ef8ffa3c7da67ebf84421690770824ea.ppt
- Количество слайдов: 51

Exploring Peritumoral White Matter Fibers for Neurosurgical Planning Sonia Pujol, Ph. D. Ron Kikinis, M. D.

3 D Slicer • An end-user application for image analysis • An open-source environment for software development • A software platform that is both easy to use for clinical researchers and easy to extend for programmers White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011

Download Slicer 3. 6 • Download and install the Slicer 3. 6. 3 release version software from the Slicer web site http: //www. slicer. org/pages/Special: Slicer. Downloads Disclaimer It is the responsibility of the user of 3 DSlicer to comply with both the terms of the license and with the applicable laws, regulations and rules. White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011

Pre-Requisite • This course supposes that you have taken the “Slicer 3 Data Loading and Visualization” tutorial http: //www. slicer. org/slicer. Wiki/index. php/Slicer 3. 6: Training#Software_tutorials White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
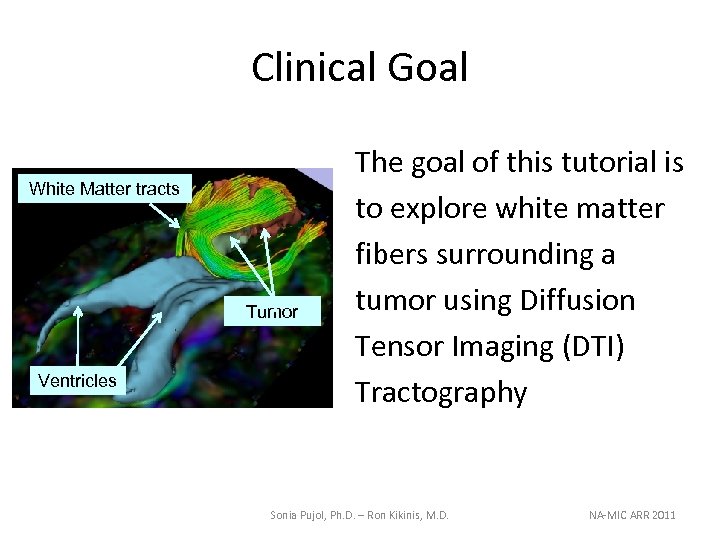
Clinical Goal White Matter tracts Tumor Ventricles The goal of this tutorial is to explore white matter fibers surrounding a tumor using Diffusion Tensor Imaging (DTI) Tractography Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
