234fb4df484691a43017b349d3defb79.ppt
- Количество слайдов: 47

I 529 - Lab 6 GO+MEME + Twin. Scan AI : Kwangmin Choi

Today’s topics • Gene Ontology prediction/mapping – Ami. Go • http: //amigo. geneontology. org/cgi-bin/amigo/go. cgi – PFP • http: //dragon. bio. purdue. edu/pfp/ – GOtcha • http: //www. compbio. dundee. ac. uk/gotcha/ • Pathway prediction/mapping – KAAS • http: //www. genome. jp/kegg/kaas • MEME • Twin. Scan
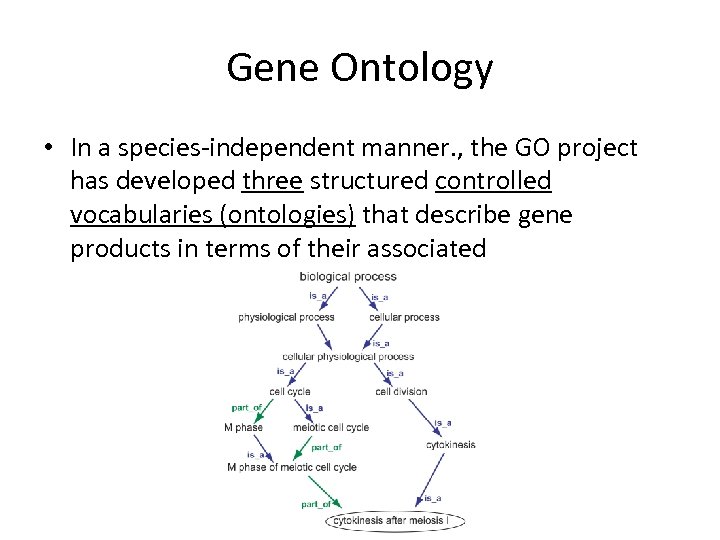
Gene Ontology • In a species-independent manner. , the GO project has developed three structured controlled vocabularies (ontologies) that describe gene products in terms of their associated

GO: biological process • A biological process is series of events accomplished by one or more ordered assemblies of molecular functions. – E. g. cellular physiological process or signal transduction. – E. g. pyrimidine metabolic process or alpha-glucoside transport. • It can be difficult to distinguish between a biological process and a molecular function, but the general rule is that a process must have more than one distinct steps. • A biological process is not equivalent to a pathway; at present, GO does not try to represent the dynamics or dependencies that would be required to fully describe a pathway.

GO: molecular functions • Molecular function describes activities, such as catalytic or binding activities, that occur at the molecular level. • GO molecular function terms represent activities rather than the entities (molecules or complexes) that perform the actions, • GO milecular function terms do not specify where or when, or in what context, the action takes place. – E. . g. (general) catalytic activity, transporter activity, or binding etc. – E. g. (specific) adenylate cyclase activity, Toll receptor binding etc.

GO: cellular components • A cellular component is just that, a component of a cell, but with the proviso that it is part of some larger object; • Less informative • This may be an anatomical structure – e. g. rough endoplasmic reticulum or nucleus • or a gene product group – e. g. ribosome, proteasome or a protein dimer

Ami. GO • URL http: //amigo. geneontology. org/cgi-bin/amigo/go. cgi • Ami. GO is the official tool for searching and browsing the Gene Ontology database • Simple blast search is provided (not useful) • Ami. GO consists of a controlled vocabulary of terms covering biological concepts, and a large number of genes or gene products whose attributes have been annotated using GO terms.

PFP (Automated Protein Function Prediction Server) • Hawkins, T. , Luban, S. and Kihara, D. 2006. Enhanced Automated Function Prediction Using Distantly Related Sequences and Contextual Association by PFP. Protein Science 15: 1550 -6. • The PFP algorithm has been shown to increase coverage of sequence-based function annotation more than fivefold by extending a PSI-BLAST search to extract and score GO terms individually • It applies the Function Association Matrix (FAM), to score significantly associating pairs of annotations.

