4331c77ecbd08a2444ef5f513ffbc670.ppt
- Количество слайдов: 77
 Gene-Environment Case-Control Studies Raymond J. Carroll Department of Statistics Center for Statistical Bioinformatics Institute for Applied Mathematics and Computational Science Texas A&M University http: //stat. tamu. edu/~carroll
Gene-Environment Case-Control Studies Raymond J. Carroll Department of Statistics Center for Statistical Bioinformatics Institute for Applied Mathematics and Computational Science Texas A&M University http: //stat. tamu. edu/~carroll
 Advertising • Training We are finishing Year 08 of an NCI: funded R 25 T training program • http: //www. stat. tamu. edu/b 3 nc • We train statistically and computationally oriented post-docs in the biology of nutrition and cancer • Active seminar series
Advertising • Training We are finishing Year 08 of an NCI: funded R 25 T training program • http: //www. stat. tamu. edu/b 3 nc • We train statistically and computationally oriented post-docs in the biology of nutrition and cancer • Active seminar series
 Outline • Problem Case-Control Studies with Gene: Environment relationships • Efficient formulation when genes are observed • Haplotype modeling and Robustness • Applications
Outline • Problem Case-Control Studies with Gene: Environment relationships • Efficient formulation when genes are observed • Haplotype modeling and Robustness • Applications
 Acknowledgment • This work is joint with Nilanjan Chatterjee (NCI) and Yi-Hau Chen (Academia Sinica)
Acknowledgment • This work is joint with Nilanjan Chatterjee (NCI) and Yi-Hau Chen (Academia Sinica)
 Software • SAS and Matlab Programs Available at my web site under the software button http: //tat. tamu. edu/~carroll s • Examples are given in the programs • Paper are in Biometrika (2005), Genetic Epidemiology (2006), Biostatistics (2007), Biometrics (2008) and JASA (2009) • R programs available from the NCI
Software • SAS and Matlab Programs Available at my web site under the software button http: //tat. tamu. edu/~carroll s • Examples are given in the programs • Paper are in Biometrika (2005), Genetic Epidemiology (2006), Biostatistics (2007), Biometrics (2008) and JASA (2009) • R programs available from the NCI
 Basic Problem Formalized • Gene and Environment • Question For women who carry the BRCA 1/2 : mutation does oral contraceptive use , provide any protection against ovarian cancer?
Basic Problem Formalized • Gene and Environment • Question For women who carry the BRCA 1/2 : mutation does oral contraceptive use , provide any protection against ovarian cancer?
 Basic Problem Formalized • Gene and Environment • Question For people carrying a particular : haplotype in the VDR pathway , does higher levels of serum Vitamin D protect against prostate cancer?
Basic Problem Formalized • Gene and Environment • Question For people carrying a particular : haplotype in the VDR pathway , does higher levels of serum Vitamin D protect against prostate cancer?
 Basic Problem Formalized • Gene and Environment • Question If you are a current smokerare you : , protected against colorectal adenoma if you carry a particular haplotype in the NAT 2 smoking metabolism region?
Basic Problem Formalized • Gene and Environment • Question If you are a current smokerare you : , protected against colorectal adenoma if you carry a particular haplotype in the NAT 2 smoking metabolism region?
 Prospective and Retrospective Studies • D = disease status (binary) • X = environmental variables • Smoking status • Vitamin D • Oral contraceptive use • G = gene status • Mutation or not • Multiple or single SNP • Haplotypes
Prospective and Retrospective Studies • D = disease status (binary) • X = environmental variables • Smoking status • Vitamin D • Oral contraceptive use • G = gene status • Mutation or not • Multiple or single SNP • Haplotypes
 Prospective and Retrospective Studies • Prospective Classic random sampling of a : population • You measure gene and environment on a cohort • You then follow up people for disease occurrence
Prospective and Retrospective Studies • Prospective Classic random sampling of a : population • You measure gene and environment on a cohort • You then follow up people for disease occurrence
 Prospective and Retrospective Studies • Prospective Studies : • Expensive disease states are rare, so large : sample sizes needed • Time-consumingyou have to wait for disease : to develop • They Exist Framingham Heart Study, NIH-AARP : Diet and Health Study, Women’s Health Initiative, etc.
Prospective and Retrospective Studies • Prospective Studies : • Expensive disease states are rare, so large : sample sizes needed • Time-consumingyou have to wait for disease : to develop • They Exist Framingham Heart Study, NIH-AARP : Diet and Health Study, Women’s Health Initiative, etc.
 Prospective and Retrospective Studies • Prospective Studies : • Daunting Task. Only very large, very expensive : prospective studies can find gene-environment interactions • Data Access to the Framingham Heart : Study requires a university commitment to security
Prospective and Retrospective Studies • Prospective Studies : • Daunting Task. Only very large, very expensive : prospective studies can find gene-environment interactions • Data Access to the Framingham Heart : Study requires a university commitment to security
 Prospective and Retrospective Studies • Retrospective Studies : Usually called casecontrol studies • Find a population of cases i. e. , people with a , disease, and sample from it. • Find a population of controls i. e. , people , without the disease, and sample from it.
Prospective and Retrospective Studies • Retrospective Studies : Usually called casecontrol studies • Find a population of cases i. e. , people with a , disease, and sample from it. • Find a population of controls i. e. , people , without the disease, and sample from it.
 Prospective and Retrospective Studies • Retrospective Studies : Because the gene G and the environment X are sample after disease status is ascertained • Microarray studies on humans : most are casecontrol studies • Genome Wide Association Studies (GWAS) : most are case-control studies
Prospective and Retrospective Studies • Retrospective Studies : Because the gene G and the environment X are sample after disease status is ascertained • Microarray studies on humans : most are casecontrol studies • Genome Wide Association Studies (GWAS) : most are case-control studies
 Prospective and Retrospective Studies • Case-control Studies : • Fast: no need to wait for disease to develop • Cheap sample sizes are much smaller : • Subtle The controls need to be representative : of the population of people without the disease.
Prospective and Retrospective Studies • Case-control Studies : • Fast: no need to wait for disease to develop • Cheap sample sizes are much smaller : • Subtle The controls need to be representative : of the population of people without the disease.
