2347754cb61159cea00392b9ebcfb735.ppt
- Количество слайдов: 42



 Human Chromosomes Male Xy X y Female XX Xy Daughter Son
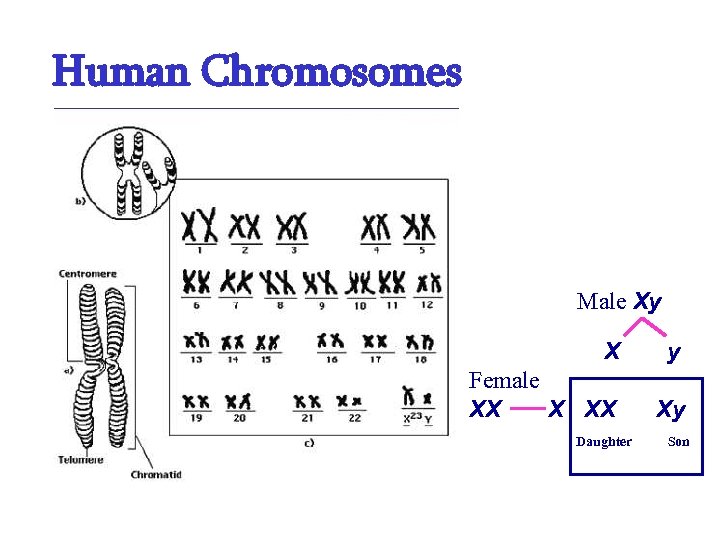
Human Chromosomes Male Xy X y Female XX Xy Daughter Son
 Gene, Allele, Genotype, Phenotype Chromosomes from Father Mother Gene A, with two alleles A and a Genotype Phenotype AA AA Height 185 182 IQ 100 104 Aa Aa 175 171 103 102 aa aa 155 152 101 103
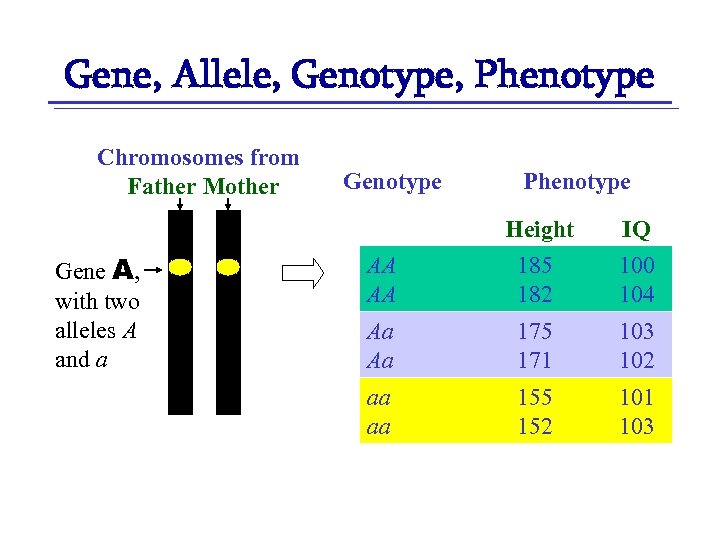
Gene, Allele, Genotype, Phenotype Chromosomes from Father Mother Gene A, with two alleles A and a Genotype Phenotype AA AA Height 185 182 IQ 100 104 Aa Aa 175 171 103 102 aa aa 155 152 101 103
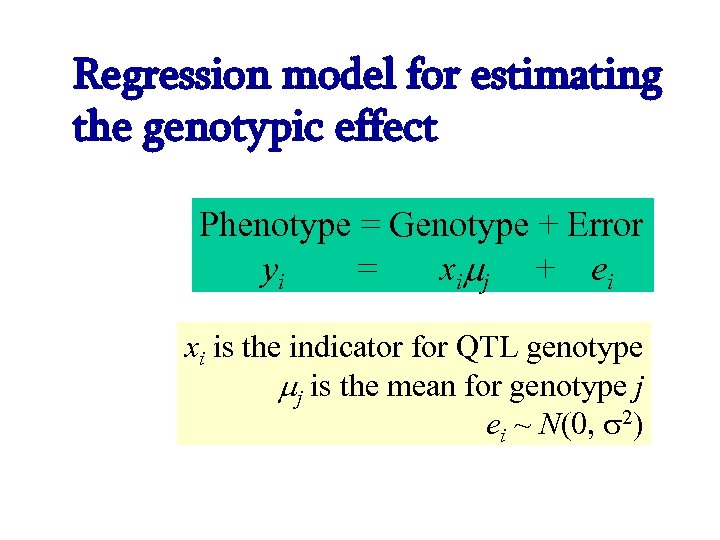 Regression model for estimating the genotypic effect Phenotype = Genotype + Error yi = x i j + e i xi is the indicator for QTL genotype j is the mean for genotype j ei ~ N(0, 2)
Regression model for estimating the genotypic effect Phenotype = Genotype + Error yi = x i j + e i xi is the indicator for QTL genotype j is the mean for genotype j ei ~ N(0, 2)
 Uniqueness for our genetic problem M 1 M 2 M 3. . . Mm QTL The genotypes for the trait are not observable and should be predicted from linked neutral molecular markers (M) The genes that lead to the phenotypic variation are called Quantitative Trait Loci (QTL) Our task is to construct a statistical model that connects the QTL genotypes and marker genotypes through observed phenotypes
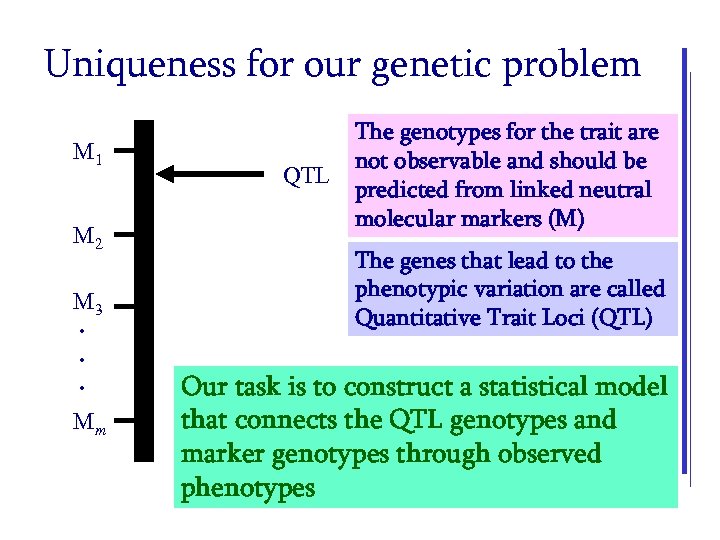
Uniqueness for our genetic problem M 1 M 2 M 3. . . Mm QTL The genotypes for the trait are not observable and should be predicted from linked neutral molecular markers (M) The genes that lead to the phenotypic variation are called Quantitative Trait Loci (QTL) Our task is to construct a statistical model that connects the QTL genotypes and marker genotypes through observed phenotypes
