ce04f7de4f9128ab013952812e5169c4.ppt
- Количество слайдов: 67

EPIGENETIKA SUMMER 2011 Petr Svoboda mail: tel: svobodap@img. cas. cz 241063147

DNA METHYLATION I (DNA METHYLATION AND ITS DETECTION)

EPIGENETICS Epigenetics deals with heritable information which is not encoded in the DNA sequence Such information can be encoded in: • structure and chromatin modifications • DNA modifications • RNA molecules
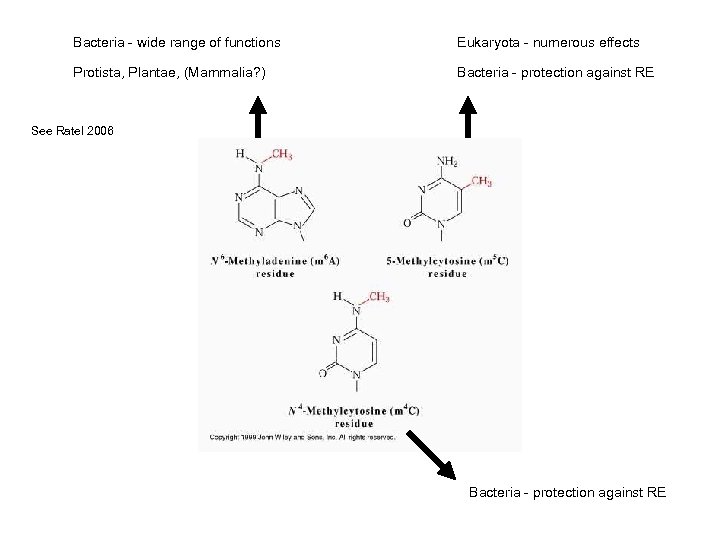
Bacteria - wide range of functions Eukaryota - numerous effects Protista, Plantae, (Mammalia? ) Bacteria - protection against RE See Ratel 2006 Bacteria - protection against RE
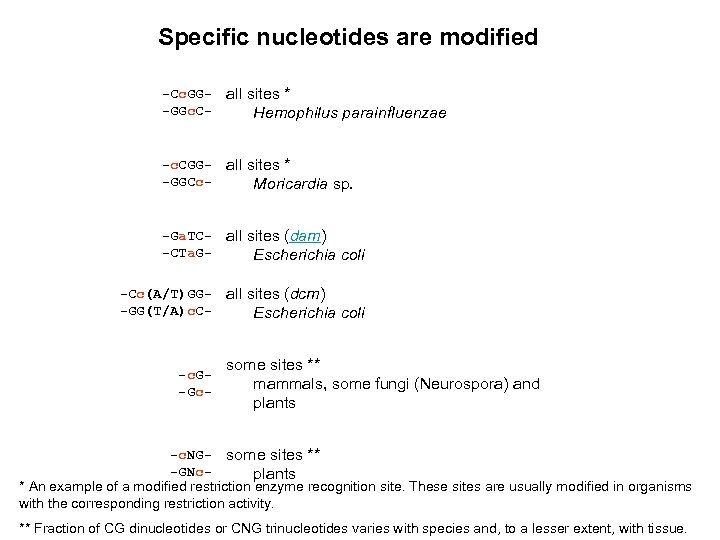

Specific nucleotides are modified -Cc. GG-GGc. C- all sites * Hemophilus parainfluenzae -c. CGG-GGCc- all sites * Moricardia sp. -Ga. TC-CTa. G- all sites (dam) Escherichia coli -Cc(A/T)GG-GG(T/A)c. C- all sites (dcm) Escherichia coli -c. G-Gc- some sites ** mammals, some fungi (Neurospora) and plants -c. NG- some sites ** -GNcplants * An example of a modified restriction enzyme recognition site. These sites are usually modified in organisms with the corresponding restriction activity. ** Fraction of CG dinucleotides or CNG trinucleotides varies with species and, to a lesser extent, with tissue.
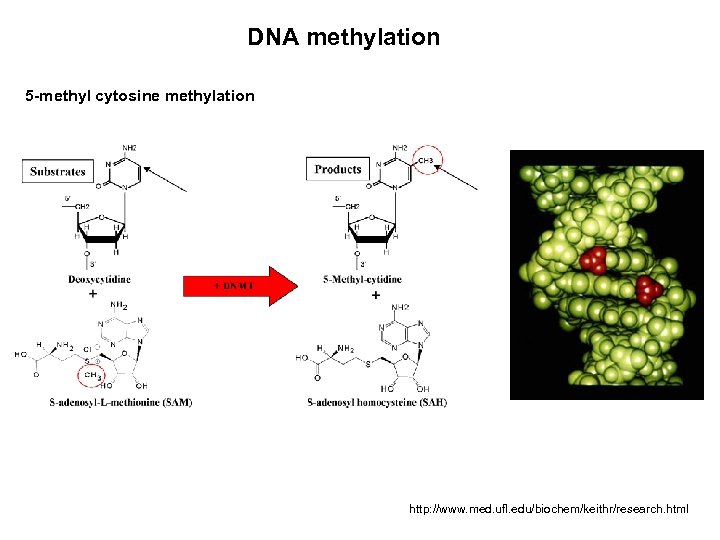
DNA methylation 5 -methyl cytosine methylation http: //www. med. ufl. edu/biochem/keithr/research. html
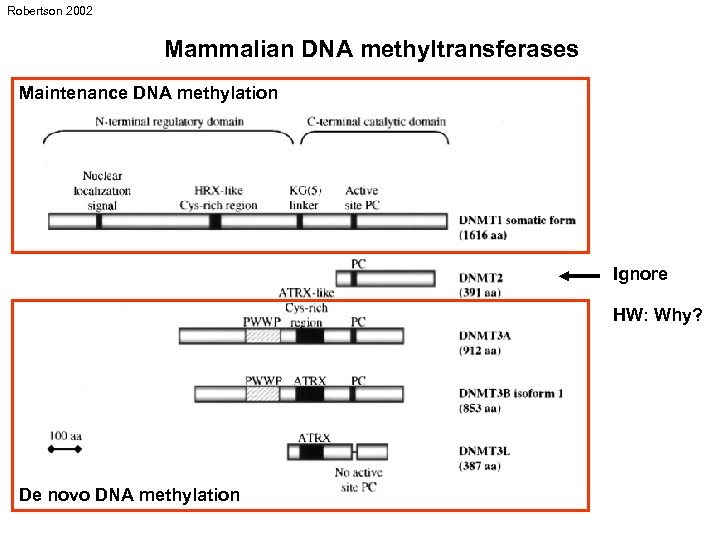
Robertson 2002 Mammalian DNA methyltransferases Maintenance DNA methylation Ignore HW: Why? De novo DNA methylation

