9e598c061ccb12f720a15542cbd9f473.ppt
- Количество слайдов: 52
 Evolution, Adaptation and Spread of Influenza Viruses: What Have We Learned? Nancy J. Cox, Ph. D. Chief, Influenza Branch National Center for Infectious Diseases Centers for Disease Control and Prevention
Evolution, Adaptation and Spread of Influenza Viruses: What Have We Learned? Nancy J. Cox, Ph. D. Chief, Influenza Branch National Center for Infectious Diseases Centers for Disease Control and Prevention
 Influenza Historical Background • Epidemics of influenza have occurred in humans since ancient times – recorded by Hippocrates in 412 BC • “Influenza” – term dates from 15 th century Italy when epidemics were attributed to the influence of the stars • First pandemic clearly described in 1580
Influenza Historical Background • Epidemics of influenza have occurred in humans since ancient times – recorded by Hippocrates in 412 BC • “Influenza” – term dates from 15 th century Italy when epidemics were attributed to the influence of the stars • First pandemic clearly described in 1580
 Background, continued • Influenza A virus isolated in ferrets in 1933 by Smith et al. • Virus first grown in embryonated eggs in 1936 • Antigenic differences detected between viruses in 1937 • Influenza B virus isolated in 1940 by Francis and Mc. Gill (independently) • Inactivated influenza vaccine found to be effective in 1944 (U. S. military)
Background, continued • Influenza A virus isolated in ferrets in 1933 by Smith et al. • Virus first grown in embryonated eggs in 1936 • Antigenic differences detected between viruses in 1937 • Influenza B virus isolated in 1940 by Francis and Mc. Gill (independently) • Inactivated influenza vaccine found to be effective in 1944 (U. S. military)
 Influenza Viruses • Enveloped RNA viruses, 80 -100 nm in diameter • 8 single-stranded negative sense gene segments encoding 10 proteins • Aerosol and droplet transmission • Incubation period 1 -4 days • Viral shedding • Adults <5 days • Young children up to 10 -14 days • Two virus types: A and B, based on NP and M 1 • Three A subtypes: H 1 N 1, H 1 N 2 & H 3 N 2, based on spike-like surface glyproteins
Influenza Viruses • Enveloped RNA viruses, 80 -100 nm in diameter • 8 single-stranded negative sense gene segments encoding 10 proteins • Aerosol and droplet transmission • Incubation period 1 -4 days • Viral shedding • Adults <5 days • Young children up to 10 -14 days • Two virus types: A and B, based on NP and M 1 • Three A subtypes: H 1 N 1, H 1 N 2 & H 3 N 2, based on spike-like surface glyproteins
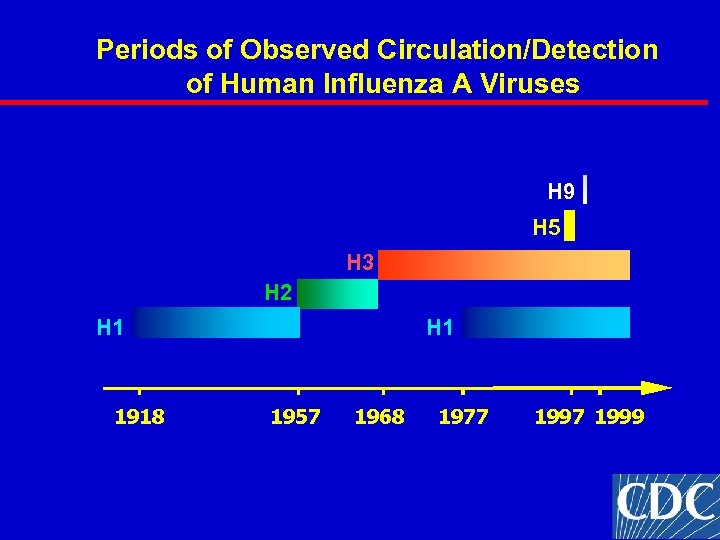 Periods of Observed Circulation/Detection of Human Influenza A Viruses H 9 H 5 H 3 H 2 H 1 1918 H 1 1957 1968 1977 1999
Periods of Observed Circulation/Detection of Human Influenza A Viruses H 9 H 5 H 3 H 2 H 1 1918 H 1 1957 1968 1977 1999
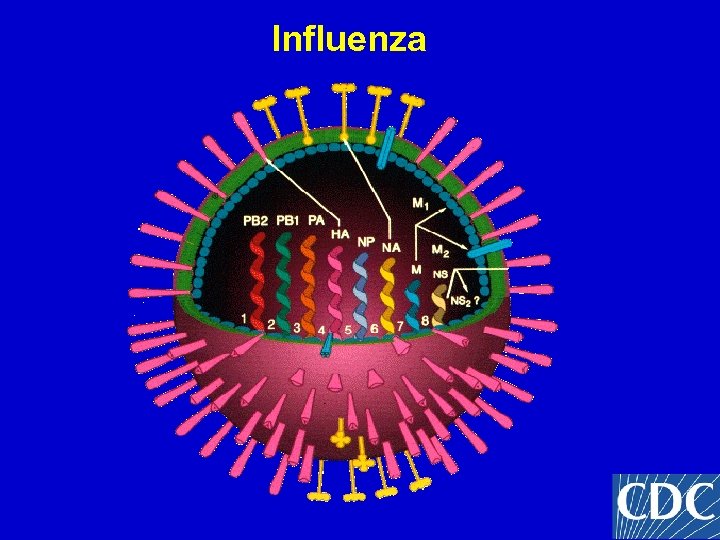 Influenza
Influenza
 Antigenic Change • Antigenic ‘drift” occurs in HA and NA • Associated with seasonal epidemics • Continual development of new strains secondary to genetic mutations • A viruses >> B viruses • Antigenic “shift” occurs in HA and NA • Associated with pandemics • Appearance of novel influenza A viruses bearing new HA or HA & NA
Antigenic Change • Antigenic ‘drift” occurs in HA and NA • Associated with seasonal epidemics • Continual development of new strains secondary to genetic mutations • A viruses >> B viruses • Antigenic “shift” occurs in HA and NA • Associated with pandemics • Appearance of novel influenza A viruses bearing new HA or HA & NA
 Infection and Disease • Highest infection & disease rates in children • Highest rates of serious complications in elderly and persons with certain underlying chronic conditions, e. g. : • Cardiopulmonary disease • Metabolic disease (e. g. diabetes) • Immune deficiency
