f0933fa697f3f1e2303e6040b665158f.ppt
- Количество слайдов: 26
 Genomic ORFans: Past, Present and Future Naomi Siew and Daniel Fischer Ben-Gurion University Be’er-Sheva, Israel
Genomic ORFans: Past, Present and Future Naomi Siew and Daniel Fischer Ben-Gurion University Be’er-Sheva, Israel
 1995: The Genomic Revolution • Dozens of genomes were fully sequenced • Dozens more are underway ORF – Open Reading Frame start codon ……… stop codon
1995: The Genomic Revolution • Dozens of genomes were fully sequenced • Dozens more are underway ORF – Open Reading Frame start codon ……… stop codon
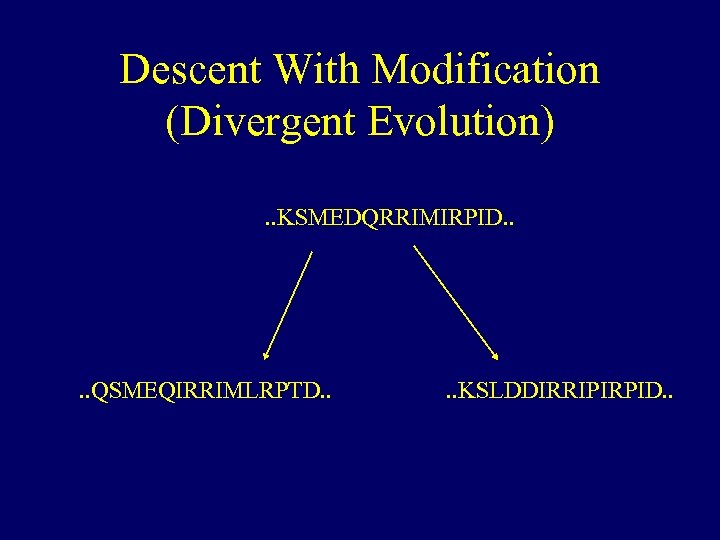 Descent With Modification (Divergent Evolution). . KSMEDQRRIMIRPID. . QSMEQIRRIMLRPTD. . KSLDDIRRIPIRPID. .
Descent With Modification (Divergent Evolution). . KSMEDQRRIMIRPID. . QSMEQIRRIMLRPTD. . KSLDDIRRIPIRPID. .
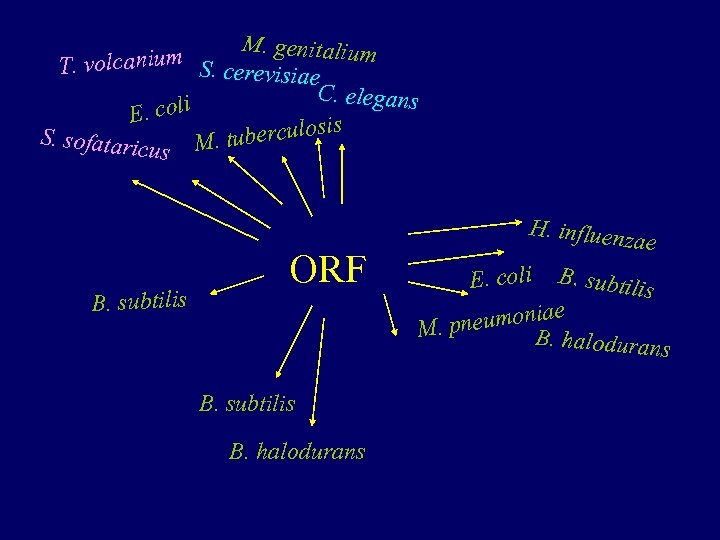 M. genitalium T. volcanium S. cerevisiae C. elegans oli E. c erculosis b S. sofatari cus M. tu B. subtilis ORF B. subtilis B. halodurans H. influenz ae E. coli B. subtilis e neumonia M. p B. halodurans
M. genitalium T. volcanium S. cerevisiae C. elegans oli E. c erculosis b S. sofatari cus M. tu B. subtilis ORF B. subtilis B. halodurans H. influenz ae E. coli B. subtilis e neumonia M. p B. halodurans
 Orphan ORFs = ORFans (Fischer and Eisenberg, Bioinformatics, 15(9), 1999) Singleton ORFan : An ORF that has no sequence similarity to any other sequence in the databases. Little can be inferred about ORFans using bioinformatic tools.
Orphan ORFs = ORFans (Fischer and Eisenberg, Bioinformatics, 15(9), 1999) Singleton ORFan : An ORF that has no sequence similarity to any other sequence in the databases. Little can be inferred about ORFans using bioinformatic tools.
 20 -30% of ORFs in each new genome are singleton ORFans.