Methods to study DNA methylation Global methylation analysis HPLC (complete hydrolysis, AP) TLC (complete hydrolysis, Msp. I, 32 P labeling with PNK) Sequence-specific methylation analysis Methylation sensitive restriction enzymes - a number of methods Bisulfite sequencing Me. DIP For more details, see Oakeley, 1999
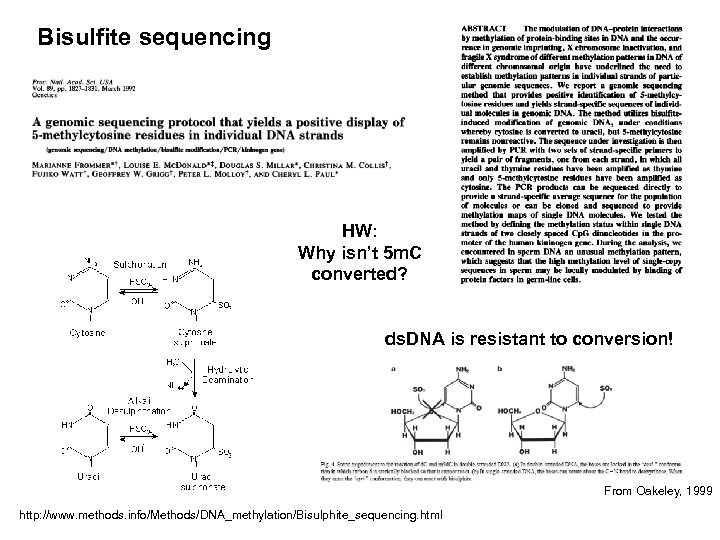
Bisulfite sequencing HW: Why isn’t 5 m. C converted? ds. DNA is resistant to conversion! From Oakeley, 1999 http: //www. methods. info/Methods/DNA_methylation/Bisulphite_sequencing. html
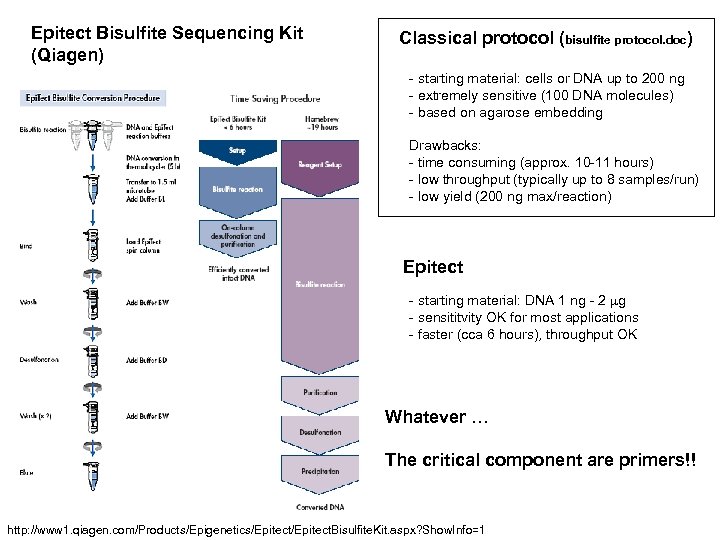
Epitect Bisulfite Sequencing Kit (Qiagen) Classical protocol (bisulfite protocol. doc) - starting material: cells or DNA up to 200 ng - extremely sensitive (100 DNA molecules) - based on agarose embedding Drawbacks: - time consuming (approx. 10 -11 hours) - low throughput (typically up to 8 samples/run) - low yield (200 ng max/reaction) Epitect - starting material: DNA 1 ng - 2 mg - sensititvity OK for most applications - faster (cca 6 hours), throughput OK Whatever … The critical component are primers!! http: //www 1. qiagen. com/Products/Epigenetics/Epitect. Bisulfite. Kit. aspx? Show. Info=1
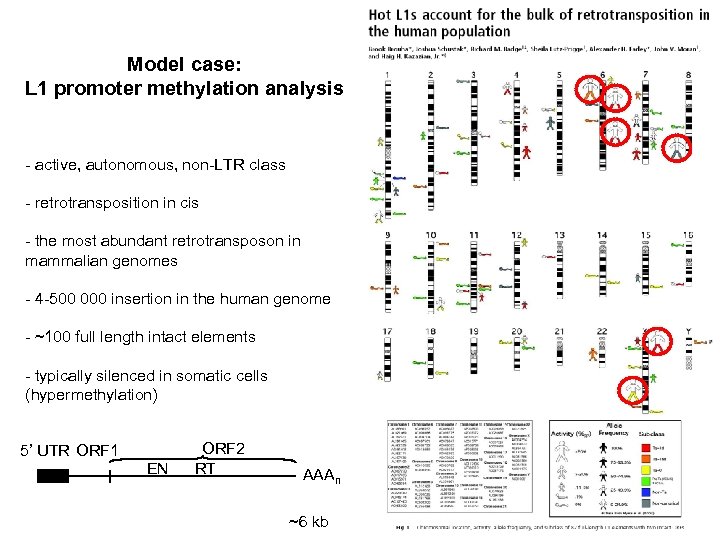
Model case: L 1 promoter methylation analysis - active, autonomous, non-LTR class - retrotransposition in cis - the most abundant retrotransposon in mammalian genomes - 4 -500 000 insertion in the human genome - ~100 full length intact elements - typically silenced in somatic cells (hypermethylation) 5’ UTR ORF 1 EN ORF 2 RT AAAn ~6 kb

BISULFITE SEQUENCING STEP BY STEP 1) Find your sequence – NCBI Genbank and Pubmed >L 1 5’ UTR GGGGGGAGGAGCCAAGATGGCCGAATAGGAACAGCTCCGGTCTACAGCTCCCAGCGTGA GCGACGCAGAAGACGGTGATTTCTGCATTTCCATCTGAGGTACCGGGTTCATCTCACTA GGGAGTGCCAGACAGTGGGCGCAGGCCAGTGTGCGCACCGTGCGCGAGCCGAAGC AGGGCGAGGCATTGCCTCACCTGGGAAGCGCAAGGGGTCAGGGAGTTCCCTTTCCGAGT CAAAGGGGTGACGCACCTGGAAAATCGGGTCACTCCCACCCGAATATTGCG CTTTTCAGACCGGCTTAAGAAACGGCGCACCACGAGACTATATCCCACACCTGGCTCGG AGGGTCCTACGCCCACGGAATCTCGCTGATTGCTAGCACAGCAGTCTGAGATCAAACTG CAAGGCGGCAACGAGGCTGGGGGAGGGGCGCCATTGCCCAGGCTTAGGTAA ACAAAGCAGCAGGGAAGCTCGAACTGGGTGGAGCCCACCACAGCTCAAGGAGGCCTGCCTCTGTAGGCTCCACCTCTGGGGGCACAGACAAAAAGACAGCAGTAA CCTCTGCAGACTTAAGTGTCCCTGTCTGACAGCTTTGAAGAGAGCAGTGGTTCTCCCAG CACGCAGCTGGAGATCTGAGAACGGGCAGACTGCCTCCTCAAGTGGGTCCCTGACCCCT GACCCCCGAGCAGCCTAACTGGGAGGCACCCCCCAGCAGGGGCACACTGACACCTCACA CGGCAGGGTATTCCAACAGACCTGCAGCTGAGGGTCCTGTTAGAAGGAAAACTAA CAACCAGAAAGGACATCTACACCGAAAACCCATCTGTACATCACCATCATCAAAGACCA AAAGTAGATAAAACCACAAAG