Infection and Disease • Highest infection & disease rates in children • Highest rates of serious complications in elderly and persons with certain underlying chronic conditions, e. g. : • Cardiopulmonary disease • Metabolic disease (e. g. diabetes) • Immune deficiency
 Influenza Viruses Infect Several Animal Species • All influenza A subtypes found in wild birds • Fecal transmission among wild birds • Usually no illness • Other animal species • Domestic poultry (chickens and quail) • Humans, swine, horses, seals, whales • Humans usually infected by human influenza viruses
Influenza Viruses Infect Several Animal Species • All influenza A subtypes found in wild birds • Fecal transmission among wild birds • Usually no illness • Other animal species • Domestic poultry (chickens and quail) • Humans, swine, horses, seals, whales • Humans usually infected by human influenza viruses

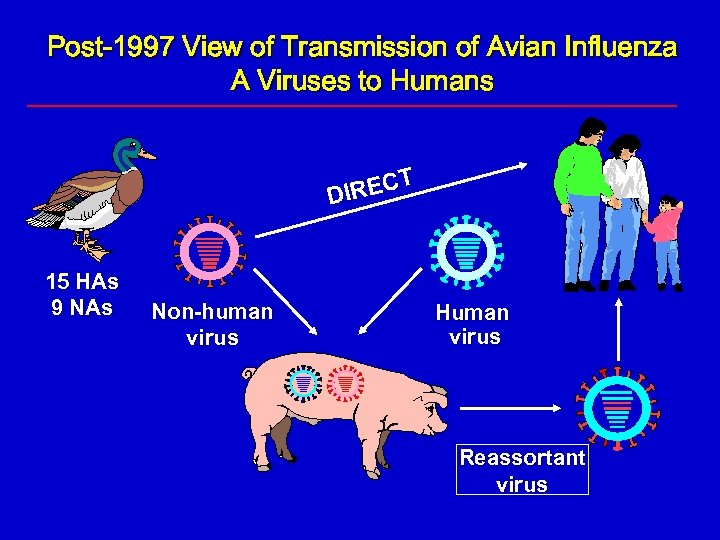 Post-1997 View of Transmission of Avian Influenza A Viruses to Humans T REC DI 15 HAs 9 NAs Non-human virus Human virus Reassortant virus
Post-1997 View of Transmission of Avian Influenza A Viruses to Humans T REC DI 15 HAs 9 NAs Non-human virus Human virus Reassortant virus
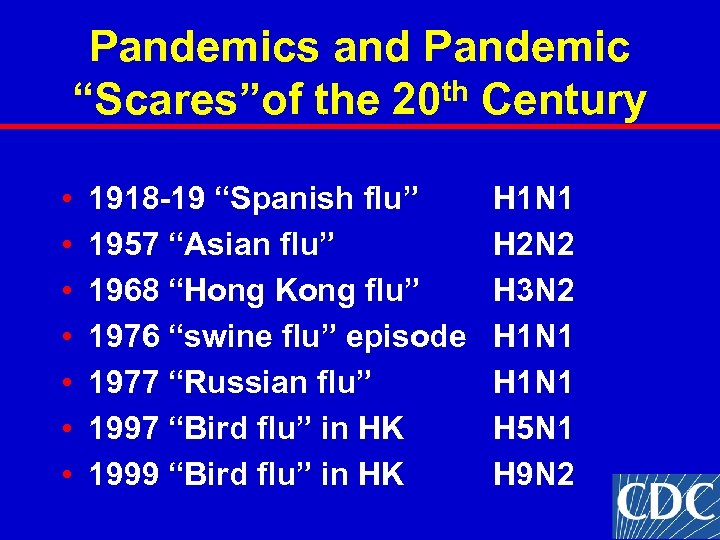 Pandemics and Pandemic “Scares”of the 20 th Century • • 1918 -19 “Spanish flu” 1957 “Asian flu” 1968 “Hong Kong flu” 1976 “swine flu” episode 1977 “Russian flu” 1997 “Bird flu” in HK 1999 “Bird flu” in HK H 1 N 1 H 2 N 2 H 3 N 2 H 1 N 1 H 5 N 1 H 9 N 2
Pandemics and Pandemic “Scares”of the 20 th Century • • 1918 -19 “Spanish flu” 1957 “Asian flu” 1968 “Hong Kong flu” 1976 “swine flu” episode 1977 “Russian flu” 1997 “Bird flu” in HK 1999 “Bird flu” in HK H 1 N 1 H 2 N 2 H 3 N 2 H 1 N 1 H 5 N 1 H 9 N 2
 1918 -19 Pandemic • • • Scattered outbreaks in U. S. in spring Relatively low activity in summer Peak of first wave in October Peak of second wave in mid-winter U. S. ~550, 000 deaths • nearly 1/2 in persons 20 -40 years old • Worldwide ~ 40 million deaths
1918 -19 Pandemic • • • Scattered outbreaks in U. S. in spring Relatively low activity in summer Peak of first wave in October Peak of second wave in mid-winter U. S. ~550, 000 deaths • nearly 1/2 in persons 20 -40 years old • Worldwide ~ 40 million deaths
 1957 -58 and 1968 -69 Pandemics • 1957 • Emergence of influenza A (H 2 N 2) virus • 66, 000 U. S. deaths • 36% of deaths (24, 000) in persons <65 years • 1968 • Emergence of influenza A (H 3 N 2) virus • 28, 100 U. S. deaths • 41% of deaths (11, 600) in persons <65 years
1957 -58 and 1968 -69 Pandemics • 1957 • Emergence of influenza A (H 2 N 2) virus • 66, 000 U. S. deaths • 36% of deaths (24, 000) in persons <65 years • 1968 • Emergence of influenza A (H 3 N 2) virus • 28, 100 U. S. deaths • 41% of deaths (11, 600) in persons <65 years
 The Next Pandemic • Another pandemic is expected. • According to our CDC model, the projected health impact in the U. S. is: • 89, 000 – 207, 000 …. . deaths • 314, 000 – 734, 000. . …hospitalizations • 18 million – 42 million…clinic visits
The Next Pandemic • Another pandemic is expected. • According to our CDC model, the projected health impact in the U. S. is: • 89, 000 – 207, 000 …. . deaths • 314, 000 – 734, 000. . …hospitalizations • 18 million – 42 million…clinic visits
 Grand Challenge of Modeling: Spread of an Influenza Pandemic • • • Susceptibility of the population Person-to-person spread Mixing of global populations Traffic patterns, especially global travel Would we have time to produce vaccine and to ramp up production of antiviral drugs?
Grand Challenge of Modeling: Spread of an Influenza Pandemic • • • Susceptibility of the population Person-to-person spread Mixing of global populations Traffic patterns, especially global travel Would we have time to produce vaccine and to ramp up production of antiviral drugs?