 Parents AA aa Data Structure Marker (M) Subject 1 2 3 4 5 6 7 8 M 1 AA(2) Aa(1) aa(0) M 2 … BB(2) Bb(1) bb(0) …. . . . … F 1 F 2 Aa AA ¼ Aa ½ aa ¼ Genotype frequency Mm Phenotype QQ(2) (y) ¼ y 1 y y 2 y 3 y 4 y 5 6 y y 7 8 ¼ ¼ ¼ ¼ Qq(1) ½ qq(0) ¼ ½ ½ ½ ½ ¼ ¼ ¼ ¼ n = n 22 + n 21 + n 20 + n 12 + n 00 + n 02 + n 01 + n 00
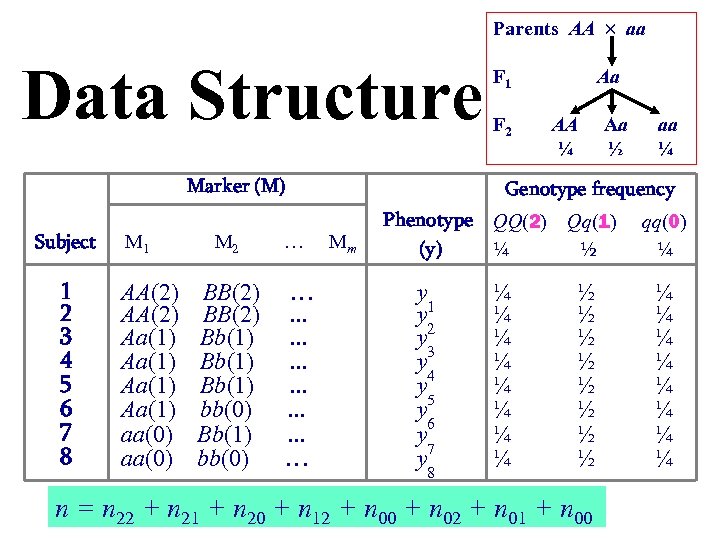
Parents AA aa Data Structure Marker (M) Subject 1 2 3 4 5 6 7 8 M 1 AA(2) Aa(1) aa(0) M 2 … BB(2) Bb(1) bb(0) …. . . . … F 1 F 2 Aa AA ¼ Aa ½ aa ¼ Genotype frequency Mm Phenotype QQ(2) (y) ¼ y 1 y y 2 y 3 y 4 y 5 6 y y 7 8 ¼ ¼ ¼ ¼ Qq(1) ½ qq(0) ¼ ½ ½ ½ ½ ¼ ¼ ¼ ¼ n = n 22 + n 21 + n 20 + n 12 + n 00 + n 02 + n 01 + n 00
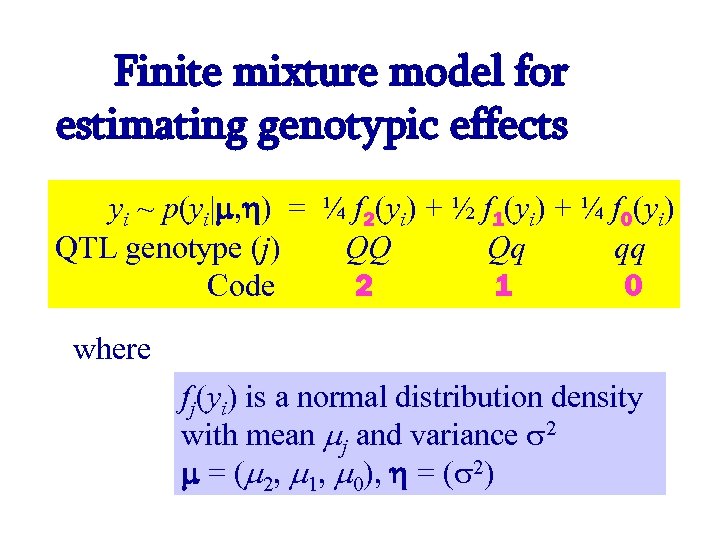 Finite mixture model for estimating genotypic effects yi ~ p(yi| , ) = ¼ f 2(yi) + ½ f 1(yi) + ¼ f 0(yi) QTL genotype (j) QQ Qq qq Code 2 1 0 where fj(yi) is a normal distribution density with mean j and variance 2 = ( 2, 1, 0), = ( 2)
Finite mixture model for estimating genotypic effects yi ~ p(yi| , ) = ¼ f 2(yi) + ½ f 1(yi) + ¼ f 0(yi) QTL genotype (j) QQ Qq qq Code 2 1 0 where fj(yi) is a normal distribution density with mean j and variance 2 = ( 2, 1, 0), = ( 2)
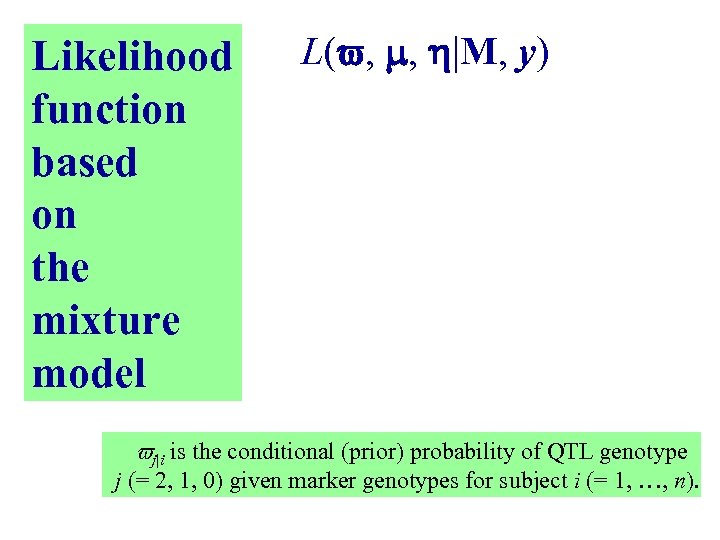 Likelihood function based on the mixture model L( , , |M, y) j|i is the conditional (prior) probability of QTL genotype j (= 2, 1, 0) given marker genotypes for subject i (= 1, …, n).
Likelihood function based on the mixture model L( , , |M, y) j|i is the conditional (prior) probability of QTL genotype j (= 2, 1, 0) given marker genotypes for subject i (= 1, …, n).