 Basic Problem Formalized • Case control sample: D = disease • Gene expression: G • Environment, can include strata: X • We are interested in main effects for G and X along with their interaction as they affect development of disease
Basic Problem Formalized • Case control sample: D = disease • Gene expression: G • Environment, can include strata: X • We are interested in main effects for G and X along with their interaction as they affect development of disease
 Basic Problem Formalized • 99. 9999% analyses of case-control data use of logistic regression • Closely related to Fisher’s Linear Discriminant Analysis (LDA) • Difference we want to understand what targets : affect disease, not just predict disease
Basic Problem Formalized • 99. 9999% analyses of case-control data use of logistic regression • Closely related to Fisher’s Linear Discriminant Analysis (LDA) • Difference we want to understand what targets : affect disease, not just predict disease
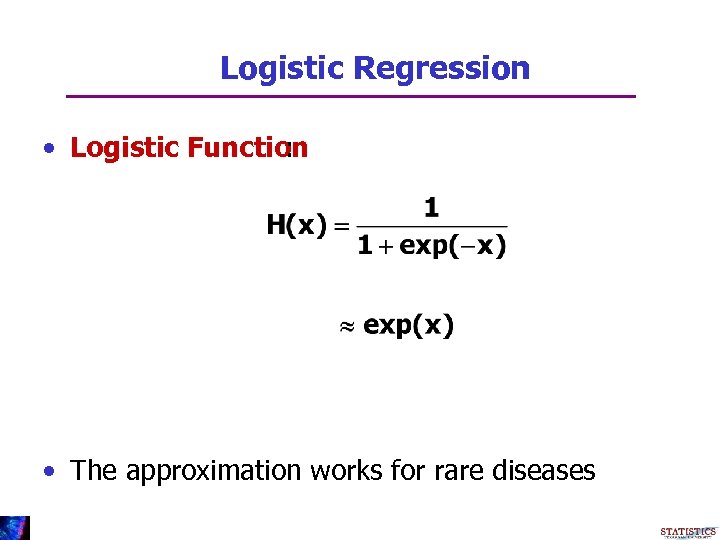 Logistic Regression • Logistic Function : • The approximation works for rare diseases
Logistic Regression • Logistic Function : • The approximation works for rare diseases
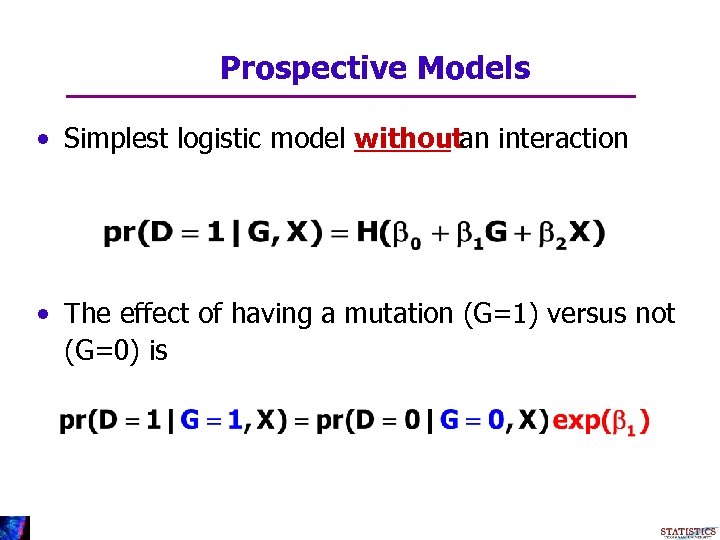 Prospective Models • Simplest logistic model withoutan interaction • The effect of having a mutation (G=1) versus not (G=0) is
Prospective Models • Simplest logistic model withoutan interaction • The effect of having a mutation (G=1) versus not (G=0) is
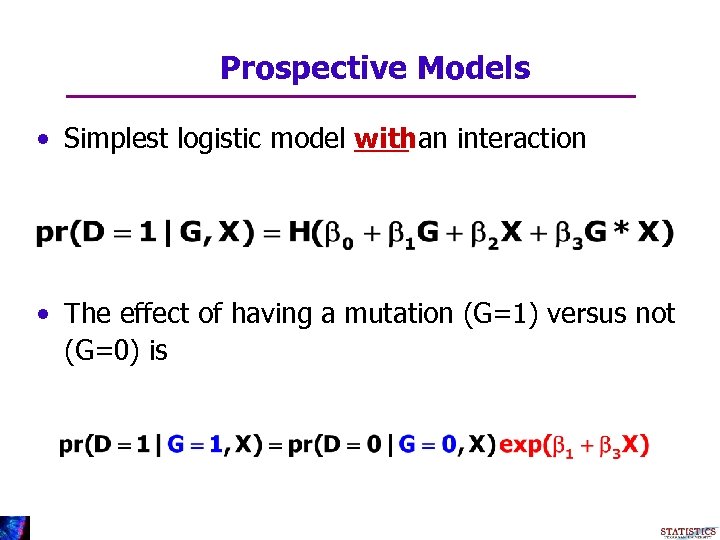 Prospective Models • Simplest logistic model withan interaction • The effect of having a mutation (G=1) versus not (G=0) is
Prospective Models • Simplest logistic model withan interaction • The effect of having a mutation (G=1) versus not (G=0) is
 Empirical Observations • Logistic regression is in every statistical package • Unfortunately, logistic regression is not efficient for understanding interactions • Much larger sample sizes are required for interactions that for just gene effects • Most gene-environment interaction case-control studiesfail for this reason
Empirical Observations • Logistic regression is in every statistical package • Unfortunately, logistic regression is not efficient for understanding interactions • Much larger sample sizes are required for interactions that for just gene effects • Most gene-environment interaction case-control studiesfail for this reason
 Empirical Observations • Statistical Theory. There is a lovely statistical : theory available • It says: ignore the fact that you have a casecontrol sample, and pretend you have a prospective study • It all works outdon’t worry, be happy! :
Empirical Observations • Statistical Theory. There is a lovely statistical : theory available • It says: ignore the fact that you have a casecontrol sample, and pretend you have a prospective study • It all works outdon’t worry, be happy! :
 Empirical Observations • Statistical Theory. Ordinary logistic regression : applied to a case-control study makes no assumptions about the population distribution of (G, X) • Remember we do not have a sample from a : population, only a case-control sample • Logistic regression is robust: to assumptions about the population distribution of (G, X)
Empirical Observations • Statistical Theory. Ordinary logistic regression : applied to a case-control study makes no assumptions about the population distribution of (G, X) • Remember we do not have a sample from a : population, only a case-control sample • Logistic regression is robust: to assumptions about the population distribution of (G, X)
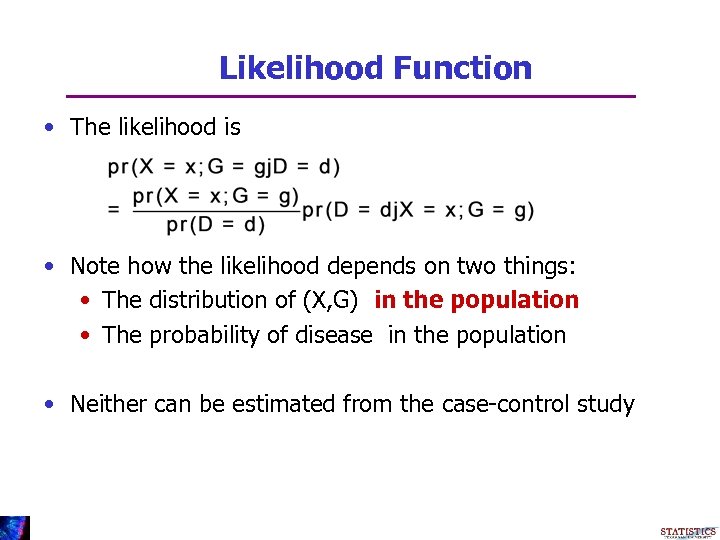 Likelihood Function • The likelihood is • Note how the likelihood depends on two things: • The distribution of (X, G) in the population • The probability of disease in the population • Neither can be estimated from the case-control study
Likelihood Function • The likelihood is • Note how the likelihood depends on two things: • The distribution of (X, G) in the population • The probability of disease in the population • Neither can be estimated from the case-control study
 When G is observed • Logistic regression is thus robust to any modeling assumptions about the covariates in the population • Unfortunately it is not very efficient for understanding interactions
When G is observed • Logistic regression is thus robust to any modeling assumptions about the covariates in the population • Unfortunately it is not very efficient for understanding interactions
 Gene-Environment Independence • In many situations, it may be reasonable to assume G and X are independently distributed in the underlying population , possibly after conditioning on strata • This assumption is often used in geneenvironment interaction studies
Gene-Environment Independence • In many situations, it may be reasonable to assume G and X are independently distributed in the underlying population , possibly after conditioning on strata • This assumption is often used in geneenvironment interaction studies
 G-E Independence • Does not always hold! • Example polymorphisms in the smoking : metabolism pathway may affect the degree of addiction
G-E Independence • Does not always hold! • Example polymorphisms in the smoking : metabolism pathway may affect the degree of addiction
 Gene-Environment Independence • If you’re willing to make assumptions about the distributions of the covariates in the population, more efficiency be obtained. can • This is NOT TRUEfor prospective studies, only true for retrospective studies.