20 -30% of ORFs in each new genome are singleton ORFans.
 ORFans May Be… • New, previously unseen proteins, (with new function, new structure) unique to one organism (species-specific). • Distant relatives of known families (similar function, similar 3 D structure) whose sequence diverged beyond recognition by sequence comparison tools.
ORFans May Be… • New, previously unseen proteins, (with new function, new structure) unique to one organism (species-specific). • Distant relatives of known families (similar function, similar 3 D structure) whose sequence diverged beyond recognition by sequence comparison tools.
 The Puzzle of ORFans • If new ORFs, where did they come from? How did they evolve? • If distant relatives, why aren’t there similar sequences? Where are the intermediates?
The Puzzle of ORFans • If new ORFs, where did they come from? How did they evolve? • If distant relatives, why aren’t there similar sequences? Where are the intermediates?
 Census and Dynamics of ORFans • Built a database of fully sequenced genomes. • Added genomes one by one in chronological order of publication. • For each ORF, ran BLAST: if there is a match non-ORFan if there is no match ORFan Previous ORFans can become non-ORFans.
Census and Dynamics of ORFans • Built a database of fully sequenced genomes. • Added genomes one by one in chronological order of publication. • For each ORF, ran BLAST: if there is a match non-ORFan if there is no match ORFan Previous ORFans can become non-ORFans.
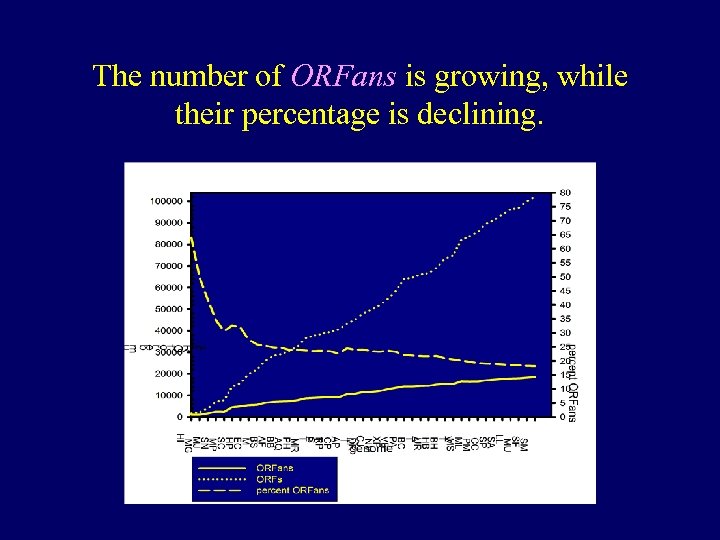 The number of ORFans is growing, while their percentage is declining.
The number of ORFans is growing, while their percentage is declining.