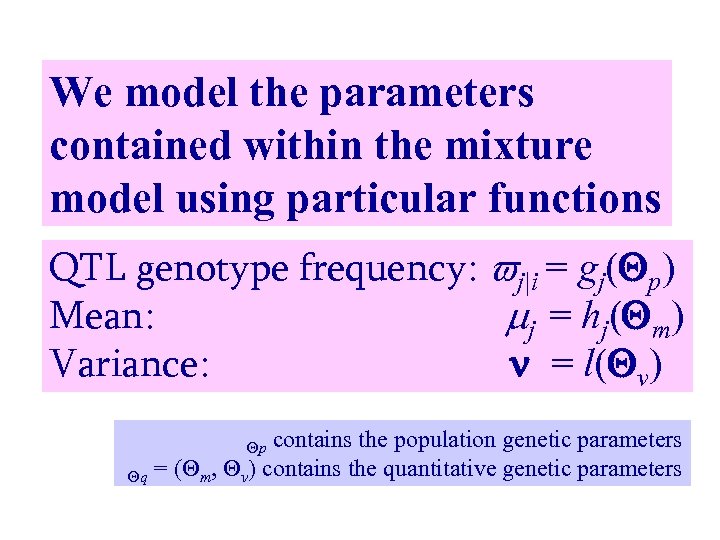 We model the parameters contained within the mixture model using particular functions QTL genotype frequency: j|i = gj( p) Mean: j = hj( m) Variance: = l( v) contains the population genetic parameters q = ( m, v) contains the quantitative genetic parameters p
We model the parameters contained within the mixture model using particular functions QTL genotype frequency: j|i = gj( p) Mean: j = hj( m) Variance: = l( v) contains the population genetic parameters q = ( m, v) contains the quantitative genetic parameters p
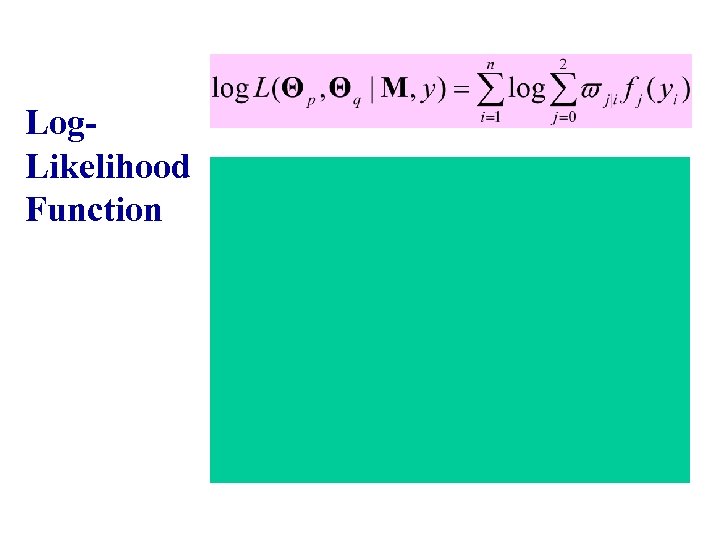 Log. Likelihood Function
Log. Likelihood Function
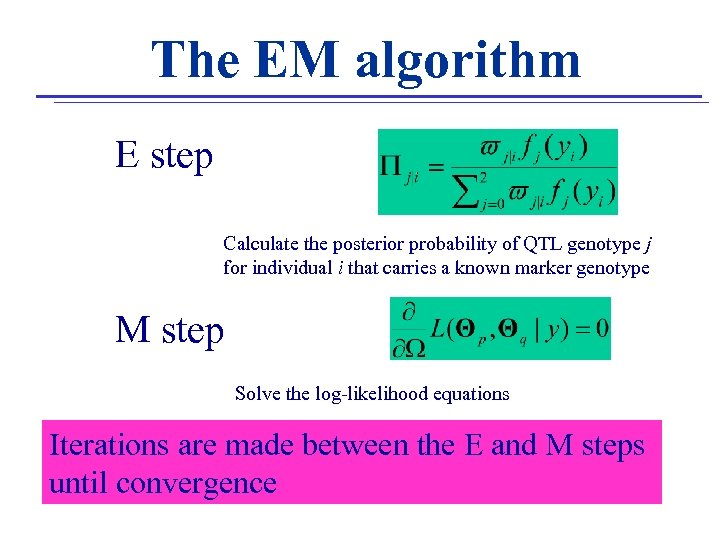 The EM algorithm E step Calculate the posterior probability of QTL genotype j for individual i that carries a known marker genotype M step Solve the log-likelihood equations Iterations are made between the E and M steps until convergence
The EM algorithm E step Calculate the posterior probability of QTL genotype j for individual i that carries a known marker genotype M step Solve the log-likelihood equations Iterations are made between the E and M steps until convergence
 Three statistical issues Modeling mixture proportions, i. e. , genotype frequencies at a putative QTL Modeling the mean vector Modeling the (co)variance matrix
Three statistical issues Modeling mixture proportions, i. e. , genotype frequencies at a putative QTL Modeling the mean vector Modeling the (co)variance matrix
 Functional Mapping An innovative model for genetic dissection of complex traits by incorporating mathematical aspects of biological principles into a mapping framework Provides a tool for cutting-edge research at the interplay between gene action and development
Functional Mapping An innovative model for genetic dissection of complex traits by incorporating mathematical aspects of biological principles into a mapping framework Provides a tool for cutting-edge research at the interplay between gene action and development