Overview of the analysis pipeline • Part 1: Loading &Visualization of Diffusion Data • Part 1: Segmentation of the ventricles, and solid and cystic parts of the tumor • Part 2: Tractography reconstruction of the white matter fibers in the peri-tumoral volume • Part 3: Tractography exploration of the ipsilateral and contralateral fibers tracts Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
Part 1: Loading and Visualization of Diffusion Data White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
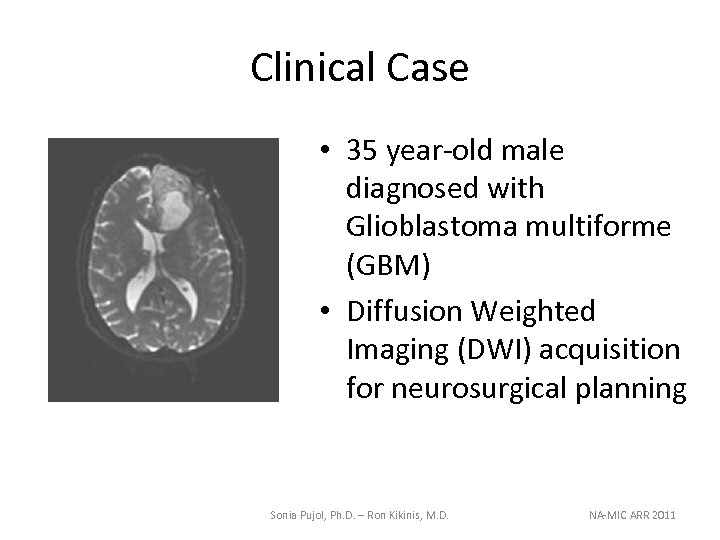
Clinical Case • 35 year-old male diagnosed with Glioblastoma multiforme (GBM) • Diffusion Weighted Imaging (DWI) acquisition for neurosurgical planning Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
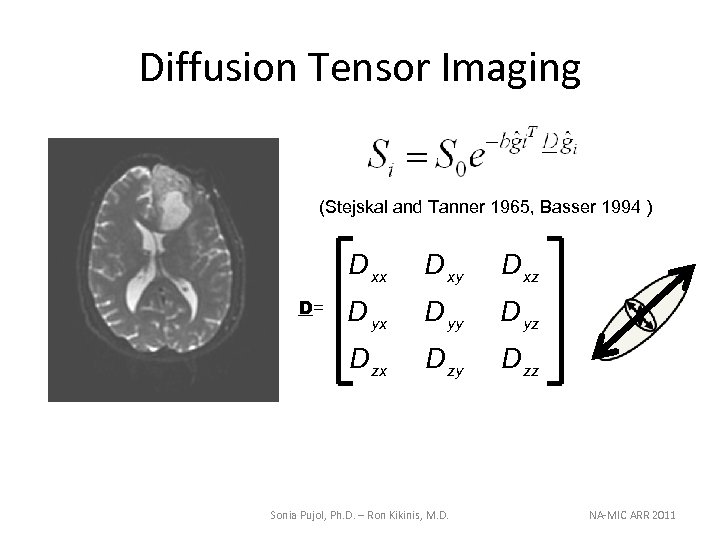
Diffusion Tensor Imaging (Stejskal and Tanner 1965, Basser 1994 ) D xx D xz D yx D yy D yz D zx D= D xy D zz Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
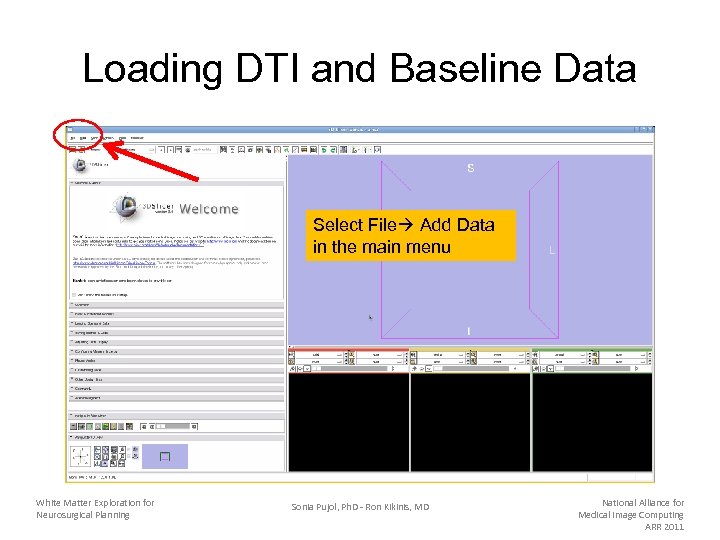
Loading DTI and Baseline Data Select File Add Data in the main menu White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
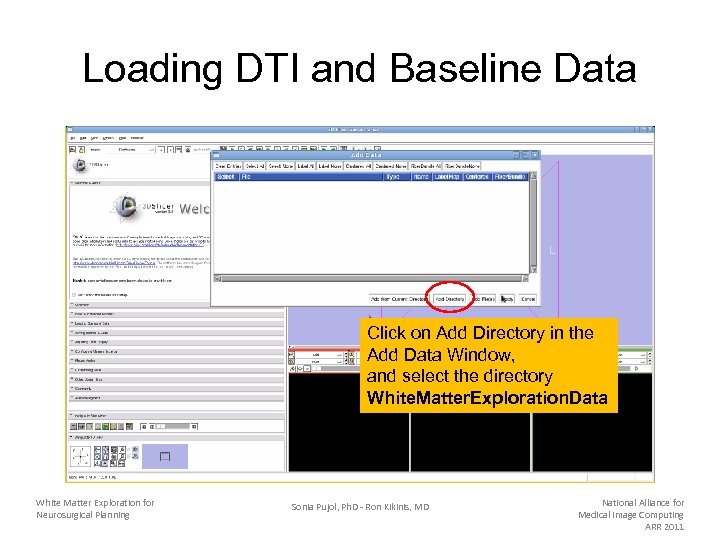
Loading DTI and Baseline Data Click on Add Directory in the Add Data Window, and select the directory White. Matter. Exploration. Data White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
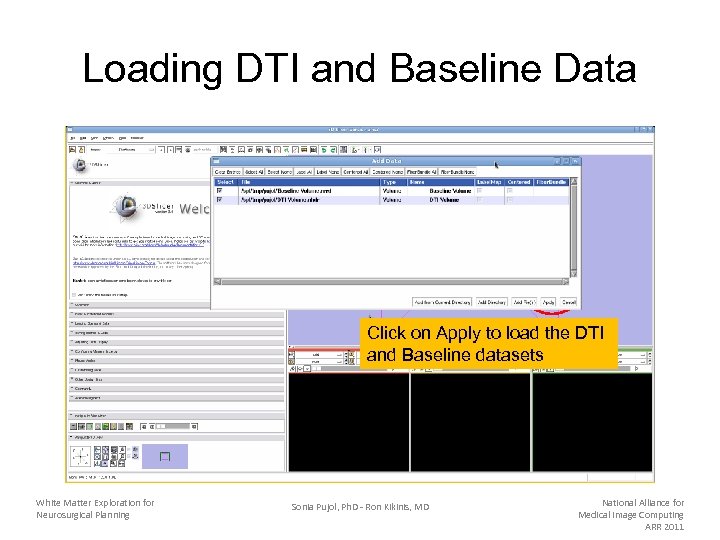
Loading DTI and Baseline Data Click on Apply to load the DTI and Baseline datasets White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
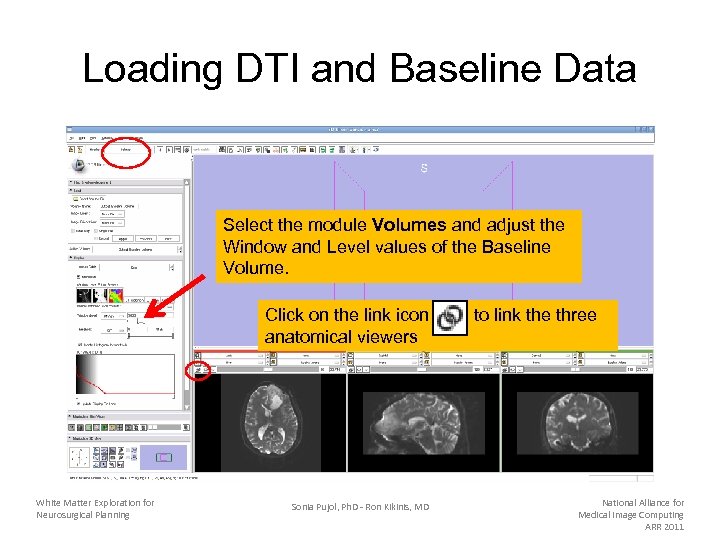