BISULFITE SEQUENCING STEP BY STEP 1) Find your sequence >L 1 5’ UTR GGGGGGAGGAGCCAAGATGGCCGAATAGGAACAGCTCCGGTCTACAGCTCCCAGCGTGA GCGACGCAGAAGACGGTGATTTCTGCATTTCCATCTGAGGTACCGGGTTCATCTCACTA GGGAGTGCCAGACAGTGGGCGCAGGCCAGTGTGCGCACCGTGCGCGAGCCGAAGC AGGGCGAGGCATTGCCTCACCTGGGAAGCGCAAGGGGTCAGGGAGTTCCCTTTCCGAGT CAAAGGGGTGACGCACCTGGAAAATCGGGTCACTCCCACCCGAATATTGCG CTTTTCAGACCGGCTTAAGAAACGGCGCACCACGAGACTATATCCCACACCTGGCTCGG AGGGTCCTACGCCCACGGAATCTCGCTGATTGCTAGCACAGCAGTCTGAGATCAAACTG CAAGGCGGCAACGAGGCTGGGGGAGGGGCGCCATTGCCCAGGCTTAGGTAA ACAAAGCAGCAGGGAAGCTCGAACTGGGTGGAGCCCACCACAGCTCAAGGAGGCCTGCCTCTGTAGGCTCCACCTCTGGGGGCACAGACAAAAAGACAGCAGTAA CCTCTGCAGACTTAAGTGTCCCTGTCTGACAGCTTTGAAGAGAGCAGTGGTTCTCCCAG CACGCAGCTGGAGATCTGAGAACGGGCAGACTGCCTCCTCAAGTGGGTCCCTGACCCCT GACCCCCGAGCAGCCTAACTGGGAGGCACCCCCCAGCAGGGGCACACTGACACCTCACA CGGCAGGGTATTCCAACAGACCTGCAGCTGAGGGTCCTGTTAGAAGGAAAACTAA CAACCAGAAAGGACATCTACACCGAAAACCCATCTGTACATCACCATCATCAAAGACCA AAAGTAGATAAAACCACAAAG HW: look at Cp. G density of: EGFP, Bluescript, Actin promoter and transcribed region, IAP LTR, L 1 ORF 2

BISULFITE SEQUENCING STEP BY STEP 2) Word -> REPLACE CG WITH XY >L 1 5’ UTR GGGGGGAGGAGCCAAGATGGCXYAATAGGAACAGCTCXYGTCTACAGCTCCCAGXYTGA GXYAXYCAGAAGAXYGTGATTTCTGCATTTCCATCTGAGGTACXYGGTTCATCTCACTA GGGAGTGCCAGACAGTGGGXYCAGGCCAGTGTGXYCACXYTGXYXYAGCXYAAGC AGGGXYAGGCATTGCCTCACCTGGGAAGXYCAAGGGGTCAGGGAGTTCCCTTTCXYAGT CAAAGGGGTGAXYCACCTGGAAAATXYGGTCACTCCCACCXYAATATTGXY CTTTTCAGACXYGCTTAAGAAAXYGXYCACCAXYAGACTATATCCCACACCTGGCTXYG AGGGTCCTAXYCCCAXYGAATCTXYCTGATTGCTAGCACAGCAGTCTGAGATCAAACTG CAAGGXYGCAAXYAGGCTGGGGGAGGGGXYCCATTGCCCAGGCTTAGGTAA ACAAAGCAGCAGGGAAGCTXYAACTGGGTGGAGCCCACCACAGCTCAAGGAGGCCTGCCTCTGTAGGCTCCACCTCTGGGGGCACAGACAAAAAGACAGCAGTAA CCTCTGCAGACTTAAGTGTCCCTGTCTGACAGCTTTGAAGAGAGCAGTGGTTCTCCCAG CAXYCAGCTGGAGATCTGAGAAXYGGCAGACTGCCTCCTCAAGTGGGTCCCTGACCCCT GACCCCXYAGCAGCCTAACTGGGAGGCACCCCCCAGCAGGGGCACACTGACACCTCACA XYGCAGGGTATTCCAACAGACCTGCAGCTGAGGGTCCTGTTAGAAGGAAAACTAA CAACCAGAAAGGACATCTACACXYAAAACCCATCTGTACATCACCATCATCAAAGACCA AAAGTAGATAAAACCACAAAG

BISULFITE SEQUENCING STEP BY STEP 3) Word -> REPLACE C WITH T >L 1 5’ UTR GGGGGGAGGAGTTAAGATGGTXYAATAGGAATAGTTTXYGTTTATAGTTTTTAGXYTGA GXYAXYTAGAAGAXYGTGATTTTTGTATTTTTATTTGAGGTATXYGGTTTATTA GGGAGTGTTAGATAGTGGGXYTAGGTTAGTGTGXYTATXYTGXYXYAGTXYAAGT AGGGXYAGGTATTGTTTTATTTGGGAAGXYTAAGGGGTTAGGGAGTTTTTXYAGT TAAAGGGGTGAXYTATTTGGAAAATXYGGTTATTTTTATTXYAATATTGXY TTTTTTAGATXYGTTTAAGAAAXYGXYTATTAXYAGATTATATTTGGTTXYG AGGGTTTTAXYGAATTTXYTTGATTGTTAGTAGTTTGAGATTAAATTG TAAGGXYGTAAXYAGGTTGGGGGAGGGGXYTTATTGTTTAGGTAA ATAAAGTAGTAGGGAAGTTXYAATTGGGTGGAGTTTATTATAGTTTAAGGAGGTTTGTTTTTGTAGGTTTTATTTTTGGGGGTATAGATAAAAAGATAGTAGTAA TTTTTGTAGATTTAAGTGTTTTTGATAGTTTTGAAGAGAGTAGTGGTTTTTTTAG TAXYTAGTTGGAGATTTGAGAAXYGGTAGATTGTTTTTTTAAGTGGGTTTTTGATTTTT GATTTTXYAGTAGTTTAATTGGGAGGTATTTTTTAGTAGGGGTATATTGATATTTTATA XYGTAGGGTATTTTAATAGATTTGTAGTTGAGGGTTTTGTTAGAAGGAAAATTAA TAATTAGAAAGGATATTTATATXYAAAATTTGTATATTATTAAAGATTA AAAGTAGATAAAATTATAAAG

BISULFITE SEQUENCING STEP BY STEP 4) Word -> REPLACE XY WITH CG >L 1 5’ UTR GGGGGGAGGAGTTAAGATGGTCGAATAGTTTCGGTTTATAGTTTTTAGCGTGA GCGACGTAGAAGACGGTGATTTTTGTATTTTTATTTGAGGTATCGGGTTTATTA GGGAGTGTTAGATAGTGGGCGTAGGTTAGTGTGCGTATCGTGCGCGAGTCGAAGT AGGGCGAGGTATTGTTTTATTTGGGAAGCGTAAGGGGTTAGGGAGTTTTTCGAGT TAAAGGGGTGACGTATTTGGAAAATCGGGTTATTTTTATTCGAATATTGCG TTTTTTAGATCGGTTTAAGAAACGGCGTATTACGAGATTATATTTGGTTCGG AGGGTTTTACGGAATTTCGTTGATTGTTAGTAGTTTGAGATTAAATTG TAAGGCGGTAACGAGGTTGGGGGAGGGGCGTTATTGTTTAGGTAA ATAAAGTAGTAGGGAAGTTCGAATTGGGTGGAGTTTATTATAGTTTAAGGAGGTTTGTTTTTGTAGGTTTTATTTTTGGGGGTATAGATAAAAAGATAGTAGTAA TTTTTGTAGATTTAAGTGTTTTTGATAGTTTTGAAGAGAGTAGTGGTTTTTTTAG TACGTAGTTGGAGATTTGAGAACGGGTAGATTGTTTTTTTAAGTGGGTTTTTGATTTTT GATTTTCGAGTAGTTTAATTGGGAGGTATTTTTTAGTAGGGGTATATTGATATTTTATA CGGTAGGGTATTTTAATAGATTTGTAGTTGAGGGTTTTGTTAGAAGGAAAATTAA TAATTAGAAAGGATATTTATATCGAAAATTTGTATATTATTAAAGATTA AAAGTAGATAAAATTATAAAG