PFP method • PFP uses a scoring scheme to rank GO annotations assigned to all of the most similar sequences according to – (1) their frequency of occurrence in those sequences – (2) the degree of similarity of the originating sequence to the query. • This is similar to the scoring basis for the R-value used by the GOtcha method to score annotations from pairwise alignment matches (Martin et al. 2004)
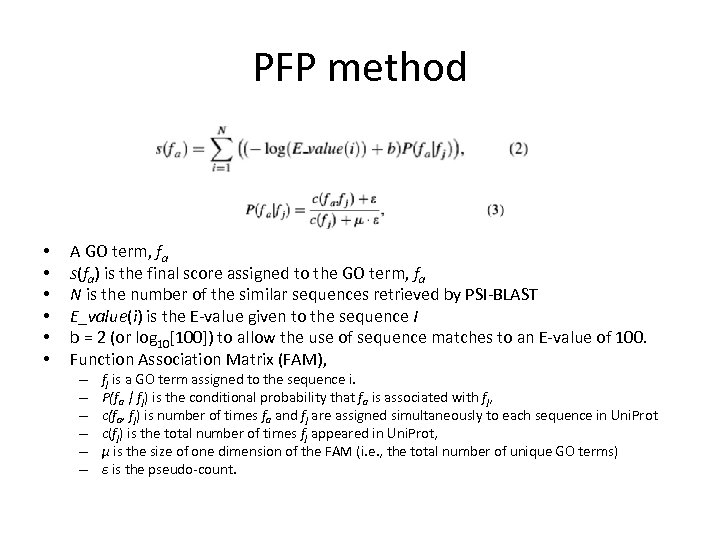

PFP method • • • A GO term, fa s(fa) is the final score assigned to the GO term, fa N is the number of the similar sequences retrieved by PSI-BLAST E_value(i) is the E-value given to the sequence I b = 2 (or log 10[100]) to allow the use of sequence matches to an E-value of 100. Function Association Matrix (FAM), – – – fj is a GO term assigned to the sequence i. P(fa | fj) is the conditional probability that fa is associated with fj, c(fa, fj) is number of times fa and fj are assigned simultaneously to each sequence in Uni. Prot c(fj) is the total number of times fj appeared in Uni. Prot, μ is the size of one dimension of the FAM (i. e. , the total number of unique GO terms) ɛ is the pseudo-count.

PFP • Web server http: //dragon. bio. purdue. edu/pfp/queue/116 8_kw. f. result. html • Local installation – http: //dragon. bio. purdue. edu/pfp/dist – You need to specify the path of blastp – And also need BLOSUM 62

PFP (Automated Protein Function Prediction Server) • PFP output – http: //darwin. informatics. indiana. edu/col/courses/I 529 -11/Lab 5/Data/pfp_data/0008. out • Columns – – – 1: predicted GO term 2: GO category (f/p/c) 3: raw term score 4: term p-value 5: rank (by p-value) 6: confidence to be exact match 7: rank (by column 7) 8: confidence within 2 edges on the GO DAG 9: rank (by column 8) 10: confidence within 4 edges on the GO DAG 11: rank (by column 10) 12: GO term short definition

GOtcha • The GOtcha method – Martin et al. BMC Bioinformatics (2004) 5: 178. • GOtcha assigns functional terms transitively based upon sequence similarity. • These terms are ranked by probability and displayed graphically on a subtree of Gene Ontology.

