sambrano Bob Sinkovits Joshua Li Brandi Williams Karen Heichman Paul Bartel RNA preparation Keng-Mean Lin Robert Hsueh Zhen Yan Joella Grossoehme Read Pierce Jason Polasek Jody Girouard
feeef08274202ed07688f776257594bb.ppt