 Average Annual Impact of Influenza on U. S. • >20, 000 deaths • >110, 000 hospitalizations • Currently >90% deaths in persons >65 years • Higher numbers of deaths and hospitalizations in A(H 3 N 2) years
Average Annual Impact of Influenza on U. S. • >20, 000 deaths • >110, 000 hospitalizations • Currently >90% deaths in persons >65 years • Higher numbers of deaths and hospitalizations in A(H 3 N 2) years
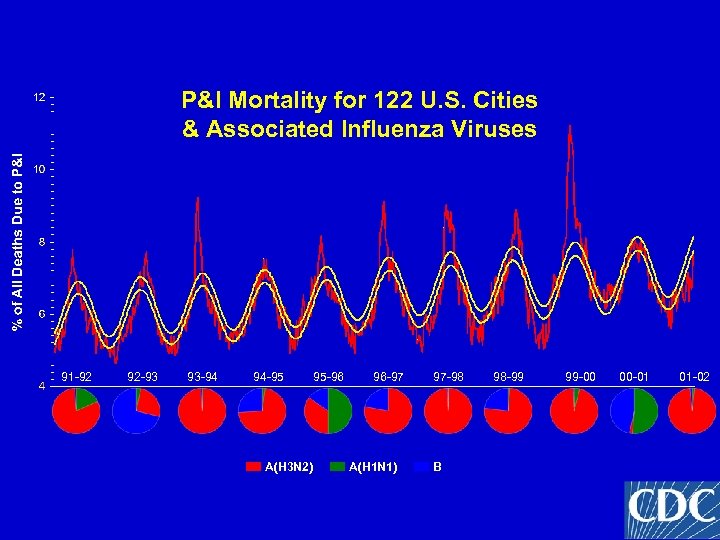 P&I Mortality for 122 U. S. Cities & Associated Influenza Viruses 91 -92 92 -93 93 -94 94 -95 A(H 3 N 2) 95 -96 96 -97 A(H 1 N 1) 97 -98 B 98 -99 99 -00 00 -01 01 -02
P&I Mortality for 122 U. S. Cities & Associated Influenza Viruses 91 -92 92 -93 93 -94 94 -95 A(H 3 N 2) 95 -96 96 -97 A(H 1 N 1) 97 -98 B 98 -99 99 -00 00 -01 01 -02
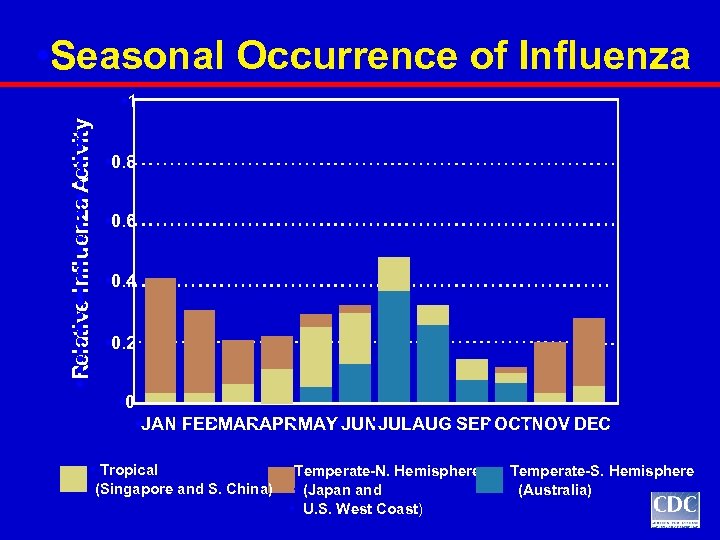 • Seasonal Occurrence of Influenza • Re • a • • ve • • n • • uenza • Ac • • v • • y • • l • ti • • I • fl • • ti • it • 1 • 0. 8 • 0. 6 • 0. 4 • 0. 2 • 0 • JAN • FEBMARAPR MAY • JUN JUL AUG • SEP OCTNOV • DEC • • Tropical (Singapore and S. China) Temperate-N. Hemisphere • Temperate-S. Hemisphere • (Japan and • (Australia) • U. S. West Coast)
• Seasonal Occurrence of Influenza • Re • a • • ve • • n • • uenza • Ac • • v • • y • • l • ti • • I • fl • • ti • it • 1 • 0. 8 • 0. 6 • 0. 4 • 0. 2 • 0 • JAN • FEBMARAPR MAY • JUN JUL AUG • SEP OCTNOV • DEC • • Tropical (Singapore and S. China) Temperate-N. Hemisphere • Temperate-S. Hemisphere • (Japan and • (Australia) • U. S. West Coast)
 WHO Recommendations for Vaccine Strains Formal recommendations for strains to be included in the currently licensed inactivated influenza vaccine are made each year during February for the following influenza season in the Northern Hemisphere and in September (since 1998) for the following season in the Southern Hemisphere.
WHO Recommendations for Vaccine Strains Formal recommendations for strains to be included in the currently licensed inactivated influenza vaccine are made each year during February for the following influenza season in the Northern Hemisphere and in September (since 1998) for the following season in the Southern Hemisphere.
 Considerations for Recommendations • Are there new antigenic variants (HI and gene sequencing data)? • Are new variants spreading and causing disease? • Are current vaccines able to induce antibodies to the new variants? • Are any new variants useful for vaccine production?
Considerations for Recommendations • Are there new antigenic variants (HI and gene sequencing data)? • Are new variants spreading and causing disease? • Are current vaccines able to induce antibodies to the new variants? • Are any new variants useful for vaccine production?
 Why Sequence HA Genes? • Sequence analysis provides information on the molecular basis for antigenic drift. – – – Rates of change at nucleotide/amino acid levels Types of amino acid changes Positions of amino acid changes Predictability in patterns of change Molecular correlates with epidemiology • Sequence analysis provides precise data on evolutionary relationships between HA genes of Influenza viruses. • Sequence data provide adjunct to HI data for vaccine strain selection.
Why Sequence HA Genes? • Sequence analysis provides information on the molecular basis for antigenic drift. – – – Rates of change at nucleotide/amino acid levels Types of amino acid changes Positions of amino acid changes Predictability in patterns of change Molecular correlates with epidemiology • Sequence analysis provides precise data on evolutionary relationships between HA genes of Influenza viruses. • Sequence data provide adjunct to HI data for vaccine strain selection.
 What Viruses are Sequenced? • Variant viruses (HI test) • Typical viruses (HI test) • Geographic distribution (emphasis on viruses from Asia) • Temporal distribution S. Hemisphere N. Hemisphere
What Viruses are Sequenced? • Variant viruses (HI test) • Typical viruses (HI test) • Geographic distribution (emphasis on viruses from Asia) • Temporal distribution S. Hemisphere N. Hemisphere
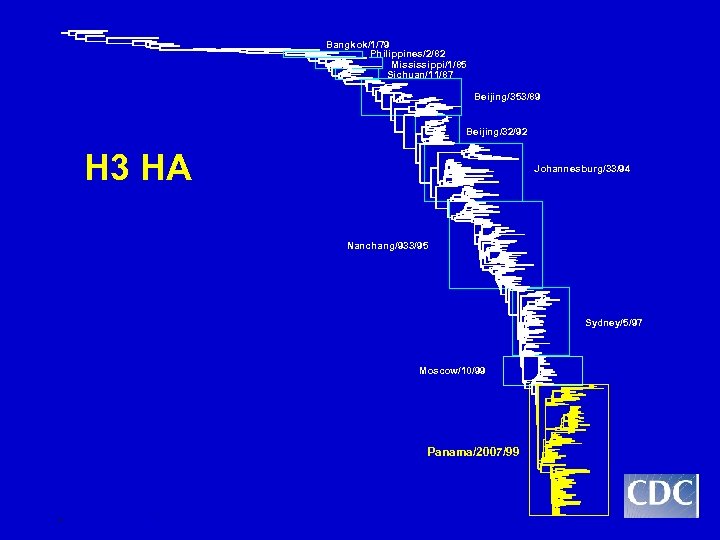 Bangkok/1/79 Philippines/2/82 Mississippi/1/85 Sichuan/11/87 Beijing/353/89 Beijing/32/92 H 3 HA Johannesburg/33/94 Nanchang/933/95 Sydney/5/97 Moscow/10/99 Panama/2007/99