Gotcha method • GOtcha performs a BLAST search of the query sequence against individual well annotated genomes. • Annotations are transitively assigned from all hits, with a score corresponding to the Evalue, individual GO-terms receiving cumulative scores from multiple sequence similarity matches. • Cumulative scores are normalized and, for each term, two scores are obtained – – • the I-score which is normalized to the root node, the C-score which is the cumulative score at the root node. For each GO-term a precomputed scoring table is used to establish the assignment likelihood for that term given that I-score and that C-score. This is represented as a probability Results: http: //darwin. informatics. indiana. edu/col/courses/ I 529 -11/Lab 5/Data/Gotcha/gotcha. 4451/

KAAS • KAAS (KEGG Automatic Annotation Server) provides functional annotation of genes in a genome by BLAST comparisons against a manually curated set of ortholog groups in KEGG GENES. • The result contains KO (KEGG Orthology) assignments and automatically generated KEGG pathways. • Moriya, Y. , Itoh, M. , Okuda, S. , Yoshizawa, A. , and Kanehisa, M. ; KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, W 182 -W 185 (2007). [NAR]

KAAS • Web server: http: //www. genome. jp/kegg/kaas/ • KAAS works best when a complete set of genes in a genome is known. Prepare query amino acid sequences and use the BBH (bidirectional best hit) method to assign orthologs. • KAAS can also be used for a limited number of genes. Prepare query amino acid sequences and use the SBH (single-directional best hit) method to assign orthologs. • When ESTs are comprehensive enough, a set of consensus contigs can be generated by the EGassembler server and used as a gene set for KAAS with the BBH method. Otherwise, use ESTs as they are with the SBH method.
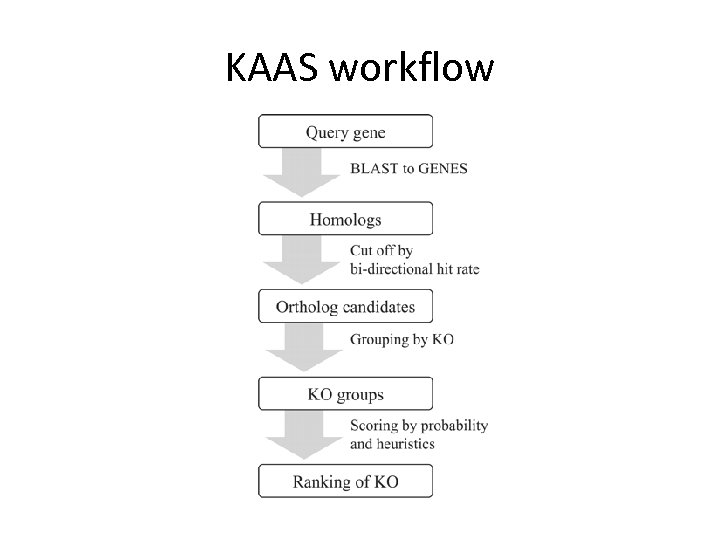
KAAS workflow

Pathway mapping • KAAS returns – KO list – KEGG Atlas Metabolism map [Create atlas] – Pathway maps [Create all maps] – Hierarchy files • You can highlight KEGG maps using KEGG API – http: //www. genome. jp/kegg/soap/doc/keggapi_man ual. html – See: color_pathway_by_objects


MEME

• Discover (conserved) motifs in a group of unaligned and related sequences (DNA or protein) • Unsupervised: Automatically choose the following (with little or no prior knowledge) – Best width of motifs – Number of occurrences in each sequence – Composition of each motif 2 1

MEME • MEME (Multiple EM for Motif Elicitation) discovers patterns in nucleotide and amino acid sequences. • MEME paper • Few tutorials - 1( Pg 6 ), 2, 3, 4 • MEME represents motifs as position-dependent letter-probability matrices which describe the probability of each possible letter at each position in the pattern. – MEME uses both alternately EM to find model parameters that maximize the likelihood of the model • Individual MEME motifs do not contain gaps. Patterns with variablelength gaps are split by MEME into two or more separate motifs.

Motif finding using the EM algorithm MEME (Bailey & Elkan 1995) http: //meme. sdsc. edu/meme/intro. html • MEME works by iteratively refining matrix and identifying sites: – Estimate motif model • Start with an n-mer seed (random or specified) • Build a matrix by incorporating some of background frequencies – Identify examples of the model • For every n-mer in the input set, identify its probability given the matrix model – Re-estimate the motif model • Calculate a new matrix, based on the weighted frequencies of all nmers in the set – Iteratively refine the matrix and identify sites until convergence.

