8f401c8261665a99d06065b0368d02d0.ppt
- Количество слайдов: 27

Genome Bioinformatics Tyler Alioto Center for Genomic Regulation Barcelona, Spain Jul-01 -0806/16/08 Bioinformatics Workshop - Malaga

Node 1 of the INB GN 1 Bioinformática y Genómica Genome Bioinformatic Lab, CRG Roderic Guigó (PI) Jul-01 -08 Bioinformatics Workshop - Malaga

Themes Gene prediction Genome feature visualization gff 2 ps Alternative splicing ab initio => Gene. ID dual-genome => SGP 2 u 12 introns => Gene. ID v 1. 3 and U 12 DB combiner => Gene. PC ASTALAVISTA Gene expression regulatory elements meta and mmeta alignment Jul-01 -08 Bioinformatics Workshop - Malaga
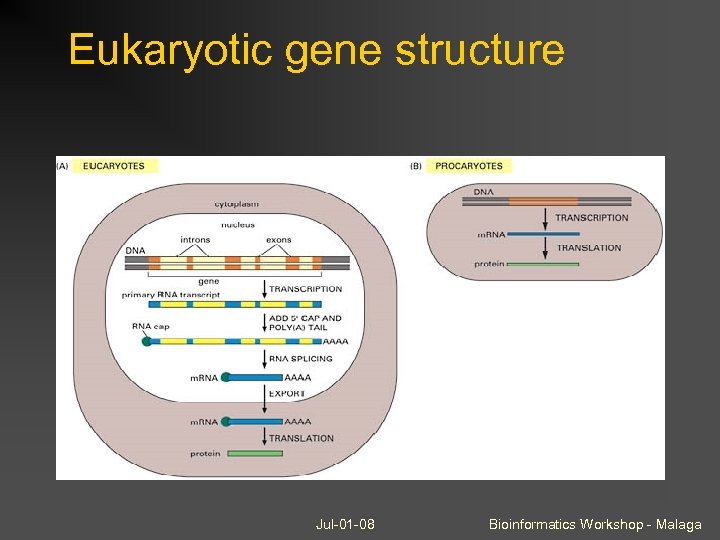
Eukaryotic gene structure Jul-01 -08 Bioinformatics Workshop - Malaga
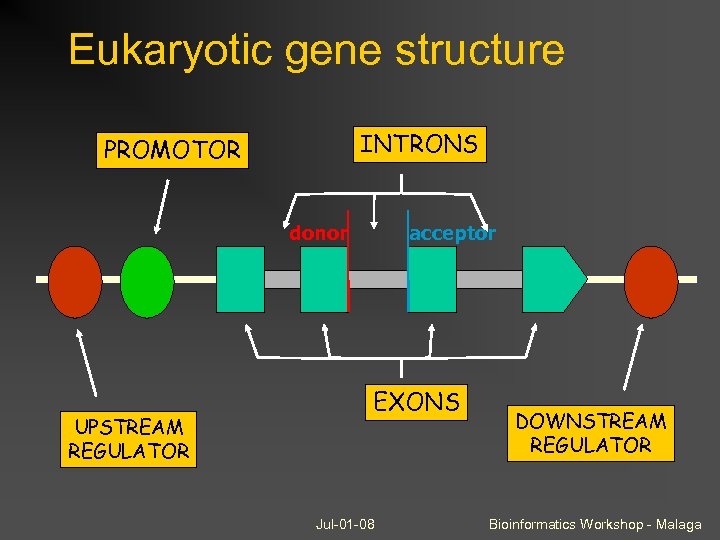
Eukaryotic gene structure INTRONS PROMOTOR donor UPSTREAM REGULATOR acceptor EXONS Jul-01 -08 DOWNSTREAM REGULATOR Bioinformatics Workshop - Malaga
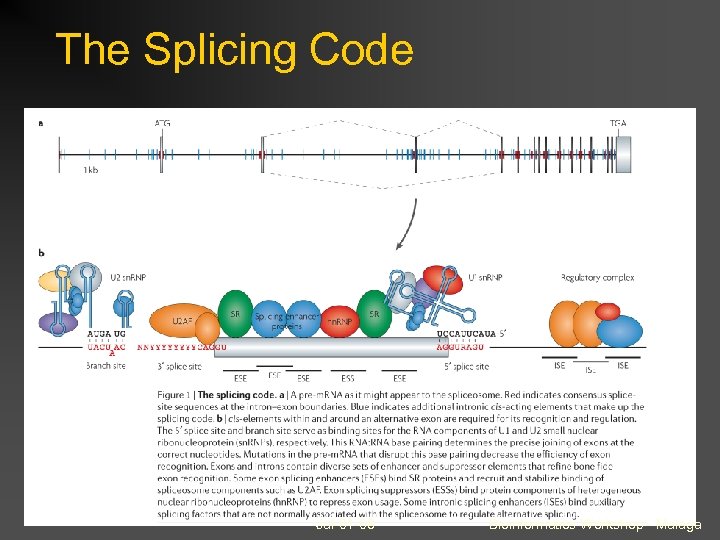
The Splicing Code Jul-01 -08 Bioinformatics Workshop - Malaga

