a013693c592213ed320d5b2c9f4b9294.ppt
- Количество слайдов: 16

Gene 3 D, Orthology and Homology-Based Inheritance of Protein-Protein Interactions Corin Yeats yeats@biochem. ucl. ac. uk http: //gene 3 d. biochem. ucl. ac. uk/

The Gene 3 D Protein Family and Annotation Resource: (1) Identify sequence homologues of CATH domains - HMMs & hit resolution protocol Domain. Finder. Uni. Prot, Ref. Seq, Ensembl (with generous help of SIMAP at MIPS). (2) Integrate with sequence annotation resources. - Pfam, GO, KEGG, Uni. Prot annotation, Int. Act, String Flexible cross-resource comparisons, including CATH PDB domains. (3) Import sequence families - In-house Ortho. Fams, HAMAP, SIMAP clusters.
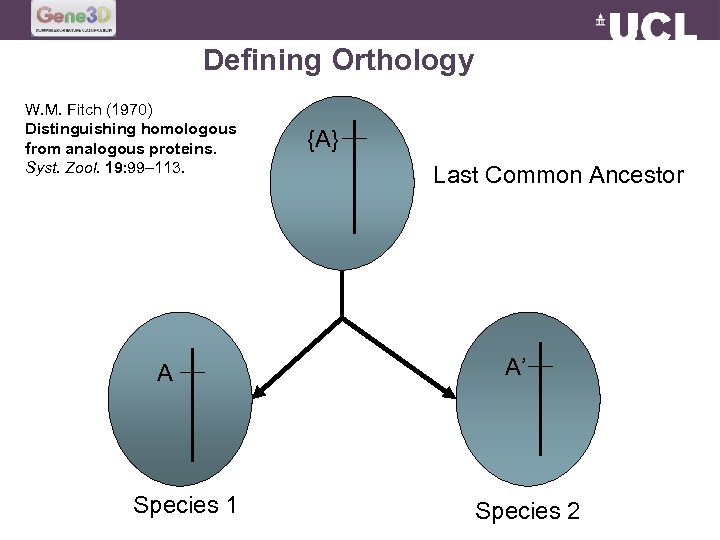
Defining Orthology W. M. Fitch (1970) Distinguishing homologous from analogous proteins. Syst. Zool. 19: 99– 113. A Species 1 {A} Last Common Ancestor A’ Species 2
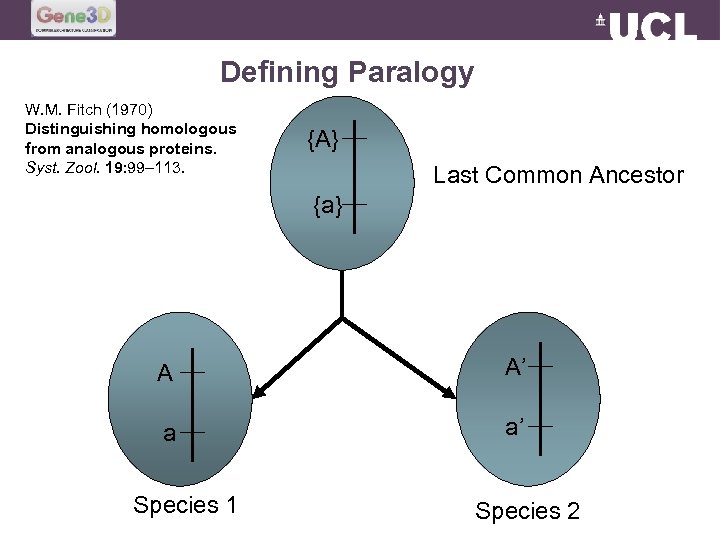
Defining Paralogy W. M. Fitch (1970) Distinguishing homologous from analogous proteins. Syst. Zool. 19: 99– 113. {A} Last Common Ancestor {a} A A’ a a’ Species 1 Species 2
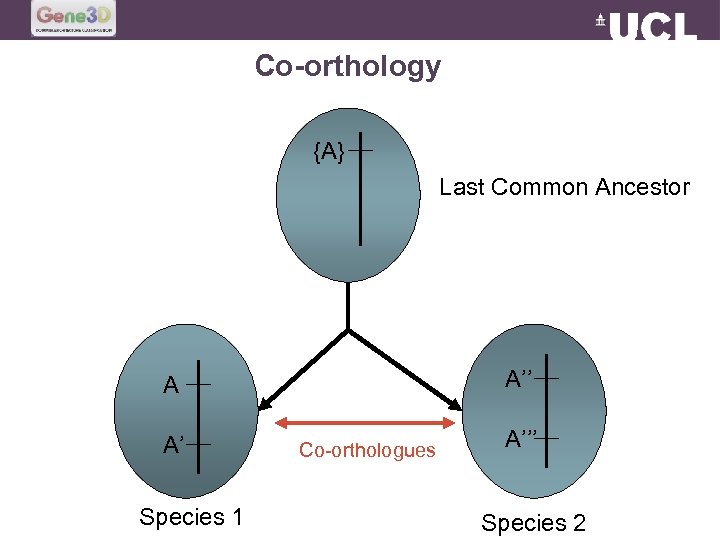
Co-orthology {A} Last Common Ancestor A’’ A A’ Species 1 Co-orthologues A’’’ Species 2