Loading DTI and Baseline Data Select the module Volumes and adjust the Window and Level values of the Baseline Volume. Click on the link icon anatomical viewers White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD to link the three National Alliance for Medical Image Computing ARR 2011
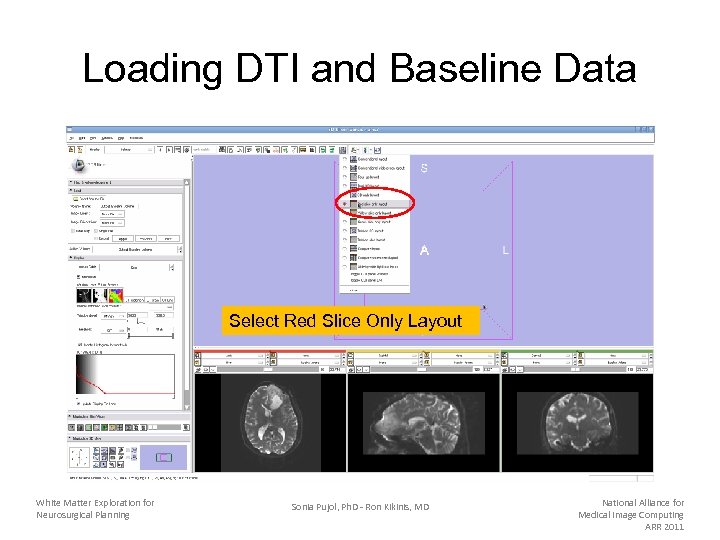
Loading DTI and Baseline Data Select Red Slice Only Layout White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
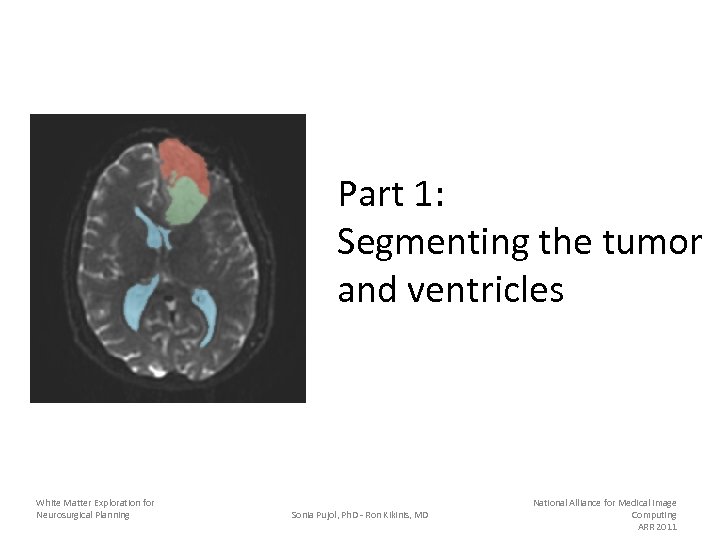
Part 1: Segmenting the tumor and ventricles White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
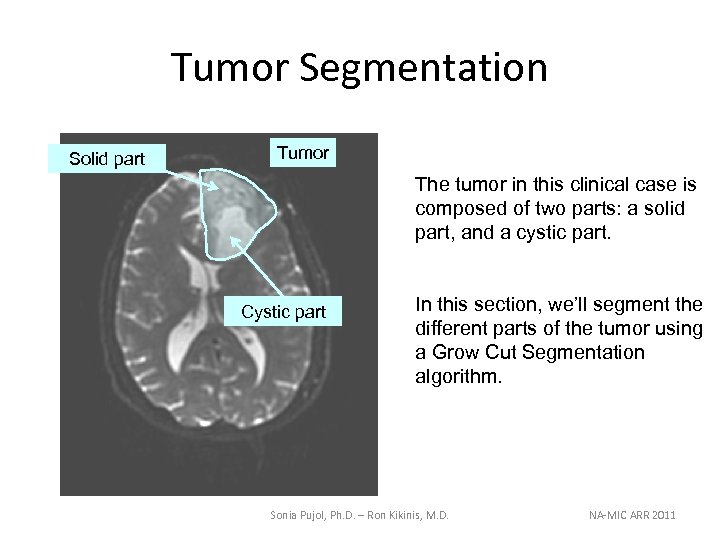
Tumor Segmentation Solid part Tumor The tumor in this clinical case is composed of two parts: a solid part, and a cystic part. Cystic part In this section, we’ll segment the different parts of the tumor using a Grow Cut Segmentation algorithm. Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
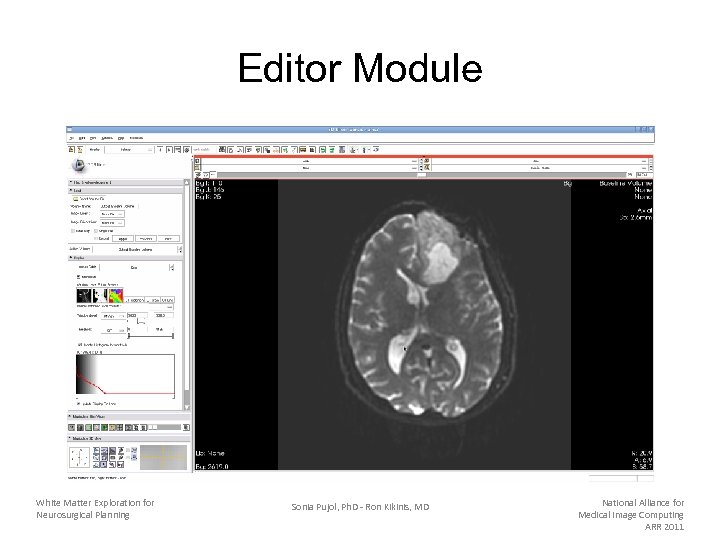
Editor Module White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
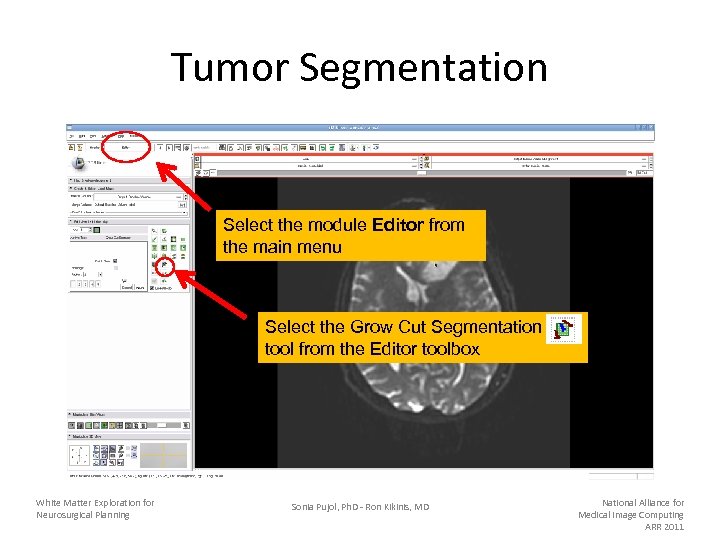
Tumor Segmentation Select the module Editor from the main menu Select the Grow Cut Segmentation tool from the Editor toolbox White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
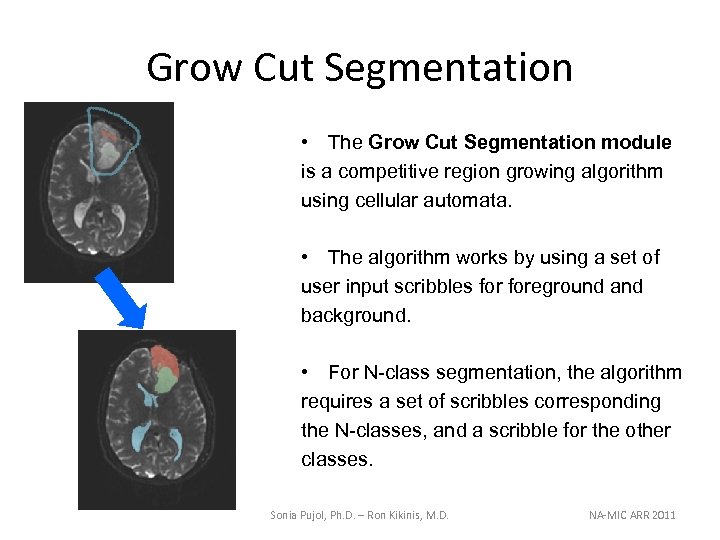
Grow Cut Segmentation • The Grow Cut Segmentation module is a competitive region growing algorithm using cellular automata. • The algorithm works by using a set of user input scribbles foreground and background. • For N-class segmentation, the algorithm requires a set of scribbles corresponding the N-classes, and a scribble for the other classes. Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