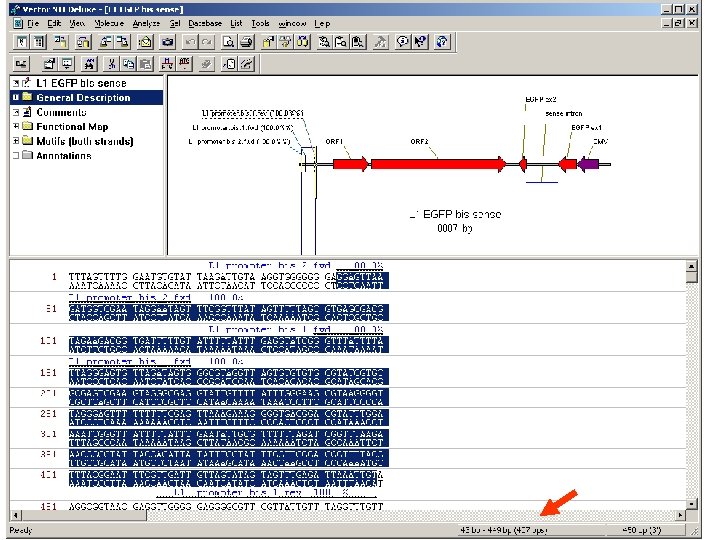
BISULFITE SEQUENCING STEP BY STEP 5) Design primers - size typically up to 500 bp, ideally around 300 bp - avoid low complexity sequences - Tm around 55 o. C - select the right region! >L 1 5’ UTR GGGGGGAGGAGTTAAGATGGTCGAATAGTTTCGGTTTATAGTTTTTAGCGTGA GCGACGTAGAAGACGGTGATTTTTGTATTTTTATTTGAGGTATCGGGTTTATTA GGGAGTGTTAGATAGTGGGCGTAGGTTAGTGTGCGTATCGTGCGCGAGTCGAAGT AGGGCGAGGTATTGTTTTATTTGGGAAGCGTAAGGGGTTAGGGAGTTTTTCGAGT TAAAGGGGTGACGTATTTGGAAAATCGGGTTATTTTTATTCGAATATTGCG TTTTTTAGATCGGTTTAAGAAACGGCGTATTACGAGATTATATTTGGTTCGG AGGGTTTTACGGAATTTCGTTGATTGTTAGTAGTTTGAGATTAAATTG TAAGGCGGTAACGAGGTTGGGGGAGGGGCGTTATTGTTTAGGTAA ATAAAGTAGTAGGGAAGTTCGAATTGGGTGGAGTTTATTATAGTTTAAGGAGGTTTGTTTTTGTAGGTTTTATTTTTGGGGGTATAGATAAAAAGATAGTAGTAA TTTTTGTAGATTTAAGTGTTTTTGATAGTTTTGAAGAGAGTAGTGGTTTTTTTAG TACGTAGTTGGAGATTTGAGAACGGGTAGATTGTTTTTTTAAGTGGGTTTTTGATTTTT GATTTTCGAGTAGTTTAATTGGGAGGTATTTTTTAGTAGGGGTATATTGATATTTTATA CGGTAGGGTATTTTAATAGATTTGTAGTTGAGGGTTTTGTTAGAAGGAAAATTAA TAATTAGAAAGGATATTTATATCGAAAATTTGTATATTATTAAAGATTA AAAGTAGATAAAATTATAAAG


BISULFITE SEQUENCING STEP BY STEP 5) Design primers - size typically up to 500 bp, ideally around 300 bp - avoid low complexity sequences - Tm around 55 o. C - select the right region! >L 1 5’ UTR GGGGGGAGGAGTTAAGATGGTCGAATAGTTTCGGTTTATAGTTTTTAGCGTGA GCGACGTAGAAGACGGTGATTTTTGTATTTTTATTTGAGGTATCGGGTTTATTA GGGAGTGTTAGATAGTGGGCGTAGGTTAGTGTGCGTATCGTGCGCGAGTCGAAGT AGGGCGAGGTATTGTTTTATTTGGGAAGCGTAAGGGGTTAGGGAGTTTTTCGAGT TAAAGGGGTGACGTATTTGGAAAATCGGGTTATTTTTATTCGAATATTGCG TTTTTTAGATCGGTTTAAGAAACGGCGTATTACGAGATTATATTTGGTTCGG AGGGTTTTACGGAATTTCGTTGATTGTTAGTAGTTTGAGATTAAATTG TAAGGCGGTAACGAGGTTGGGGGAGGGGCGTTATTGTTTAGGTAA ATAAAGTAGTAGGGAAGTTCGAATTGGGTGGAGTTTATTATAGTTTAAGGAGGTTTGTTTTTGTAGGTTTTATTTTTGGGGGTATAGATAAAAAGATAGTAGTAA TTTTTGTAGATTTAAGTGTTTTTGATAGTTTTGAAGAGAGTAGTGGTTTTTTTAG TACGTAGTTGGAGATTTGAGAACGGGTAGATTGTTTTTTTAAGTGGGTTTTTGATTTTT GATTTTCGAGTAGTTTAATTGGGAGGTATTTTTTAGTAGGGGTATATTGATATTTTATA CGGTAGGGTATTTTAATAGATTTGTAGTTGAGGGTTTTGTTAGAAGGAAAATTAA TAATTAGAAAGGATATTTATATCGAAAATTTGTATATTATTAAAGATTA AAAGTAGATAAAATTATAAAG
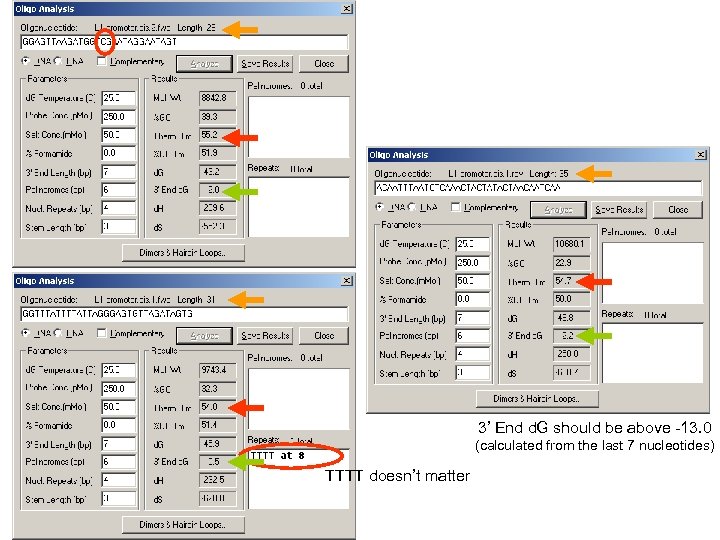
3’ End d. G should be above -13. 0 (calculated from the last 7 nucleotides) TTTT doesn’t matter
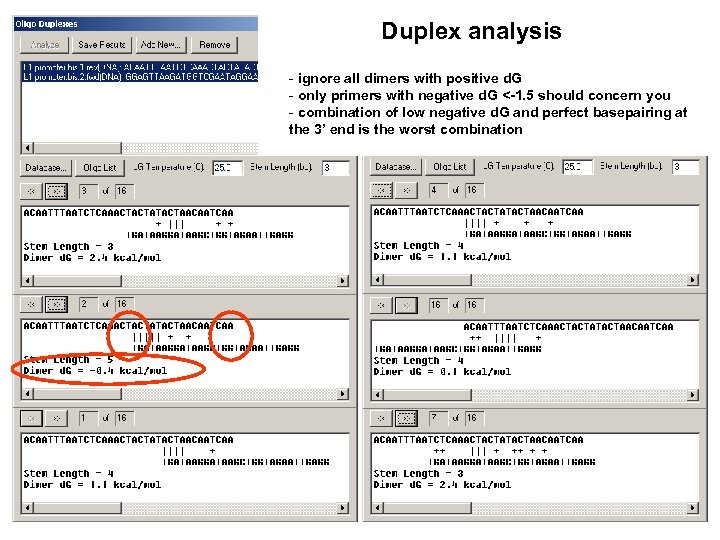
Duplex analysis - ignore all dimers with positive d. G - only primers with negative d. G <-1. 5 should concern you - combination of low negative d. G and perfect basepairing at the 3’ end is the worst combination
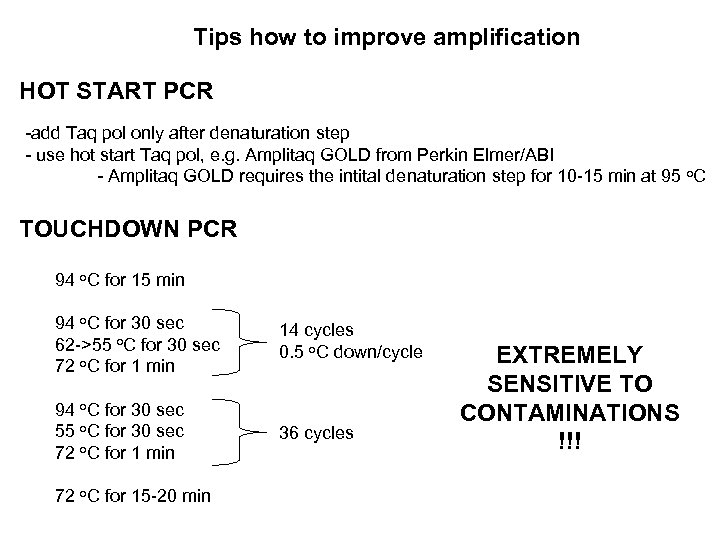