Expectation Maximization 1. Expectation step – initial guess about the location of a (variable) sequence pattern in a set of sequences 2. Maximization step – improve/update pattern. Iterative search 24
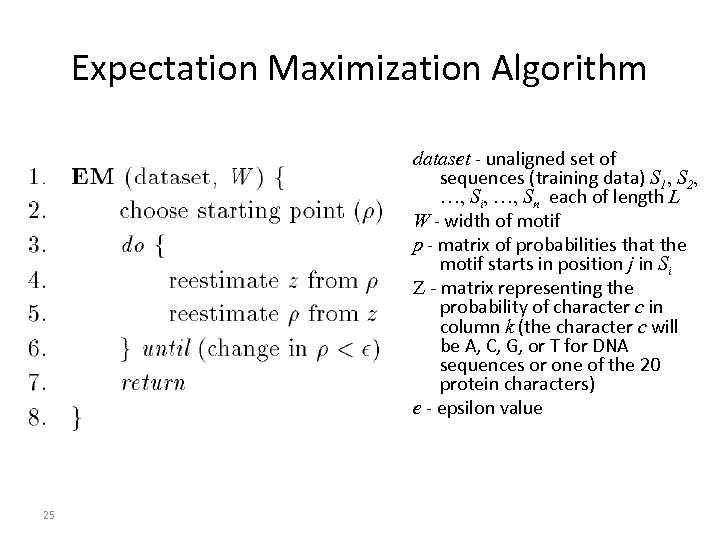

Expectation Maximization Algorithm dataset - unaligned set of sequences (training data) S 1, S 2, …, Si, …, Sn each of length L W - width of motif p - matrix of probabilities that the motif starts in position j in Si Z - matrix representing the probability of character c in column k (the character c will be A, C, G, or T for DNA sequences or one of the 20 protein characters) e - epsilon value 25

MEME Algorithm 26
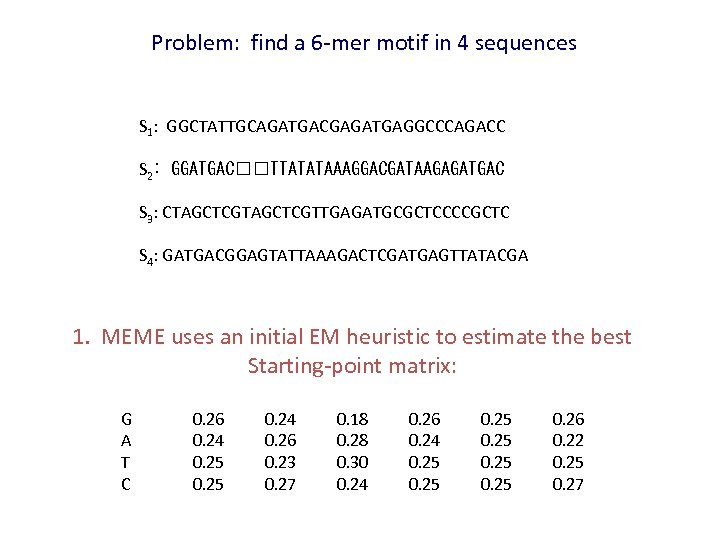

Problem: find a 6 -mer motif in 4 sequences S 1: GGCTATTGCAGATGACGAGATGAGGCCCAGACC S 2: GGATGAC TTATATAAAGGACGATAAGAGATGAC S 3: CTAGCTCGTTGAGATGCGCTCCCCGCTC S 4: GATGACGGAGTATTAAAGACTCGATGAGTTATACGA 1. MEME uses an initial EM heuristic to estimate the best Starting-point matrix: G A T C 0. 26 0. 24 0. 25 0. 24 0. 26 0. 23 0. 27 0. 18 0. 28 0. 30 0. 24 0. 26 0. 24 0. 25 0. 26 0. 22 0. 25 0. 27