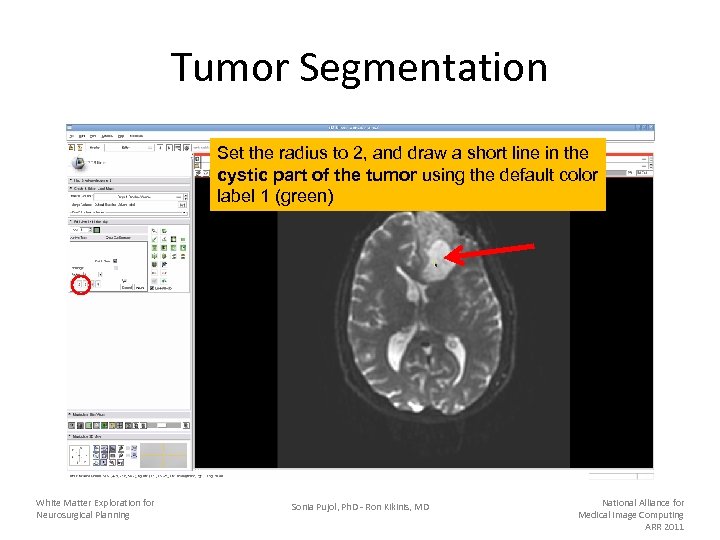
Tumor Segmentation Set the radius to 2, and draw a short line in the cystic part of the tumor using the default color label 1 (green) White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
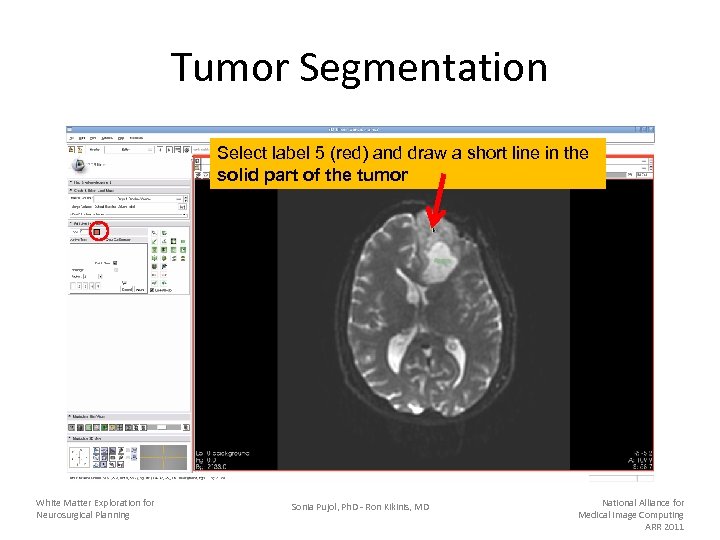
Tumor Segmentation Select label 5 (red) and draw a short line in the solid part of the tumor White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
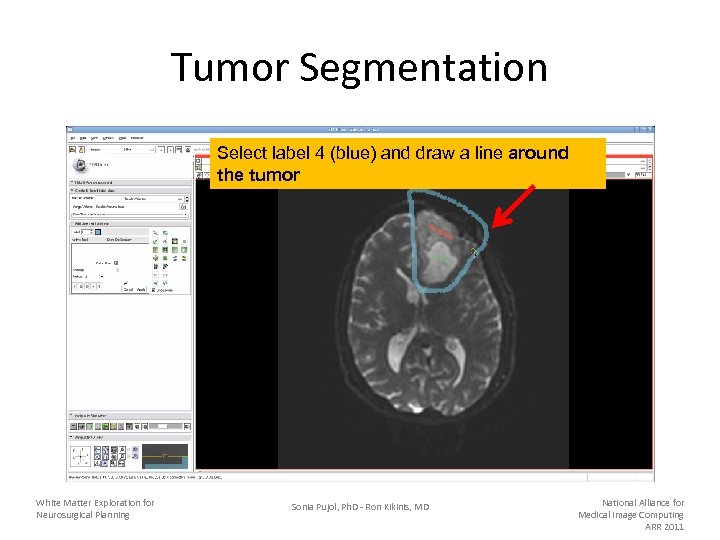
Tumor Segmentation Select label 4 (blue) and draw a line around the tumor White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
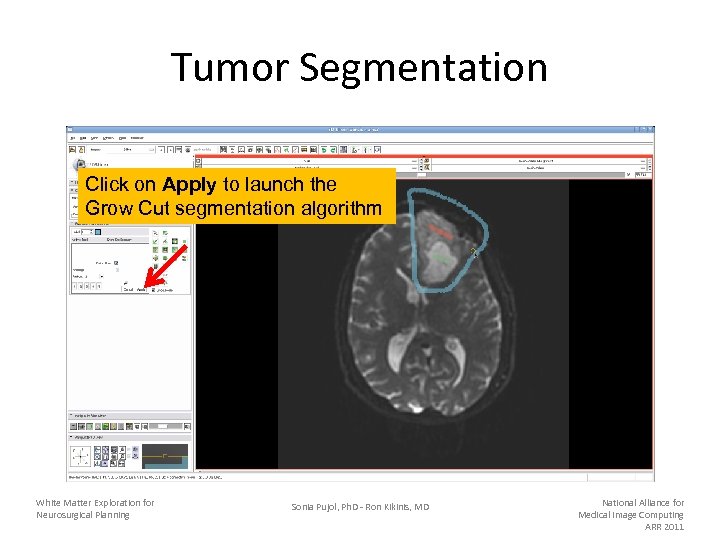
Tumor Segmentation Click on Apply to launch the Grow Cut segmentation algorithm White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
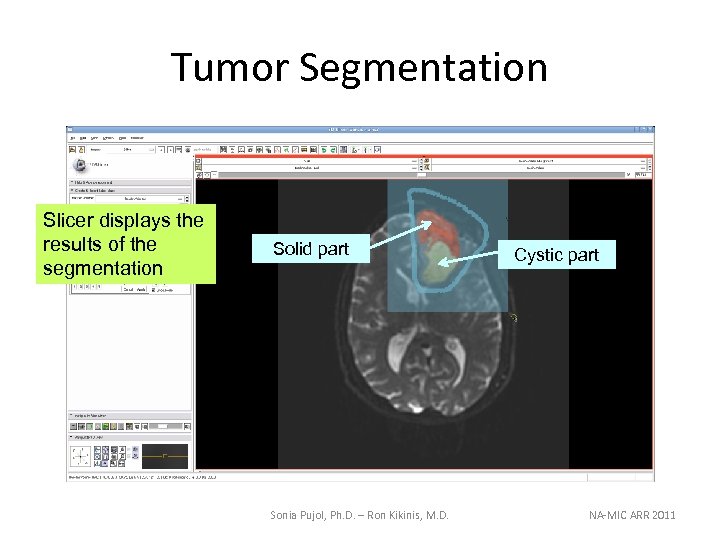
Tumor Segmentation Slicer displays the results of the segmentation Solid part Sonia Pujol, Ph. D. – Ron Kikinis, M. D. Cystic part NA-MIC ARR 2011
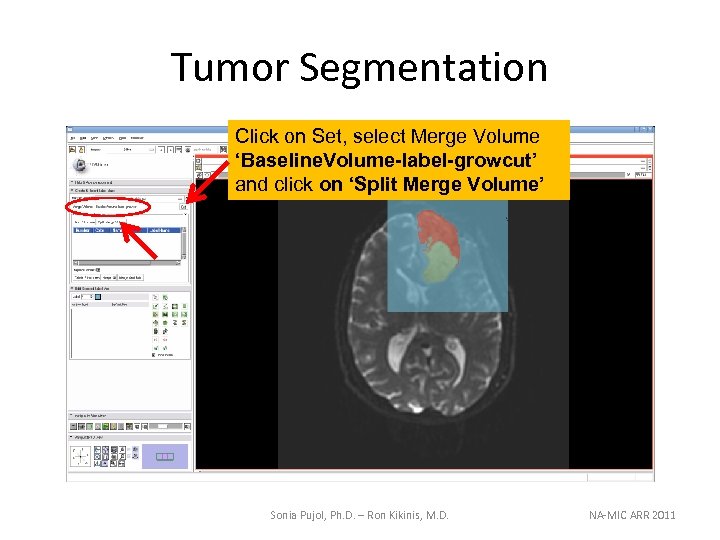
Tumor Segmentation Click on Set, select Merge Volume ‘Baseline. Volume-label-growcut’ and click on ‘Split Merge Volume’ Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
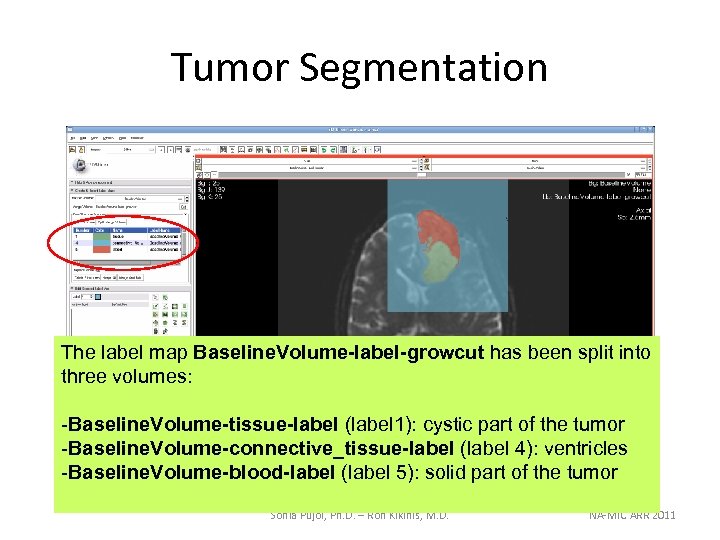