Gene-Environment Independence • If you’re willing to make assumptions about the distributions of the covariates in the population, more efficiency be obtained. can • This is NOT TRUEfor prospective studies, only true for retrospective studies.
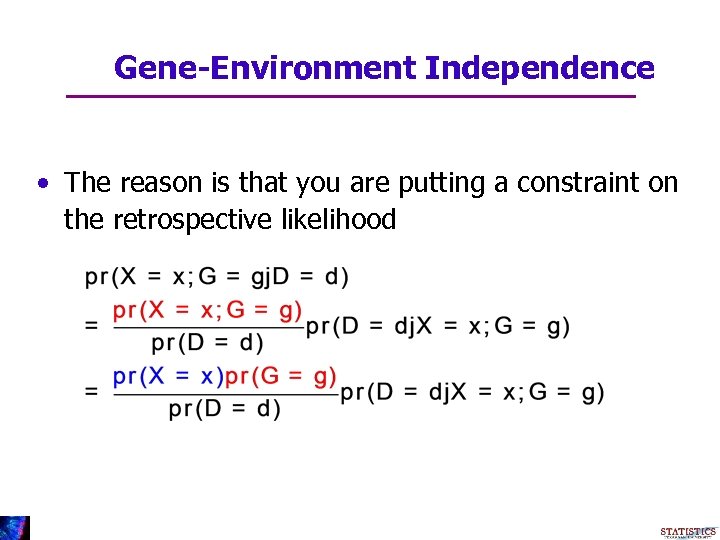 Gene-Environment Independence • The reason is that you are putting a constraint on the retrospective likelihood
Gene-Environment Independence • The reason is that you are putting a constraint on the retrospective likelihood
 Gene-Environment Independence • Our Methodology. Is far more general than : assuming that genetic status and environment are independent • We have developed capacity for modeling the distribution of genetic status given strata and environmental factors • I will skip this and just pretend G-E independence here
Gene-Environment Independence • Our Methodology. Is far more general than : assuming that genetic status and environment are independent • We have developed capacity for modeling the distribution of genetic status given strata and environmental factors • I will skip this and just pretend G-E independence here
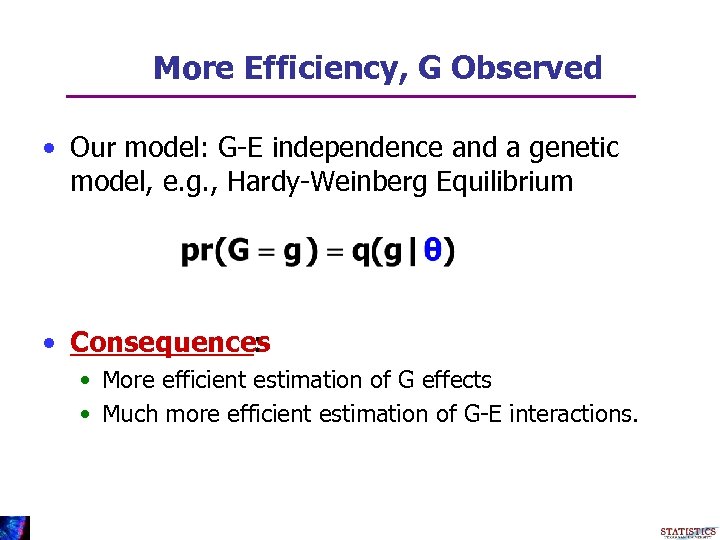 More Efficiency, G Observed • Our model: G-E independence and a genetic model, e. g. , Hardy-Weinberg Equilibrium • Consequences : • More efficient estimation of G effects • Much more efficient estimation of G-E interactions.
More Efficiency, G Observed • Our model: G-E independence and a genetic model, e. g. , Hardy-Weinberg Equilibrium • Consequences : • More efficient estimation of G effects • Much more efficient estimation of G-E interactions.
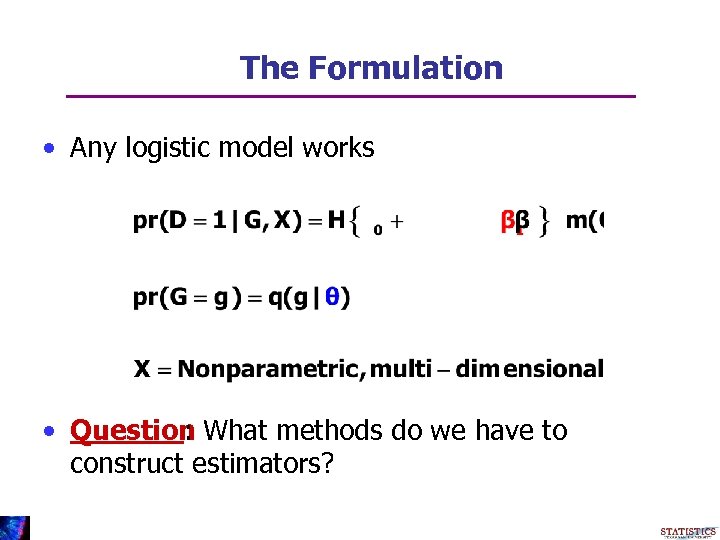 The Formulation • Any logistic model works • Question What methods do we have to : construct estimators?
