dd5bc9a4a09570ea6dc45558068f331d.ppt
- Количество слайдов: 25

Extending FISH Analysis of Paediatric Tumours Rachel Newby Trainee Cytogeneticist Northern Genetics Service

Paediatric Tumours n Alveolar Rhabdomyosarcomas n Ewing’s Tumour n Diagnosis - critical for correct treatment - specific translocations - classical cytogenetics problematic n FISH and RT-PCR are pivotal in getting CORRECT diagnosis
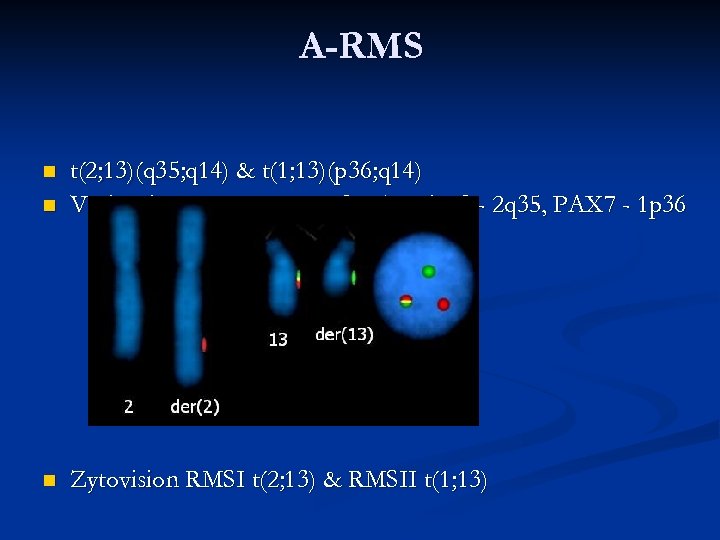
A-RMS n t(2; 13)(q 35; q 14) & t(1; 13)(p 36; q 14) Vysis FKHR (FOXO 1) - 13 q 14, PAX 3 - 2 q 35, PAX 7 - 1 p 36 n Zytovision RMSI t(2; 13) & RMSII t(1; 13) n

Problems with current A-RMS FISH probes n FKHR not always split! - other rearrangements involving PAX genes but not FKHR exist - AFX 1 -PAX 3 fusion (AFX 1 Xq 13) - NCOA 1 -PAX 3 - 2 p 23 in t(2; 2)(q 35; p 23) n Risk of FALSE negative results

Solution? ‘breakapart’ probes - PAX 3 (2 q 35) and PAX 7 (1 p 36) n Identify rearrangements NOT involving FKHR. n BACs - Ensembl -> FISH probes n Verified by FISH and PCR n
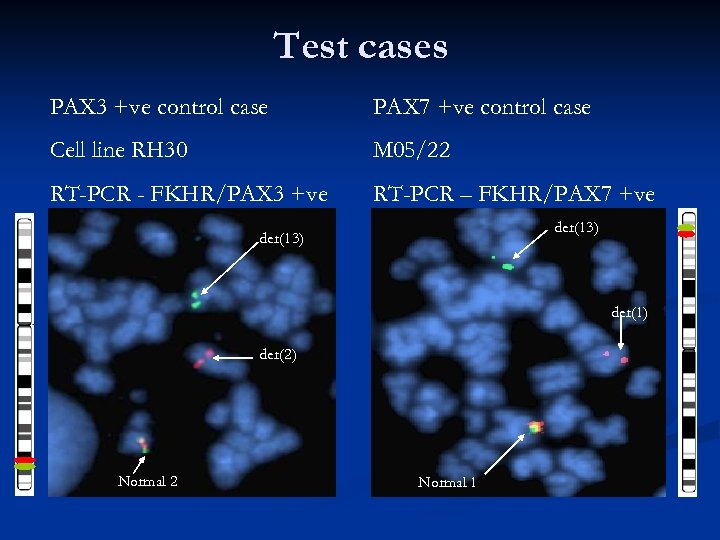
Test cases PAX 3 +ve control case PAX 7 +ve control case Cell line RH 30 M 05/22 RT-PCR - FKHR/PAX 3 +ve RT-PCR – FKHR/PAX 7 +ve der(13) der(1) der(2) Normal 2 Normal 1