Gene Prediction Strategies Expressed Sequence (c. DNA) or protein sequence available? Yes Spliced alignment BLAT, Exonerate, est_genome, spidey, GMAP, Genewise No Integrated gene prediction Informant genome(s) available? Yes Dual or n-genome de novo predictors: SGP 2, Twinscan, NSCAN, (Genomescan – same or cross genome protein blastx) No ab initio predictors geneid, genscan, augustus, fgenesh, genemark, etc. Many newer gene predictors can run in multiple modes depending on the evidence available. Jul-01 -08 Bioinformatics Workshop - Malaga
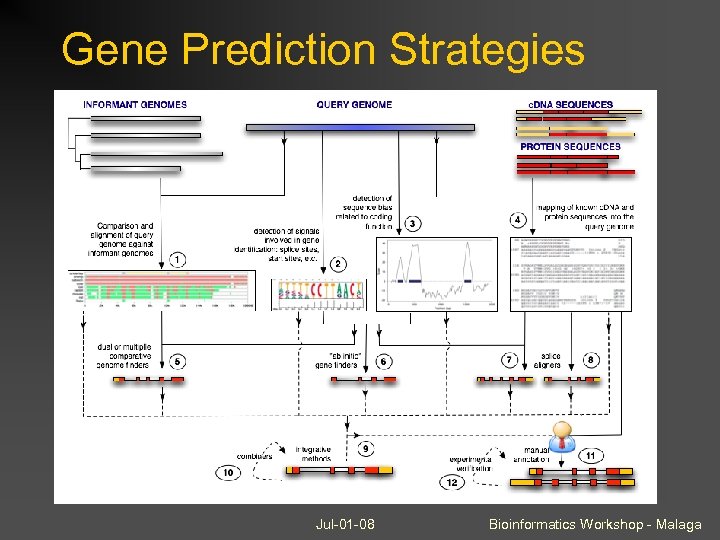
Gene Prediction Strategies Jul-01 -08 Bioinformatics Workshop - Malaga

Frameworks for gene prediction Hierarchical exon-buliding and chaining Hidden Markov Models (many flavors) HMM, GPHMM, Phylo-HMM Conditional Random Fields (new!) Conrad, Contrast. . . and, no doubt, more to come All of them involve parsing the optimal path of exons using dynamic programming (e. g. Gen. Amic, Viterbi algorithms) Jul-01 -08 Bioinformatics Workshop - Malaga