The Formulation • Any logistic model works • Question What methods do we have to : construct estimators?
 Methodology • I won’t give you the full methodology, but it works as follows. • Case-control studies are very close to a prospective (random sampling) study, with the exception that sometimes you do not observe people
Methodology • I won’t give you the full methodology, but it works as follows. • Case-control studies are very close to a prospective (random sampling) study, with the exception that sometimes you do not observe people
 Pretend Missing Data Formulation • Suppose you have a large but finite population of size N • Then, there are • There are with the disease without the disease
Pretend Missing Data Formulation • Suppose you have a large but finite population of size N • Then, there are • There are with the disease without the disease
 Pretend Missing Data Formulation • In a case-control sample, we randomly select n 1 with the disease, and n 0 without. • The fraction of people with disease status D=d that we observe is
Pretend Missing Data Formulation • In a case-control sample, we randomly select n 1 with the disease, and n 0 without. • The fraction of people with disease status D=d that we observe is
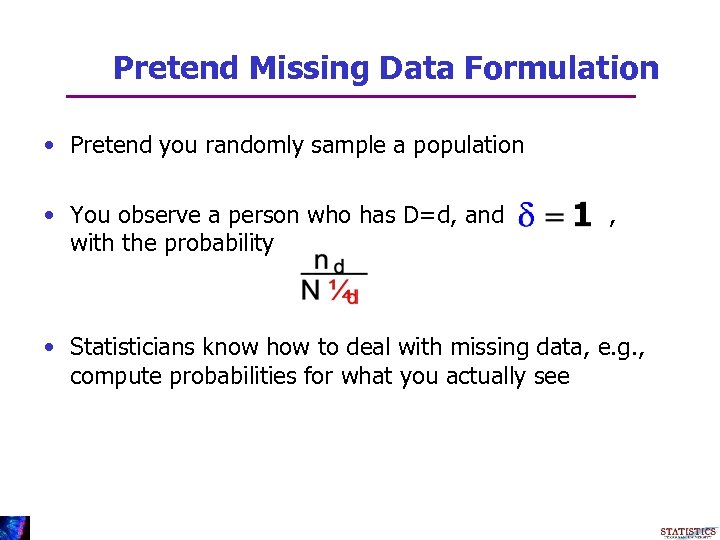 Pretend Missing Data Formulation • Pretend you randomly sample a population • You observe a person who has D=d, and with the probability , • Statisticians know how to deal with missing data, e. g. , compute probabilities for what you actually see
Pretend Missing Data Formulation • Pretend you randomly sample a population • You observe a person who has D=d, and with the probability , • Statisticians know how to deal with missing data, e. g. , compute probabilities for what you actually see
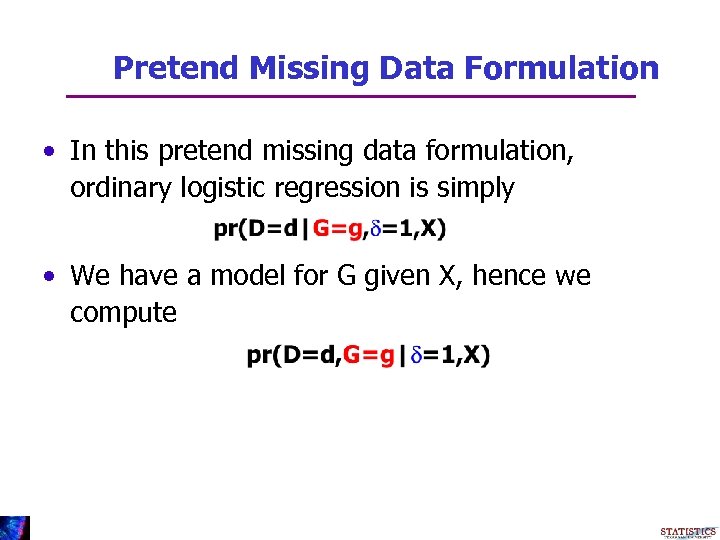 Pretend Missing Data Formulation • In this pretend missing data formulation, ordinary logistic regression is simply • We have a model for G given X, hence we compute
Pretend Missing Data Formulation • In this pretend missing data formulation, ordinary logistic regression is simply • We have a model for G given X, hence we compute
 Methodology • Our method has an explicit form, i. e. , no integrals or anything nasty • It is easy to program the method to estimate the logistic model • It is likelihood based. Technically, a semiparametric profile likelihood
Methodology • Our method has an explicit form, i. e. , no integrals or anything nasty • It is easy to program the method to estimate the logistic model • It is likelihood based. Technically, a semiparametric profile likelihood
 Methodology • We can handle missing gene data • We can handle error in genotyping • We can handle measurement errors in environmental variables, e. g. , diet
Methodology • We can handle missing gene data • We can handle error in genotyping • We can handle measurement errors in environmental variables, e. g. , diet
 Methodology • Our method results in much more efficient statistical inference
Methodology • Our method results in much more efficient statistical inference
 More Data • What does More efficient statistical inference mean? • It means, effectively, that you have more data • In cases that G is a simple mutation, our method is typically equivalent to having 3 times more data
More Data • What does More efficient statistical inference mean? • It means, effectively, that you have more data • In cases that G is a simple mutation, our method is typically equivalent to having 3 times more data
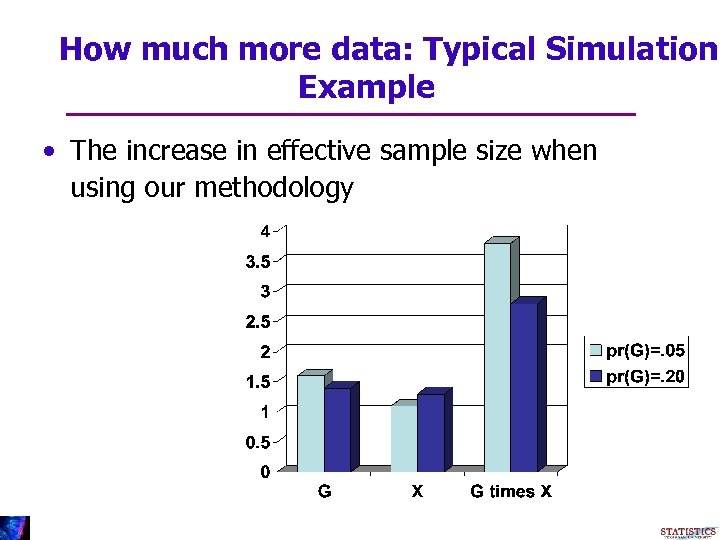 How much more data: Typical Simulation Example • The increase in effective sample size when using our methodology
How much more data: Typical Simulation Example • The increase in effective sample size when using our methodology
 Real Data Complexities • The Israeli Ovarian Cancer Study • G = BRCA 1/2 mutation (very deadly) • X includes • age, • ethnic status (below), • parity, • oral contraceptive use • Family history • Smoking • Etc.