Interesting Case 1 Case received from Nottingham n Pathology – A-RMS n Complex karyotype n No visible t(2; 13) or t(1; 13) translocations n Vysis FKHR ‘breakapart’ probe NOT split n n Rearrangement which does not involve FKHR?
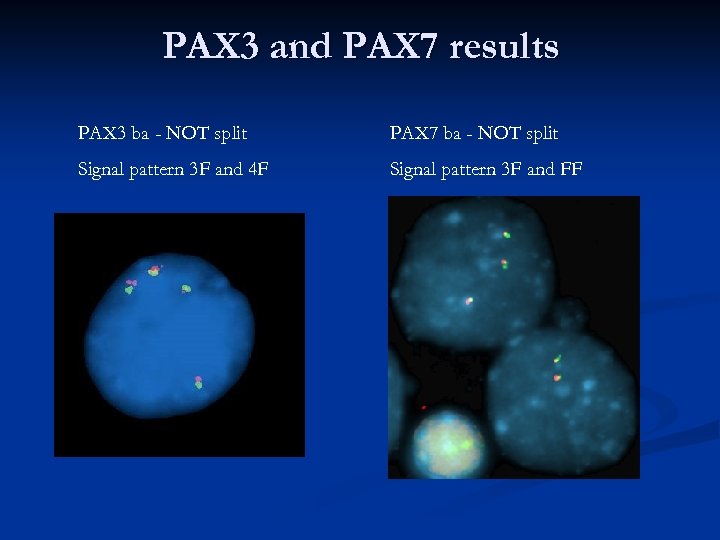
PAX 3 and PAX 7 results PAX 3 ba - NOT split PAX 7 ba - NOT split Signal pattern 3 F and 4 F Signal pattern 3 F and FF

Interesting Case 2 Case NG – 5 year old boy - ? A-RMS n FISH – FKHR NOT split 100% cells n RT-PCR – NO PAX 3 -FKHR or PAX 7 -FKHR fusion transcript n n Rearrangement which does not involve FKHR?
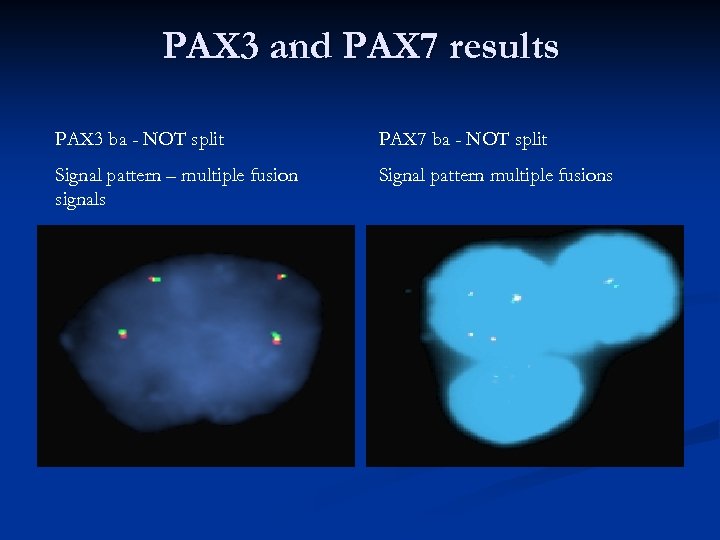
PAX 3 and PAX 7 results PAX 3 ba - NOT split PAX 7 ba - NOT split Signal pattern – multiple fusion signals Signal pattern multiple fusions

A-RMS results n No novel PAX 3 and PAX 7 rearrangements discovered yet. . .
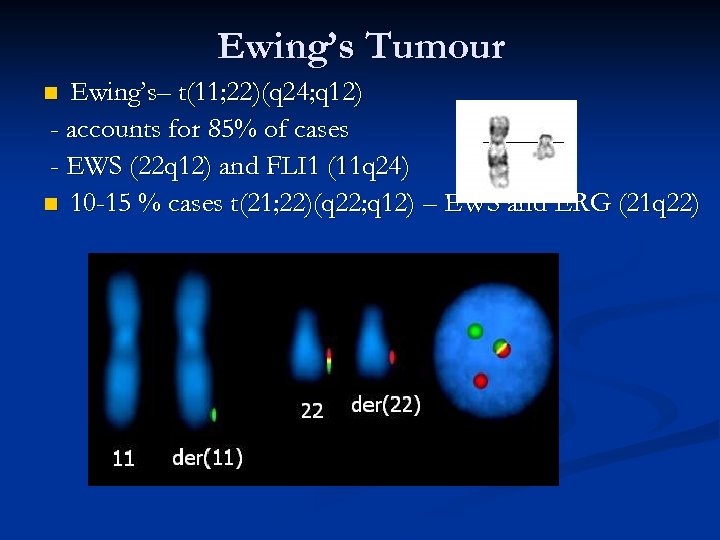
Ewing’s Tumour Ewing’s– t(11; 22)(q 24; q 12) - accounts for 85% of cases - EWS (22 q 12) and FLI 1 (11 q 24) n 10 -15 % cases t(21; 22)(q 22; q 12) – EWS and ERG (21 q 22) n