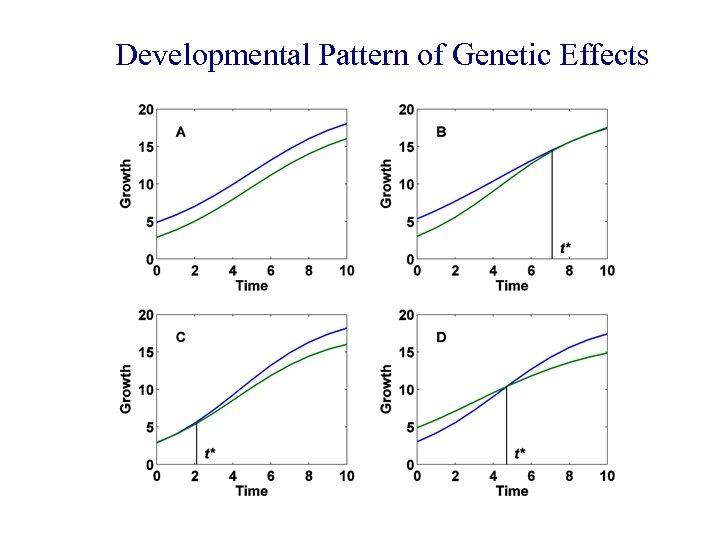 Developmental Pattern of Genetic Effects
Developmental Pattern of Genetic Effects
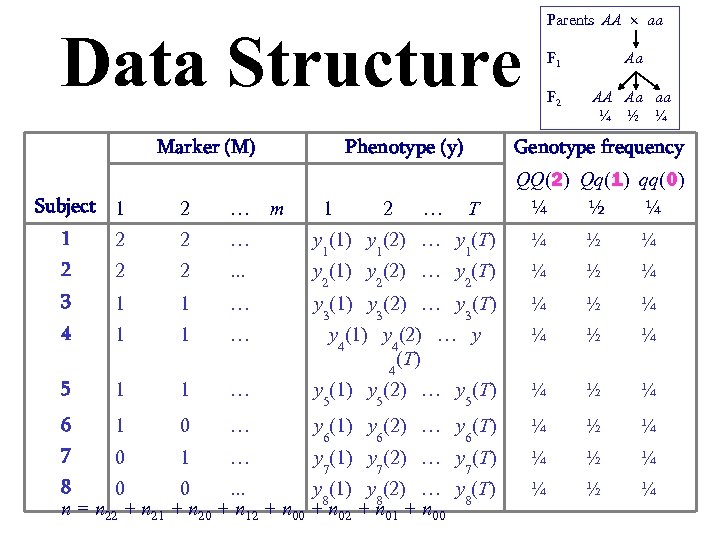 Parents AA aa Data Structure Marker (M) Subject 1 2 … m Phenotype (y) 1 2 … F 1 Aa F 2 AA Aa aa ¼ ½ ¼ Genotype frequency T QQ(2) Qq(1) qq(0) ¼ ½ ¼ 1 2 2 … ¼ ½ ¼ 2 y 1(1) y 1(2) … y 1(T) 2 2 . . . ¼ ½ ¼ 3 y 2(1) y 2(2) … y 2(T) 1 1 … ¼ ½ ¼ 4 y 3(1) y 3(2) … y 3(T) 1 1 … ¼ ½ ¼ 5 y 4(1) y 4(2) … y (T) 4 1 1 … y 5(1) y 5(2) … y 5(T) ¼ ½ ¼ 6 1 0 … ¼ ½ ¼ 7 y 6(1) y 6(2) … y 6(T) 0 1 … y 7(1) y 7(2) … y 7(T) ¼ ½ ¼ 8 0 0. . . y 8(1) y 8(2) … y 8(T) n = n 22 + n 21 + n 20 + n 12 + n 00 + n 02 + n 01 + n 00
Parents AA aa Data Structure Marker (M) Subject 1 2 … m Phenotype (y) 1 2 … F 1 Aa F 2 AA Aa aa ¼ ½ ¼ Genotype frequency T QQ(2) Qq(1) qq(0) ¼ ½ ¼ 1 2 2 … ¼ ½ ¼ 2 y 1(1) y 1(2) … y 1(T) 2 2 . . . ¼ ½ ¼ 3 y 2(1) y 2(2) … y 2(T) 1 1 … ¼ ½ ¼ 4 y 3(1) y 3(2) … y 3(T) 1 1 … ¼ ½ ¼ 5 y 4(1) y 4(2) … y (T) 4 1 1 … y 5(1) y 5(2) … y 5(T) ¼ ½ ¼ 6 1 0 … ¼ ½ ¼ 7 y 6(1) y 6(2) … y 6(T) 0 1 … y 7(1) y 7(2) … y 7(T) ¼ ½ ¼ 8 0 0. . . y 8(1) y 8(2) … y 8(T) n = n 22 + n 21 + n 20 + n 12 + n 00 + n 02 + n 01 + n 00
![The Finite Mixture Model Observation vector, yi = [yi(1), …, yi(T)] ~ MVN(uj, ) The Finite Mixture Model Observation vector, yi = [yi(1), …, yi(T)] ~ MVN(uj, )](https://present5.com/presentation/2347754cb61159cea00392b9ebcfb735/image-18.jpg) The Finite Mixture Model Observation vector, yi = [yi(1), …, yi(T)] ~ MVN(uj, ) Mean vector, uj = [uj(1), uj(2), …, uj(T)], (Co)variance matrix,
The Finite Mixture Model Observation vector, yi = [yi(1), …, yi(T)] ~ MVN(uj, ) Mean vector, uj = [uj(1), uj(2), …, uj(T)], (Co)variance matrix,