Real Data Complexities • The Israeli Ovarian Cancer Study • G = BRCA 1/2 mutation (very deadly) • X includes • age, • ethnic status (below), • parity, • oral contraceptive use • Family history • Smoking • Etc.
 Real Data Complexities • In the Israeli Study, G is missing in 50% of the controls, and 10% of the cases • Also, among Jewish citizens, Israel has two dominant ethnic types • Ashkenazi (European) • Shephardic (North African)
Real Data Complexities • In the Israeli Study, G is missing in 50% of the controls, and 10% of the cases • Also, among Jewish citizens, Israel has two dominant ethnic types • Ashkenazi (European) • Shephardic (North African)
 Real Data Complexities • The gene mutation BRCA 1/2 if frequent among the Ashkenazi, but rare among the Shephardic • Thus, if one component of X is ethnic status, then pr(G =1 | X)depends on X • Gene-Environment independence fails here • What can be done? Model pr(G =1 | X)as binary with different probabilities!
Real Data Complexities • The gene mutation BRCA 1/2 if frequent among the Ashkenazi, but rare among the Shephardic • Thus, if one component of X is ethnic status, then pr(G =1 | X)depends on X • Gene-Environment independence fails here • What can be done? Model pr(G =1 | X)as binary with different probabilities!
 Israeli Ovarian Cancer Study • Question Can carriers of the BRCA 1/2 mutation : be protected via OC-use?
Israeli Ovarian Cancer Study • Question Can carriers of the BRCA 1/2 mutation : be protected via OC-use?
 Typical Empirical Example
Typical Empirical Example
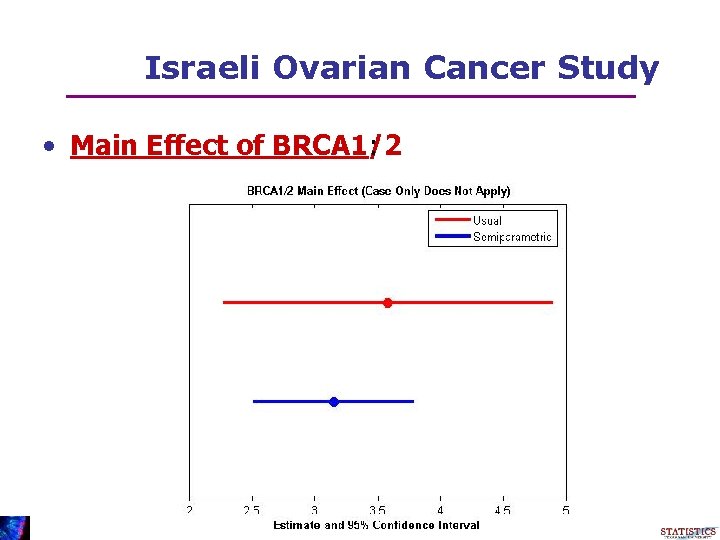 Israeli Ovarian Cancer Study • Main Effect of BRCA 1/2 :
Israeli Ovarian Cancer Study • Main Effect of BRCA 1/2 :
 Israeli Ovarian Cancer Study • Odds ratio for OC use among carriers = 1. 04 (0. 98, 1. 09) • No evidence for protective effect • Not available from case-only analysis • Length of interval is ½ the length of the usual analysis
Israeli Ovarian Cancer Study • Odds ratio for OC use among carriers = 1. 04 (0. 98, 1. 09) • No evidence for protective effect • Not available from case-only analysis • Length of interval is ½ the length of the usual analysis
 Haplotypes • Haplotypes consist of what we get from our mother and father at more than one site • Mother gives us the haplotype hm = (Am, Bm) • Father gives us the haplotype hf = (af, bf) • Our diplotype is Hdip = {(Am, Bm), (af, bf)}
Haplotypes • Haplotypes consist of what we get from our mother and father at more than one site • Mother gives us the haplotype hm = (Am, Bm) • Father gives us the haplotype hf = (af, bf) • Our diplotype is Hdip = {(Am, Bm), (af, bf)}
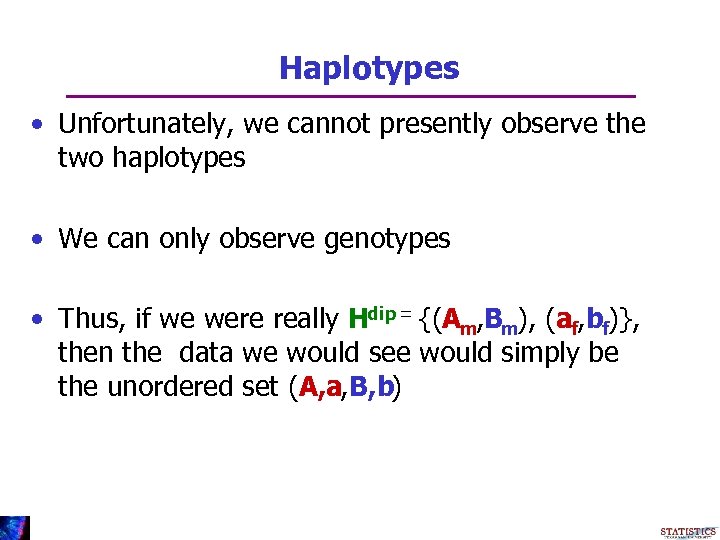 Haplotypes • Unfortunately, we cannot presently observe the two haplotypes • We can only observe genotypes • Thus, if we were really Hdip = {(Am, Bm), (af, bf)}, then the data we would see would simply be the unordered set (A, a, B, b)
Haplotypes • Unfortunately, we cannot presently observe the two haplotypes • We can only observe genotypes • Thus, if we were really Hdip = {(Am, Bm), (af, bf)}, then the data we would see would simply be the unordered set (A, a, B, b)
 Missing Haplotypes • Thus, if we were really Hdip = {(Am, Bm), (af, bf)}, then the data we would see would simply be the unordered set (A, a, B, b) • However, this is also consistent with a different diplotype, namely Hdip = {(am, Bm), (Af, bf)} • Note that the number of copies of the (a, b) haplotype differs in these two cases • The true diploid = haplotype pair is missing
Missing Haplotypes • Thus, if we were really Hdip = {(Am, Bm), (af, bf)}, then the data we would see would simply be the unordered set (A, a, B, b) • However, this is also consistent with a different diplotype, namely Hdip = {(am, Bm), (Af, bf)} • Note that the number of copies of the (a, b) haplotype differs in these two cases • The true diploid = haplotype pair is missing
 Missing Haplotypes • Our methods handle unphased diplotyes (missing haplotypes) with no problem. • Standard EM-algorithm calculations can be used • We assume that the haplotypes are in HWE, and have extended to cases of non-HWE
Missing Haplotypes • Our methods handle unphased diplotyes (missing haplotypes) with no problem. • Standard EM-algorithm calculations can be used • We assume that the haplotypes are in HWE, and have extended to cases of non-HWE
 Robustness • Robustness We are making assumptions to gain : efficiency = “get more data” • What happens if the assumptions are wrong? • Biases, incorrect conclusions, etc. • How can we gain efficiency when it is warranted, and yet have valid inferences?