Problems with current EWS FISH EWS probe splits - which other chromosome is involved? n EWS is not always split! n Risk of false negative result! - detrimental effect on treatment & patient prognosis. n

Solution? Fusion probes for common EWS partners - EWS-FLI 1 - EWS-ERG n ERG ‘breakapart’ probe for rearrangements of 21 q 22. n Useful when no metaphases or EWS NOT split n
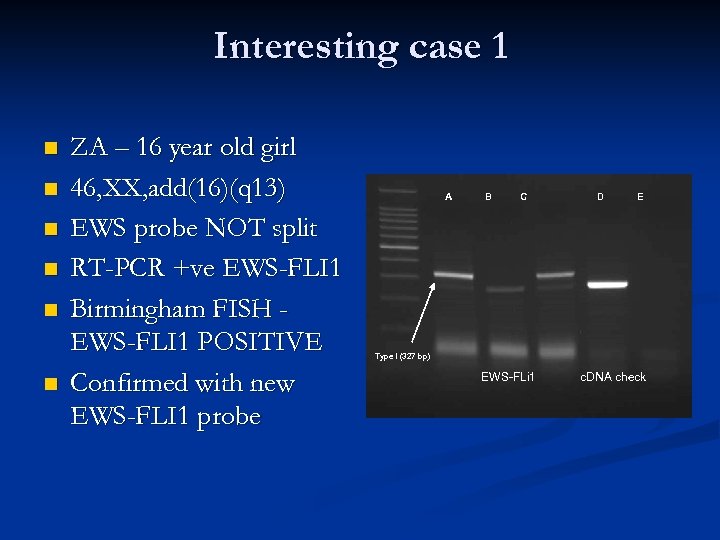
Interesting case 1 n n n ZA – 16 year old girl 46, XX, add(16)(q 13) EWS probe NOT split RT-PCR +ve EWS-FLI 1 Birmingham FISH EWS-FLI 1 POSITIVE Confirmed with new EWS-FLI 1 probe A B C D E Type I (327 bp) EWS-FLi 1 c. DNA check
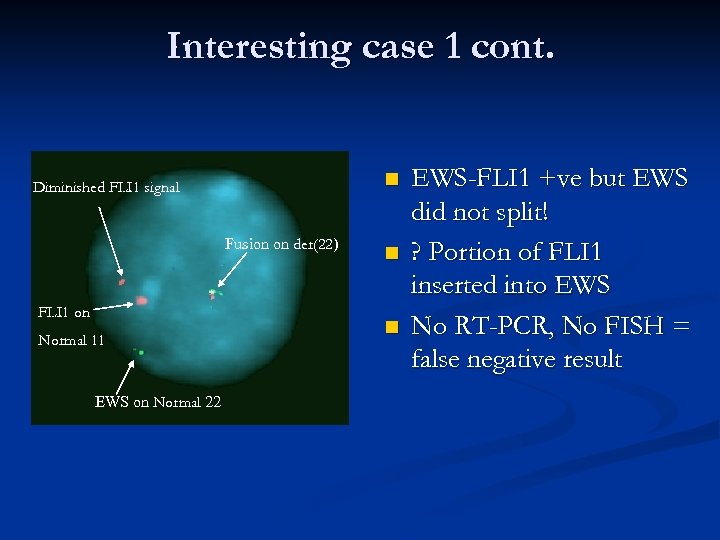