How does Gene. ID approach gene prediction? Jul-01 -0806/16/08 Bioinformatics Workshop - Malaga
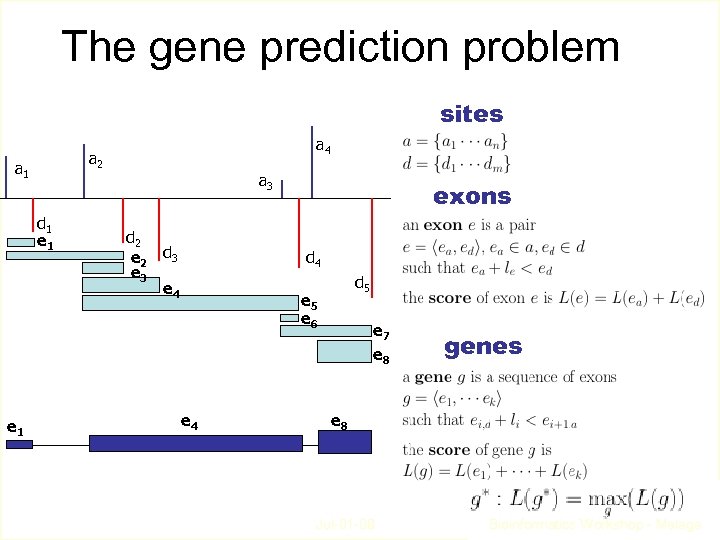
The gene prediction problem sites a 2 a 1 d 1 e 1 a 4 a 3 d 2 e 2 d 3 e 4 exons d 4 d 5 e 6 e 4 e 7 e 8 genes e 8 Jul-01 -08 Bioinformatics Workshop - Malaga
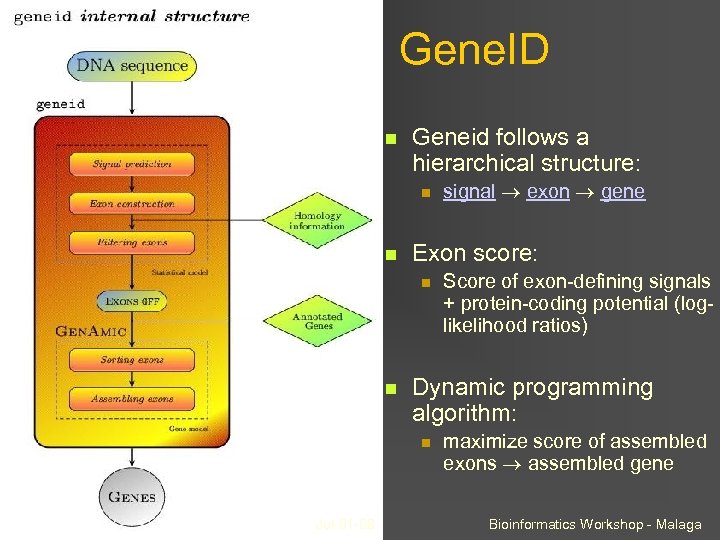
Gene. ID Geneid follows a hierarchical structure: Exon score: Score of exon-defining signals + protein-coding potential (loglikelihood ratios) Dynamic programming algorithm: Jul-01 -08 signal exon gene maximize score of assembled exons assembled gene Bioinformatics Workshop - Malaga
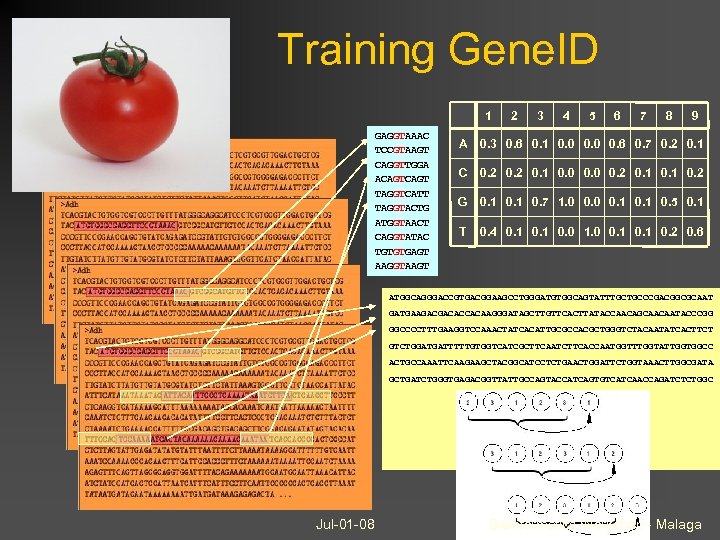