 Modeling the Mean Vector • Parametric approach Growth trajectories – Logistic curve HIV dynamics – Bi-exponential function Biological clock – Van Der Pol equation Drug response – Emax model • Nonparametric approach Lengedre function (orthogonal polynomial) B-spline (Xueli Liu & R. Wu: Genetics, to be submitted)
Modeling the Mean Vector • Parametric approach Growth trajectories – Logistic curve HIV dynamics – Bi-exponential function Biological clock – Van Der Pol equation Drug response – Emax model • Nonparametric approach Lengedre function (orthogonal polynomial) B-spline (Xueli Liu & R. Wu: Genetics, to be submitted)
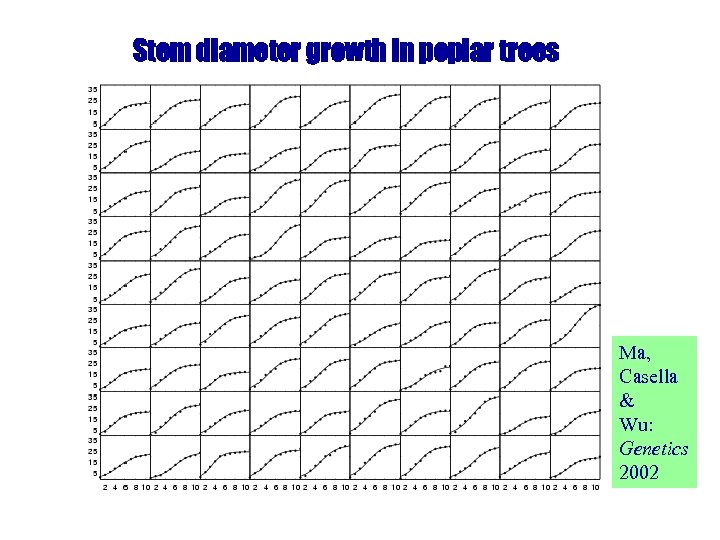 Stem diameter growth in poplar trees Ma, Casella & Wu: Genetics 2002
Stem diameter growth in poplar trees Ma, Casella & Wu: Genetics 2002
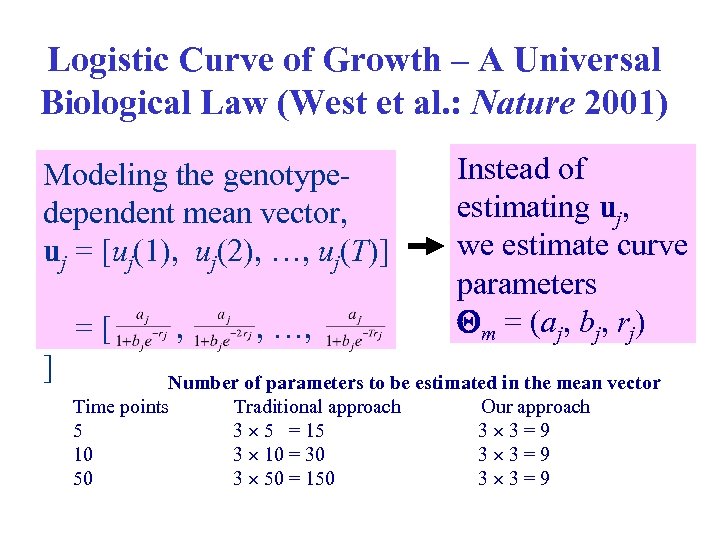 Logistic Curve of Growth – A Logistic of Growth – A Universal Biological Law (West et al. : Nature 2001) Universal Biological Law Modeling the genotypedependent mean vector, uj = [uj(1), uj(2), …, uj(T)] =[ ] , , …, Instead of estimating uj, we estimate curve parameters m = (aj, bj, rj) Number of parameters to be estimated in the mean vector Time points Traditional approach Our approach 5 3 5 = 15 3 3=9 10 3 10 = 30 3 3=9 50 3 50 = 150 3 3=9
Logistic Curve of Growth – A Logistic of Growth – A Universal Biological Law (West et al. : Nature 2001) Universal Biological Law Modeling the genotypedependent mean vector, uj = [uj(1), uj(2), …, uj(T)] =[ ] , , …, Instead of estimating uj, we estimate curve parameters m = (aj, bj, rj) Number of parameters to be estimated in the mean vector Time points Traditional approach Our approach 5 3 5 = 15 3 3=9 10 3 10 = 30 3 3=9 50 3 50 = 150 3 3=9
 Modeling the Variance Matrix Stationary parametric approach Autoregressive (AR) model Nonstationary parameteric approach Structured antedependence (SAD) model Ornstein-Uhlenbeck (OU) process Nonparametric approach Lengendre function
