7f3a22643dee893a02db5924d57fdca6.ppt
- Количество слайдов: 42
 High throughput gene synthesis and cloning of polyketide synthase modules Kosan Biosciences Sarah Reisinger
High throughput gene synthesis and cloning of polyketide synthase modules Kosan Biosciences Sarah Reisinger
 Kosan Business Technology platform polyketide alteration & production High value pharmaceuticals
Kosan Business Technology platform polyketide alteration & production High value pharmaceuticals
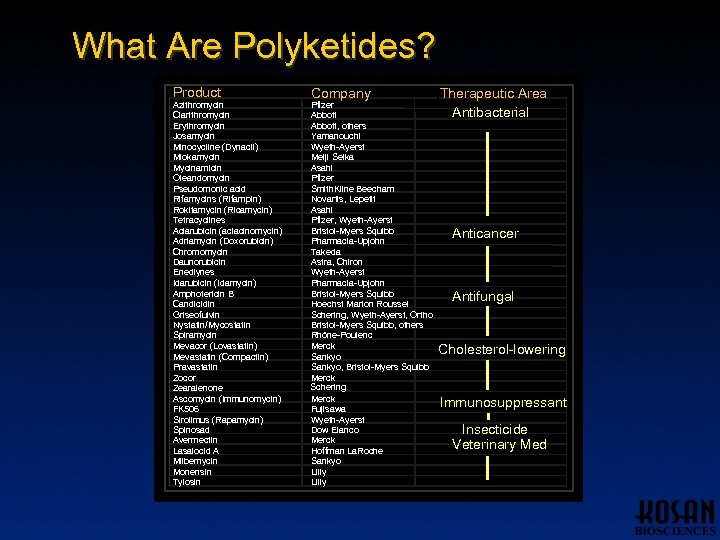 What Are Polyketides? Product Azithromycin Clarithromycin Erythromycin Josamycin Minocycline (Dynacil) Miokamycin Mycinamicin Oleandomycin Pseudomonic acid Rifamycins (Rifampin) Rokitamycin (Ricamycin) Tetracyclines Aclarubicin (aclacinomycin) Adriamycin (Doxorubicin) Chromomycin Daunorubicin Enediynes Idarubicin (Idamycin) Amphotericin B Candicidin Griseofulvin Nystatin/Mycostatin Spiramycin Mevacor (Lovastatin) Mevastatin (Compactin) Pravastatin Zocor Zearalenone Ascomycin (Immunomycin) FK 506 Sirolimus (Rapamycin) Spinosad Avermectin Lasalocid A Milbemycin Monensin Tylosin Company Pfizer Abbott, others Yamanouchi Wyeth-Ayerst Meiji Seika Asahi Pfizer Smith. Kline Beecham Novartis, Lepetit Asahi Pfizer, Wyeth-Ayerst Bristol-Myers Squibb Pharmacia-Upjohn Takeda Astra, Chiron Wyeth-Ayerst Pharmacia-Upjohn Bristol-Myers Squibb Hoechst Marion Roussel Schering, Wyeth-Ayerst, Ortho Bristol-Myers Squibb, others Rhône-Poulenc Merck Sankyo, Bristol-Myers Squibb Merck Schering Merck Fujisawa Wyeth-Ayerst Dow Elanco Merck Hoffman La. Roche Sankyo Lilly Therapeutic Area Antibacterial Anticancer Antifungal Cholesterol-lowering Immunosuppressant Insecticide Veterinary Med
What Are Polyketides? Product Azithromycin Clarithromycin Erythromycin Josamycin Minocycline (Dynacil) Miokamycin Mycinamicin Oleandomycin Pseudomonic acid Rifamycins (Rifampin) Rokitamycin (Ricamycin) Tetracyclines Aclarubicin (aclacinomycin) Adriamycin (Doxorubicin) Chromomycin Daunorubicin Enediynes Idarubicin (Idamycin) Amphotericin B Candicidin Griseofulvin Nystatin/Mycostatin Spiramycin Mevacor (Lovastatin) Mevastatin (Compactin) Pravastatin Zocor Zearalenone Ascomycin (Immunomycin) FK 506 Sirolimus (Rapamycin) Spinosad Avermectin Lasalocid A Milbemycin Monensin Tylosin Company Pfizer Abbott, others Yamanouchi Wyeth-Ayerst Meiji Seika Asahi Pfizer Smith. Kline Beecham Novartis, Lepetit Asahi Pfizer, Wyeth-Ayerst Bristol-Myers Squibb Pharmacia-Upjohn Takeda Astra, Chiron Wyeth-Ayerst Pharmacia-Upjohn Bristol-Myers Squibb Hoechst Marion Roussel Schering, Wyeth-Ayerst, Ortho Bristol-Myers Squibb, others Rhône-Poulenc Merck Sankyo, Bristol-Myers Squibb Merck Schering Merck Fujisawa Wyeth-Ayerst Dow Elanco Merck Hoffman La. Roche Sankyo Lilly Therapeutic Area Antibacterial Anticancer Antifungal Cholesterol-lowering Immunosuppressant Insecticide Veterinary Med
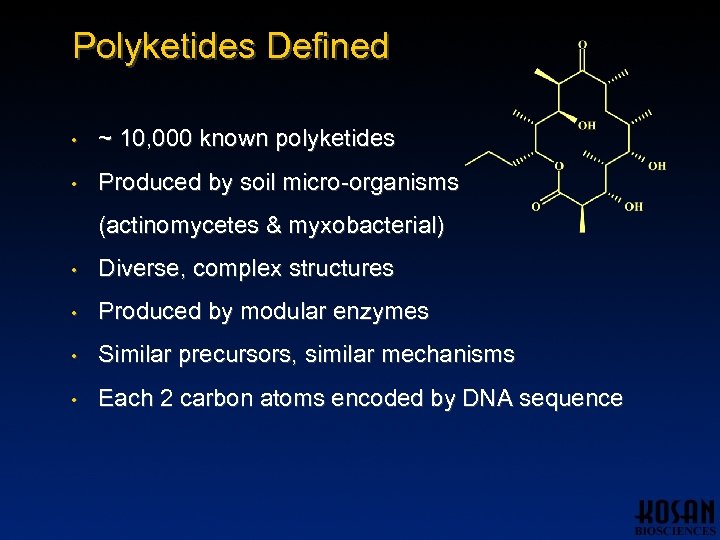 Polyketides Defined • ~ 10, 000 known polyketides • Produced by soil micro-organisms (actinomycetes & myxobacterial) • Diverse, complex structures • Produced by modular enzymes • Similar precursors, similar mechanisms • Each 2 carbon atoms encoded by DNA sequence
Polyketides Defined • ~ 10, 000 known polyketides • Produced by soil micro-organisms (actinomycetes & myxobacterial) • Diverse, complex structures • Produced by modular enzymes • Similar precursors, similar mechanisms • Each 2 carbon atoms encoded by DNA sequence
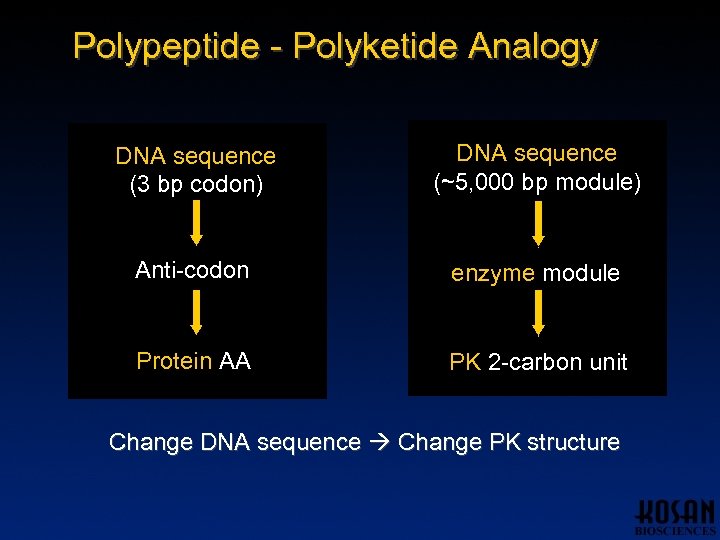 Polypeptide - Polyketide Analogy DNA sequence (3 bp codon) DNA sequence (~5, 000 bp module) Anti-codon enzyme module Protein AA PK 2 -carbon unit Change DNA sequence Change PK structure
Polypeptide - Polyketide Analogy DNA sequence (3 bp codon) DNA sequence (~5, 000 bp module) Anti-codon enzyme module Protein AA PK 2 -carbon unit Change DNA sequence Change PK structure
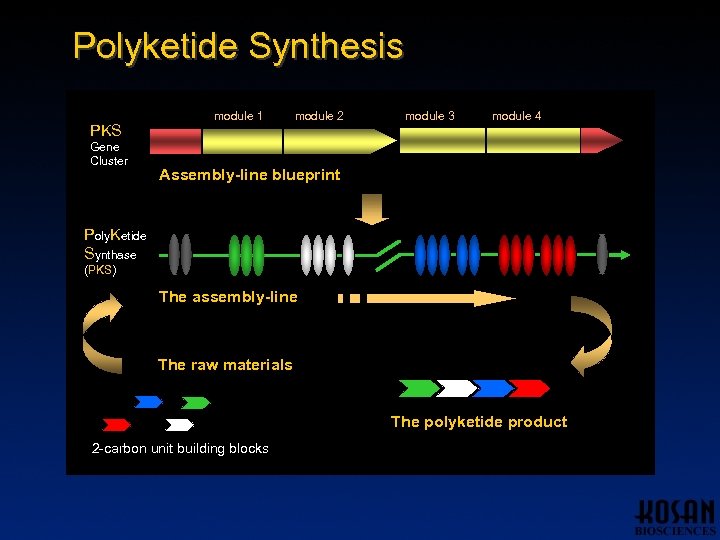 Polyketide Synthesis PKS Gene Cluster module 1 module 2 module 3 module 4 Assembly-line blueprint Poly. Ketide Synthase (PKS) The assembly-line The raw materials The polyketide product 2 -carbon unit building blocks
Polyketide Synthesis PKS Gene Cluster module 1 module 2 module 3 module 4 Assembly-line blueprint Poly. Ketide Synthase (PKS) The assembly-line The raw materials The polyketide product 2 -carbon unit building blocks