2. MEME scores the match of all 6 -mers to current matrix GGCTATTGCATATGACGAGATGAGGCCCAGACC GGATGAC TT AGGACCGTGATAAGAGATTAC ATATAA CTAGCTCGTTGAGATGCGCTCCCCGCTC GATGACGGAGTATTAAAGACTCGATGAGTTATACGA Here, just consider the underlined 6 -mers, Although in reality all 6 -mers are scored
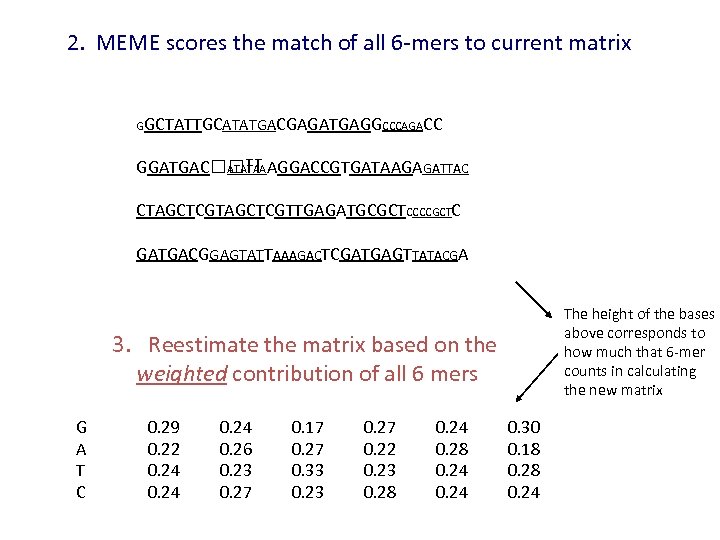
2. MEME scores the match of all 6 -mers to current matrix GGCTATTGCATATGACGAGATGAGGCCCAGACC GGATGAC TT AGGACCGTGATAAGAGATTAC ATATAA CTAGCTCGTTGAGATGCGCTCCCCGCTC GATGACGGAGTATTAAAGACTCGATGAGTTATACGA The height of the bases above corresponds to how much that 6 -mer counts in calculating the new matrix 3. Reestimate the matrix based on the weighted contribution of all 6 mers G A T C 0. 29 0. 22 0. 24 0. 26 0. 23 0. 27 0. 17 0. 27 0. 33 0. 27 0. 22 0. 23 0. 28 0. 24 0. 30 0. 18 0. 24
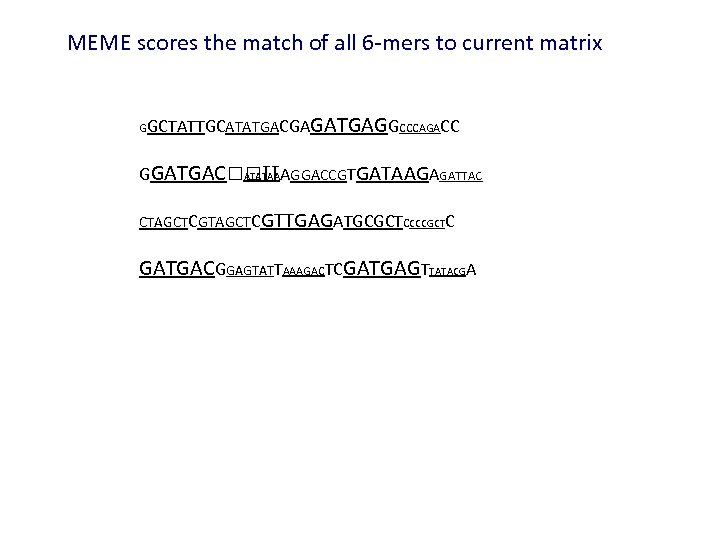

MEME scores the match of all 6 -mers to current matrix GGCTATTGCATATGACGA GATGAGGCCCAGACC GGATGAC TTAGGACCGTGATAAGAGATTAC ATATAA CTAGCTCGTTGAGATGCGCTCCCCGCTC GATGACGGAGTATTAAAGACTCGATGAGTTATACGA