Tips how to improve amplification HOT START PCR -add Taq pol only after denaturation step - use hot start Taq pol, e. g. Amplitaq GOLD from Perkin Elmer/ABI - Amplitaq GOLD requires the intital denaturation step for 10 -15 min at 95 o. C TOUCHDOWN PCR 94 o. C for 15 min 94 o. C for 30 sec 62 ->55 o. C for 30 sec 72 o. C for 1 min 14 cycles 0. 5 o. C down/cycle 94 o. C for 30 sec 55 o. C for 30 sec 72 o. C for 1 min 36 cycles 72 o. C for 15 -20 min EXTREMELY SENSITIVE TO CONTAMINATIONS !!!
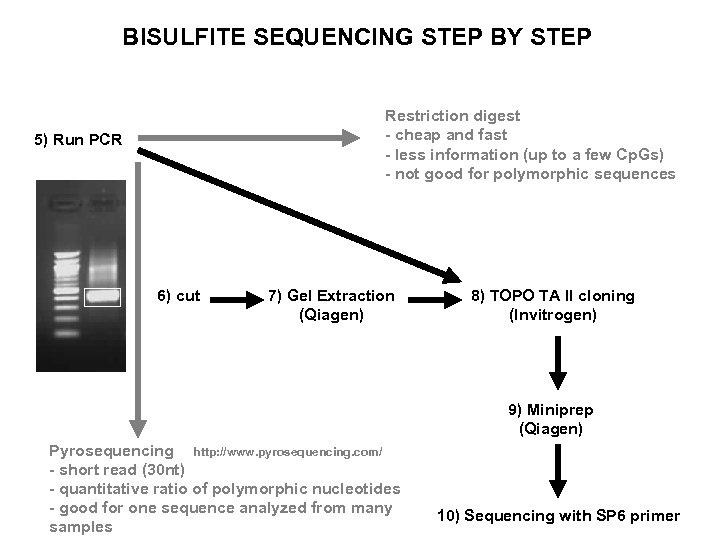
BISULFITE SEQUENCING STEP BY STEP Restriction digest - cheap and fast - less information (up to a few Cp. Gs) - not good for polymorphic sequences 5) Run PCR 6) cut 7) Gel Extraction (Qiagen) 8) TOPO TA II cloning (Invitrogen) 9) Miniprep (Qiagen) Pyrosequencing http: //www. pyrosequencing. com/ - short read (30 nt) - quantitative ratio of polymorphic nucleotides - good for one sequence analyzed from many samples 10) Sequencing with SP 6 primer
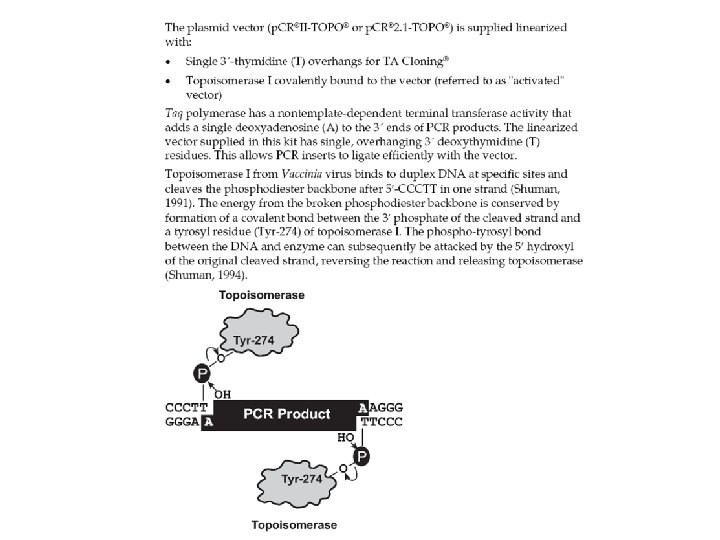
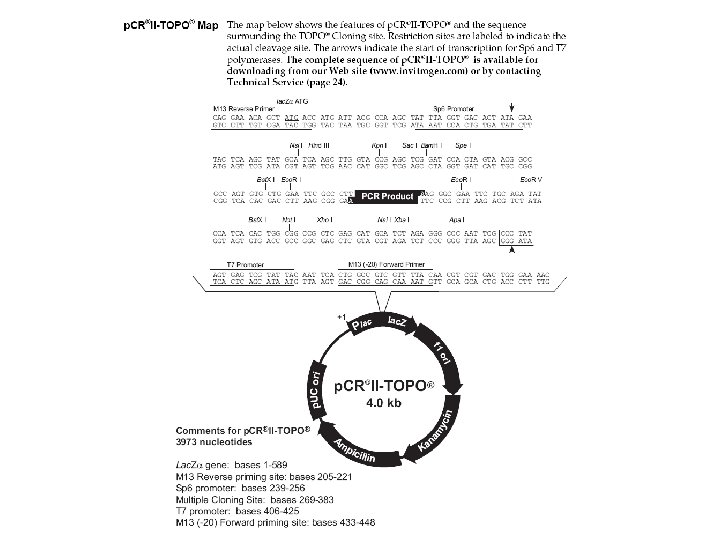