Updating The Terminology: * E. L. L. Sonnhammer & E. V. Koonin (2002) Orthology, paralogy and proposed classification for paralog subtypes. Ti. G 18: 619 -620. • In. Paralogues: – “paralogs in a given lineage that all evolved by gene duplications that happened after the radiation (speciation) event that separated the given lineage from the other lineage under consideration” • Out. Paralogues: – “paralogs in the given lineage that evolved by gene duplications that happened before the radiation (speciation) event”

Defining “Ortholog Families”: • Strict Definition: – Families split at every duplication event. – Many small families. • Normal Definition: – Set root at appropriate level of interest. – Accept inparalogues. – More useful for function prediction.
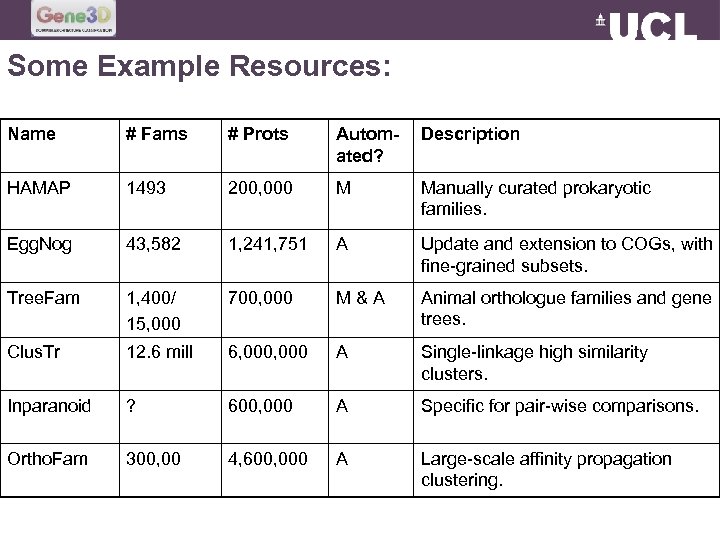
Some Example Resources: Name # Fams # Prots Automated? Description HAMAP 1493 200, 000 M Manually curated prokaryotic families. Egg. Nog 43, 582 1, 241, 751 A Update and extension to COGs, with fine-grained subsets. Tree. Fam 1, 400/ 15, 000 700, 000 M&A Animal orthologue families and gene trees. Clus. Tr 12. 6 mill 6, 000 A Single-linkage high similarity clusters. Inparanoid ? 600, 000 A Specific for pair-wise comparisons. Ortho. Fam 300, 00 4, 600, 000 A Large-scale affinity propagation clustering.
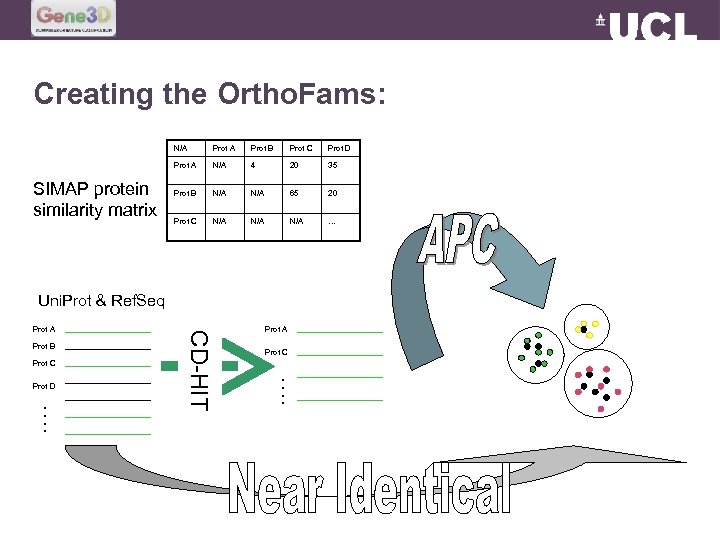
Creating the Ortho. Fams: N/A Prot B Prot C Prot D Prot A SIMAP protein similarity matrix Prot A N/A 4 20 35 Prot B N/A 65 20 Prot C N/A N/A … Uni. Prot & Ref. Seq Prot B Prot C Prot A Prot C …. Prot D CD-HIT Prot A

A Simple Test of the Ortho. Fams: • 99. 9% Ortho. Fams map to one HAMAP family in bacteria. • Each HAMAP family tends to map to several Ortho. Fams => Too conservative? • >80% map to a single KEGG Orthologue term.