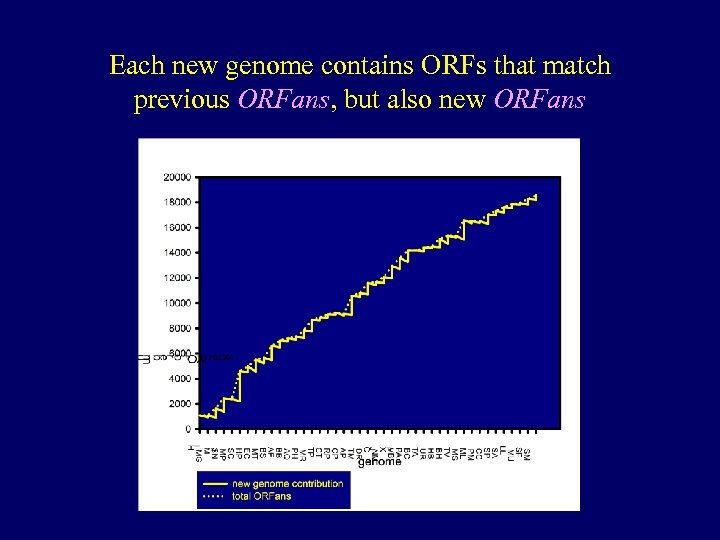 Each new genome contains ORFs that match previous ORFans, but also new ORFans
Each new genome contains ORFs that match previous ORFans, but also new ORFans
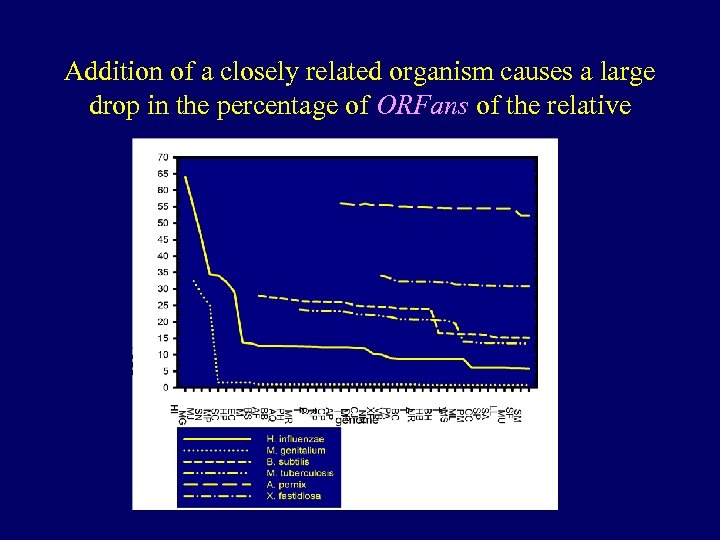 Addition of a closely related organism causes a large drop in the percentage of ORFans of the relative
Addition of a closely related organism causes a large drop in the percentage of ORFans of the relative
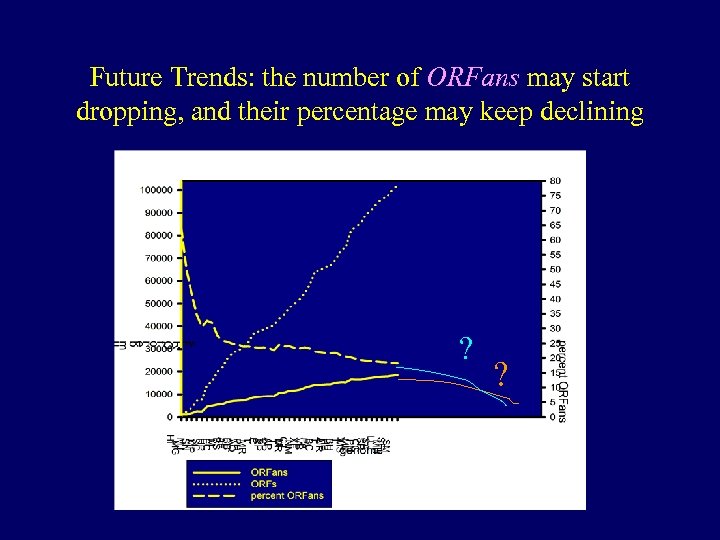 Future Trends: the number of ORFans may start dropping, and their percentage may keep declining ? ?
Future Trends: the number of ORFans may start dropping, and their percentage may keep declining ? ?
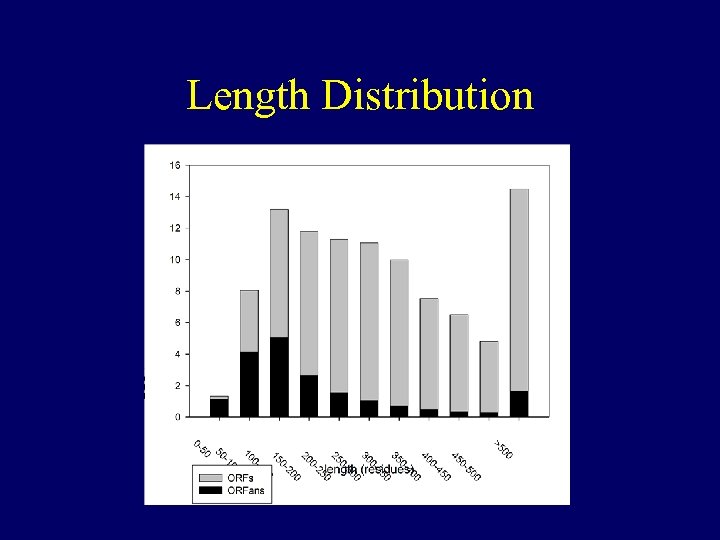 Length Distribution
Length Distribution
 Length Bias • Bias among short sequences for ORFans. (almost half of short sequences are ORFans) • Bias among ORFans for short sequences. (half of ORFans are short)
Length Bias • Bias among short sequences for ORFans. (almost half of short sequences are ORFans) • Bias among ORFans for short sequences. (half of ORFans are short)
 Separate dynamics analyses of short and long ORFans show different behaviors • Percentage of short ORFans is declining more slowly. Possible explanations: not expressed; frame shifts; wrong stop codons; technical limitations. • Percentage of long ORFans is declining faster. Possible explanations: more conserved; ORFan modules.
Separate dynamics analyses of short and long ORFans show different behaviors • Percentage of short ORFans is declining more slowly. Possible explanations: not expressed; frame shifts; wrong stop codons; technical limitations. • Percentage of long ORFans is declining faster. Possible explanations: more conserved; ORFan modules.