GGCTATTGCATATGACGA GATGAGGCCCAGACC GGATGAC TTAGGACCGTGATAAGAGATTAC ATATAA CTAGCTCGTTGAGATGCGCTCCCCGCTC GATGACGGAGTATTAAAGACTCGATGAGTTATACGA The height of the bases above corresponds to how much that 6 -mer counts in calculating the new matrix Reestimate the matrix based on the weighted contribution of all 6 mers G A T C 0. 40 0. 30 0. 15 0. 20 0. 30 0. 20 0. 15 0. 20 0. 42 0. 24 0. 16 0. 24 0. 46 0. 15 0. 30 0. 18 0. 24

MEME scores the match of all 6 -mers to current matrix GGCTATTGCATATGACGA GATGAGGCCCAGACC GGATGAC GGACCGTGATAAGAGATTAC TTATATAAA CTAGCTCGTTGAGATGCGCTCCCCGCTC GATGACGGAGTATTAAAGACTCGATGAGT A TATACG Iterations continue until convergence (ie. numbers don’t change much between iterations)
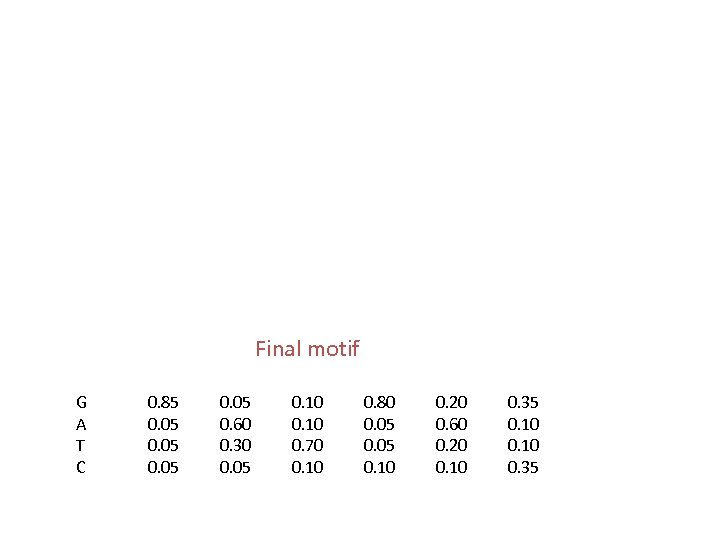
Final motif G A T C 0. 85 0. 05 0. 60 0. 30 0. 05 0. 10 0. 70 0. 10 0. 80 0. 05 0. 10 0. 20 0. 60 0. 20 0. 10 0. 35
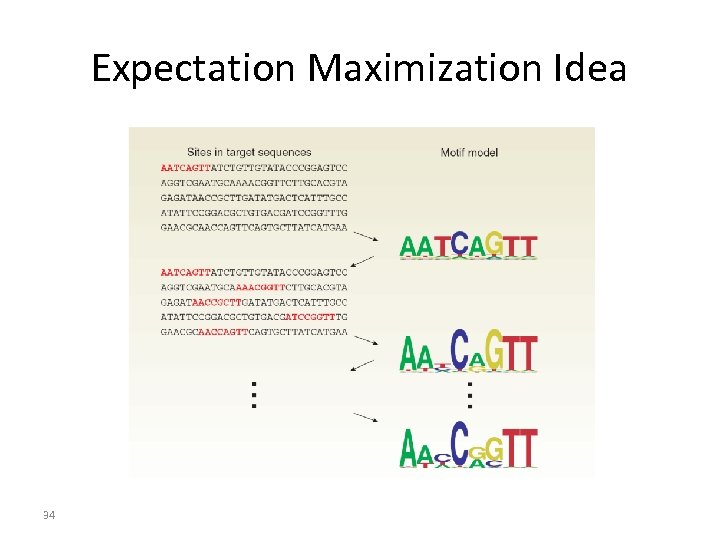
Expectation Maximization Idea 34