Tumor Segmentation The label map Baseline. Volume-label-growcut has been split into three volumes: -Baseline. Volume-tissue-label (label 1): cystic part of the tumor -Baseline. Volume-connective_tissue-label (label 4): ventricles -Baseline. Volume-blood-label (label 5): solid part of the tumor Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
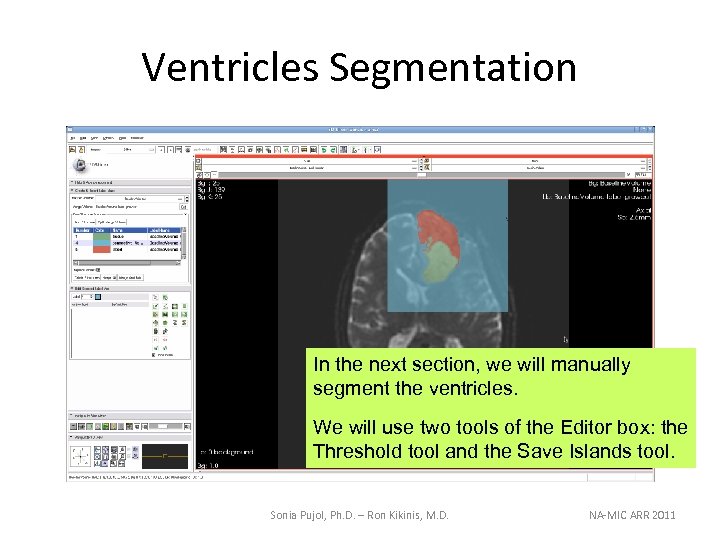
Ventricles Segmentation In the next section, we will manually segment the ventricles. We will use two tools of the Editor box: the Threshold tool and the Save Islands tool. Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
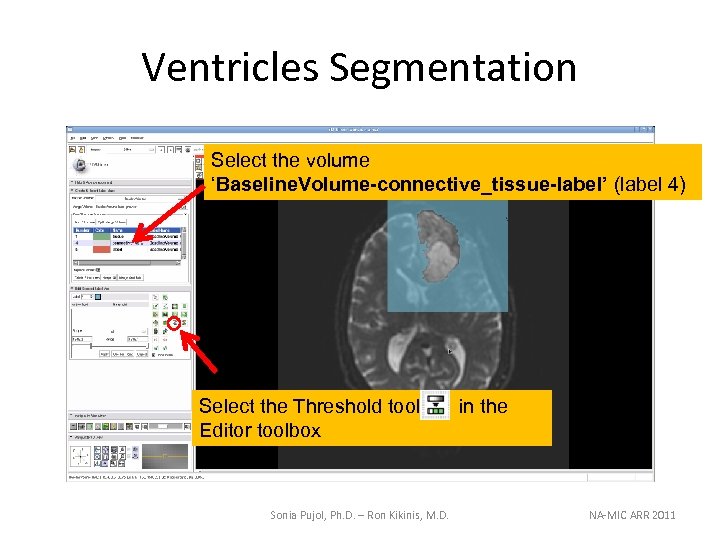
Ventricles Segmentation Select the volume ‘Baseline. Volume-connective_tissue-label’ (label 4) Select the Threshold tool Editor toolbox Sonia Pujol, Ph. D. – Ron Kikinis, M. D. in the NA-MIC ARR 2011
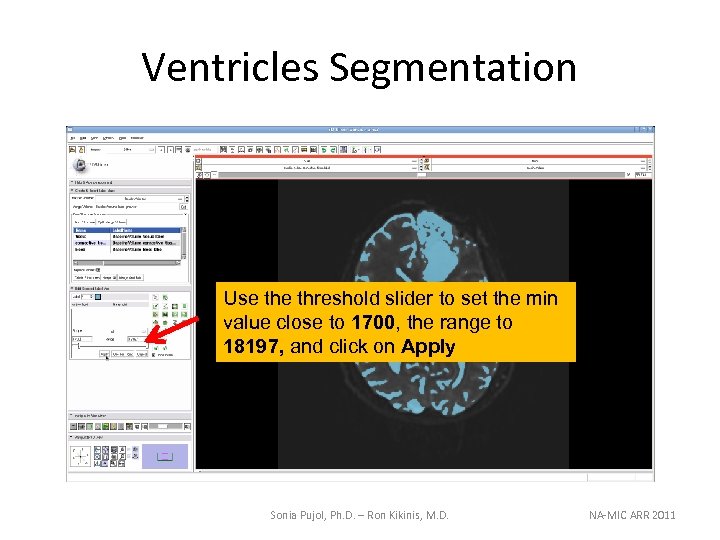
Ventricles Segmentation Use threshold slider to set the min value close to 1700, the range to 18197, and click on Apply Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
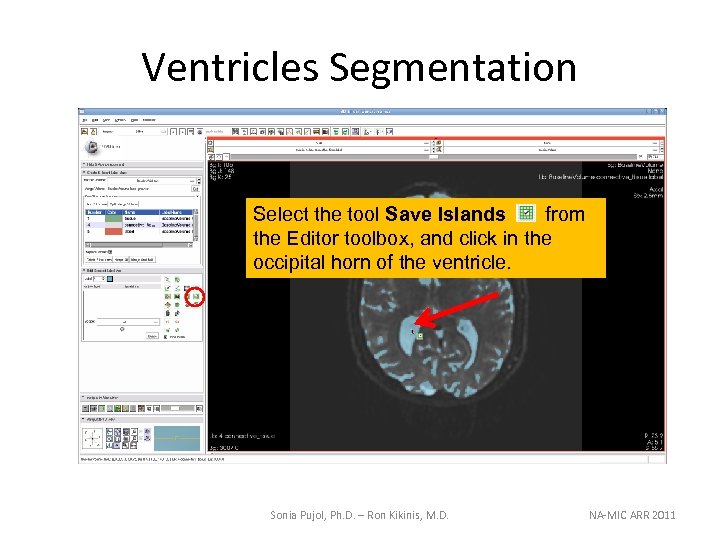
Ventricles Segmentation Select the tool Save Islands from the Editor toolbox, and click in the occipital horn of the ventricle. Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
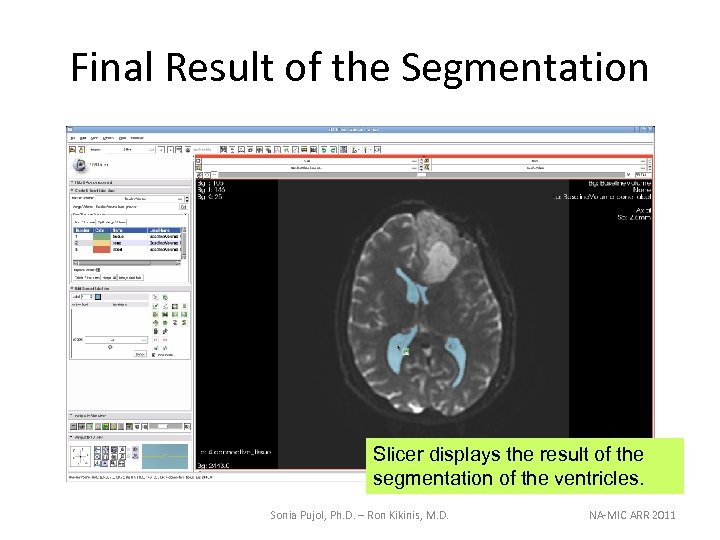
Final Result of the Segmentation Slicer displays the result of the segmentation of the ventricles. Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
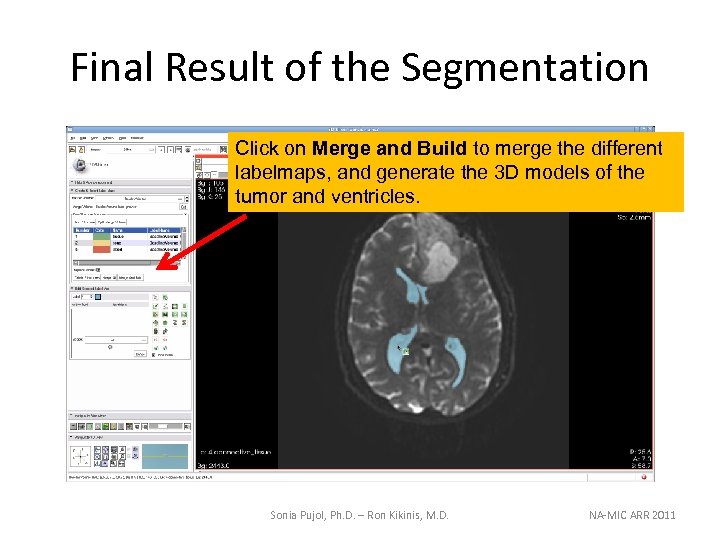
Final Result of the Segmentation Click on Merge and Build to merge the different labelmaps, and generate the 3 D models of the tumor and ventricles. Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