Bangkok/1/79 Philippines/2/82 Mississippi/1/85 Sichuan/11/87 Beijing/353/89 Beijing/32/92 H 3 HA Johannesburg/33/94 Nanchang/933/95 Sydney/5/97 Moscow/10/99 Panama/2007/99
 Conclusions from Prediction Attempts • We identified 18 codons in HA 1 that are positively selected; sites A, B and RBP • Strains with more mutations in these codons = progenitors of successful new lineages – retrospective testing • Alternate hypotheses less successful in predicting • Monitoring change in + selected codons may predict course of evolution and be useful for vaccine strain selection
Conclusions from Prediction Attempts • We identified 18 codons in HA 1 that are positively selected; sites A, B and RBP • Strains with more mutations in these codons = progenitors of successful new lineages – retrospective testing • Alternate hypotheses less successful in predicting • Monitoring change in + selected codons may predict course of evolution and be useful for vaccine strain selection
 Antigenic Drift in H 3 N 2 Viruses Demonstrated in an HI Assay Post-infection ferret antisera raised against Viruses Joh/33/94 Nan/933/95 Syd/5/97 A/Johannesburg/33/94 640 40 <10 A/Nanchang/933/95 40 1280 20 A/Sydney/5/97 <10 40 640
Antigenic Drift in H 3 N 2 Viruses Demonstrated in an HI Assay Post-infection ferret antisera raised against Viruses Joh/33/94 Nan/933/95 Syd/5/97 A/Johannesburg/33/94 640 40 <10 A/Nanchang/933/95 40 1280 20 A/Sydney/5/97 <10 40 640
 Signature Amino Acid Changes Observed Between the HAs of A/Johannesburg/33/94 and A/Nanchang/933/95 -like Viruses Joh/33/94 G 135 T, N 145 K, N 262 S Nan/933/95
Signature Amino Acid Changes Observed Between the HAs of A/Johannesburg/33/94 and A/Nanchang/933/95 -like Viruses Joh/33/94 G 135 T, N 145 K, N 262 S Nan/933/95
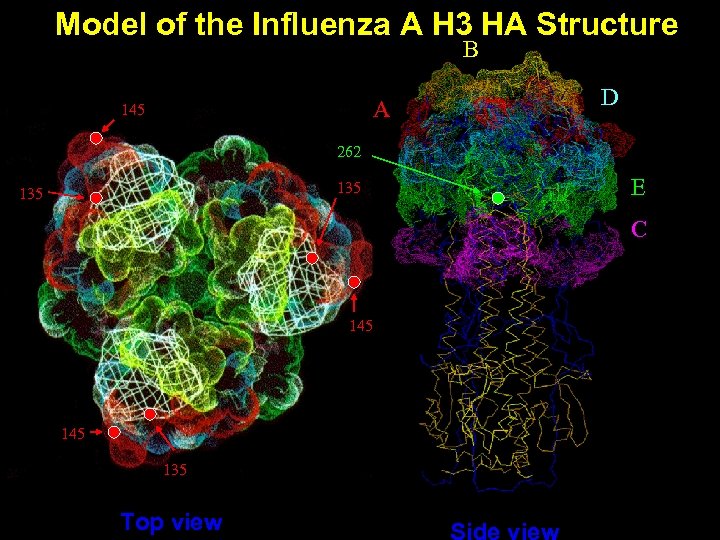 Model of the Influenza A H 3 HA Structure B A 145 D 262 135 E C 145 135 Top view
Model of the Influenza A H 3 HA Structure B A 145 D 262 135 E C 145 135 Top view
 Approach Ø Generate transfectant viruses with the Joh/HA containing – single mutations alone • G 135 T • N 145 K • N 262 S and – combinations of double mutations • G 135 T/ N 145 K • G 135 T/ N 262 S • N 145 K/ N 262 S Ø Determine antigenicity of each mutant virus using postinfection ferret sera in an HI assay
Approach Ø Generate transfectant viruses with the Joh/HA containing – single mutations alone • G 135 T • N 145 K • N 262 S and – combinations of double mutations • G 135 T/ N 145 K • G 135 T/ N 262 S • N 145 K/ N 262 S Ø Determine antigenicity of each mutant virus using postinfection ferret sera in an HI assay
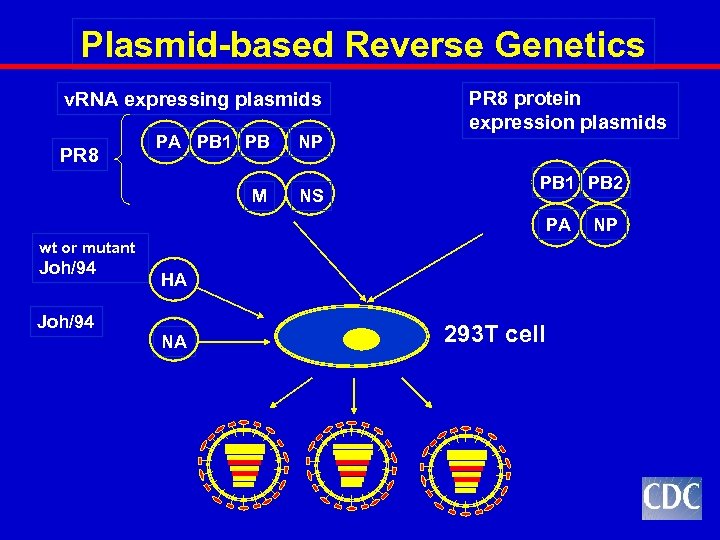 Plasmid-based Reverse Genetics v. RNA expressing plasmids PR 8 PA PB 1 PB 2 NP M NS PR 8 protein expression plasmids PB 1 PB 2 PA wt or mutant Joh/94 HA NA 293 T cell NP
Plasmid-based Reverse Genetics v. RNA expressing plasmids PR 8 PA PB 1 PB 2 NP M NS PR 8 protein expression plasmids PB 1 PB 2 PA wt or mutant Joh/94 HA NA 293 T cell NP
 Summary of Reverse Genetics Experiments Ø The presence of each mutation alone was insufficient to recreate the antigenic difference observed between Johannesburg/33/94 and Nanchang/933/95 -like viruses. Ø But, the combination of mutations G 135 T and N 145 K in site A of the Joh 94 HA were sufficient to re-create this example of antigenic drift in the H 3 HA. Ø Reverse genetics techniques allowed us to identify contribution of specific aa residues to antigenic drift in the H 3 HA. Look at other examples and at contributions of 18 codons.
Summary of Reverse Genetics Experiments Ø The presence of each mutation alone was insufficient to recreate the antigenic difference observed between Johannesburg/33/94 and Nanchang/933/95 -like viruses. Ø But, the combination of mutations G 135 T and N 145 K in site A of the Joh 94 HA were sufficient to re-create this example of antigenic drift in the H 3 HA. Ø Reverse genetics techniques allowed us to identify contribution of specific aa residues to antigenic drift in the H 3 HA. Look at other examples and at contributions of 18 codons.
 The Flu Brew: Is it Right?
The Flu Brew: Is it Right?