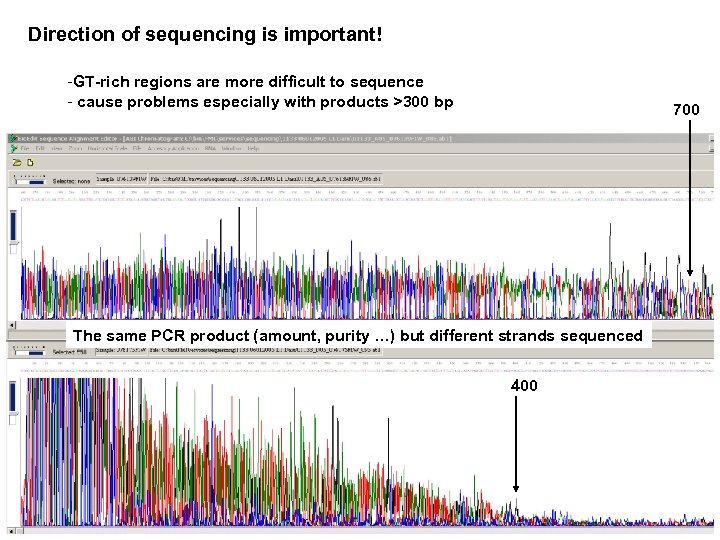
Direction of sequencing is important! -GT-rich regions are more difficult to sequence - cause problems especially with products >300 bp 700 The same PCR product (amount, purity …) but different strands sequenced 400
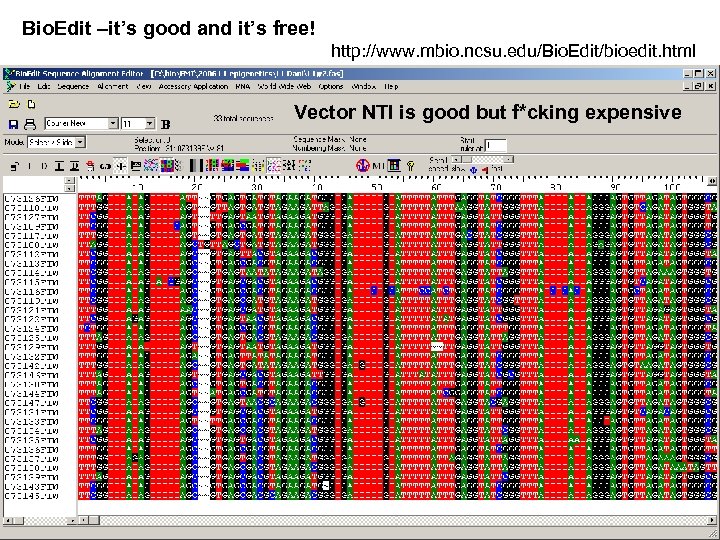
Bio. Edit –it’s good and it’s free! http: //www. mbio. ncsu. edu/Bio. Edit/bioedit. html Vector NTI is good but f*cking expensive
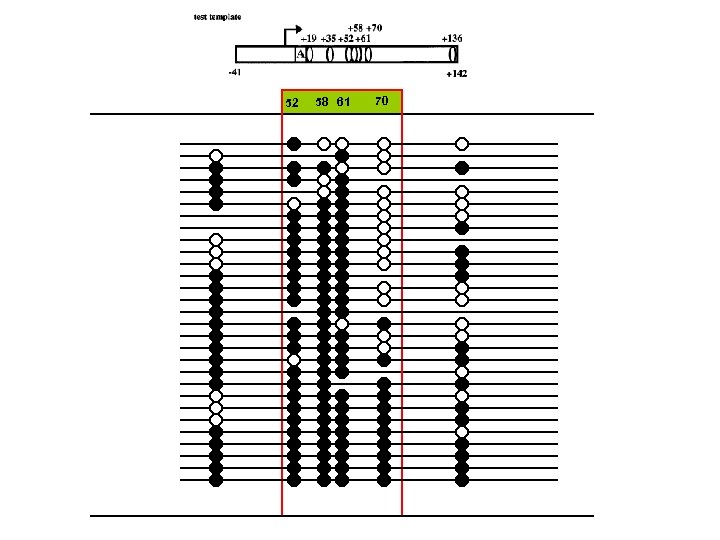
52 58 61 70
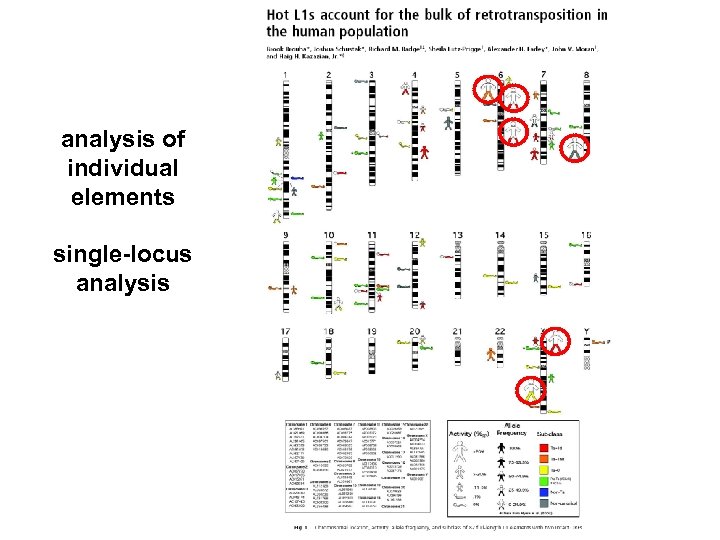
analysis of individual elements single-locus analysis
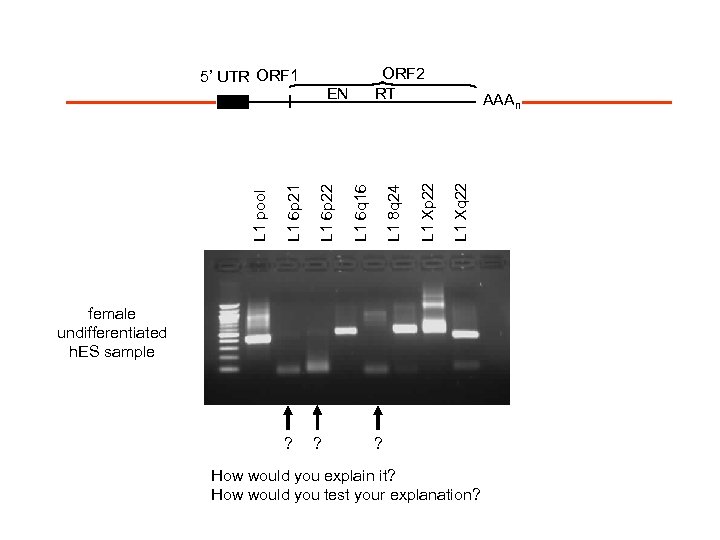
AAAn L 1 Xq 22 L 1 Xp 22 L 1 6 p 22 ? L 1 6 q 16 L 1 6 p 21 EN ? L 1 pool ORF 2 RT L 1 8 q 24 5’ UTR ORF 1 female undifferentiated h. ES sample ? How would you explain it? How would you test your explanation?
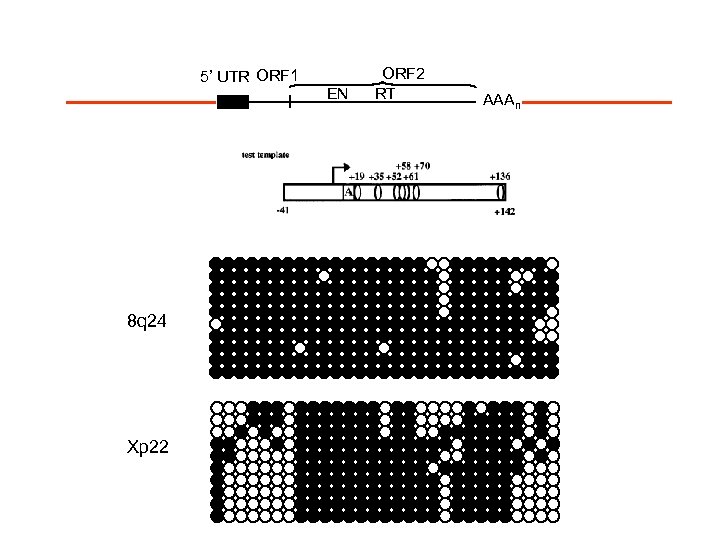
5’ UTR ORF 1 8 q 24 Xp 22 EN ORF 2 RT AAAn
HOMEWORK Oct-4 promoter analysis -PCR cloned in p. CR II -sequenced with SP 6
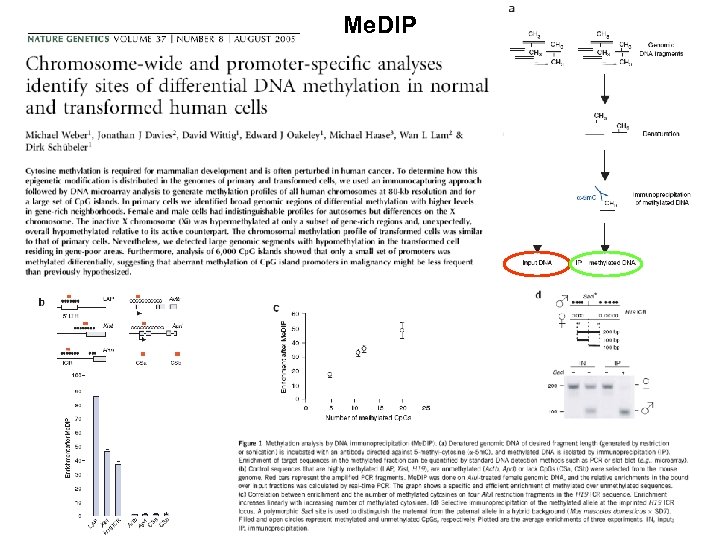
Me. DIP