MEME web server • MEME website • Protein motif • Sample Input • Fill in email address • Keep all else default • DNA motif • Sample Input • Minimum motif width: 4 • Number of different motifs: 2
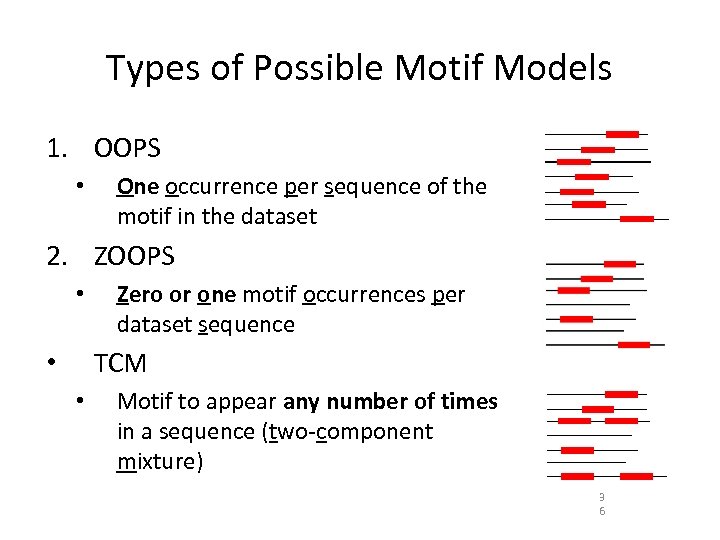

Types of Possible Motif Models 1. OOPS • One occurrence per sequence of the motif in the dataset 2. ZOOPS • Zero or one motif occurrences per dataset sequence TCM • • Motif to appear any number of times in a sequence (two-component mixture) 3 6

A simple DNA example meme dna. fasta -dna -mod oops -pal -o output_dir • MEME looks for a single motif in the file dna. fasta which contains DNA sequences in FASTA format. • The OOPS model is used so MEME assumes that every sequence contains exactly one occurrence of the motif. • The palindrome switch is given so the motif model (PSPM) is converted into a palindrome by combining corresponding frequency columns. • • • If MEME decides that a motif is a palindrome, it averages the letter frequencies in corresponding columns together. For instance, if the width of the motif is 10, columns 1 and 10, 2 and 9, 3 and 8, etc. , are averaged together. MEME automatically chooses the best width for the motif in this example since no width was specified.

Searching for motifs on both DNA strands meme dna. fasta -dna -mod oops -revcomp -o output_dir • The -revcomp switch tells MEME to consider both DNA strands, and • The -pal switch is absent so the palindrome conversion is omitted. • When DNA uses both DNA strands, motif occurrences on the two strands may not overlap. That is, any position in the sequence given in the training set may be contained in an occurrence of a motif on the positive strand or the negative strand, but not both.

A simple protein example meme protein. fasta -mod oops -maxw 20 -nmotifs 2 -o output_dir • The -dna switch is absent, so MEME assumes the input file as protein sequences. • Each motif is assumed to occur in each of the sequences because the OOPS model is specified. • Specifying -maxw 20 makes MEME run faster since it does not have to consider motifs longer than 20. • MEME searches for two motifs each of width less than or equal to 20.

MEME on Big. Red • MEME is installed on Big. Red. • KB: http: //kb. iu. edu/data/awwa. html

TWINSCAN

Twin. Scan • Twin. Scan finds genes in a "target" genomic sequence by simultaneously maximizing the probability of the gene structure in the target and the evolutionary conservation derived from "informant" genomic sequences. • The target sequence (i. e. the sequence to be annotated) should generally be of draft or finished quality. • The informant can range from a single sequence to a whole genome in any condition from raw shotgun reads to finished assembly. • References – http: //mblab. wustl. edu/media/publications/Hu-Brent-2003 -Proof. pdf – http: //mrw. interscience. wiley. com/emrw/9780471250951/cp/cpbi/article/bi 0 408/current/abstract – http: //bioinformatics. oxfordjournals. org/cgi/content/abstract/17/suppl_1/S 1 40