Training Gene. ID 1 GAGGTAAAC TCCGTAAGT CAGGTTGGA ACAGT TAGGTCATT TAGGTACTG ATGGTAACT CAGGTATAC TGTGTGAGT AAGGTAAGT 2 3 4 5 6 7 8 9 A 0. 3 0. 6 0. 1 0. 0 0. 6 0. 7 0. 2 0. 1 C 0. 2 0. 1 0. 0 0. 2 0. 1 0. 2 G 0. 1 0. 7 1. 0 0. 1 0. 5 0. 1 T 0. 4 0. 1 0. 0 1. 0 0. 1 0. 2 0. 6 ATGGCAGGGACCGTGACGGAAGCCTGGGATGTGGCAGTATTTGCTGCCCGACGGCGCAAT GATGAAGACGACACCACAAGGGATAGCTTGTTCACTTATACCAACAGCAACAATACCCGG GGCCCCTTTGAAGGTCCAAACTATCACATTGCGCCACGCTGGGTCTACAATATCACTTCT GTCTGGATGATTTTTGTGGTCATCGCTTCAATCTTCACCAATGGTTTGGTATTGGTGGCC ACTGCCAAATTCAAGAAGCTACGGCATCCTCTGAACTGGATTCTGGTAAACTTGGCGATA GCTGATCTGGGTGAGACGGTTATTGCCAGTACCATCAGTGTCATCAACCAGATCTCTGGC Jul-01 -08 Bioinformatics Workshop - Malaga
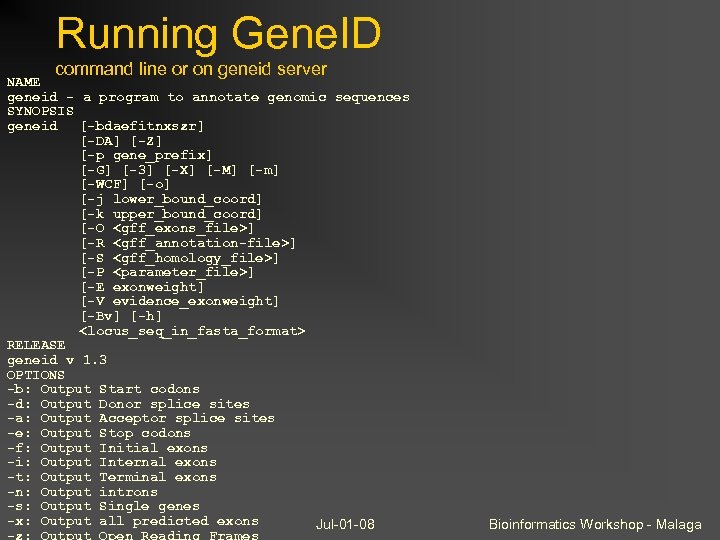
Running Gene. ID command line or on geneid server NAME geneid - a program to annotate genomic sequences SYNOPSIS geneid [-bdaefitnxszr] [-DA] [-Z] [-p gene_prefix] [-G] [-3] [-X] [-M] [-m] [-WCF] [-o] [-j lower_bound_coord] [-k upper_bound_coord] [-O <gff_exons_file>] [-R <gff_annotation-file>] [-S <gff_homology_file>] [-P <parameter_file>] [-E exonweight] [-V evidence_exonweight] [-Bv] [-h] <locus_seq_in_fasta_format> RELEASE geneid v 1. 3 OPTIONS -b: Output Start codons -d: Output Donor splice sites -a: Output Acceptor splice sites -e: Output Stop codons -f: Output Initial exons -i: Output Internal exons -t: Output Terminal exons -n: Output introns -s: Output Single genes -x: Output all predicted exons Jul-01 -08 -z: Output Open Reading Frames Bioinformatics Workshop - Malaga
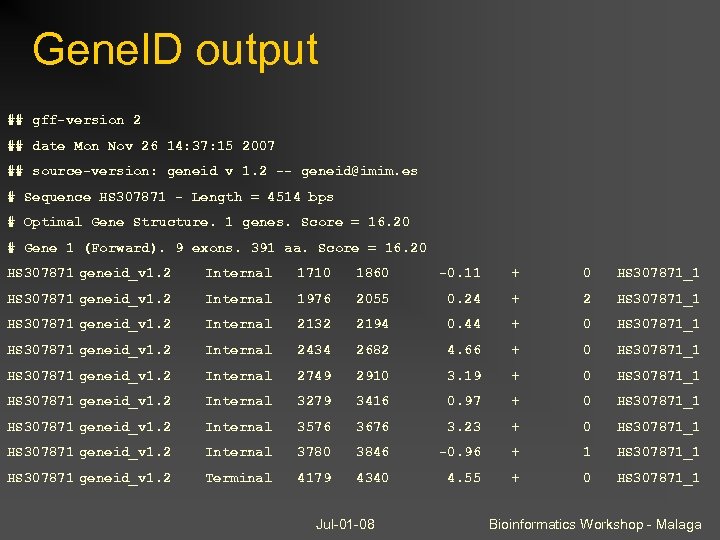
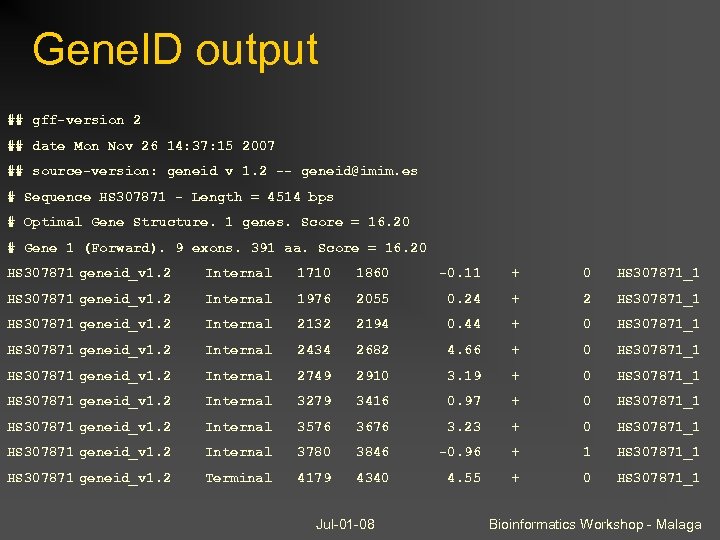
Gene. ID output ## gff-version 2 ## date Mon Nov 26 14: 37: 15 2007 ## source-version: geneid v 1. 2 -- geneid@imim. es # Sequence HS 307871 - Length = 4514 bps # Optimal Gene Structure. 1 genes. Score = 16. 20 # Gene 1 (Forward). 9 exons. 391 aa. Score = 16. 20 HS 307871 geneid_v 1. 2 Internal 1710 1860 -0. 11 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 1976 2055 0. 24 + 2 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 2132 2194 0. 44 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 2434 2682 4. 66 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 2749 2910 3. 19 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 3279 3416 0. 97 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 3576 3676 3. 23 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 3780 3846 -0. 96 + 1 HS 307871_1 HS 307871 geneid_v 1. 2 Terminal 4179 4340 4. 55 + 0 HS 307871_1 Jul-01 -08 Bioinformatics Workshop - Malaga