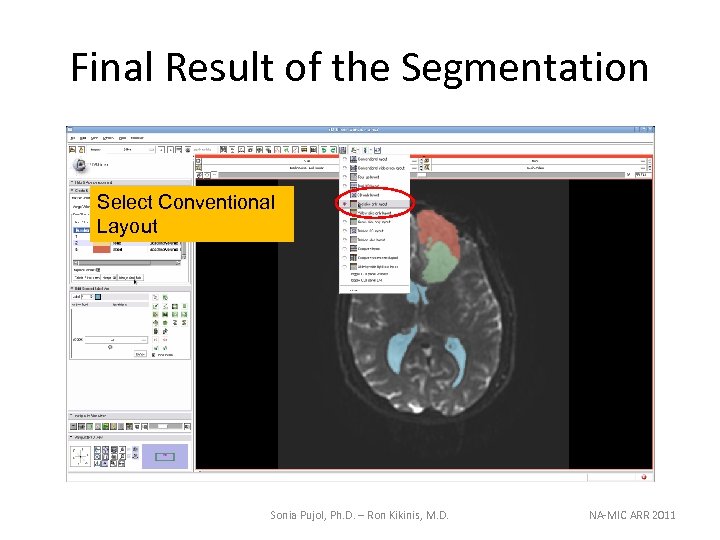
Final Result of the Segmentation Select Conventional Layout Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
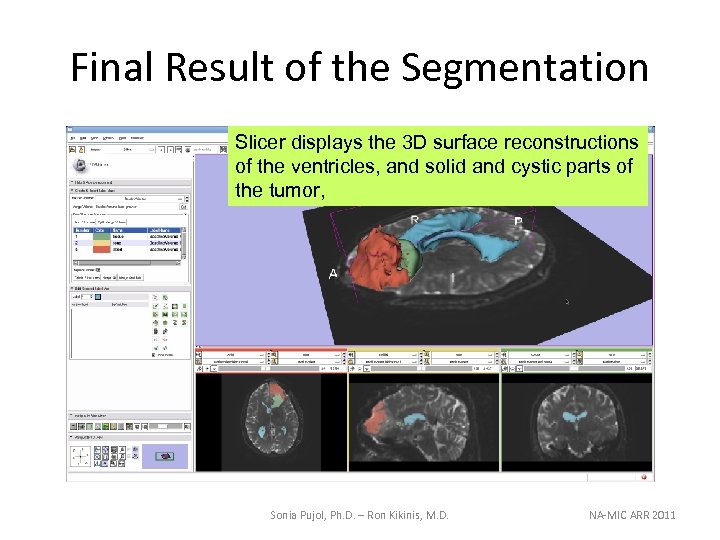
Final Result of the Segmentation Slicer displays the 3 D surface reconstructions of the ventricles, and solid and cystic parts of the tumor, Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
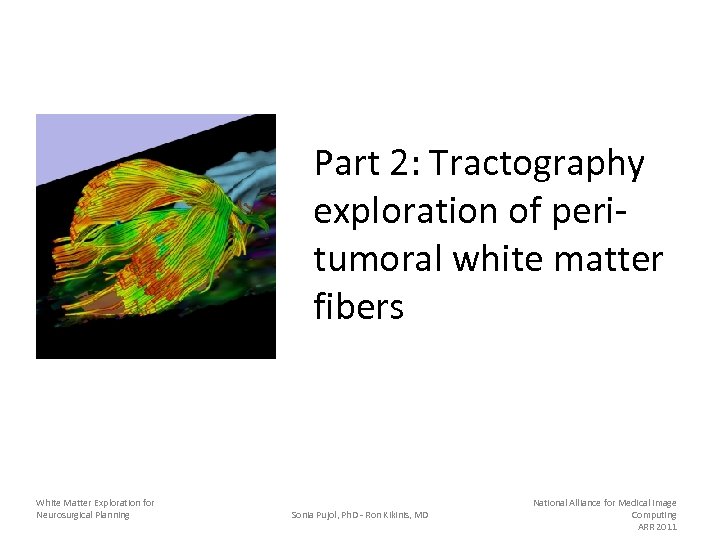
Part 2: Tractography exploration of peritumoral white matter fibers White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
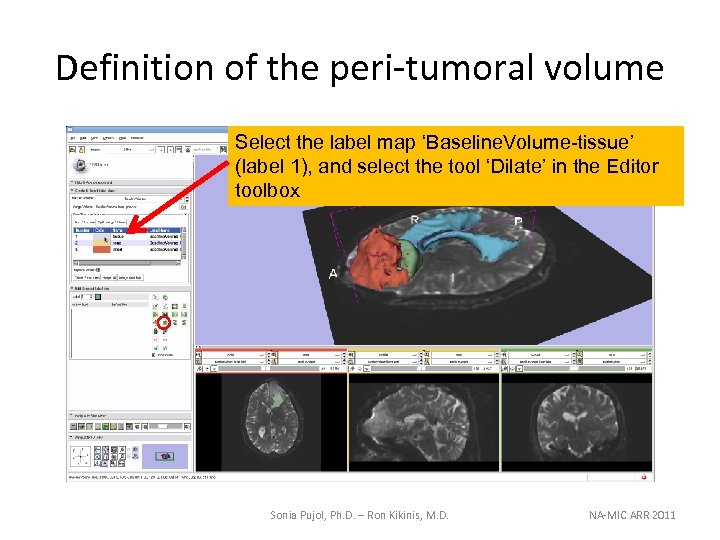
Definition of the peri-tumoral volume Select the label map ‘Baseline. Volume-tissue’ (label 1), and select the tool ‘Dilate’ in the Editor toolbox Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
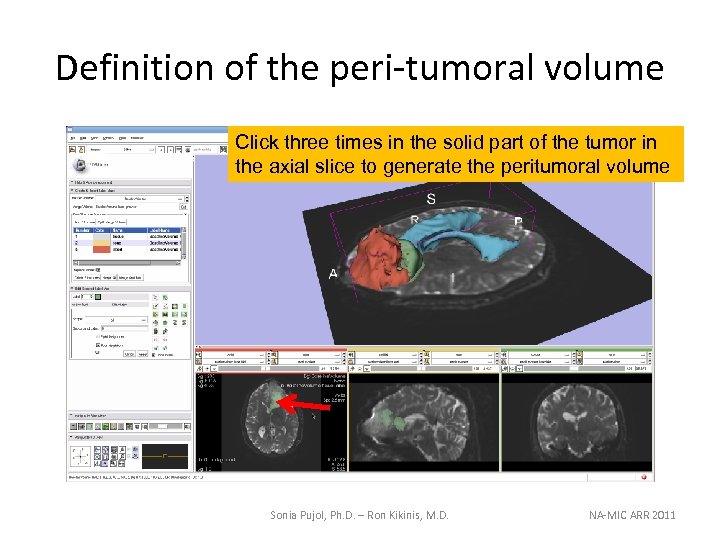
Definition of the peri-tumoral volume Click three times in the solid part of the tumor in the axial slice to generate the peritumoral volume Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
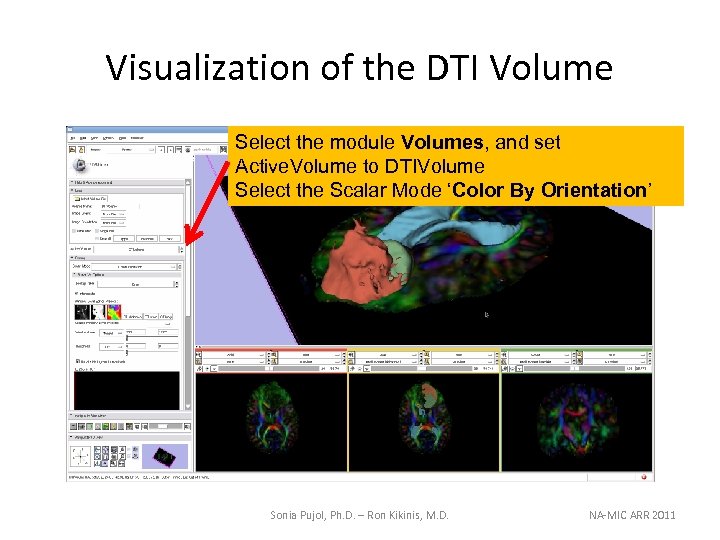
Visualization of the DTI Volume Select the module Volumes, and set Active. Volume to DTIVolume Select the Scalar Mode ‘Color By Orientation’ Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
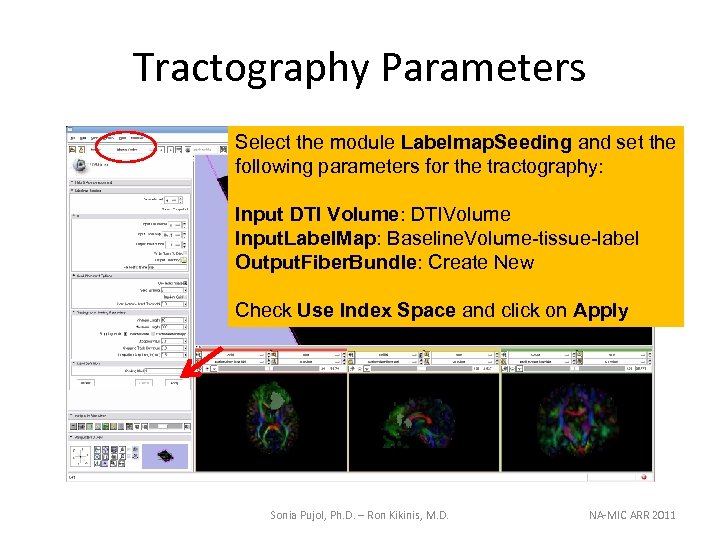