Inheriting Protein-Protein Interactions: • Protein-protein interactions (including mechanism) can be conserved after gene duplication and speciation events. • Some interactions are ancient and well conserved, many are not. • Interactions within species are better conserved between homologues than between species. • Interactions are not binary, but are based on affinity • Not all detectable interactions are biologically relevant. Refs: Mika & Rost 2006, Shoemaker & Panchenko 2007

Interaction Inheritance Approaches: • Homology-based approaches have struggled… – Mika & Rost, 2007 • Problems: – High coverage or high quality input, not both. – Interaction networks re-arrange rapidly – No simple universal accurate sequence identity threshold can be found. • Need to separate those that can be inherited reliably, and those that can’t.

The hi. PPI Idea: homology inferred Protein-Protein Interactions (1) Assume Ortho. Fams provide more reliable functional groupings than simple similarity measures. (2) Assume high affinity ~= high conservation ~= low experimental false positive rate. (3) Require more than one piece of supporting evidence.
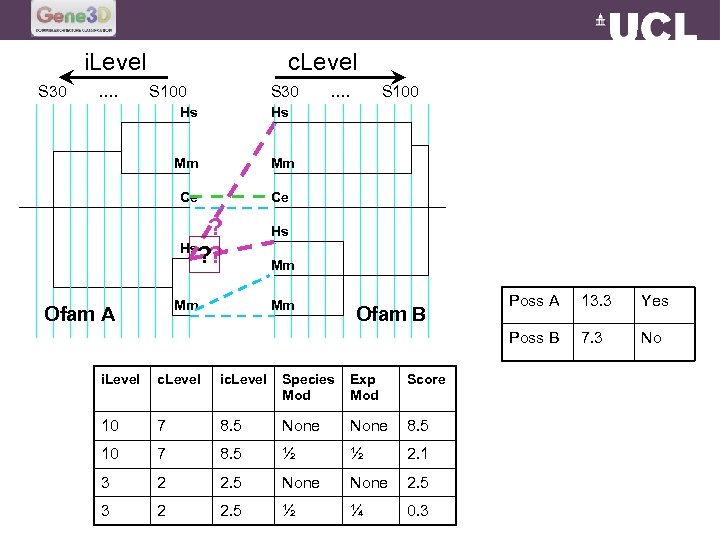
i. Level S 30 …. c. Level S 100 S 30 Hs Mm Ce S 100 Hs Mm …. Ce ? Hs ? ? Hs Mm Ofam A Mm Ofam B i. Level c. Level ic. Level Species Mod Exp Mod Score 10 7 8. 5 None 8. 5 10 7 8. 5 ½ ½ 2. 1 3 2 2. 5 None 2. 5 3 2 2. 5 ½ ¼ 0. 3 Poss A 13. 3 Yes Poss B Mm 7. 3 No
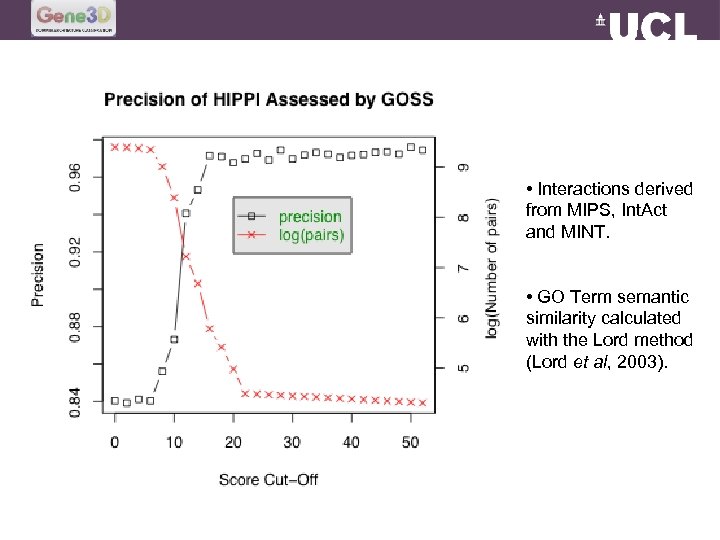
• Interactions derived from MIPS, Int. Act and MINT. • GO Term semantic similarity calculated with the Lord method (Lord et al, 2003).

Links and References http: //gene 3 d. biochem. ucl. ac. uk/ “Gene 3 D: comprehensive structural and functional annotation of genomes” Corin Yeats, Jonathan Lees, Adam Reid, Paul Kellam, Nigel Martin, Xinhui Liu, and Christine Orengo NAR (2008) 36: D 414–D 418.
a013693c592213ed320d5b2c9f4b9294.ppt