 ORFan Modules MGTGDKFCKDKIECAPL KFSRDKIECAFLHGRFCGDGSP GEISFLIGGRYL ORFan Module: A segment of a sequence that has no matches with other sequences.
ORFan Modules MGTGDKFCKDKIECAPL KFSRDKIECAFLHGRFCGDGSP GEISFLIGGRYL ORFan Module: A segment of a sequence that has no matches with other sequences.
 Interim Conclusions • Evolution has left us with two types of sequences: homologs and ORFans. • The number of singleton ORFans has been growing. • Their percentage is diminishing.
Interim Conclusions • Evolution has left us with two types of sequences: homologs and ORFans. • The number of singleton ORFans has been growing. • Their percentage is diminishing.
 Interim Conclusions II • There is a bias towards short sequences among singleton ORFans, and vice versa. • Most longer singleton ORFans may disappear with time. • New genomes of closely related organisms will have fewer singleton ORFans.
Interim Conclusions II • There is a bias towards short sequences among singleton ORFans, and vice versa. • Most longer singleton ORFans may disappear with time. • New genomes of closely related organisms will have fewer singleton ORFans.
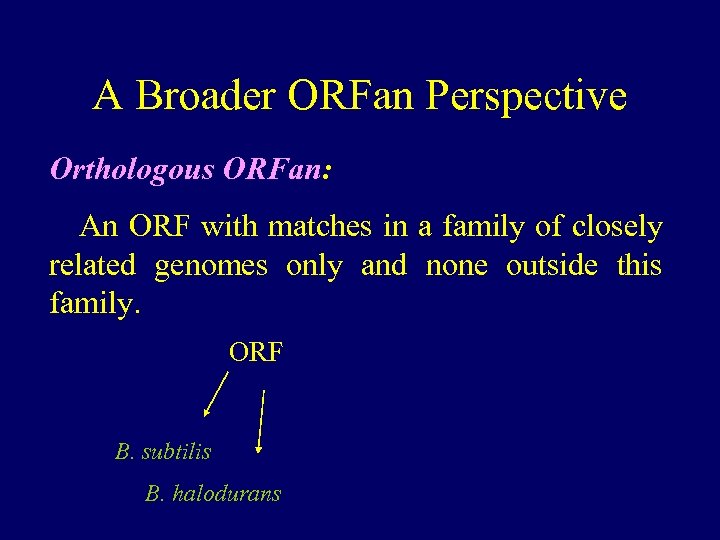 A Broader ORFan Perspective Orthologous ORFan: An ORF with matches in a family of closely related genomes only and none outside this family. ORF B. subtilis B. halodurans
A Broader ORFan Perspective Orthologous ORFan: An ORF with matches in a family of closely related genomes only and none outside this family. ORF B. subtilis B. halodurans
 • Currently orthologous ORFans are counted as non-ORFans. • Family-specific? • Most probably expressed proteins.
• Currently orthologous ORFans are counted as non-ORFans. • Family-specific? • Most probably expressed proteins.
 Paralogous ORFan: An ORF with matches in the same genome only and none outside the genome.
Paralogous ORFan: An ORF with matches in the same genome only and none outside the genome.
 • Currently paralogous ORFans are counted as non-ORFans. • Species-specific? • Most probably expressed proteins.
• Currently paralogous ORFans are counted as non-ORFans. • Species-specific? • Most probably expressed proteins.
 Future and On-Going Work • • • Study the other types of ORFans (orthologous, paralogous, modules). Try to assign distantly related ORFans to known families: * in silico: using more sensitive bioinformatic tools such as fold recognition. * In the lab: determining the 3 D structure of selected ORFans. However, even if all ORFans were assigned to known families, the puzzle of their evolution will still remain.
Future and On-Going Work • • • Study the other types of ORFans (orthologous, paralogous, modules). Try to assign distantly related ORFans to known families: * in silico: using more sensitive bioinformatic tools such as fold recognition. * In the lab: determining the 3 D structure of selected ORFans. However, even if all ORFans were assigned to known families, the puzzle of their evolution will still remain.
 Ongoing in silico/experimental ORFan studies in BGU • Mini-structural genomics project to study selected paralogous ORFans in the archeon Halobacterium NRC-15. Bioinformatics (our group) Archea biology (Dr. Gerry Eichler) Crystallography (Prof. Boaz Sha’anan)
Ongoing in silico/experimental ORFan studies in BGU • Mini-structural genomics project to study selected paralogous ORFans in the archeon Halobacterium NRC-15. Bioinformatics (our group) Archea biology (Dr. Gerry Eichler) Crystallography (Prof. Boaz Sha’anan)
 Acknowledgements Prof. Joel Bernstein Department of Chemistry, BGU
Acknowledgements Prof. Joel Bernstein Department of Chemistry, BGU