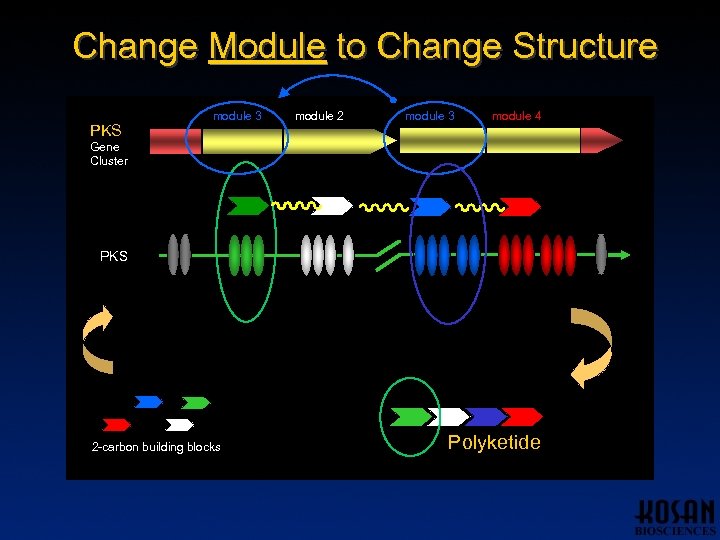 Change Module to Change Structure PKS module 1 module 3 module 2 module 3 module 4 Gene Cluster PKS 2 -carbon building blocks Polyketide
Change Module to Change Structure PKS module 1 module 3 module 2 module 3 module 4 Gene Cluster PKS 2 -carbon building blocks Polyketide
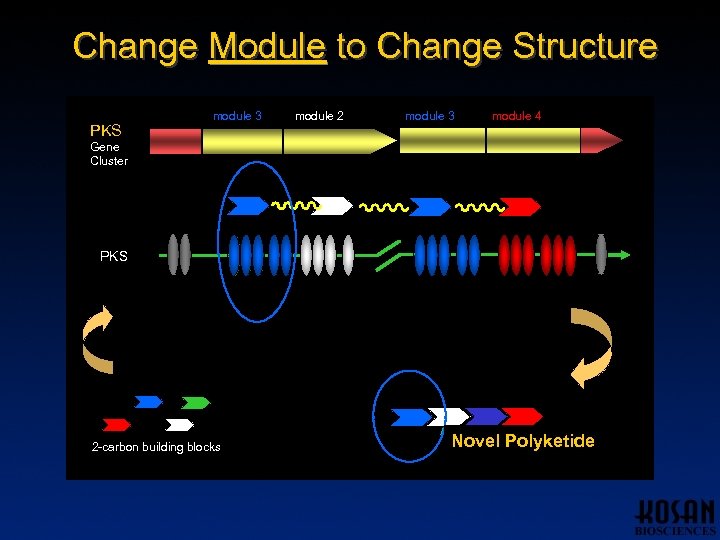 Change Module to Change Structure PKS module 1 module 3 module 2 module 3 module 4 Gene Cluster PKS 2 -carbon building blocks Novel Polyketide
Change Module to Change Structure PKS module 1 module 3 module 2 module 3 module 4 Gene Cluster PKS 2 -carbon building blocks Novel Polyketide
 Morphing • In theory, could sew PKS modules together to make any or many polyketides • In practice, difficult to obtain functional PKS module interactions
Morphing • In theory, could sew PKS modules together to make any or many polyketides • In practice, difficult to obtain functional PKS module interactions
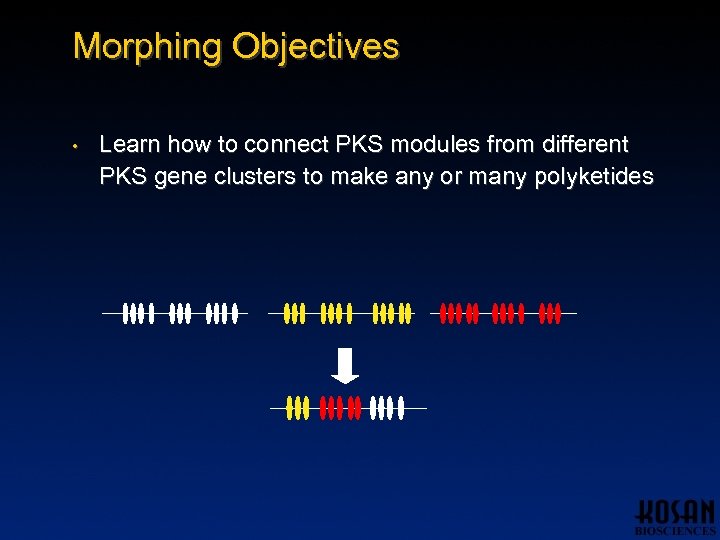 Morphing Objectives • Learn how to connect PKS modules from different PKS gene clusters to make any or many polyketides
Morphing Objectives • Learn how to connect PKS modules from different PKS gene clusters to make any or many polyketides
 Morphing Toolbox Objectives: • Develop a library of modules to express in genetic host • Connect modules in all permutations • Determine which module sets produce products • Learn how to correct inefficient module sets
Morphing Toolbox Objectives: • Develop a library of modules to express in genetic host • Connect modules in all permutations • Determine which module sets produce products • Learn how to correct inefficient module sets
 Develop a Library of Modules Possibilities: • Natural modules – Pros • Already exist • Synthetic genes – Pros • Control of G+C content; fewer expression problems • Designer restriction sites; simple to mobilize module/domains – Cons • Requires isolated genes • High G+C content; possible expression problems • No convenient restriction sites – Cons • Huge effort to create synthetic genes (100 modules = 500 kbp)