GFF: a standard annotation format Stands for: Designed as a single line record for describing features on DNA sequence -- originally used for gene prediction output 9 tab-delimited fields common to all versions Gene Finding Format -or- General Feature Format seq source feature begin end score strand frame group The group field differs between versions, but in every case no tabs are allowed GFF 2: group is a unique description, usually the gene name. GFF 2. 5 / GTF (Gene Transfer Format): tag-value pairs introduced, start_codon and stop_codon are required features for CDS NCOA 1 transcript_id “NM_056789” ; gene_id “NCOA 1” GFF 3: Capitalized tags follow Sequence Ontology (SO) relationships, FASTA seqs can be embedded ID=NM_056789_exon 1; Parent=NM_056789; note=“ 5’ UTR exon” Jul-01 -08 Bioinformatics Workshop - Malaga
Gene. ID output ## gff-version 2 ## date Mon Nov 26 14: 37: 15 2007 ## source-version: geneid v 1. 2 -- geneid@imim. es # Sequence HS 307871 - Length = 4514 bps # Optimal Gene Structure. 1 genes. Score = 16. 20 # Gene 1 (Forward). 9 exons. 391 aa. Score = 16. 20 HS 307871 geneid_v 1. 2 Internal 1710 1860 -0. 11 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 1976 2055 0. 24 + 2 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 2132 2194 0. 44 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 2434 2682 4. 66 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 2749 2910 3. 19 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 3279 3416 0. 97 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 3576 3676 3. 23 + 0 HS 307871_1 HS 307871 geneid_v 1. 2 Internal 3780 3846 -0. 96 + 1 HS 307871_1 HS 307871 geneid_v 1. 2 Terminal 4179 4340 4. 55 + 0 HS 307871_1 Jul-01 -08 Bioinformatics Workshop - Malaga
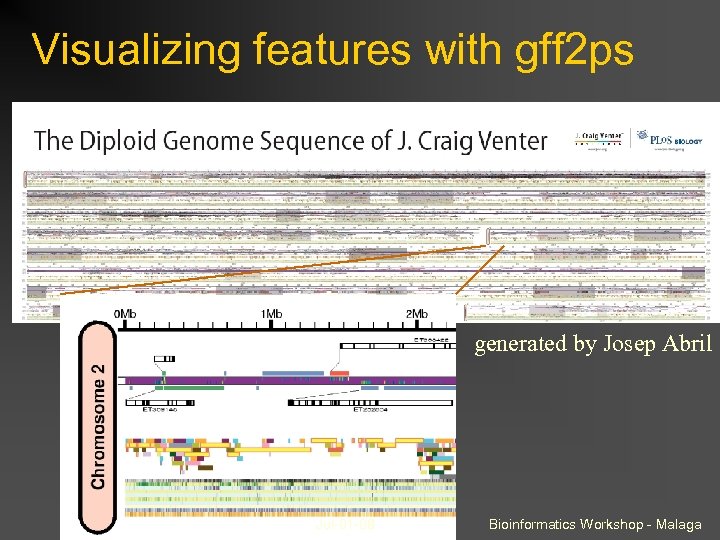
Visualizing features with gff 2 ps generated by Josep Abril Jul-01 -08 Bioinformatics Workshop - Malaga