Robustness • Robustness We are making assumptions to gain : efficiency = “get more data” • What happens if the assumptions are wrong? • Biases, incorrect conclusions, etc. • How can we gain efficiency when it is warranted, and yet have valid inferences?
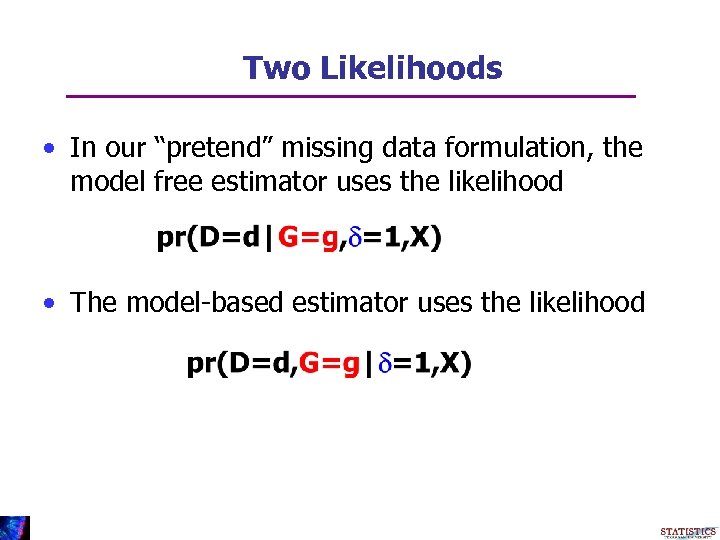 Two Likelihoods • In our “pretend” missing data formulation, the model free estimator uses the likelihood • The model-based estimator uses the likelihood
Two Likelihoods • In our “pretend” missing data formulation, the model free estimator uses the likelihood • The model-based estimator uses the likelihood
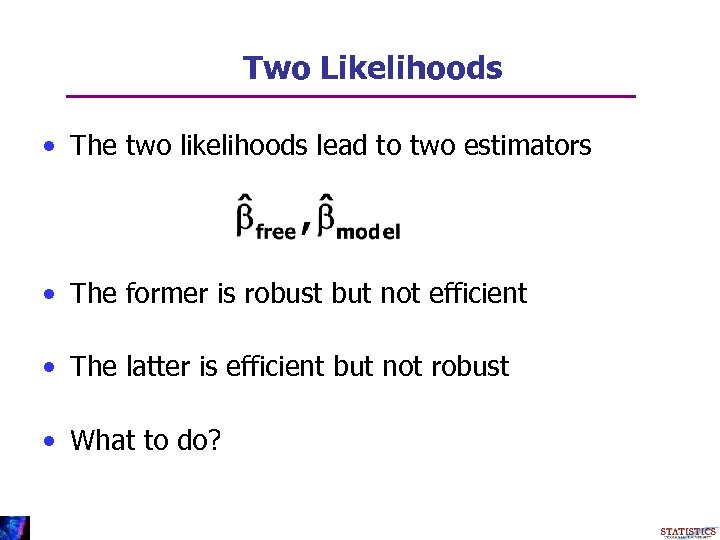 Two Likelihoods • The two likelihoods lead to two estimators • The former is robust but not efficient • The latter is efficient but not robust • What to do?
Two Likelihoods • The two likelihoods lead to two estimators • The former is robust but not efficient • The latter is efficient but not robust • What to do?
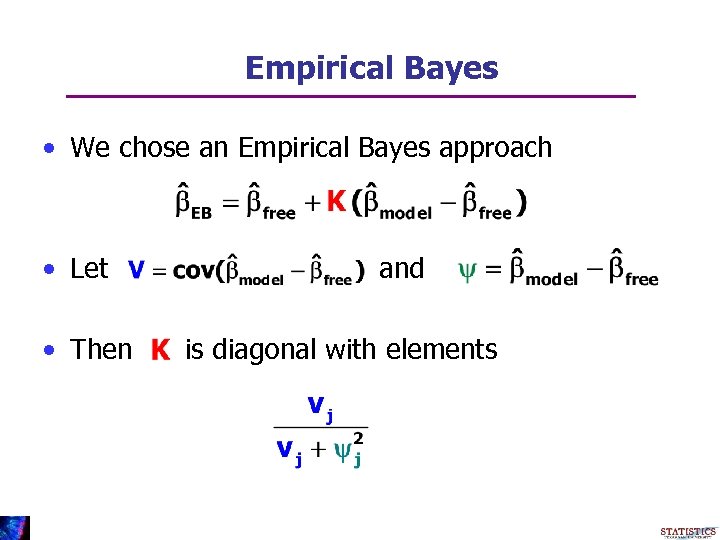 Empirical Bayes • We chose an Empirical Bayes approach • Let • Then and is diagonal with elements
Empirical Bayes • We chose an Empirical Bayes approach • Let • Then and is diagonal with elements
 Comments on Empirical Bayes • If the model fails, then the estimator converges to the model-free estimator • If the model holds, the estimator estimates the right thing, but is much more efficient than the model-free estimator
Comments on Empirical Bayes • If the model fails, then the estimator converges to the model-free estimator • If the model holds, the estimator estimates the right thing, but is much more efficient than the model-free estimator
 Simulations • Various simulations show the following • If the model holds, EB is • slightly less efficient that model-based • much more efficient than model-free • If the model fails, • Model-based is badly biased • EB and shrinkage eliminate most bias, at least as efficient as model-free
Simulations • Various simulations show the following • If the model holds, EB is • slightly less efficient that model-based • much more efficient than model-free • If the model fails, • Model-based is badly biased • EB and shrinkage eliminate most bias, at least as efficient as model-free
 Example 1: Prostate Cancer • G = SNPs in the Vitamin D Pathway • X = Serum-level biomarker of vitamin D (diet and sun) • The VDR gene is downstream in the pathway, hence unlikely to influence the level of X • Gene-environment independence likely
Example 1: Prostate Cancer • G = SNPs in the Vitamin D Pathway • X = Serum-level biomarker of vitamin D (diet and sun) • The VDR gene is downstream in the pathway, hence unlikely to influence the level of X • Gene-environment independence likely
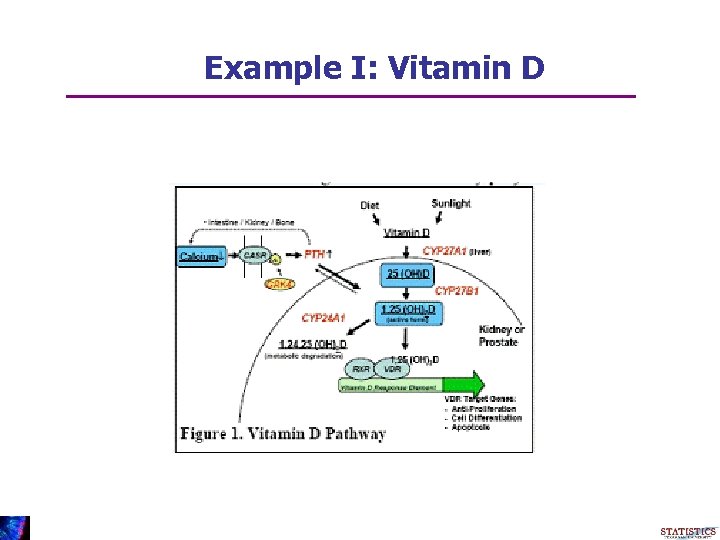 Example I: Vitamin D
Example I: Vitamin D
 Example 2: Colorectal Adenoma • G = SNPs in the NAT 2 gene, which is important in the metabolism of • X =Various measures of smoking history • The NAT 2 gene may make smokers more addicted • Gene-environment independence unlikely
Example 2: Colorectal Adenoma • G = SNPs in the NAT 2 gene, which is important in the metabolism of • X =Various measures of smoking history • The NAT 2 gene may make smokers more addicted • Gene-environment independence unlikely
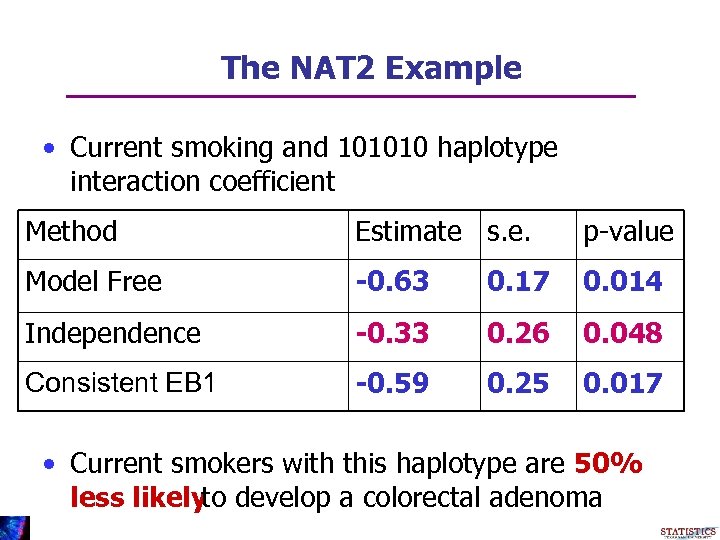 The NAT 2 Example • Current smoking and 101010 haplotype interaction coefficient Method Estimate s. e. p-value Model Free -0. 63 0. 17 0. 014 Independence -0. 33 0. 26 0. 048 Consistent EB 1 -0. 59 0. 25 0. 017 • Current smokers with this haplotype are 50% less likely develop a colorectal adenoma to