Develop a Library of Modules Possibilities: • Natural modules – Pros • Already exist • Synthetic genes – Pros • Control of G+C content; fewer expression problems • Designer restriction sites; simple to mobilize module/domains – Cons • Requires isolated genes • High G+C content; possible expression problems • No convenient restriction sites – Cons • Huge effort to create synthetic genes (100 modules = 500 kbp)
 High Throughput Gene Synthesis
High Throughput Gene Synthesis
 Objective To develop a fully automated process to quickly and efficiently synthesize and engineer large PKS. Input: Gene Sequence Gene Design Synthesis Output: Synthetic Gene of Interest
Objective To develop a fully automated process to quickly and efficiently synthesize and engineer large PKS. Input: Gene Sequence Gene Design Synthesis Output: Synthetic Gene of Interest
 Module Gene Design Develop a system for generating synthetic PKS modules that allows for: – Codon optimization for expression in E. coli – Common restriction sites at module and domain edges – Additional restriction sites within modules to facilitate partial domain or module swaps/replacements
Module Gene Design Develop a system for generating synthetic PKS modules that allows for: – Codon optimization for expression in E. coli – Common restriction sites at module and domain edges – Additional restriction sites within modules to facilitate partial domain or module swaps/replacements
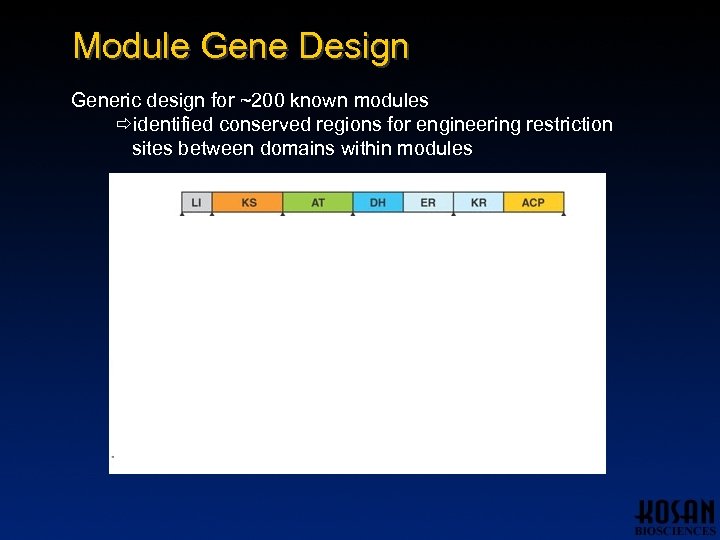 Module Gene Design Generic design for ~200 known modules identified conserved regions for engineering restriction sites between domains within modules
Module Gene Design Generic design for ~200 known modules identified conserved regions for engineering restriction sites between domains within modules
 Software Automation • Developed suite of tools for gene synthesis design and analysis – Synthetic gene design • Split gene into smaller parts, codon optimize, restriction sites • Oligo design/specificity testing/order • Automation input information – Sequence analysis – Database
Software Automation • Developed suite of tools for gene synthesis design and analysis – Synthetic gene design • Split gene into smaller parts, codon optimize, restriction sites • Oligo design/specificity testing/order • Automation input information – Sequence analysis – Database