Interesting case 1 cont. n Diminished FLI 1 signal Fusion on der(22) FLI 1 on Normal 11 EWS on Normal 22 n n EWS-FLI 1 +ve but EWS did not split! ? Portion of FLI 1 inserted into EWS No RT-PCR, No FISH = false negative result
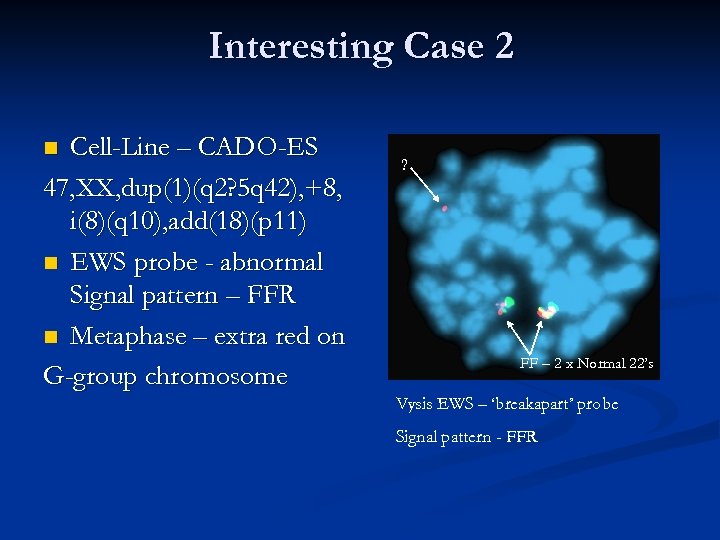
Interesting Case 2 Cell-Line – CADO-ES 47, XX, dup(1)(q 2? 5 q 42), +8, i(8)(q 10), add(18)(p 11) n EWS probe - abnormal Signal pattern – FFR n Metaphase – extra red on G-group chromosome n ? FF – 2 x Normal 22’s Vysis EWS – ‘breakapart’ probe Signal pattern - FFR
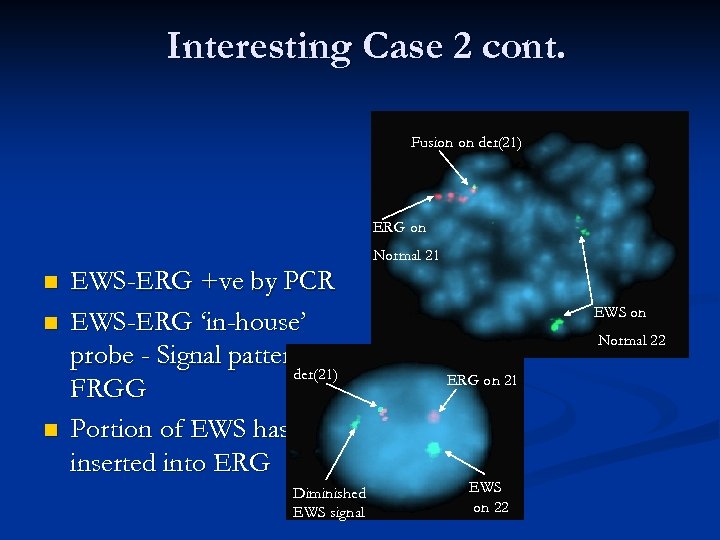
Interesting Case 2 cont. Fusion on der(21) ERG on Normal 21 n n n EWS-ERG +ve by PCR EWS-ERG ‘in-house’ probe - Signal pattern der(21) FRGG Portion of EWS has inserted into ERG Diminished EWS signal EWS on Normal 22 ERG on 21 EWS on 22