Tractography Parameters Select the module Labelmap. Seeding and set the following parameters for the tractography: Input DTI Volume: DTIVolume Input. Label. Map: Baseline. Volume-tissue-label Output. Fiber. Bundle: Create New Check Use Index Space and click on Apply Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
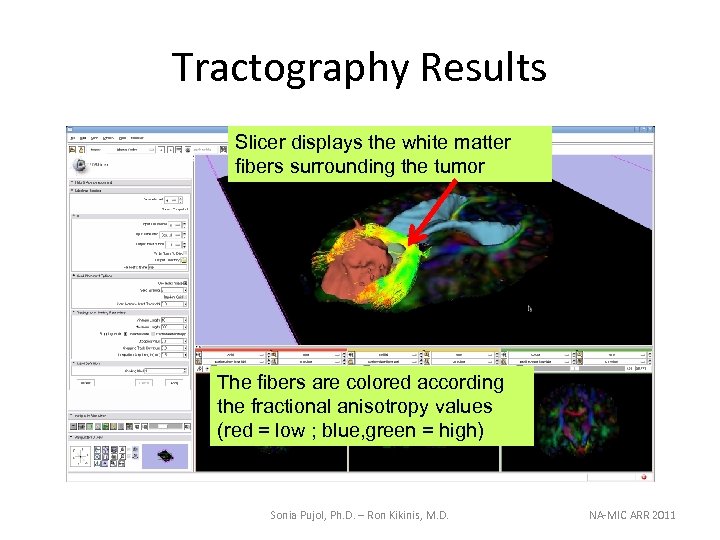
Tractography Results Slicer displays the white matter fibers surrounding the tumor The fibers are colored according the fractional anisotropy values (red = low ; blue, green = high) Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
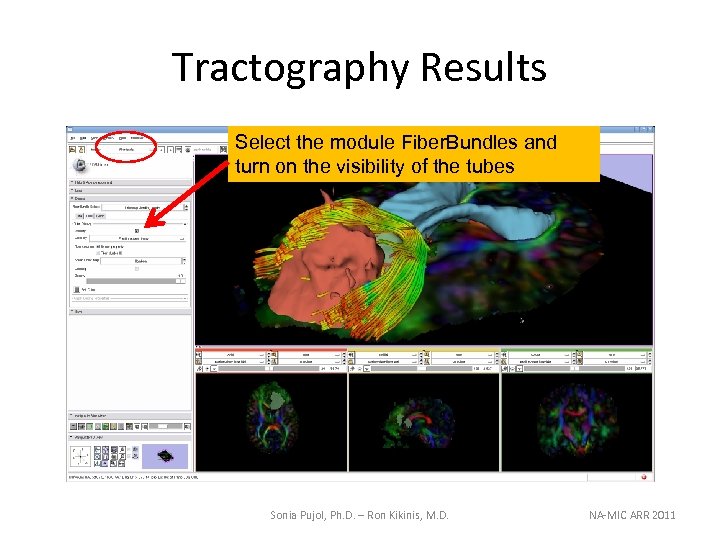
Tractography Results Select the module Fiber. Bundles and turn on the visibility of the tubes Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
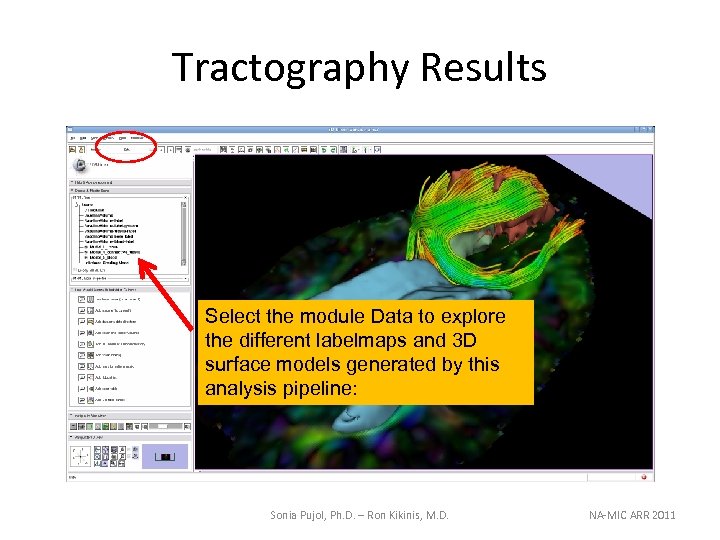
Tractography Results Select the module Data to explore the different labelmaps and 3 D surface models generated by this analysis pipeline: Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011

Part 3: Tractography exploration of the contralateral side White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
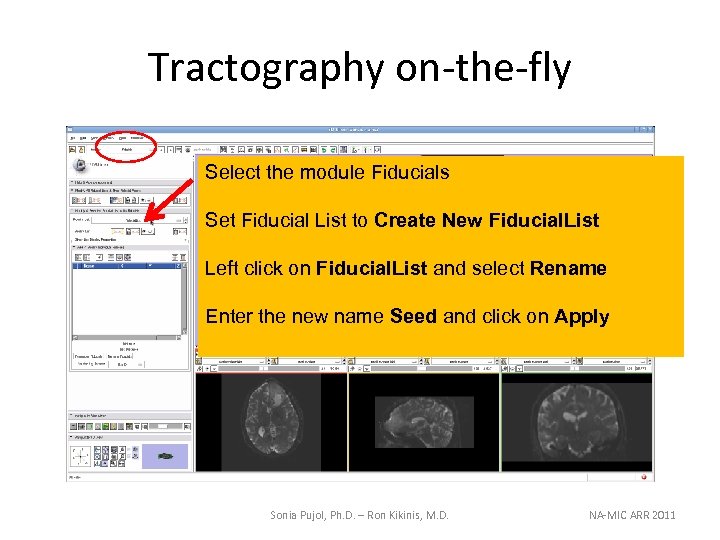
Tractography on-the-fly Select the module Fiducials Set Fiducial List to Create New Fiducial. List Left click on Fiducial. List and select Rename Enter the new name Seed and click on Apply Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
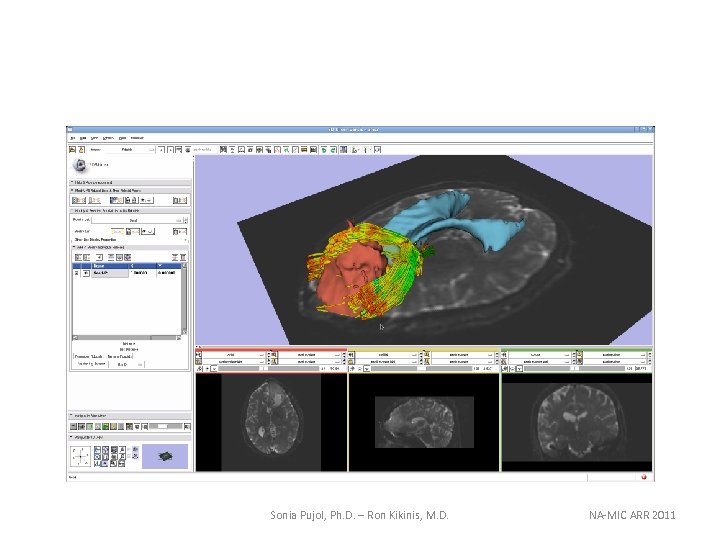
Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
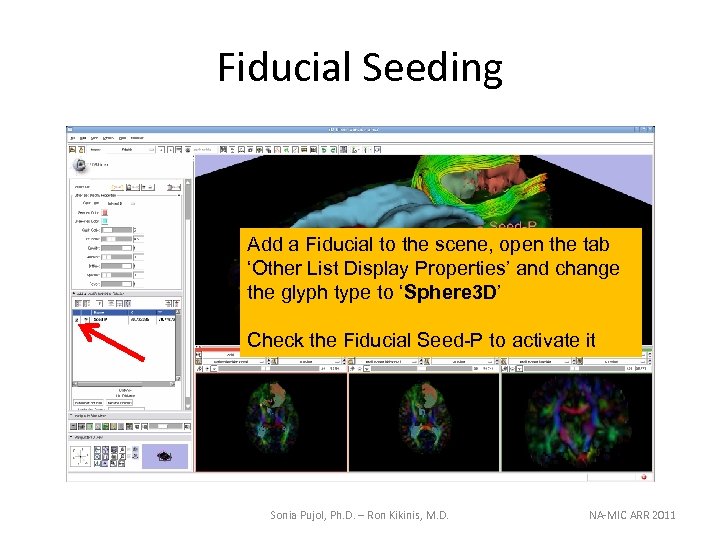