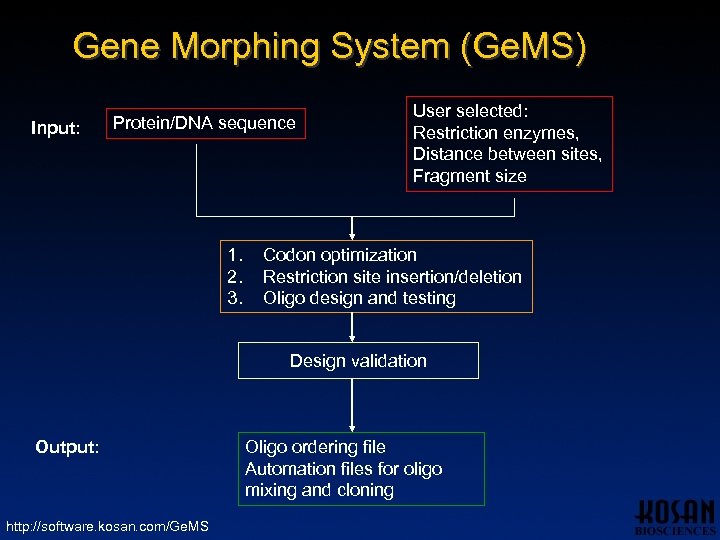 Gene Morphing System (Ge. MS) Input: Protein/DNA sequence 1. 2. 3. User selected: Restriction enzymes, Distance between sites, Fragment size Codon optimization Restriction site insertion/deletion Oligo design and testing Design validation Output: http: //software. kosan. com/Ge. MS Oligo ordering file Automation files for oligo mixing and cloning
Gene Morphing System (Ge. MS) Input: Protein/DNA sequence 1. 2. 3. User selected: Restriction enzymes, Distance between sites, Fragment size Codon optimization Restriction site insertion/deletion Oligo design and testing Design validation Output: http: //software. kosan. com/Ge. MS Oligo ordering file Automation files for oligo mixing and cloning
 Gene Synthesis: Fragment Generation Input: Oligo components of 500 bp synthons • Distribution of individual oligos to gene synthesis wells • Gene synthesis • Clone into vector • Transformation into E. coli • Isolation of colonies • DNA sequencing Output: 500 bp synthons in plasmids with correct sequence
Gene Synthesis: Fragment Generation Input: Oligo components of 500 bp synthons • Distribution of individual oligos to gene synthesis wells • Gene synthesis • Clone into vector • Transformation into E. coli • Isolation of colonies • DNA sequencing Output: 500 bp synthons in plasmids with correct sequence
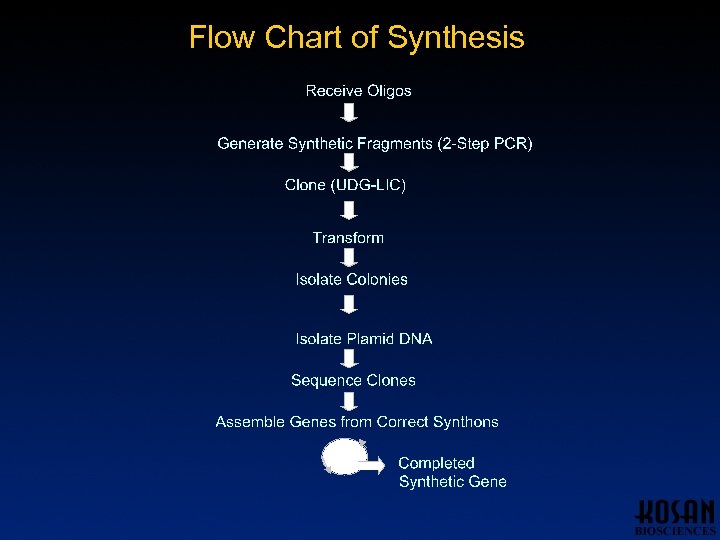 Flow Chart of Synthesis
Flow Chart of Synthesis
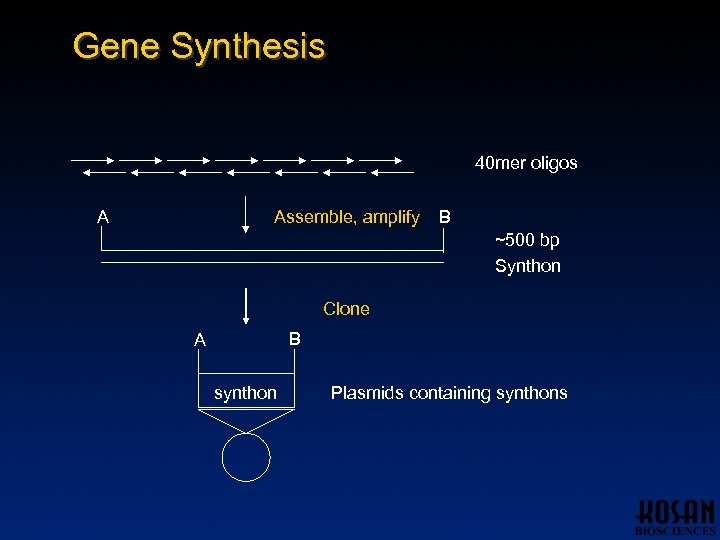 Gene Synthesis 40 mer oligos A Assemble, amplify B ~500 bp Synthon Clone B A synthon Plasmids containing synthons
Gene Synthesis 40 mer oligos A Assemble, amplify B ~500 bp Synthon Clone B A synthon Plasmids containing synthons
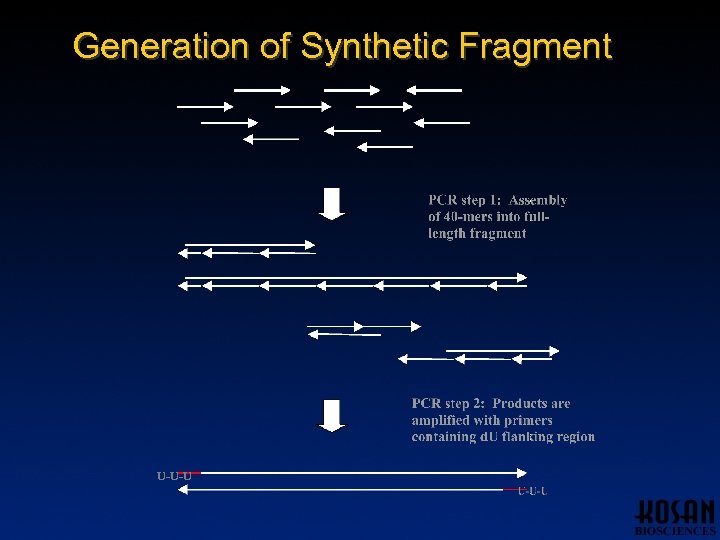 Generation of Synthetic Fragment U-U-U
Generation of Synthetic Fragment U-U-U
 HTP Cloning Criteria • Purification of PCR products unnecessary • High efficiency • Amenable to HTP automation
HTP Cloning Criteria • Purification of PCR products unnecessary • High efficiency • Amenable to HTP automation
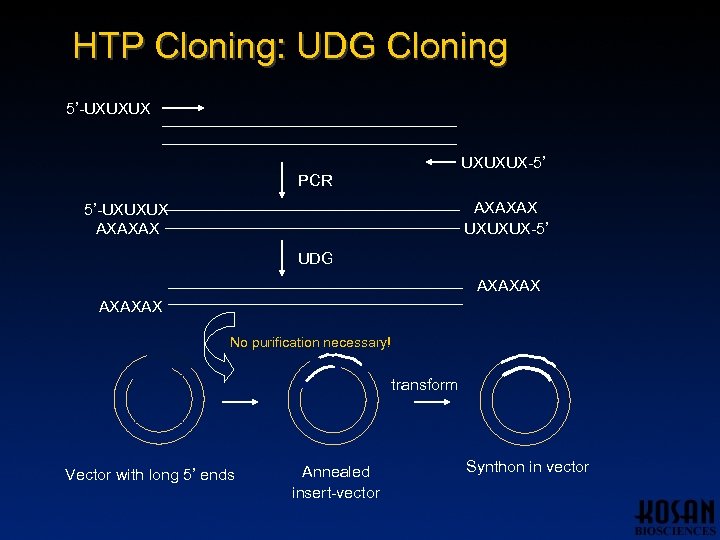 HTP Cloning: UDG Cloning 5’-UXUXUX-5’ PCR AXAXAX UXUXUX-5’ 5’-UXUXUX AXAXAX UDG AXAXAX No purification necessary! transform Vector with long 5’ ends Annealed insert-vector Synthon in vector