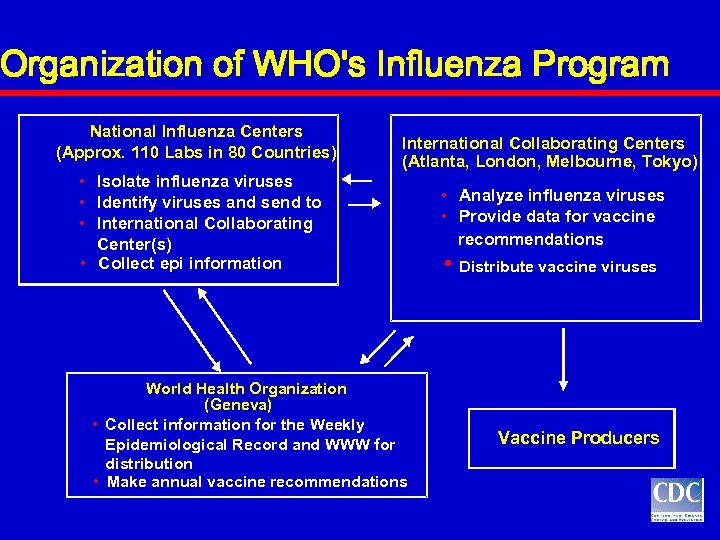 Organization of WHO's Influenza Program National Influenza Centers (Approx. 110 Labs in 80 Countries) International Collaborating Centers (Atlanta, London, Melbourne, Tokyo) • Isolate influenza viruses • Identify viruses and send to • International Collaborating Center(s) • Collect epi information World Health Organization (Geneva) • Collect information for the Weekly Epidemiological Record and WWW for distribution • Make annual vaccine recommendations • Analyze influenza viruses • Provide data for vaccine recommendations • Distribute vaccine viruses Vaccine Producers
Organization of WHO's Influenza Program National Influenza Centers (Approx. 110 Labs in 80 Countries) International Collaborating Centers (Atlanta, London, Melbourne, Tokyo) • Isolate influenza viruses • Identify viruses and send to • International Collaborating Center(s) • Collect epi information World Health Organization (Geneva) • Collect information for the Weekly Epidemiological Record and WWW for distribution • Make annual vaccine recommendations • Analyze influenza viruses • Provide data for vaccine recommendations • Distribute vaccine viruses Vaccine Producers
 Current Status of WHO System • 175, 000 isolates/yr (600 to 1200 M cases) • WHO CCs receive 6, 500 samples/yr. • WHO CCs and NICs sequence 1, 000 samples/yr. • >240 M doses of influenza vaccine produced worldwide
Current Status of WHO System • 175, 000 isolates/yr (600 to 1200 M cases) • WHO CCs receive 6, 500 samples/yr. • WHO CCs and NICs sequence 1, 000 samples/yr. • >240 M doses of influenza vaccine produced worldwide
 Enhancing Surveillance for Pandemic and Epidemic Influenza • Need higher throughput labs • Need better data collection • Lab + Epi linked • New algorithms for identifying key viruses • Increase data based size for better predictive methods
Enhancing Surveillance for Pandemic and Epidemic Influenza • Need higher throughput labs • Need better data collection • Lab + Epi linked • New algorithms for identifying key viruses • Increase data based size for better predictive methods
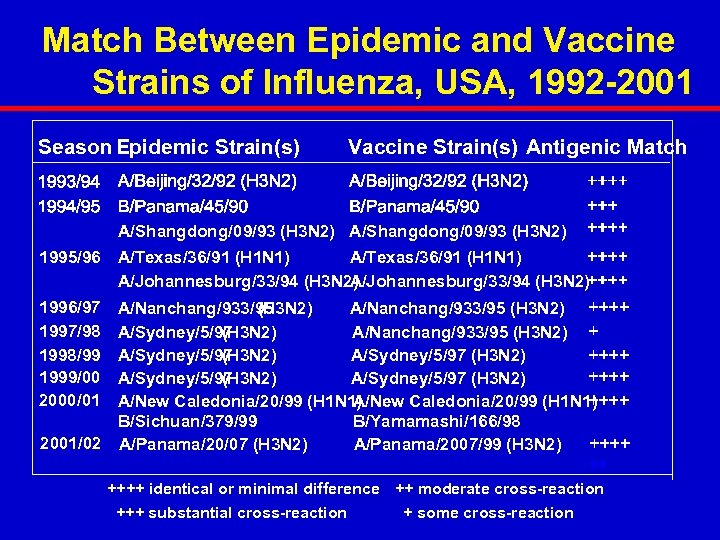 Match Between Epidemic and Vaccine Strains of Influenza, USA, 1992 -2001 Season Epidemic Strain(s) Vaccine Strain(s) Antigenic Match A/Shangdong/09/93 (H 3 N 2) 1995/96 A/Texas/36/91 (H 1 N 1) A/Johannesburg/33/94 (H 3 N 2) 1996/97 1997/98 1998/99 1999/00 2000/01 A/Nanchang/933/95 (H 3 N 2) A/Sydney/5/97 (H 3 N 2) A/Sydney/5/97 (H 3 N 2) A/New Caledonia/20/99 (H 1 N 1) B/Sichuan/379/99 B/Yamamashi/166/98 2001/02 A/Panama/20/07 (H 3 N 2) A/Panama/2007/99 (H 3 N 2) ++ ++++ identical or minimal difference ++ moderate cross-reaction +++ substantial cross-reaction + some cross-reaction
Match Between Epidemic and Vaccine Strains of Influenza, USA, 1992 -2001 Season Epidemic Strain(s) Vaccine Strain(s) Antigenic Match A/Shangdong/09/93 (H 3 N 2) 1995/96 A/Texas/36/91 (H 1 N 1) A/Johannesburg/33/94 (H 3 N 2) 1996/97 1997/98 1998/99 1999/00 2000/01 A/Nanchang/933/95 (H 3 N 2) A/Sydney/5/97 (H 3 N 2) A/Sydney/5/97 (H 3 N 2) A/New Caledonia/20/99 (H 1 N 1) B/Sichuan/379/99 B/Yamamashi/166/98 2001/02 A/Panama/20/07 (H 3 N 2) A/Panama/2007/99 (H 3 N 2) ++ ++++ identical or minimal difference ++ moderate cross-reaction +++ substantial cross-reaction + some cross-reaction
 Factors Contributing to Improved Influenza Vaccine Match • Better Worldwide Surveillance • Europe, Latin America and Asia • Improved Communications • Telephone and Fax • E-mail • Technical Advances • • Molecular Analyses • • Increasing International Cooperation • Exchange of Viruses and Serum Panels • Standardization of Reagents and Vaccines