Fiducial Seeding Add a Fiducial to the scene, open the tab ‘Other List Display Properties’ and change the glyph type to ‘Sphere 3 D’ Check the Fiducial Seed-P to activate it Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
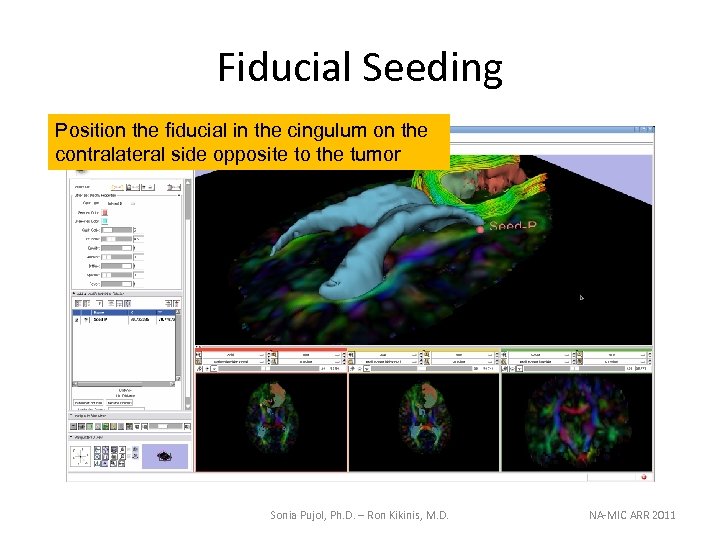
Fiducial Seeding Position the fiducial in the cingulum on the contralateral side opposite to the tumor Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
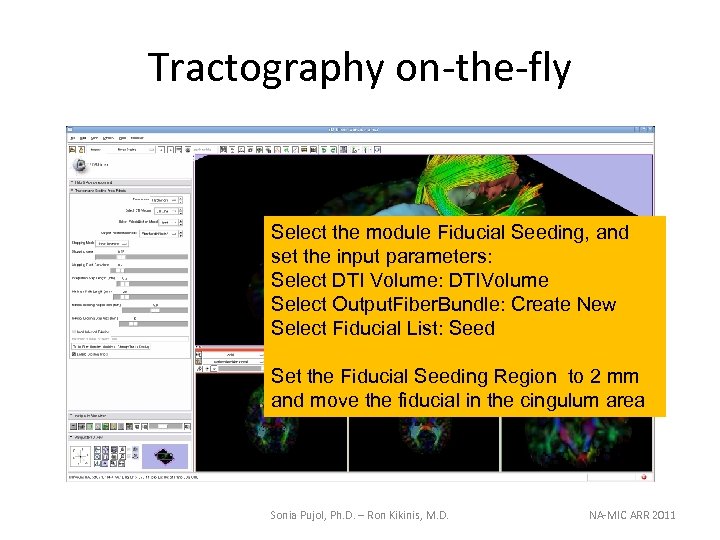
Tractography on-the-fly Select the module Fiducial Seeding, and set the input parameters: Select DTI Volume: DTIVolume Select Output. Fiber. Bundle: Create New Select Fiducial List: Seed Set the Fiducial Seeding Region to 2 mm and move the fiducial in the cingulum area Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011
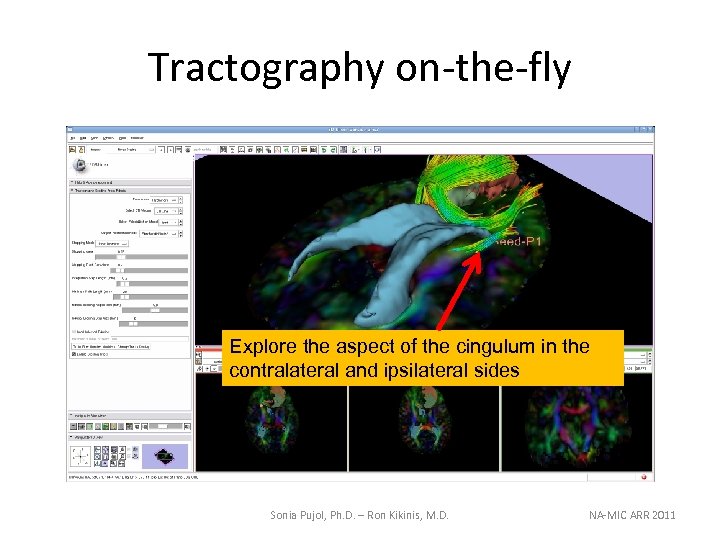
Tractography on-the-fly Explore the aspect of the cingulum in the contralateral and ipsilateral sides Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011

Conclusion • Fully integrated pipeline for semi-automated tumor segmentation and white matter tract reconstruction • 3 D interactive exploration of the white matter tracts surrounding a tumor (peri-tumoral tracts) for neurosurgical planning Sonia Pujol, Ph. D. – Ron Kikinis, M. D. NA-MIC ARR 2011

Acknowledgments • National Alliance for Medical Image Computing (NA-MIC) NIH U 54 EB 005149 • Neuroimage Analysis Center (NAC) NIH P 41 RR 013218 White Matter Exploration for Neurosurgical Planning Sonia Pujol, Ph. D - Ron Kikinis, MD National Alliance for Medical Image Computing ARR 2011
ef8ffa3c7da67ebf84421690770824ea.ppt