Ewing’s results EWS-FLI 1 and EWS-ERG fusion probes - good results on positive controls and archived cases n ERG ‘breakapart’ probe did not split! WHY? n Would expect ERG to split in ~ 10% case n

ERG 3’ to 5’ n EWS 5’ to 3’ n ERG inverted for in-frame fusion gene with EWS n More complex than a translocation n EWS or ERG translocates by insertion-invertion mechanism n ERG never split? n

Summary Probes will benefit the service we provide n PAX 3 and PAX 7 – to be used routinely on new cases n PAX probes - FKHR is not split. n n Reduce - false negative results

Summary ERG – better understanding of complexity n EWS split -> EWS-FLI 1 and EWS-ERG n Increased confidence n Microinsertions n Commercial probes - a false negative result n Without extended FISH or RT-PCR = PITFALLS
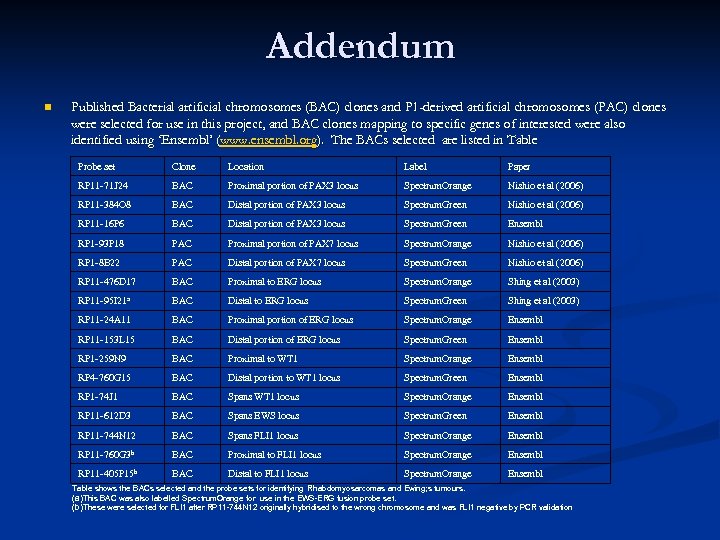
Addendum n Published Bacterial artificial chromosomes (BAC) clones and P 1 -derived artificial chromosomes (PAC) clones were selected for use in this project, and BAC clones mapping to specific genes of interested were also identified using ‘Ensembl’ (www. ensembl. org). The BACs selected are listed in Table Probe set Clone Location Label Paper RP 11 -71 J 24 BAC Proximal portion of PAX 3 locus Spectrum. Orange Nishio et al (2006) RP 11 -384 O 8 BAC Distal portion of PAX 3 locus Spectrum. Green Nishio et al (2006) RP 11 -16 P 6 BAC Distal portion of PAX 3 locus Spectrum. Green Ensembl RP 1 -93 P 18 PAC Proximal portion of PAX 7 locus Spectrum. Orange Nishio et al (2006) RP 1 -8 B 22 PAC Distal portion of PAX 7 locus Spectrum. Green Nishio et al (2006) RP 11 -476 D 17 BAC Proximal to ERG locus Spectrum. Orange Shing et al (2003) RP 11 -95 I 21 a BAC Distal to ERG locus Spectrum. Green Shing et al (2003) RP 11 -24 A 11 BAC Proximal portion of ERG locus Spectrum. Orange Ensembl RP 11 -153 L 15 BAC Distal portion of ERG locus Spectrum. Green Ensembl RP 1 -259 N 9 BAC Proximal to WT 1 Spectrum. Orange Ensembl RP 4 -760 G 15 BAC Distal portion to WT 1 locus Spectrum. Green Ensembl RP 1 -74 J 1 BAC Spans WT 1 locus Spectrum. Orange Ensembl RP 11 -612 D 3 BAC Spans EWS locus Spectrum. Green Ensembl RP 11 -744 N 12 BAC Spans FLI 1 locus Spectrum. Orange Ensembl RP 11 -760 G 3 b BAC Proximal to FLI 1 locus Spectrum. Orange Ensembl RP 11 -405 P 15 b BAC Distal to FLI 1 locus Spectrum. Orange Ensembl Table shows the BACs selected and the probe sets for identifying Rhabdomyosarcomas and Ewing; s tumours. (a)This BAC was also labelled Spectrum. Orange for use in the EWS-ERG fusion probe set. (b)These were selected for FLI 1 after RP 11 -744 N 12 originally hybridised to the wrong chromosome and was FLI 1 negative by PCR validation

Addendum n BAC’s were ordered from BACPAC CHORI (Children’s Hospital Oakland Research Institute) grown up and DNA extracted using the Qiagen plasmid preparation kit and then fluorescent labelled with either Spectrum. Orange or Spectrum. Green using the Vysis Nick translation Kit. n Ref: Danielle C. Shing, Dominic J. Mc. Mullan, Paul Roberts, Kim Smith, Suet-Feung Chin, James Nicholson, Roger M. Tillman, Pramila Ramani, Catherine Cullinane, and Nicholas Coleman. FUS/ERG Gene fusions in Ewing’s Tumours, Cancer Research 63, 4568 -4576, August 1, 2003. n n Jun Nishio, Pamel A Althof, Jacqueline M Bailey, Ming Zhou, James. R Neff, Frederic G Barr, David M Parham, Lisa Teot, Stephen J Qualman and Julia A Bridge. Use of Novel FISH assay on paraffin-embedded tissues as an adjunct to diagnosis of alveolar rhabdomyosarcoma. Laboratory Investigation (2006) 86, 547 -556. n Georges Maire, Christopher W. Brown, Jane Bayani, Carlos Pereira, Denis H. Gravel, John C. Bell, Maria Zielenska, Jeremy A. Squire. Complex rearrangement of chromosomes 19, 21, and 22 in Ewing sarcoma involving a novel reciprocal inversion-insertion mechanism of EWS-ERG fusion gene formation; a case analysis and literature review. Cancer Genetics and Cytogenetics 181 (2008) 81 -92

Acknowledgements n Thanks to - Nick Bown - Fiona Harding - Steve Hellens - Malignancy Section at the Northern Genetics Service - Meg Heath, Kate Martin & Tom Mc. Culloch Nottingham Cytogenetics lab - Dom Mc. Mullan – Birmingham Cytogenetics Lab
dd5bc9a4a09570ea6dc45558068f331d.ppt