HTP Cloning: UDG Cloning 5’-UXUXUX-5’ PCR AXAXAX UXUXUX-5’ 5’-UXUXUX AXAXAX UDG AXAXAX No purification necessary! transform Vector with long 5’ ends Annealed insert-vector Synthon in vector
 Generation of Synthetic DNA • > 500 synthetic DNA fragments generated – 100% success rate – GC content from 44 -69% – Size between 129 and 1400 bp • Over 250, 000 bp synthesized • Average error rate around 1. 5 errors/kb • Fully automated most steps in process
Generation of Synthetic DNA • > 500 synthetic DNA fragments generated – 100% success rate – GC content from 44 -69% – Size between 129 and 1400 bp • Over 250, 000 bp synthesized • Average error rate around 1. 5 errors/kb • Fully automated most steps in process
 Gene Synthesis: Module Assembly Input: 500 bp synthons in plasmids with correct sequence • Digestion • Ligation • Transformation • Isolation of colonies • Verification of correct clone • Repeat until full-length gene assembled Output: Complete module (>5 kb) in plasmid with correct sequence
Gene Synthesis: Module Assembly Input: 500 bp synthons in plasmids with correct sequence • Digestion • Ligation • Transformation • Isolation of colonies • Verification of correct clone • Repeat until full-length gene assembled Output: Complete module (>5 kb) in plasmid with correct sequence
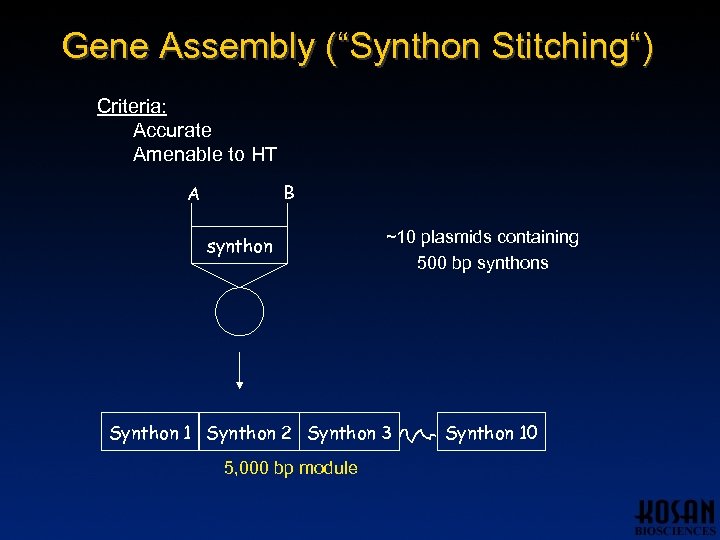 Gene Assembly (“Synthon Stitching“) Criteria: Accurate Amenable to HT B A synthon ~10 plasmids containing 500 bp synthons Synthon 1 Synthon 2 Synthon 3 5, 000 bp module Synthon 10
Gene Assembly (“Synthon Stitching“) Criteria: Accurate Amenable to HT B A synthon ~10 plasmids containing 500 bp synthons Synthon 1 Synthon 2 Synthon 3 5, 000 bp module Synthon 10
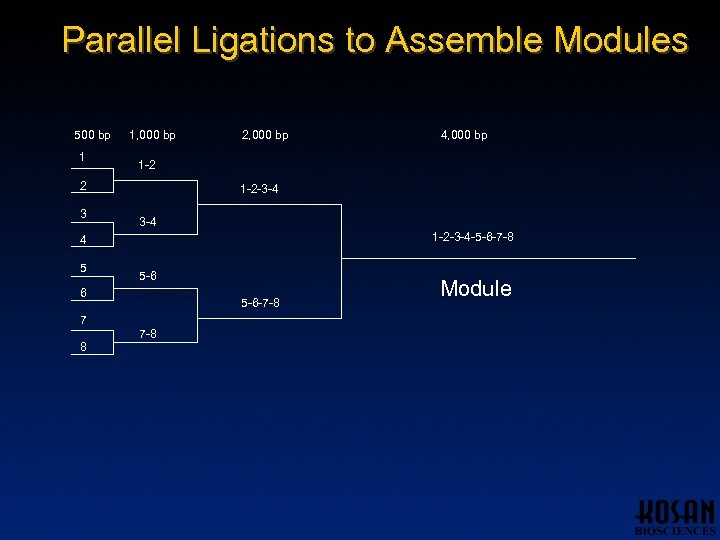 Parallel Ligations to Assemble Modules 500 bp 1 1, 000 bp 1 -2 -3 -4 -5 -6 -7 -8 4 5 5 -6 6 5 -6 -7 -8 7 8 4, 000 bp 1 -2 2 3 2, 000 bp 7 -8 Module
Parallel Ligations to Assemble Modules 500 bp 1 1, 000 bp 1 -2 -3 -4 -5 -6 -7 -8 4 5 5 -6 6 5 -6 -7 -8 7 8 4, 000 bp 1 -2 2 3 2, 000 bp 7 -8 Module