Modeling the Variance Matrix Stationary parametric approach Autoregressive (AR) model Nonstationary parameteric approach Structured antedependence (SAD) model Ornstein-Uhlenbeck (OU) process Nonparametric approach Lengendre function
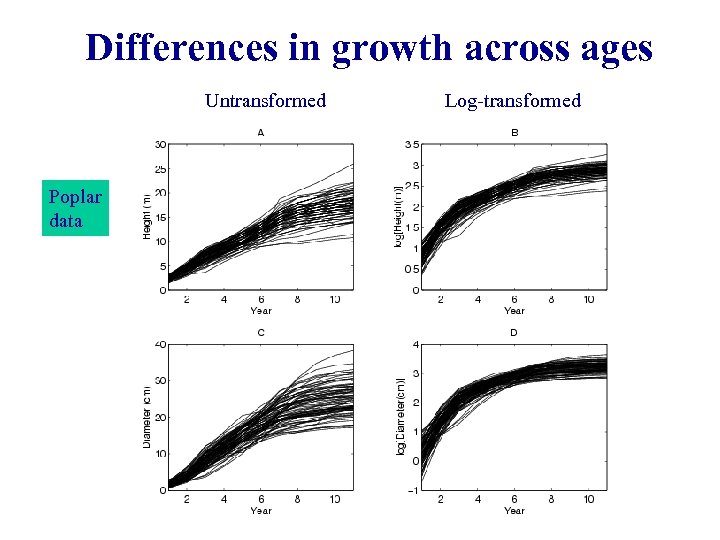 Differences in growth across ages Untransformed Poplar data Log-transformed
Differences in growth across ages Untransformed Poplar data Log-transformed
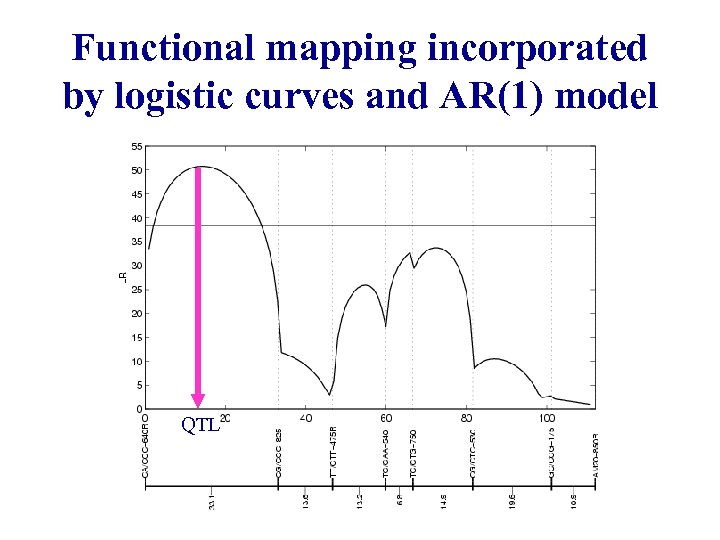 Functional mapping incorporated by logistic curves and AR(1) model QTL
Functional mapping incorporated by logistic curves and AR(1) model QTL
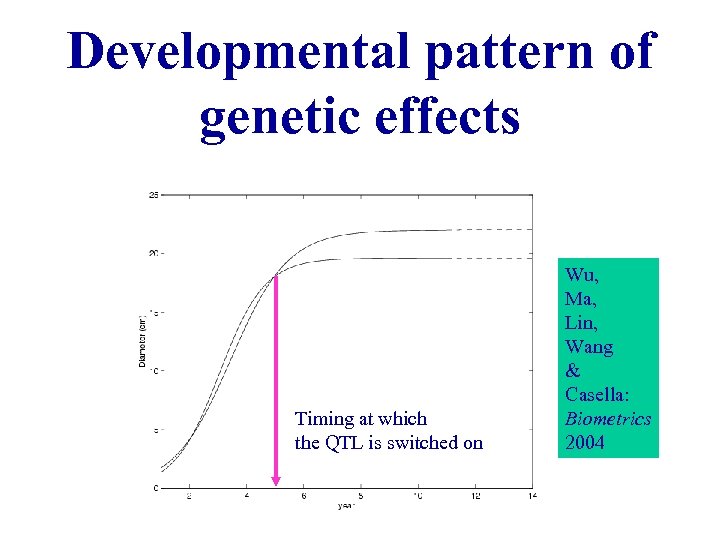 Developmental pattern of genetic effects Timing at which the QTL is switched on Wu, Ma, Lin, Wang & Casella: Biometrics 2004
Developmental pattern of genetic effects Timing at which the QTL is switched on Wu, Ma, Lin, Wang & Casella: Biometrics 2004
 The implications of functional mapping for high-dimensional biology High-dimensional biology deals with • Multiple genes – Epistatic gene-gene interactions • Multiple environments – Genotype environment interactions • Multiple traits – Trait correlations • Multiple developmental stages • Complex networks among genes, products and phenotypes
The implications of functional mapping for high-dimensional biology High-dimensional biology deals with • Multiple genes – Epistatic gene-gene interactions • Multiple environments – Genotype environment interactions • Multiple traits – Trait correlations • Multiple developmental stages • Complex networks among genes, products and phenotypes
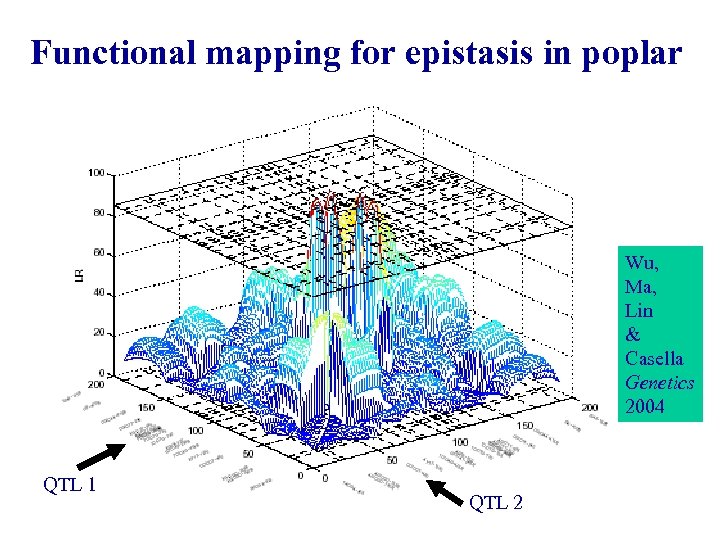 Functional mapping for epistasis in poplar Wu, Ma, Lin & Casella Genetics 2004 QTL 1 QTL 2
Functional mapping for epistasis in poplar Wu, Ma, Lin & Casella Genetics 2004 QTL 1 QTL 2
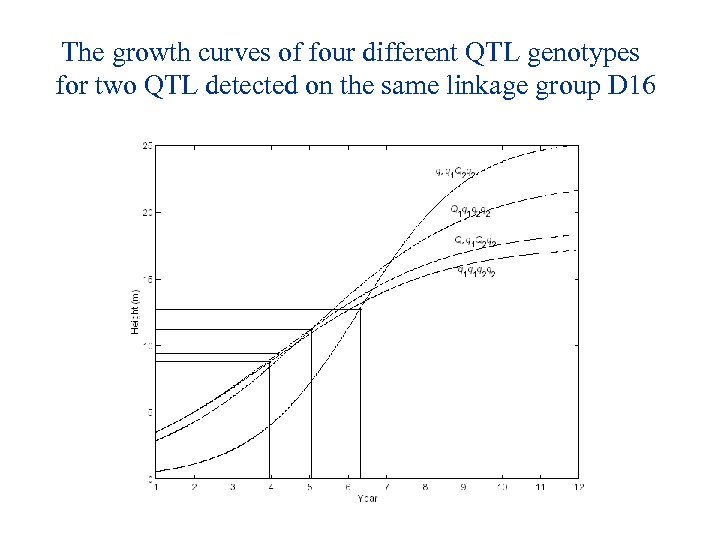 The growth curves of four different QTL genotypes for two QTL detected on the same linkage group D 16