Factors Contributing to Improved Influenza Vaccine Match • Better Worldwide Surveillance • Europe, Latin America and Asia • Improved Communications • Telephone and Fax • E-mail • Technical Advances • • Molecular Analyses • • Increasing International Cooperation • Exchange of Viruses and Serum Panels • Standardization of Reagents and Vaccines
 Success of WHO's Influenza Surveillance Network • Action-orientated network • Reagents updated annually • Annual recommendations • Collaborative spirit of participants
Success of WHO's Influenza Surveillance Network • Action-orientated network • Reagents updated annually • Annual recommendations • Collaborative spirit of participants
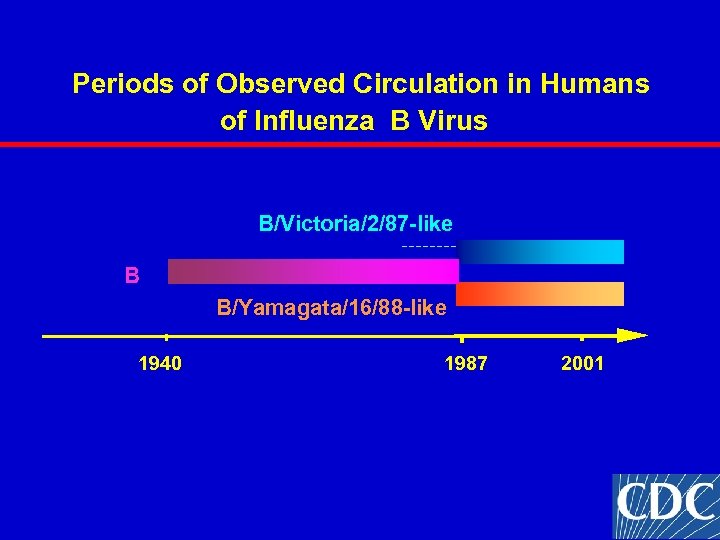 Periods of Observed Circulation in Humans of Influenza B Virus B/Victoria/2/87 -like B B/Yamagata/16/88 -like 1940 1987 2001
Periods of Observed Circulation in Humans of Influenza B Virus B/Victoria/2/87 -like B B/Yamagata/16/88 -like 1940 1987 2001
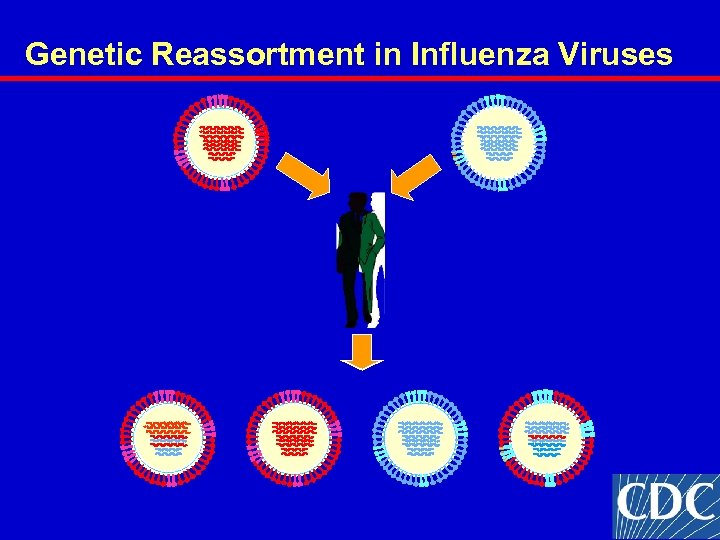 Genetic Reassortment in Influenza Viruses
Genetic Reassortment in Influenza Viruses
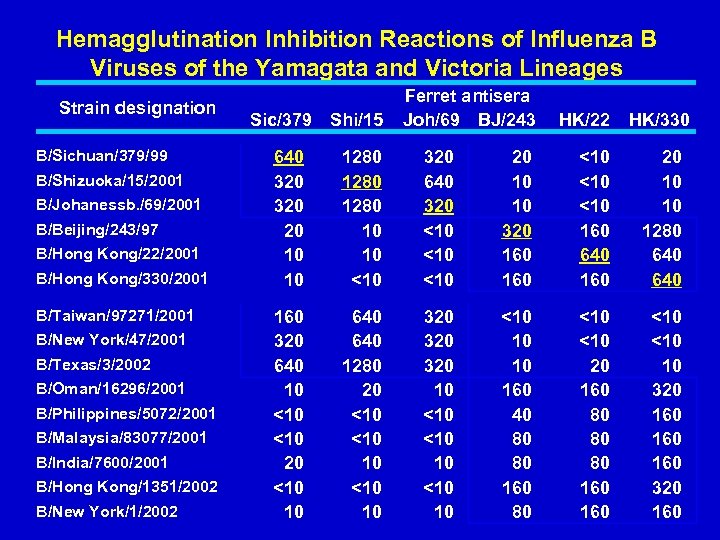 Hemagglutination Inhibition Reactions of Influenza B Viruses of the Yamagata and Victoria Lineages Strain designation B/Sichuan/379/99 B/Shizuoka/15/2001 B/Johanessb. /69/2001 B/Beijing/243/97 B/Hong Kong/22/2001 B/Hong Kong/330/2001 B/Taiwan/97271/2001 B/New York/47/2001 B/Texas/3/2002 B/Oman/16296/2001 B/Philippines/5072/2001 B/Malaysia/83077/2001 B/India/7600/2001 B/Hong Kong/1351/2002 B/New York/1/2002 Sic/379 Shi/15 Ferret antisera Joh/69 BJ/243 HK/22 HK/330 640 320 20 10 10 1280 10 10 <10 320 640 320 <10 <10 20 10 10 320 160 <10 <10 160 640 160 20 10 10 1280 640 160 320 640 10 <10 20 <10 10 640 1280 20 <10 10 320 320 10 <10 10 10 160 40 80 80 160 80 <10 20 160 80 80 80 160 <10 10 320 160 160 320 160
Hemagglutination Inhibition Reactions of Influenza B Viruses of the Yamagata and Victoria Lineages Strain designation B/Sichuan/379/99 B/Shizuoka/15/2001 B/Johanessb. /69/2001 B/Beijing/243/97 B/Hong Kong/22/2001 B/Hong Kong/330/2001 B/Taiwan/97271/2001 B/New York/47/2001 B/Texas/3/2002 B/Oman/16296/2001 B/Philippines/5072/2001 B/Malaysia/83077/2001 B/India/7600/2001 B/Hong Kong/1351/2002 B/New York/1/2002 Sic/379 Shi/15 Ferret antisera Joh/69 BJ/243 HK/22 HK/330 640 320 20 10 10 1280 10 10 <10 320 640 320 <10 <10 20 10 10 320 160 <10 <10 160 640 160 20 10 10 1280 640 160 320 640 10 <10 20 <10 10 640 1280 20 <10 10 320 320 10 <10 10 10 160 40 80 80 160 80 <10 20 160 80 80 80 160 <10 10 320 160 160 320 160
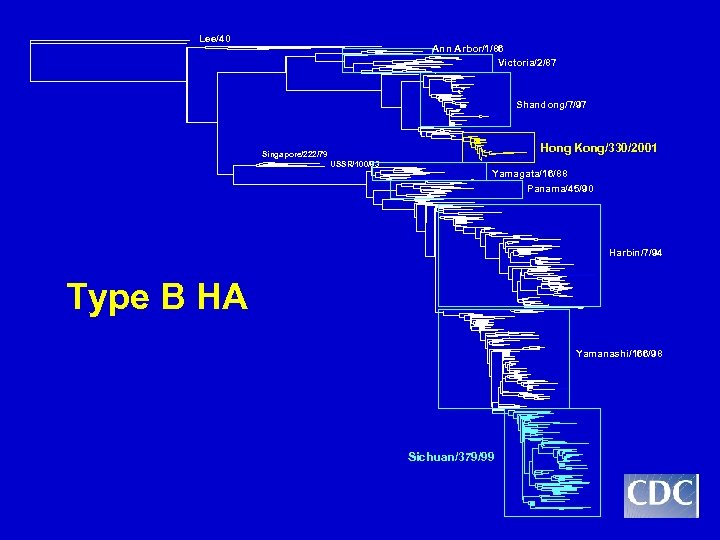 Lee/40 Ann Arbor/1/86 Victoria/2/87 Shandong/7/97 Hong Kong/330/2001 Singapore/222/79 USSR/100/83 Yamagata/16/88 Panama/45/90 Harbin/7/94 Type B HA Yamanashi/166/98 Sichuan/379/99
Lee/40 Ann Arbor/1/86 Victoria/2/87 Shandong/7/97 Hong Kong/330/2001 Singapore/222/79 USSR/100/83 Yamagata/16/88 Panama/45/90 Harbin/7/94 Type B HA Yamanashi/166/98 Sichuan/379/99
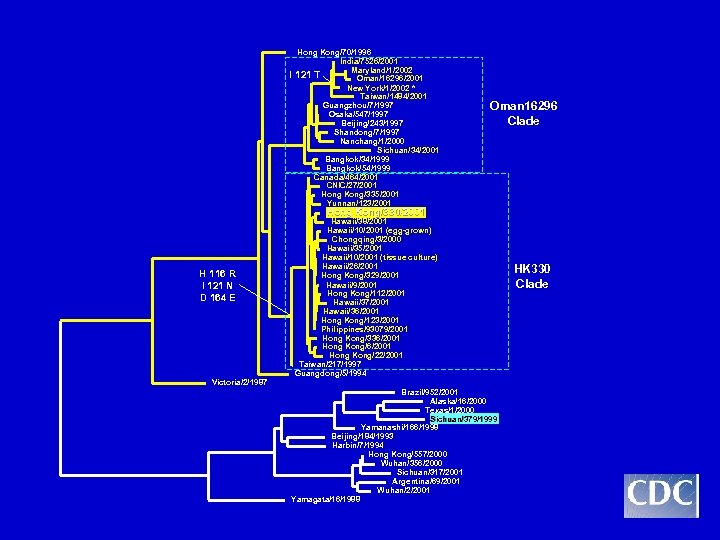 I Hong Kong/70/1996 India/7526/2001 Maryland/1/2002 121 T Oman/16296/2001 New York/1/2002 * Taiwan/1484/2001 Guangzhou/7/1997 Osaka/547/1997 Beijing/243/1997 Shandong/7/1997 Nanchang/1/2000 Sichuan/34/2001 Bangkok/34/1999 Bangkok/54/1999 Canada/464/2001 CNIC/27/2001 Hong Kong/335/2001 Yunnan/123/2001 Oman 16296 Clade Hong Kong/330/2001 H 116 R I 121 N D 164 E Victoria/2/1987 Hawaii/38/2001 Hawaii/10/2001 (egg-grown) Chongqing/3/2000 Hawaii/35/2001 Hawaii/10/2001 (tissue culture) Hawaii/26/2001 Hong Kong/329/2001 Hawaii/9/2001 Hong Kong/112/2001 Hawaii/37/2001 Hawaii/36/2001 Hong Kong/123/2001 Philippines/93079/2001 Hong Kong/336/2001 Hong Kong/22/2001 Taiwan/217/1997 Guangdong/5/1994 Brazil/952/2001 Alaska/16/2000 Texas/1/2000 Sichuan/379/1999 Yamanashi/166/1998 Beijing/184/1993 Harbin/7/1994 Hong Kong/557/2000 Wuhan/356/2000 Sichuan/317/2001 Argentina/69/2001 Wuhan/2/2001 Yamagata/16/1988 HK 330 Clade