 Synthon Stitching Method • Utilize Type IIs restriction enzymes – Cut DNA outside of recognition site Bsa I: 5´. . . G G T C (N)1^. . . 3´ 3´. . . C C A G (N)5^. . . 5´ – Use different Type IIs enzymes to create compatible overhangs – Same enzymes can be used for all synthon pairs to facilitate automation
Synthon Stitching Method • Utilize Type IIs restriction enzymes – Cut DNA outside of recognition site Bsa I: 5´. . . G G T C (N)1^. . . 3´ 3´. . . C C A G (N)5^. . . 5´ – Use different Type IIs enzymes to create compatible overhangs – Same enzymes can be used for all synthon pairs to facilitate automation
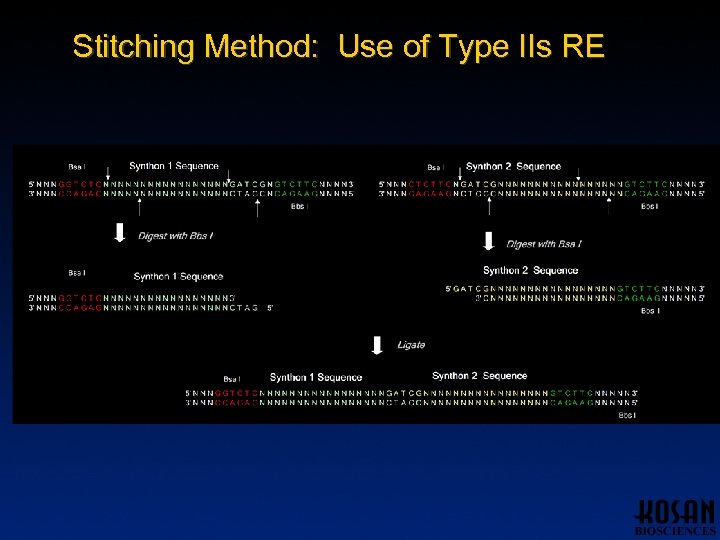 Stitching Method: Use of Type IIs RE
Stitching Method: Use of Type IIs RE
 Synthon Stitching Method • Unique selectable markers on two sister plasmids eliminates need for purification of fragments
Synthon Stitching Method • Unique selectable markers on two sister plasmids eliminates need for purification of fragments
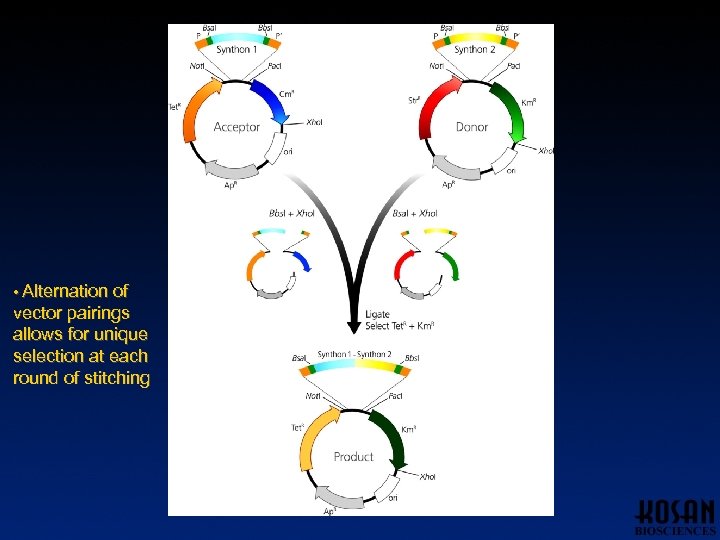 • Alternation of vector pairings allows for unique selection at each round of stitching
• Alternation of vector pairings allows for unique selection at each round of stitching
 Results of Synthon Stitching • 26 complete modules constructed • > 250 successful ligations • Selection scheme works extremely well – Majority of ligations performed gave only correct product • Use of Type IIs enzymes makes method amenable to automation
Results of Synthon Stitching • 26 complete modules constructed • > 250 successful ligations • Selection scheme works extremely well – Majority of ligations performed gave only correct product • Use of Type IIs enzymes makes method amenable to automation
 Improvements of Gene Synthesis: Designer Vectors • 3 -plasmid system for synthon stitching – Counter-selectable markers – Allows 4 -piece ligations of unpurified digests
Improvements of Gene Synthesis: Designer Vectors • 3 -plasmid system for synthon stitching – Counter-selectable markers – Allows 4 -piece ligations of unpurified digests
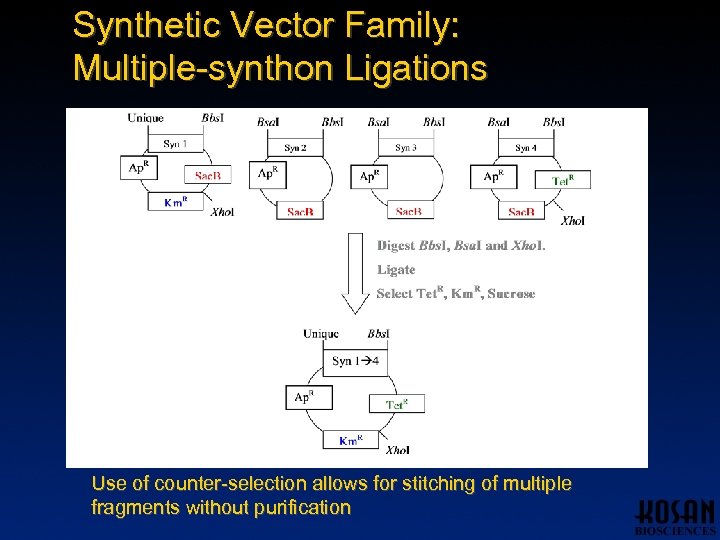 Synthetic Vector Family: Multiple-synthon Ligations Use of counter-selection allows for stitching of multiple fragments without purification
Synthetic Vector Family: Multiple-synthon Ligations Use of counter-selection allows for stitching of multiple fragments without purification