The growth curves of four different QTL genotypes for two QTL detected on the same linkage group D 16
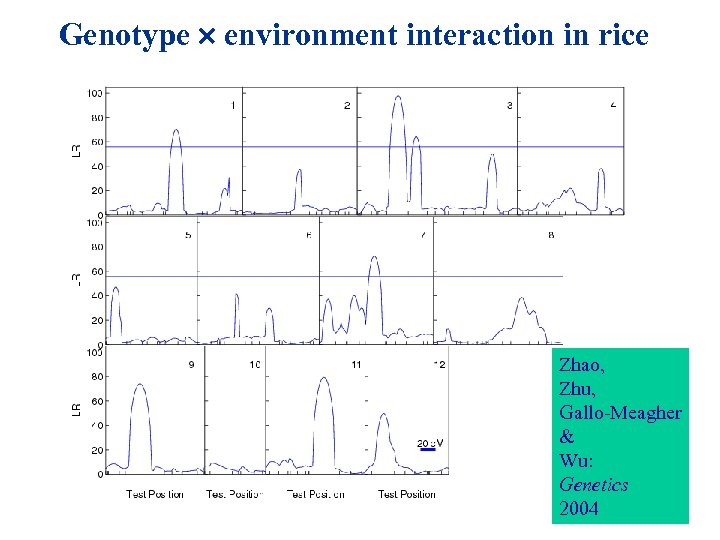 Genotype environment interaction in rice Zhao, Zhu, Gallo-Meagher & Wu: Genetics 2004
Genotype environment interaction in rice Zhao, Zhu, Gallo-Meagher & Wu: Genetics 2004
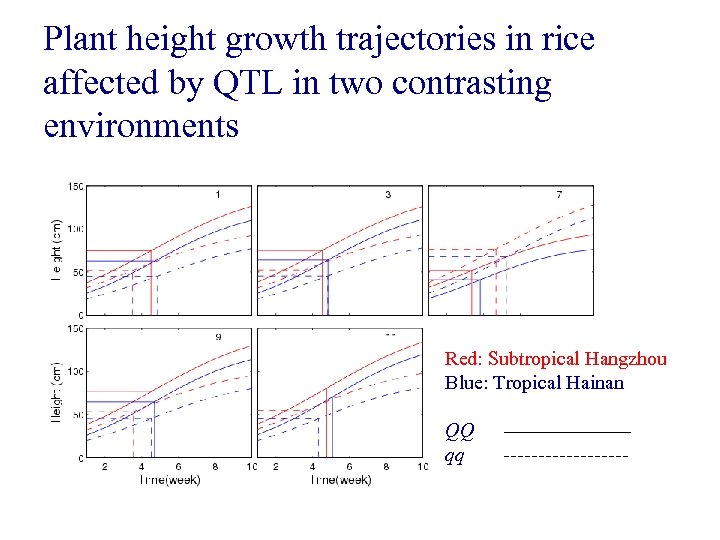 Plant height growth trajectories in rice affected by QTL in two contrasting environments Red: Subtropical Hangzhou Blue: Tropical Hainan QQ qq
Plant height growth trajectories in rice affected by QTL in two contrasting environments Red: Subtropical Hangzhou Blue: Tropical Hainan QQ qq
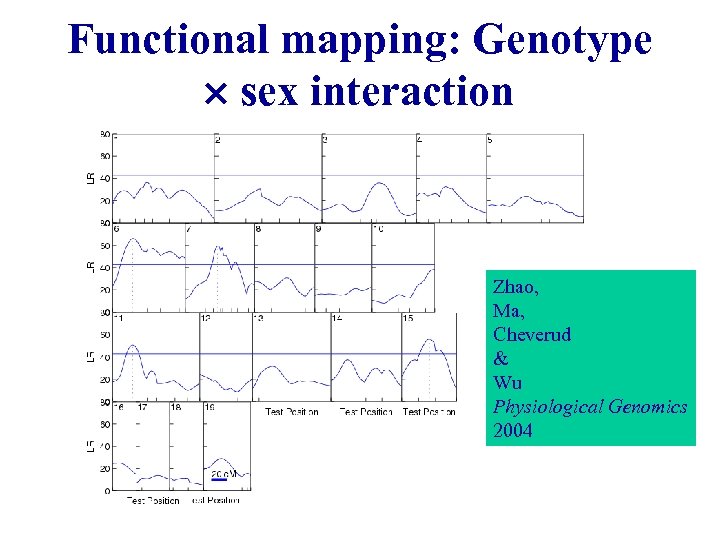 Functional mapping: Genotype sex interaction Zhao, Ma, Cheverud & Wu Physiological Genomics 2004
Functional mapping: Genotype sex interaction Zhao, Ma, Cheverud & Wu Physiological Genomics 2004
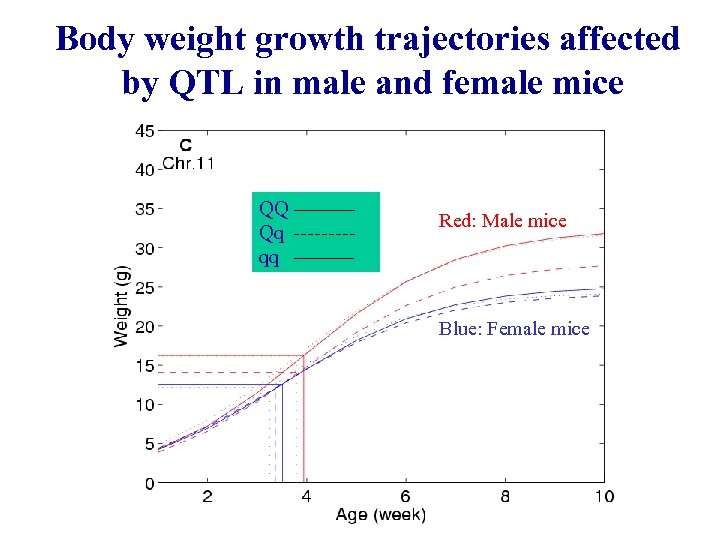 Body weight growth trajectories affected by QTL in male and female mice QQ Qq qq Red: Male mice Blue: Female mice
Body weight growth trajectories affected by QTL in male and female mice QQ Qq qq Red: Male mice Blue: Female mice
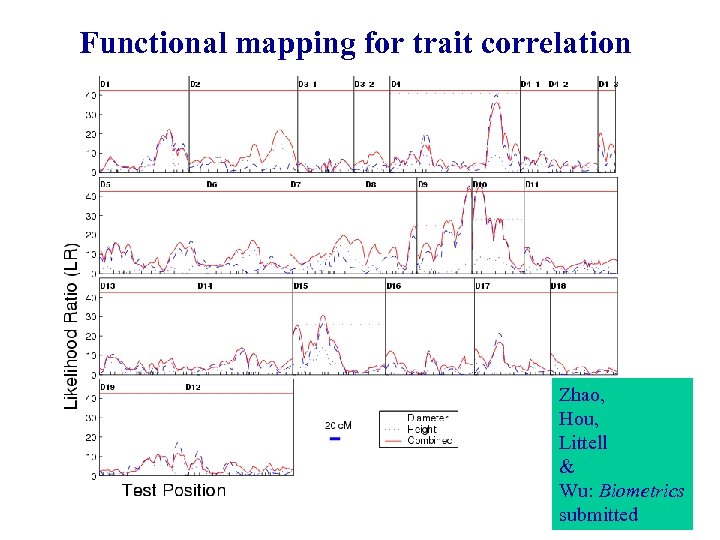 Functional mapping for trait correlation Zhao, Hou, Littell & Wu: Biometrics submitted
Functional mapping for trait correlation Zhao, Hou, Littell & Wu: Biometrics submitted
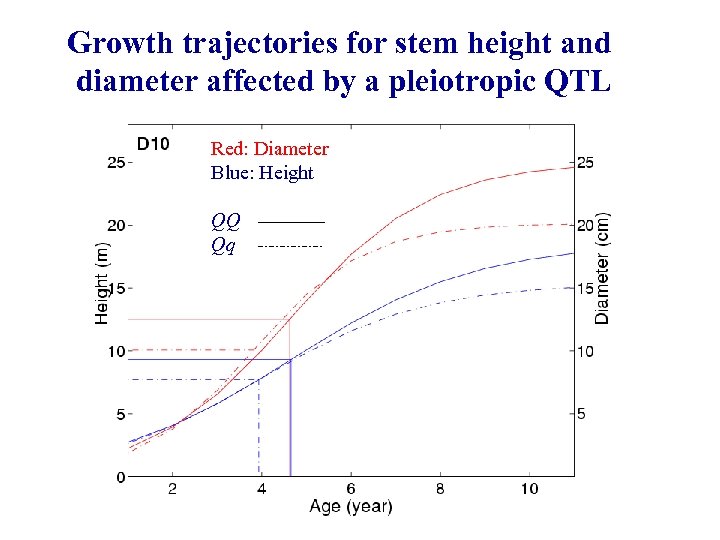 Growth trajectories for stem height and diameter affected by a pleiotropic QTL Red: Diameter Blue: Height QQ Qq
Growth trajectories for stem height and diameter affected by a pleiotropic QTL Red: Diameter Blue: Height QQ Qq
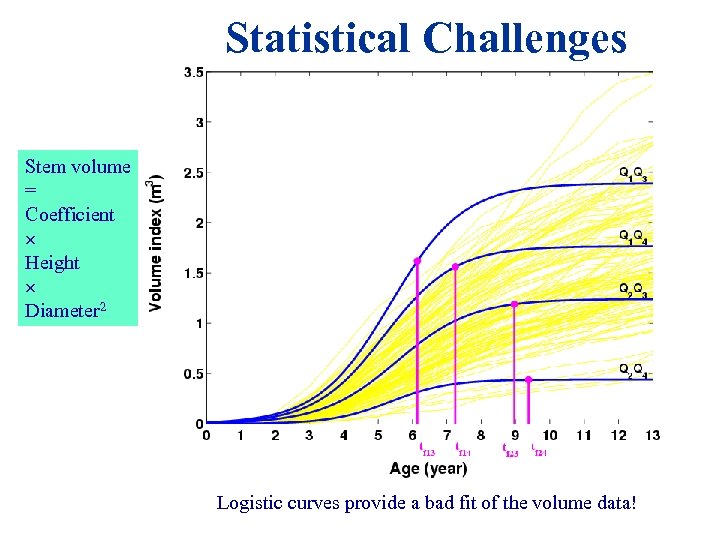 Statistical Challenges Stem volume = Coefficient Height Diameter 2 Logistic curves provide a bad fit of the volume data!
Statistical Challenges Stem volume = Coefficient Height Diameter 2 Logistic curves provide a bad fit of the volume data!
