 Haloferax sulfurifontis sp. nov. , a Halophilic Archaeon Isolated from a Low Salt Sulfide-rich Spring K. N. Savage 1, M. S. Elshahed 1, A. Oren 2, L. R. Krumholz 1 1 University of Oklahoma, Norman, OK, 2 The Hebrew University of Jerusalem, ISRAEL
Haloferax sulfurifontis sp. nov. , a Halophilic Archaeon Isolated from a Low Salt Sulfide-rich Spring K. N. Savage 1, M. S. Elshahed 1, A. Oren 2, L. R. Krumholz 1 1 University of Oklahoma, Norman, OK, 2 The Hebrew University of Jerusalem, ISRAEL
 Zodletone Spring http: //fermi. jhuapl. edu/states/maps 1/ok. gif • 8 l • min-1 • 20 m from source to creek • High sulfide concentrations (810 m. M) • 0. 2 M Na. Cl • F, Br, Li, B, Sr, Ba, SO 42 • Ba. SO 4 and Ca. CO 3 • Dissolved sulfur
Zodletone Spring http: //fermi. jhuapl. edu/states/maps 1/ok. gif • 8 l • min-1 • 20 m from source to creek • High sulfide concentrations (810 m. M) • 0. 2 M Na. Cl • F, Br, Li, B, Sr, Ba, SO 42 • Ba. SO 4 and Ca. CO 3 • Dissolved sulfur
 Archaeal Diversity
Archaeal Diversity
 Halophilic Archaea • 14 validly described genera • Isolated from hypersaline environments (i. e. The Dead Sea) • Rods, cocci and pleomorphic forms • Require high salt concentrations from 3. 5 M Na. Cl to saturation (5. 2 M) • Contain pigment material (bacteriorhodopsin and halorhodopsin) which gives them a distinct pink or red color
Halophilic Archaea • 14 validly described genera • Isolated from hypersaline environments (i. e. The Dead Sea) • Rods, cocci and pleomorphic forms • Require high salt concentrations from 3. 5 M Na. Cl to saturation (5. 2 M) • Contain pigment material (bacteriorhodopsin and halorhodopsin) which gives them a distinct pink or red color
 Isolation of Haloarchaea • 36% of the mat and 4% of the source library • Isolated halophiles from the mat using a high salt plus antibiotic medium with different salt concentrations (7, 12 and 18%) • Serially diluted material and plated onto HM plates (immediately and after incubation) • Cultures were incubated at 37ºC under light
Isolation of Haloarchaea • 36% of the mat and 4% of the source library • Isolated halophiles from the mat using a high salt plus antibiotic medium with different salt concentrations (7, 12 and 18%) • Serially diluted material and plated onto HM plates (immediately and after incubation) • Cultures were incubated at 37ºC under light
 A Novel Species of Haloferax • 16 S r. RNA sequence analysis of strain M 6 showed a 96. 7 -98. 0% similarity to other validly described species of this genus • M 6 was 89% similar to Halogeometricum borinquense the most closely related species outside the genus • DNA-DNA hybridization confirmed its novel speciation having a hybridization value of only 24% to Haloferax gibbonsii
A Novel Species of Haloferax • 16 S r. RNA sequence analysis of strain M 6 showed a 96. 7 -98. 0% similarity to other validly described species of this genus • M 6 was 89% similar to Halogeometricum borinquense the most closely related species outside the genus • DNA-DNA hybridization confirmed its novel speciation having a hybridization value of only 24% to Haloferax gibbonsii
 Phylogeny
Phylogeny
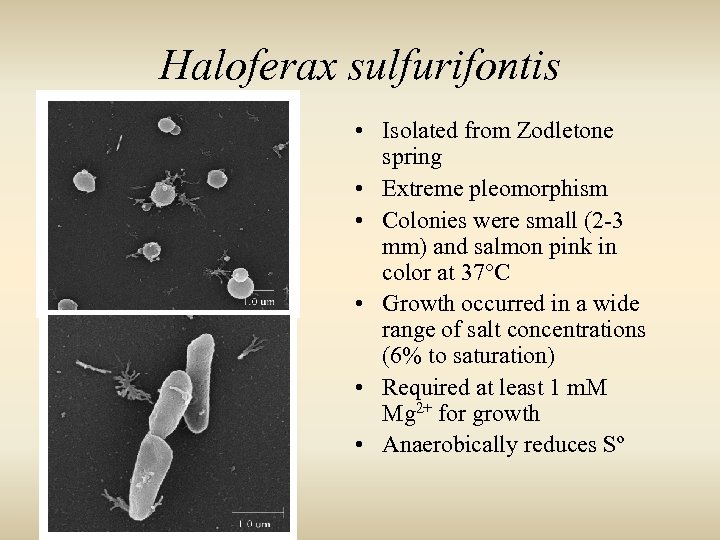 Haloferax sulfurifontis • Isolated from Zodletone spring • Extreme pleomorphism • Colonies were small (2 -3 mm) and salmon pink in color at 37°C • Growth occurred in a wide range of salt concentrations (6% to saturation) • Required at least 1 m. M Mg 2+ for growth • Anaerobically reduces Sº
Haloferax sulfurifontis • Isolated from Zodletone spring • Extreme pleomorphism • Colonies were small (2 -3 mm) and salmon pink in color at 37°C • Growth occurred in a wide range of salt concentrations (6% to saturation) • Required at least 1 m. M Mg 2+ for growth • Anaerobically reduces Sº
 Haloferax sulfurifontis
Haloferax sulfurifontis
 A Continued Search for Halophiles • Clone libraries indicated the presence of a diverse halophilic community • Originally 18 strains were isolated from the mats present at the stream • Studies indicated that these isolates were of the same species • In order to encourage the growth of different isolates the media was prepared at 3 different salt concentrations (18, 25 and 30%) and 11 different carbon sources were used instead of yeast extract
A Continued Search for Halophiles • Clone libraries indicated the presence of a diverse halophilic community • Originally 18 strains were isolated from the mats present at the stream • Studies indicated that these isolates were of the same species • In order to encourage the growth of different isolates the media was prepared at 3 different salt concentrations (18, 25 and 30%) and 11 different carbon sources were used instead of yeast extract
 Future Work • Microscopic inspection of colonies to determine cell morphologies present and trends • Further purification and isolation • Colony PCR and sequencing • Anaerobic reduction of S° • Strain characterizations
Future Work • Microscopic inspection of colonies to determine cell morphologies present and trends • Further purification and isolation • Colony PCR and sequencing • Anaerobic reduction of S° • Strain characterizations
 Acknowledgements • • Dr. Krumholz Dr. Elshahed Aharon Oren and Antonio Ventosa Roe Laboratory Funding for this project is by the National Science Foundation
Acknowledgements • • Dr. Krumholz Dr. Elshahed Aharon Oren and Antonio Ventosa Roe Laboratory Funding for this project is by the National Science Foundation