The NAT 2 Example • Current smoking and 101010 haplotype interaction coefficient Method Estimate s. e. p-value Model Free -0. 63 0. 17 0. 014 Independence -0. 33 0. 26 0. 048 Consistent EB 1 -0. 59 0. 25 0. 017 • Current smokers with this haplotype are 50% less likely develop a colorectal adenoma to
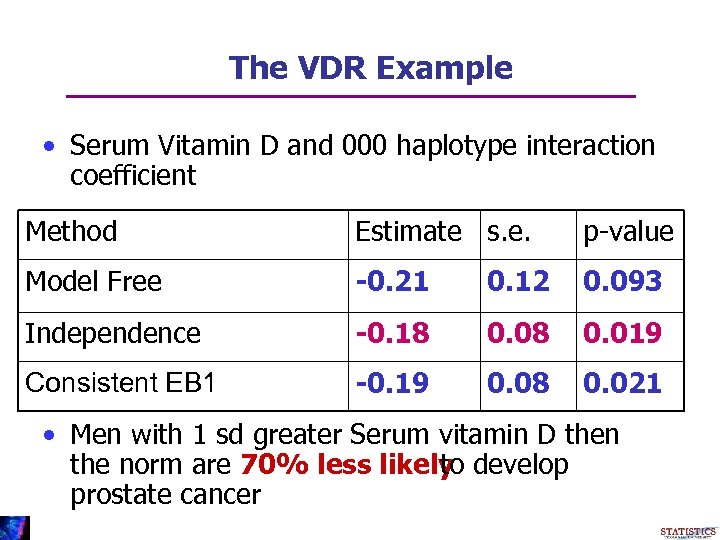 The VDR Example • Serum Vitamin D and 000 haplotype interaction coefficient Method Estimate s. e. p-value Model Free -0. 21 0. 12 0. 093 Independence -0. 18 0. 019 Consistent EB 1 -0. 19 0. 08 0. 021 • Men with 1 sd greater Serum vitamin D then the norm are 70% less likely develop to prostate cancer
The VDR Example • Serum Vitamin D and 000 haplotype interaction coefficient Method Estimate s. e. p-value Model Free -0. 21 0. 12 0. 093 Independence -0. 18 0. 019 Consistent EB 1 -0. 19 0. 08 0. 021 • Men with 1 sd greater Serum vitamin D then the norm are 70% less likely develop to prostate cancer
 Genome-Wide Association Studies • These methods can be applied to GWAS • My last two examples were actually from the PLCO GWAS • Also, can call the environment = other SNP
Genome-Wide Association Studies • These methods can be applied to GWAS • My last two examples were actually from the PLCO GWAS • Also, can call the environment = other SNP
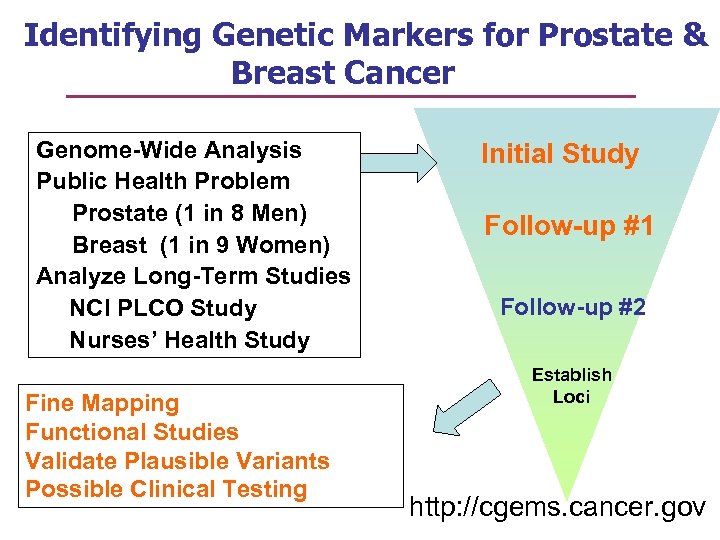 Identifying Genetic Markers for Prostate & Breast Cancer Genome-Wide Analysis Public Health Problem Prostate (1 in 8 Men) Breast (1 in 9 Women) Analyze Long-Term Studies NCI PLCO Study Nurses’ Health Study Fine Mapping Functional Studies Validate Plausible Variants Possible Clinical Testing Initial Study Follow-up #1 Follow-up #2 Establish Loci http: //cgems. cancer. gov
Identifying Genetic Markers for Prostate & Breast Cancer Genome-Wide Analysis Public Health Problem Prostate (1 in 8 Men) Breast (1 in 9 Women) Analyze Long-Term Studies NCI PLCO Study Nurses’ Health Study Fine Mapping Functional Studies Validate Plausible Variants Possible Clinical Testing Initial Study Follow-up #1 Follow-up #2 Establish Loci http: //cgems. cancer. gov
 Identifying Genetic Markers for Prostate & Breast Cancer
Identifying Genetic Markers for Prostate & Breast Cancer
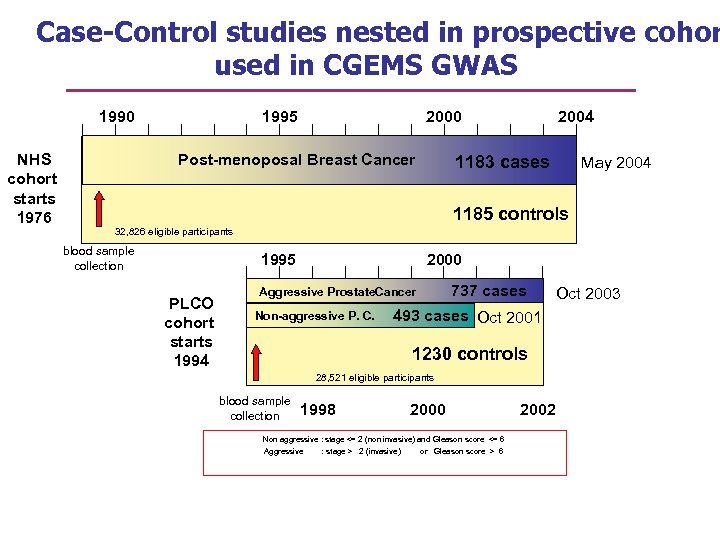 Case-Control studies nested in prospective cohor used in CGEMS GWAS 1990 NHS cohort starts 1976 1995 2000 Post-menoposal Breast Cancer 2004 1183 cases May 2004 1185 controls 32, 826 eligible participants blood sample collection 1995 PLCO cohort starts 1994 2000 Aggressive Prostate. Cancer Non-aggressive P. C. 737 cases 493 cases Oct 2001 1230 controls 28, 521 eligible participants blood sample collection 1998 2000 Non aggressive : stage <= 2 (non invasive) and Gleason score <= 6 Aggressive : stage > 2 (invasive) or Gleason score > 6 2002 Oct 2003
Case-Control studies nested in prospective cohor used in CGEMS GWAS 1990 NHS cohort starts 1976 1995 2000 Post-menoposal Breast Cancer 2004 1183 cases May 2004 1185 controls 32, 826 eligible participants blood sample collection 1995 PLCO cohort starts 1994 2000 Aggressive Prostate. Cancer Non-aggressive P. C. 737 cases 493 cases Oct 2001 1230 controls 28, 521 eligible participants blood sample collection 1998 2000 Non aggressive : stage <= 2 (non invasive) and Gleason score <= 6 Aggressive : stage > 2 (invasive) or Gleason score > 6 2002 Oct 2003
 Genome-Wide Association Studies • The methodology I will describe is now the standard gene-environment analysis at the National Cancer Institute for GWAS • There are now 500, 000 SNP in a typical GWAS, and our method is fast enough to handle this