DNA METHYLATION II (EFFECTS OF DNA METHYLATION)
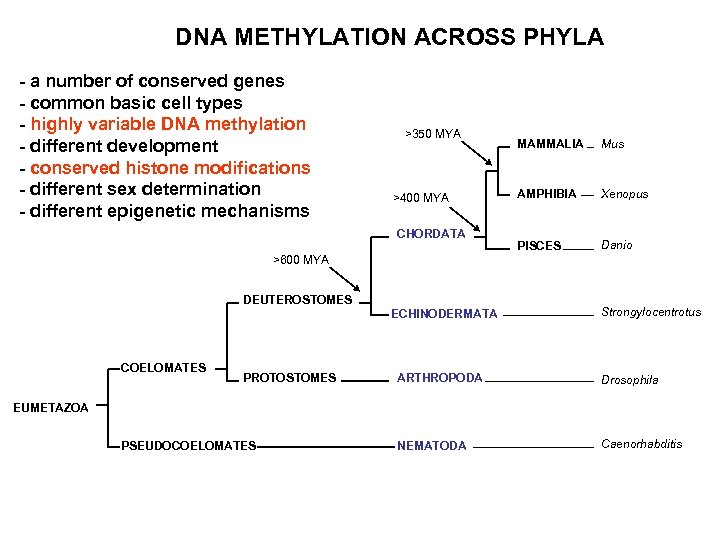
DNA METHYLATION ACROSS PHYLA - a number of conserved genes - common basic cell types - highly variable DNA methylation - different development - conserved histone modifications - different sex determination - different epigenetic mechanisms >350 MYA >400 MYA CHORDATA MAMMALIA Mus AMPHIBIA Xenopus PISCES Danio >600 MYA DEUTEROSTOMES ECHINODERMATA COELOMATES PROTOSTOMES Strongylocentrotus ARTHROPODA Drosophila NEMATODA Caenorhabditis EUMETAZOA PSEUDOCOELOMATES
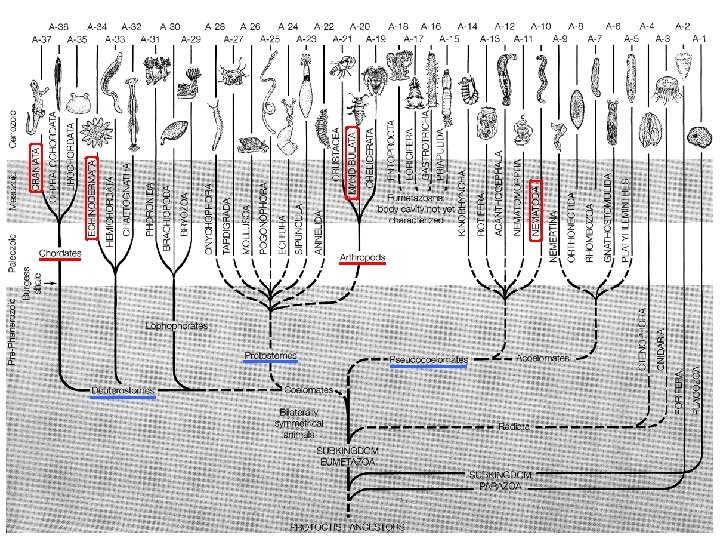
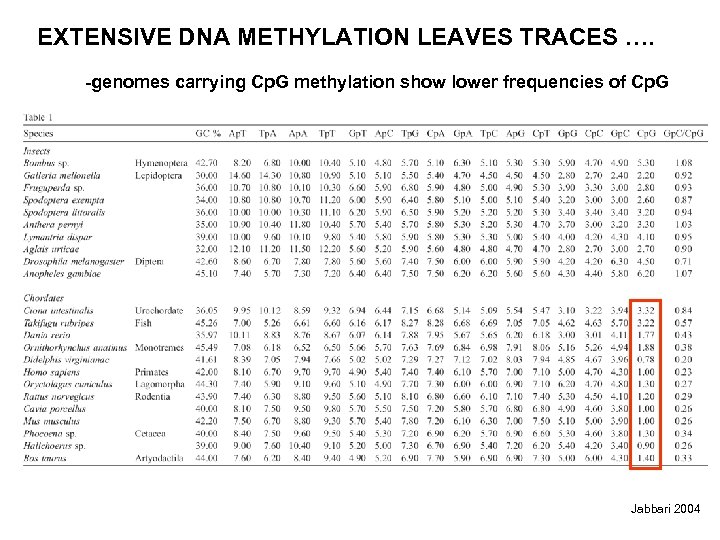
EXTENSIVE DNA METHYLATION LEAVES TRACES …. -genomes carrying Cp. G methylation show lower frequencies of Cp. G Jabbari 2004
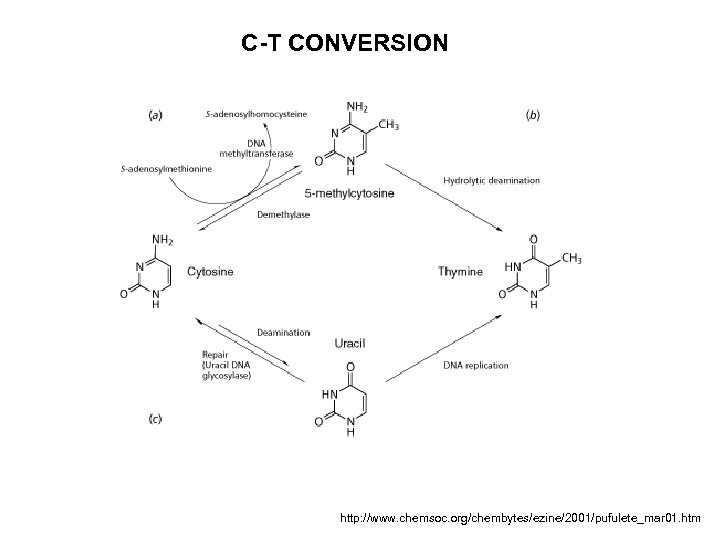
C-T CONVERSION http: //www. chemsoc. org/chembytes/ezine/2001/pufulete_mar 01. htm
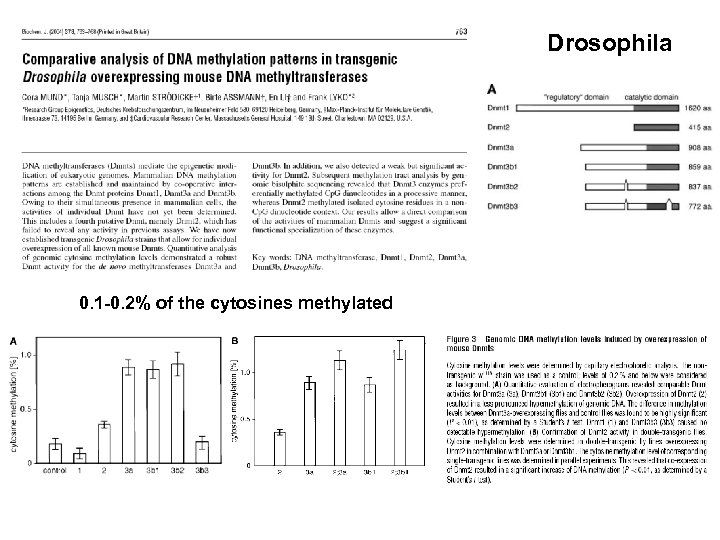
Drosophila 0. 1 -0. 2% of the cytosines methylated
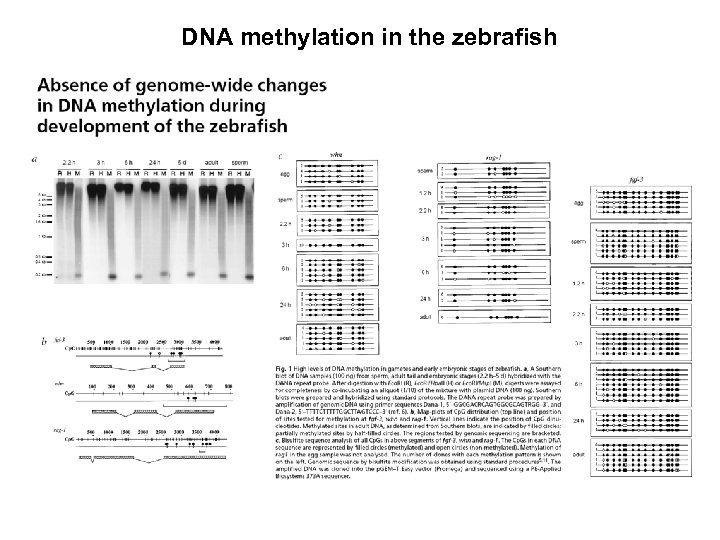
DNA methylation in the zebrafish
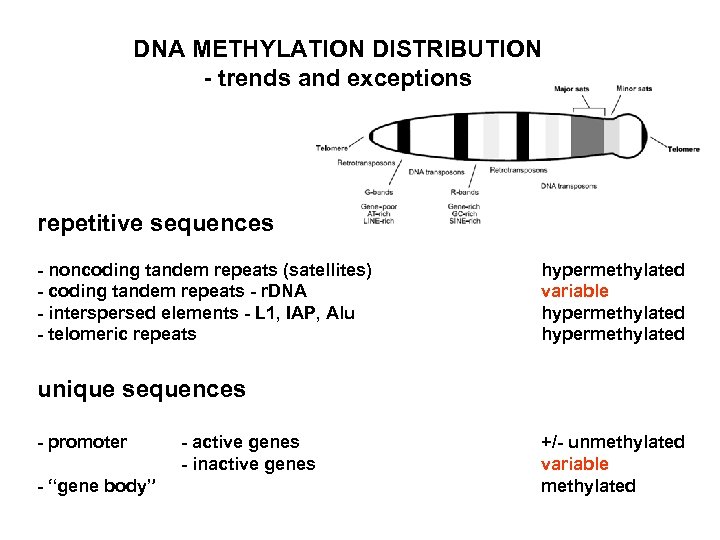