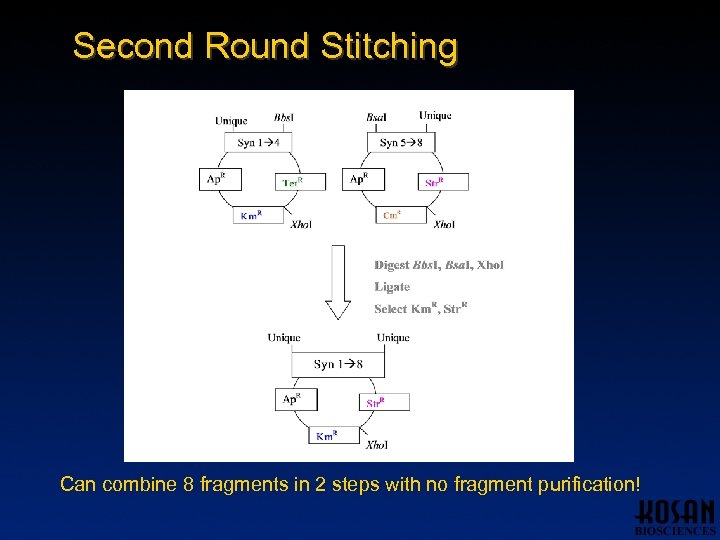 Second Round Stitching Can combine 8 fragments in 2 steps with no fragment purification!
Second Round Stitching Can combine 8 fragments in 2 steps with no fragment purification!
 Testing of Modules
Testing of Modules
 Proof of Concept • Expressed synthetic 6 -module DEBS gene cluster in E. coli • Protein subunits observed on SDS-PAGE in the soluble fraction • Product (6 -d. EB) identified by LC-MS
Proof of Concept • Expressed synthetic 6 -module DEBS gene cluster in E. coli • Protein subunits observed on SDS-PAGE in the soluble fraction • Product (6 -d. EB) identified by LC-MS
 Results of Module Testing • Tested 14 synthetic modules in 154 bimodular combinations • 72 of the 154 combinations tested produced measurable triketide lactone • All modules tested worked
Results of Module Testing • Tested 14 synthetic modules in 154 bimodular combinations • 72 of the 154 combinations tested produced measurable triketide lactone • All modules tested worked
 Summary • Successfully developed method for high throughput gene synthesis • High-throughput method for assembly of DNA fragments into larger genes (modules) developed • Populated module library and tested in bimodular cases
Summary • Successfully developed method for high throughput gene synthesis • High-throughput method for assembly of DNA fragments into larger genes (modules) developed • Populated module library and tested in bimodular cases
 Acknowledgements Kosan Biosciences – Morphing Group • Dan Santi • Ralph Reid • Kedar Patel • Sebastian Jayaraj • Hugo Menzella • Sunil Chandran
Acknowledgements Kosan Biosciences – Morphing Group • Dan Santi • Ralph Reid • Kedar Patel • Sebastian Jayaraj • Hugo Menzella • Sunil Chandran
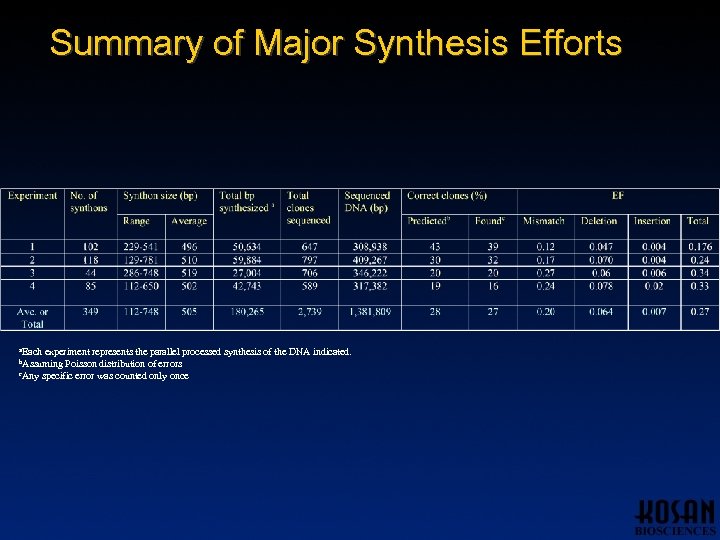 Summary of Major Synthesis Efforts a. Each experiment represents the parallel processed synthesis of the DNA indicated. Poisson distribution of errors c. Any specific error was counted only once b. Assuming
Summary of Major Synthesis Efforts a. Each experiment represents the parallel processed synthesis of the DNA indicated. Poisson distribution of errors c. Any specific error was counted only once b. Assuming