I Hong Kong/70/1996 India/7526/2001 Maryland/1/2002 121 T Oman/16296/2001 New York/1/2002 * Taiwan/1484/2001 Guangzhou/7/1997 Osaka/547/1997 Beijing/243/1997 Shandong/7/1997 Nanchang/1/2000 Sichuan/34/2001 Bangkok/34/1999 Bangkok/54/1999 Canada/464/2001 CNIC/27/2001 Hong Kong/335/2001 Yunnan/123/2001 Oman 16296 Clade Hong Kong/330/2001 H 116 R I 121 N D 164 E Victoria/2/1987 Hawaii/38/2001 Hawaii/10/2001 (egg-grown) Chongqing/3/2000 Hawaii/35/2001 Hawaii/10/2001 (tissue culture) Hawaii/26/2001 Hong Kong/329/2001 Hawaii/9/2001 Hong Kong/112/2001 Hawaii/37/2001 Hawaii/36/2001 Hong Kong/123/2001 Philippines/93079/2001 Hong Kong/336/2001 Hong Kong/22/2001 Taiwan/217/1997 Guangdong/5/1994 Brazil/952/2001 Alaska/16/2000 Texas/1/2000 Sichuan/379/1999 Yamanashi/166/1998 Beijing/184/1993 Harbin/7/1994 Hong Kong/557/2000 Wuhan/356/2000 Sichuan/317/2001 Argentina/69/2001 Wuhan/2/2001 Yamagata/16/1988 HK 330 Clade
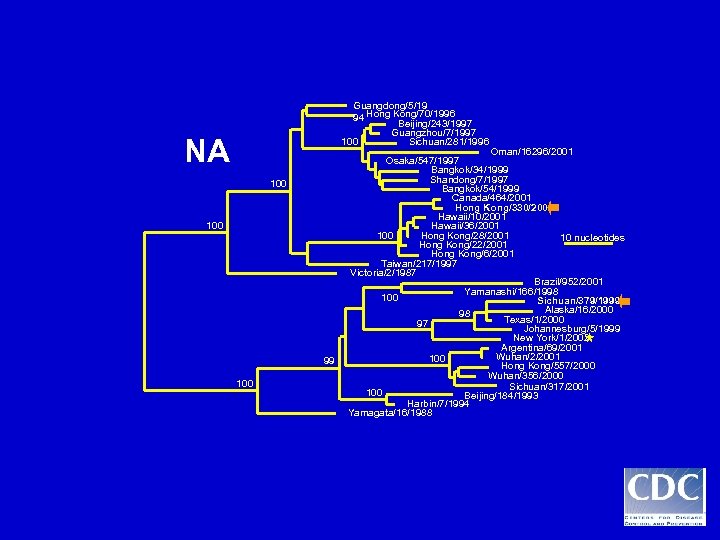 Guangdong/5/19 94 Hong Kong/70/1996 Beijing/243/1997 Guangzhou/7/1997 100 Sichuan/281/1996 NA 100 99 100 Oman/16296/2001 Osaka/547/1997 Bangkok/34/1999 Shandong/7/1997 Bangkok/54/1999 Canada/464/2001 Hong Kong/330/2001 Hawaii/10/2001 Hawaii/36/2001 Hong Kong/28/2001 100 10 nucleotides Hong Kong/22/2001 Hong Kong/6/2001 Taiwan/217/1997 Victoria/2/1987 Brazil/952/2001 Yamanashi/166/1998 100 Sichuan/379/1999 Alaska/16/2000 98 Texas/1/2000 97 Johannesburg/5/1999 New York/1/2002 Argentina/69/2001 Wuhan/2/2001 100 Hong Kong/557/2000 Wuhan/356/2000 Sichuan/317/2001 100 Beijing/184/1993 Harbin/7/1994 Yamagata/16/1988
Guangdong/5/19 94 Hong Kong/70/1996 Beijing/243/1997 Guangzhou/7/1997 100 Sichuan/281/1996 NA 100 99 100 Oman/16296/2001 Osaka/547/1997 Bangkok/34/1999 Shandong/7/1997 Bangkok/54/1999 Canada/464/2001 Hong Kong/330/2001 Hawaii/10/2001 Hawaii/36/2001 Hong Kong/28/2001 100 10 nucleotides Hong Kong/22/2001 Hong Kong/6/2001 Taiwan/217/1997 Victoria/2/1987 Brazil/952/2001 Yamanashi/166/1998 100 Sichuan/379/1999 Alaska/16/2000 98 Texas/1/2000 97 Johannesburg/5/1999 New York/1/2002 Argentina/69/2001 Wuhan/2/2001 100 Hong Kong/557/2000 Wuhan/356/2000 Sichuan/317/2001 100 Beijing/184/1993 Harbin/7/1994 Yamagata/16/1988
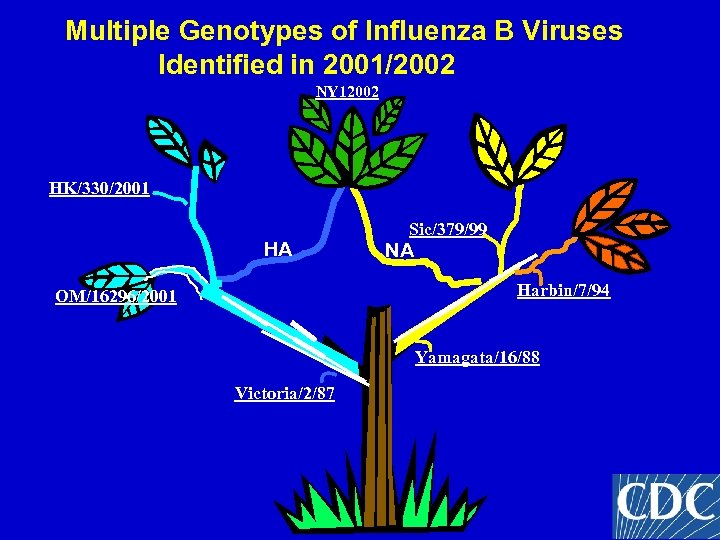 Multiple Genotypes of Influenza B Viruses Identified in 2001/2002 NY 12002 HK/330/2001 HA Sic/379/99 NA Harbin/7/94 OM/16296/2001 Yamagata/16/88 Victoria/2/87
Multiple Genotypes of Influenza B Viruses Identified in 2001/2002 NY 12002 HK/330/2001 HA Sic/379/99 NA Harbin/7/94 OM/16296/2001 Yamagata/16/88 Victoria/2/87
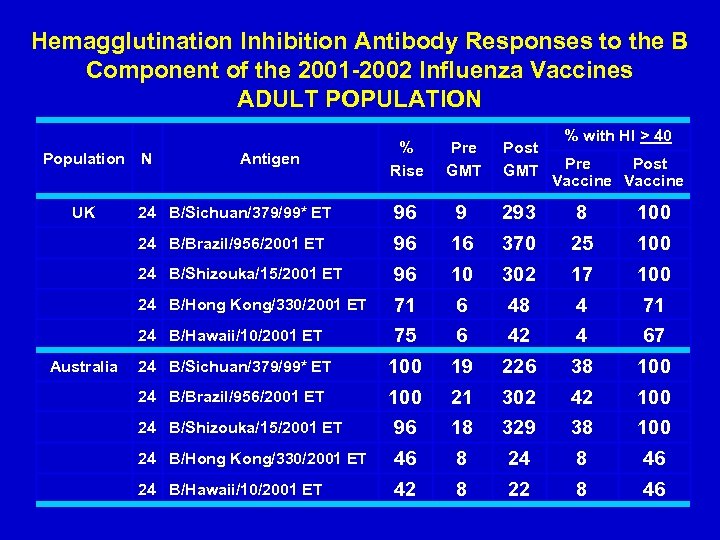 Hemagglutination Inhibition Antibody Responses to the B Component of the 2001 -2002 Influenza Vaccines ADULT POPULATION % with HI > 40 24 B/Sichuan/379/99* ET 96 9 293 8 100 96 16 370 25 100 96 10 302 17 100 24 B/Hong Kong/330/2001 ET 71 6 48 4 71 24 B/Hawaii/10/2001 ET Australia Post GMT 24 B/Shizouka/15/2001 ET UK Pre GMT 24 B/Brazil/956/2001 ET Population % Rise 75 6 42 4 67 24 B/Sichuan/379/99* ET 100 19 226 38 100 24 B/Brazil/956/2001 ET 100 21 302 42 100 24 B/Shizouka/15/2001 ET 96 18 329 38 100 24 B/Hong Kong/330/2001 ET 46 8 24 8 46 24 B/Hawaii/10/2001 ET 42 8 22 8 46 N Antigen Pre Post Vaccine