Visualizing features on UCSC genome browser (custom tracks) If “your” genome is served by UCSC, this is a good option because: browsing is dynamic access to other annotations can view DNA sequence can do complex intersections and filtering gff 2 ps is good when: your genome is not on UCSC you want more flexible layout options you want to run it ‘offline’ Jul-01 -08 Bioinformatics Workshop - Malaga

Extensions to Gene. ID Syntenic Gene Prediction (dual-genome) Evidence-based (constrained) gene prediction U 12 intron detection Combining gene predictions Selenoprotein gene prediction Jul-01 -08 Bioinformatics Workshop - Malaga
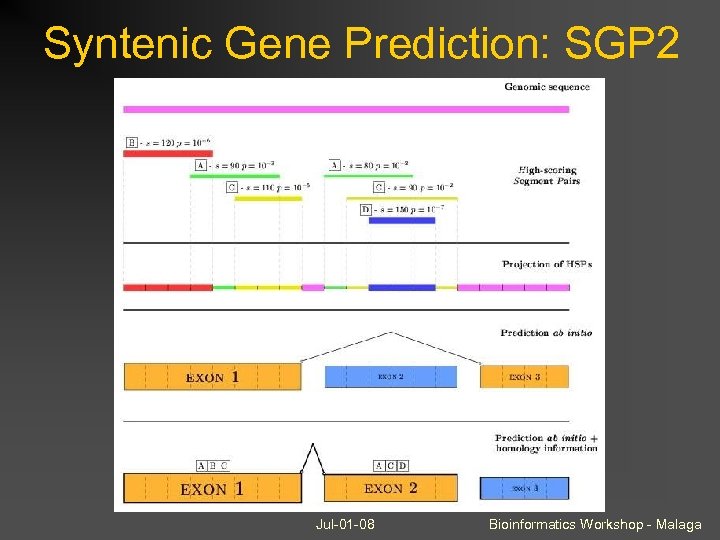
Syntenic Gene Prediction: SGP 2 Jul-01 -08 Bioinformatics Workshop - Malaga

Minor splicing and U 12 introns make up a minor proportion of all introns (~0. 33% in human, less in insects) But they can be found in 2 -3% of genes Normally ignored, but this causes annotation problems Easy to predict due to highly conserved donor and branch sites Jul-01 -08 Bioinformatics Workshop - Malaga
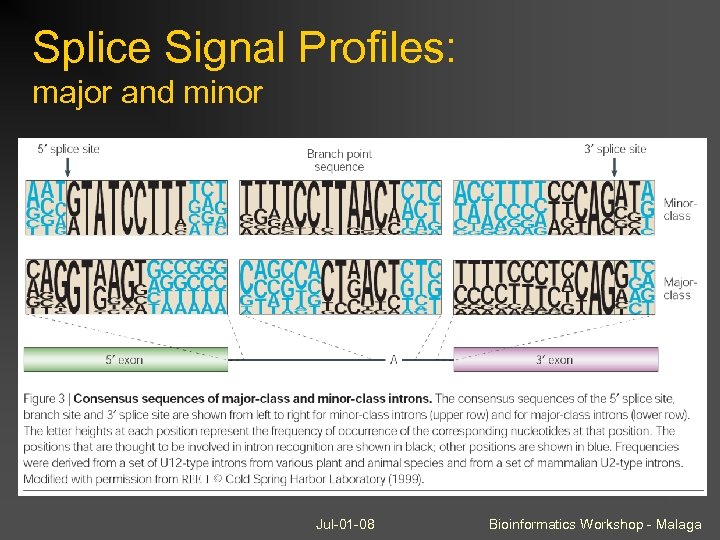