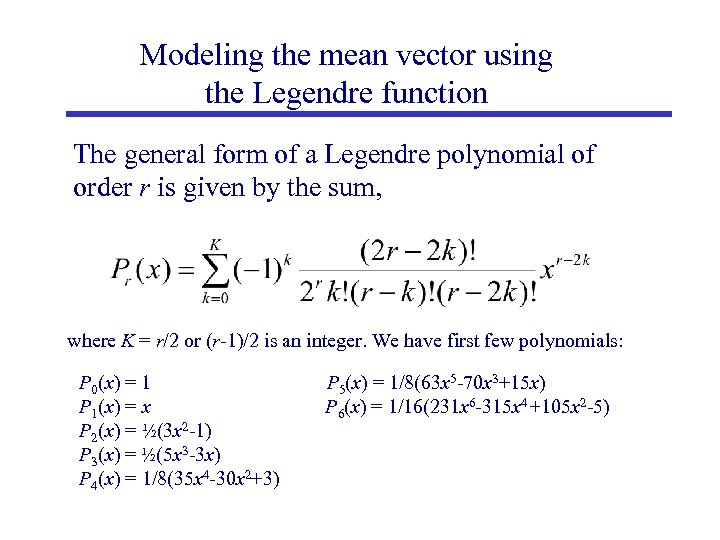 Modeling the mean vector using the Legendre function The general form of a Legendre polynomial of order r is given by the sum, where K = r/2 or (r-1)/2 is an integer. We have first few polynomials: P 0(x) = 1 P 1(x) = x P 2(x) = ½(3 x 2 -1) P 3(x) = ½(5 x 3 -3 x) P 4(x) = 1/8(35 x 4 -30 x 2+3) P 5(x) = 1/8(63 x 5 -70 x 3+15 x) P 6(x) = 1/16(231 x 6 -315 x 4+105 x 2 -5)
Modeling the mean vector using the Legendre function The general form of a Legendre polynomial of order r is given by the sum, where K = r/2 or (r-1)/2 is an integer. We have first few polynomials: P 0(x) = 1 P 1(x) = x P 2(x) = ½(3 x 2 -1) P 3(x) = ½(5 x 3 -3 x) P 4(x) = 1/8(35 x 4 -30 x 2+3) P 5(x) = 1/8(63 x 5 -70 x 3+15 x) P 6(x) = 1/16(231 x 6 -315 x 4+105 x 2 -5)
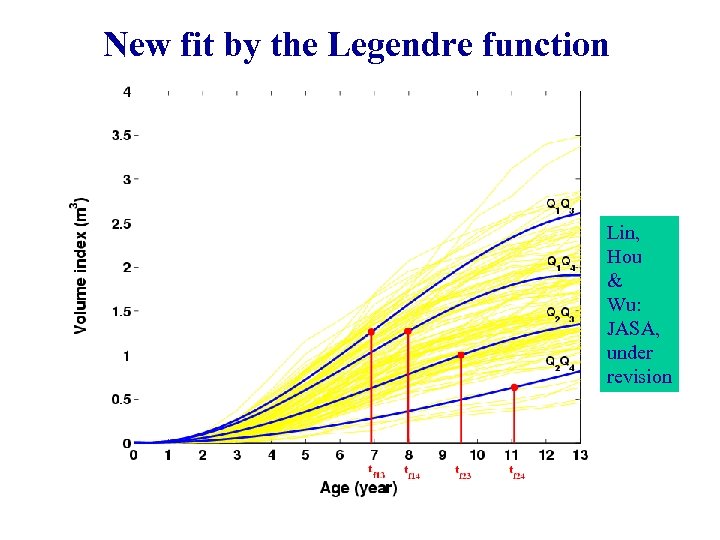 New fit by the Legendre function Lin, Hou & Wu: JASA, under revision
New fit by the Legendre function Lin, Hou & Wu: JASA, under revision
 Functional Mapping: toward high-dimensional biology • A new conceptual model for genetic mapping of complex traits • A systems approach for studying sophisticated biological problems • A framework for testing biological hypotheses at the interplay among genetics, development, physiology and biomedicine
Functional Mapping: toward high-dimensional biology • A new conceptual model for genetic mapping of complex traits • A systems approach for studying sophisticated biological problems • A framework for testing biological hypotheses at the interplay among genetics, development, physiology and biomedicine
 Functional Mapping: Simplicity from complexity • Estimating fewer biologically meaningful parameters that model the mean vector, • Modeling the structure of the variance matrix by developing powerful statistical methods, leading to few parameters to be estimated, • The reduction of dimension increases the power and precision of parameter estimation
Functional Mapping: Simplicity from complexity • Estimating fewer biologically meaningful parameters that model the mean vector, • Modeling the structure of the variance matrix by developing powerful statistical methods, leading to few parameters to be estimated, • The reduction of dimension increases the power and precision of parameter estimation
 Prospects
Prospects
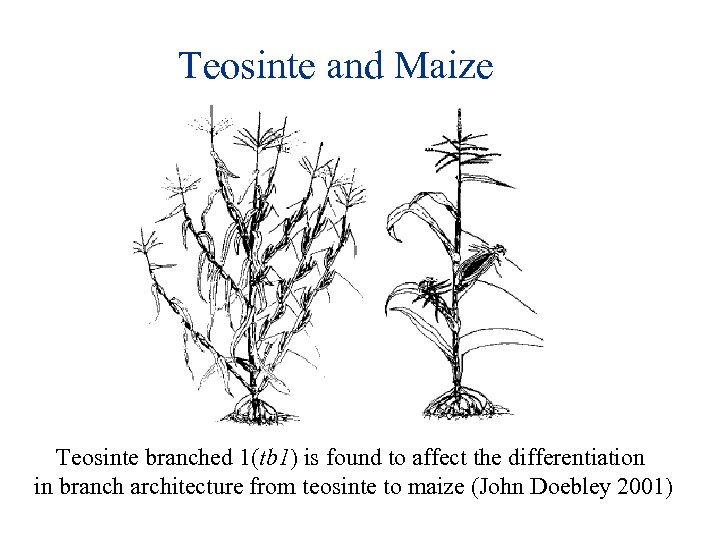 Teosinte and Maize Teosinte branched 1(tb 1) is found to affect the differentiation in branch architecture from teosinte to maize (John Doebley 2001)
Teosinte and Maize Teosinte branched 1(tb 1) is found to affect the differentiation in branch architecture from teosinte to maize (John Doebley 2001)
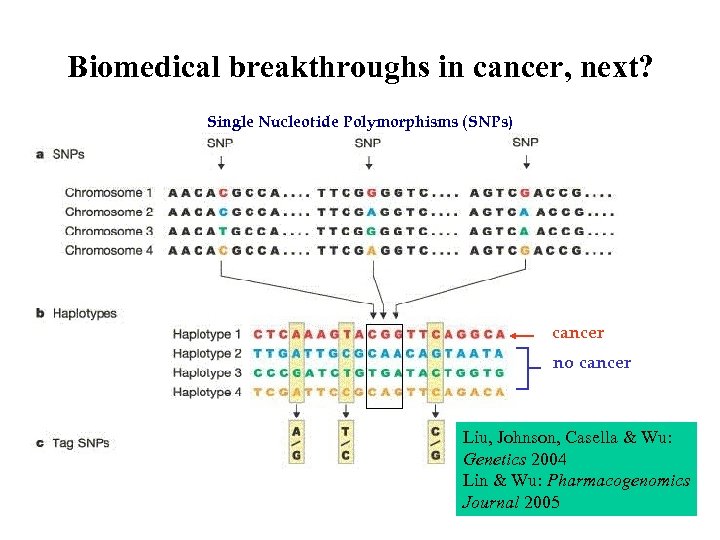 Biomedical breakthroughs in cancer, next? Single Nucleotide Polymorphisms (SNPs) cancer no cancer Liu, Johnson, Casella & Wu: Genetics 2004 Lin & Wu: Pharmacogenomics Journal 2005
Biomedical breakthroughs in cancer, next? Single Nucleotide Polymorphisms (SNPs) cancer no cancer Liu, Johnson, Casella & Wu: Genetics 2004 Lin & Wu: Pharmacogenomics Journal 2005