Hemagglutination Inhibition Antibody Responses to the B Component of the 2001 -2002 Influenza Vaccines ADULT POPULATION % with HI > 40 24 B/Sichuan/379/99* ET 96 9 293 8 100 96 16 370 25 100 96 10 302 17 100 24 B/Hong Kong/330/2001 ET 71 6 48 4 71 24 B/Hawaii/10/2001 ET Australia Post GMT 24 B/Shizouka/15/2001 ET UK Pre GMT 24 B/Brazil/956/2001 ET Population % Rise 75 6 42 4 67 24 B/Sichuan/379/99* ET 100 19 226 38 100 24 B/Brazil/956/2001 ET 100 21 302 42 100 24 B/Shizouka/15/2001 ET 96 18 329 38 100 24 B/Hong Kong/330/2001 ET 46 8 24 8 46 24 B/Hawaii/10/2001 ET 42 8 22 8 46 N Antigen Pre Post Vaccine
 Geographic Distribution of B/Victoria/02/87 Lineage Viruses April 2001 to Present Canada Netherlands Switzerland Italy USA Hawaii China Israel India Oman Japan Hong Kong Philippines Thailand Australia
Geographic Distribution of B/Victoria/02/87 Lineage Viruses April 2001 to Present Canada Netherlands Switzerland Italy USA Hawaii China Israel India Oman Japan Hong Kong Philippines Thailand Australia
 Conclusions Influenza B/Victoria/2/87 lineage viruses have reemerged in North America and Europe after a 10 year absence. Reassortment has occurred between 2 lineages of B viruses. Genetic reassortants bearing a B/Shandong/7/94 -like HA and B/Sichuan/379/99 -like NA were identified – vaccine choices. A B/Victoria lineage virus has been recommended for vaccines. Children under 10 expected to be most susceptible.
Conclusions Influenza B/Victoria/2/87 lineage viruses have reemerged in North America and Europe after a 10 year absence. Reassortment has occurred between 2 lineages of B viruses. Genetic reassortants bearing a B/Shandong/7/94 -like HA and B/Sichuan/379/99 -like NA were identified – vaccine choices. A B/Victoria lineage virus has been recommended for vaccines. Children under 10 expected to be most susceptible.
 Questions About B Viruses • Why did B/Vic lineage viruses survive only in China after B/Yamagata lineage viruses spread in the late 1980 s? • Why are B/Vic lineage viruses spreading to North America, Europe and Australia now after a 10 year absence? • Will both lineages of influenza B viruses continue to circulate?
Questions About B Viruses • Why did B/Vic lineage viruses survive only in China after B/Yamagata lineage viruses spread in the late 1980 s? • Why are B/Vic lineage viruses spreading to North America, Europe and Australia now after a 10 year absence? • Will both lineages of influenza B viruses continue to circulate?
 Other Key Questions • Can we explain the seasonality of influenza? • Why do so many epidemic variants appear first in China? • Is there real spread or only apparent spread of epidemic variants? • Is it possible to predict the next antigenic variant?
Other Key Questions • Can we explain the seasonality of influenza? • Why do so many epidemic variants appear first in China? • Is there real spread or only apparent spread of epidemic variants? • Is it possible to predict the next antigenic variant?
 Acknowledgements • WHO National Influenza Centers • WHO Collaborating Centers in London, Melbourne and Tokyo • Robin Bush and Walter Fitch • Members of the Influenza Branch (A. Klimov, H. Hall, C. Smith, K. Subbarao, S. Tong, M. Shaw and X. Xu)
Acknowledgements • WHO National Influenza Centers • WHO Collaborating Centers in London, Melbourne and Tokyo • Robin Bush and Walter Fitch • Members of the Influenza Branch (A. Klimov, H. Hall, C. Smith, K. Subbarao, S. Tong, M. Shaw and X. Xu)