Genome-Wide Association Studies • The methodology I will describe is now the standard gene-environment analysis at the National Cancer Institute for GWAS • There are now 500, 000 SNP in a typical GWAS, and our method is fast enough to handle this
 Genome-Wide Association Studies • Typically, loci are identified initially for main effects, then followed up for gene-environment interactions • My analyses have come from the PLCO study • In some cases, the “environment” is other genes on different chromosomes, i. e. , gene interactions
Genome-Wide Association Studies • Typically, loci are identified initially for main effects, then followed up for gene-environment interactions • My analyses have come from the PLCO study • In some cases, the “environment” is other genes on different chromosomes, i. e. , gene interactions
 Genome-Wide Association Studies • Despite the fact that the genes are on different chromosomes, they are not always independent • For example they might be in the same pathway
Genome-Wide Association Studies • Despite the fact that the genes are on different chromosomes, they are not always independent • For example they might be in the same pathway
 Genome-Wide Association Studies • When genes on different chromosomes are independenjt, our methods give huge gains in efficiency = “more data” = smaller standard errors • When they are not, our methods give, in effect, the robust method of ordinary logistic regression
Genome-Wide Association Studies • When genes on different chromosomes are independenjt, our methods give huge gains in efficiency = “more data” = smaller standard errors • When they are not, our methods give, in effect, the robust method of ordinary logistic regression
 Summary • Case-control studies are the backbone of epidemiology in general, and genetic epidemiology in particular • Their retrospective nature distinguishes them from random samples = prospective studies
Summary • Case-control studies are the backbone of epidemiology in general, and genetic epidemiology in particular • Their retrospective nature distinguishes them from random samples = prospective studies
 Summary • We start by assuming relationships between the genes and the “environment in the population, ” e. g. , independence • This model can be fully flexible • We also, where necessary, specify distributions for genes
Summary • We start by assuming relationships between the genes and the “environment in the population, ” e. g. , independence • This model can be fully flexible • We also, where necessary, specify distributions for genes
 Summary • We calculated a new likelihood function, leading to more much more precise inferences • The method can handle missing genes , genotyping errors , measurement errors in the environment • Calculations are straightforward via the EM algorithm
Summary • We calculated a new likelihood function, leading to more much more precise inferences • The method can handle missing genes , genotyping errors , measurement errors in the environment • Calculations are straightforward via the EM algorithm
 Summary • Forced to face the dilemma • a
Summary • Forced to face the dilemma • a
 Summary • Forced to face the dilemma • Lousy but robust method • Great but not robust method • We developed a fast, data adaptive, novel way of addressing this issue • In cases where one can predict the outcome, the EB method works as desired
Summary • Forced to face the dilemma • Lousy but robust method • Great but not robust method • We developed a fast, data adaptive, novel way of addressing this issue • In cases where one can predict the outcome, the EB method works as desired