Splice Signal Profiles: major and minor Jul-01 -08 Bioinformatics Workshop - Malaga
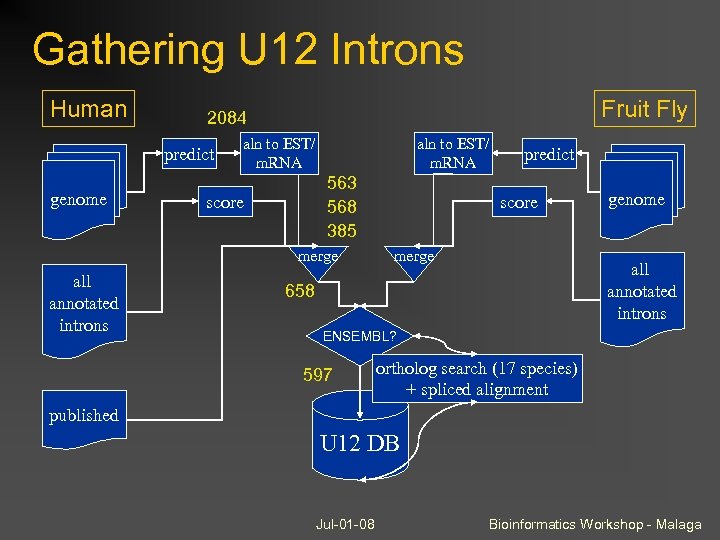
Gathering U 12 Introns Human predict genome Fruit Fly 2084 aln to EST/ m. RNA 563 568 385 score merge all annotated introns predict score merge genome all annotated introns 658 ENSEMBL? 597 ortholog search (17 species) + spliced alignment published U 12 DB Jul-01 -08 Bioinformatics Workshop - Malaga

Jul-01 -08 Bioinformatics Workshop - Malaga
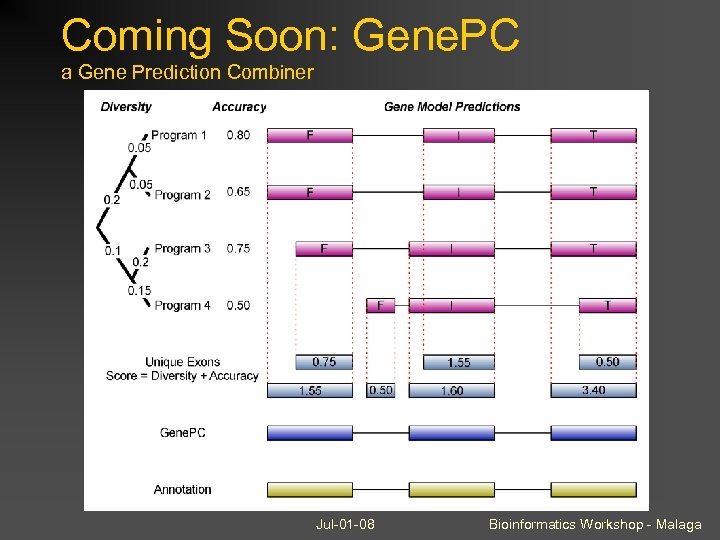
Coming Soon: Gene. PC a Gene Prediction Combiner Jul-01 -08 Bioinformatics Workshop - Malaga

Tutorial Homepage http: //genome. imim. es/courses/Malaga 08/ GBL Homepage http: //genome. imim. es/ Jul-01 -08 Bioinformatics Workshop - Malaga
8f401c8261665a99d06065b0368d02d0.ppt