Requirements • Twin. Scan/ Nscan 4. 0 executable • DNA sequence in FASTA format • Twinscan parameter file (/parameters/twinscan_parameters) – • Conservation sequence (see examples/example. conseq) – – – – • Each filename contains the name of the target organism: eg maize_twinscan. zhmm. A symbolic representation of the best alignments between the target and informant sequences. To create this conservation sequence, WU-BLAST (http: //blast. wustl. edu) is used. For NCBI BLAST, the input parameters need to be changed. (see parameters/example_blast_parameters. txt) xdformat (WU-BLAST package) formats the informant sequences to create a blast database. After running BLAST, the output must be formatted with conseq, which is included in this package. Example (using WU-BLAST) • xdformat -n informant. fa • Blast M=1 N=-1 Q=5 R=1 B=10000 V=100 -cpus=1 -warnings -lcfilter=seg filter=dust topcombo. N=1 informant. fa target. fa > blast. out • conseq target. fa blast. out > conseq. fa Note: The run. Twinscan 2 script will run these steps without user intervention (see below). EST sequence (optional) – – – EST sequence is a symbolic representation of evidence from ESTs that align to the target sequence. The estseq script included in the distribution creates EST sequence when given a DNA sequence and a (set of) BLAT reports of the ESTs aligned to the target. For downloading BLAT, go to http: //genome. ucsc. edu/FAQblat. html#blat 3

Web service and Installation • Web service – http: //mblab. wustl. edu/nscan/submit/ • Local installation – http: //mblab. wustl. edu/software/twinscan/ • Installation step $ tar xvzf iscan-4. 1. 2. tar. gz $ cd N-SCAN $ make linux $. /test-executable
![How to run • Usage twinscan <parameter file> <masked sequence file> -c=<conseq file> [e=estseq_file] How to run • Usage twinscan <parameter file> <masked sequence file> -c=<conseq file> [e=estseq_file]](https://present5.com/presentation/234fb4df484691a43017b349d3defb79/image-45.jpg)
How to run • Usage twinscan <parameter file> <masked sequence file> -c=<conseq file> [e=estseq_file] > <outputfile> • Example: . /bin/iscan. /parameters/twinscan_parameters/human_twinscan. zhmm. /examples/example. fa. masked -c=. /examples/example. conseq > my. Sequence. gtf

Running Twinscan using the run. Twinscan 2 script • In summary, there are 5 steps required to run Twinscan: – – – Step 1: Mask target sequence with Repeat. Masker Step 2: Create informant BLAST database Step 3: Run BLAST Step 4: Create conservation sequence (Step 4 b: Create EST sequence) Step 5: Run Twinscan • These five steps are all contained in the example script run. Twinscan 2 (see. /bin) • The default BLAST parameters used by run. Twinscan 2 are those for C. elegans (see parameters/blast_params/Celegans. blast. param). – This can and should be changed for any other species with the -B option to the run. Twinscan 2 script. • The file example. output in the /examples directory contains the output from run. Twincan 2 using the BLAST parameters found in the script.

Local environment seeting for run. Twinscan 2 • Example –. . /bin/run. Twinscan 2 -r. . /parameters/twinscan_parameters/human_twinscan. zhmm -d output -B. . /parameters/blast_params/Hsapiens. blast. param example. fa. masked informant. fa • After running you can find output files in the newly created /output directory. • Several programs must be installed on your system to run. Twinscan. • You may need to change run. Twinscan to point it to these programs. – – – my $REPEATMASKER my $BLASTN my $BLAT my $XDFORMAT my $PRESSDB = "Repeat. Masker"; = "blastn"; = "blat"; = "xdformat"; = "pressdb"; # Format for local environment # Format for local environment
234fb4df484691a43017b349d3defb79.ppt