DNA METHYLATION DISTRIBUTION - trends and exceptions repetitive sequences - noncoding tandem repeats (satellites) - coding tandem repeats - r. DNA - interspersed elements - L 1, IAP, Alu - telomeric repeats hypermethylated variable hypermethylated unique sequences - promoter - “gene body” - active genes - inactive genes +/- unmethylated variable methylated
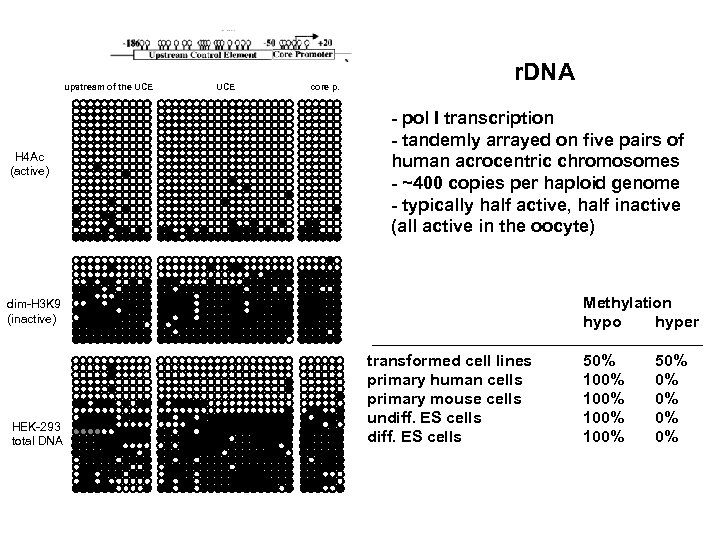
upstream of the UCE H 4 Ac (active) UCE core p. r. DNA - pol I transcription - tandemly arrayed on five pairs of human acrocentric chromosomes - ~400 copies per haploid genome - typically half active, half inactive (all active in the oocyte) Methylation hypo hyper dim-H 3 K 9 (inactive) HEK-293 total DNA transformed cell lines primary human cells primary mouse cells undiff. ES cells 50% 100% 50% 0% 0%
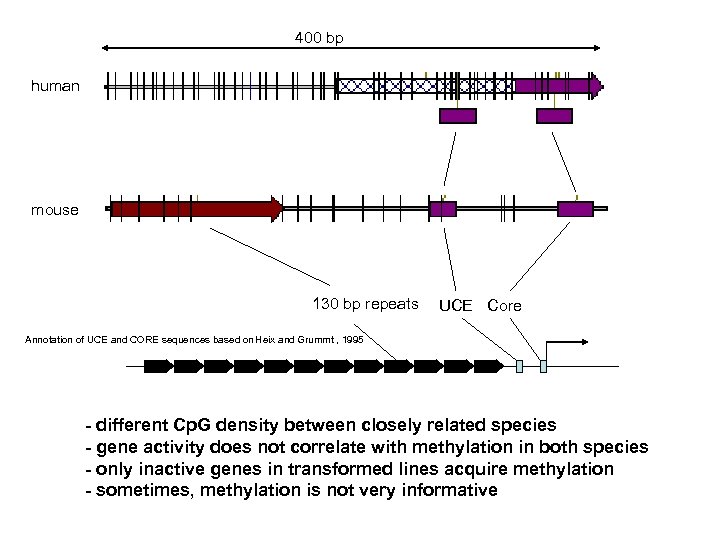
400 bp human mouse 130 bp repeats UCE Core Annotation of UCE and CORE sequences based on Heix and Grummt , 1995 - different Cp. G density between closely related species - gene activity does not correlate with methylation in both species - only inactive genes in transformed lines acquire methylation - sometimes, methylation is not very informative
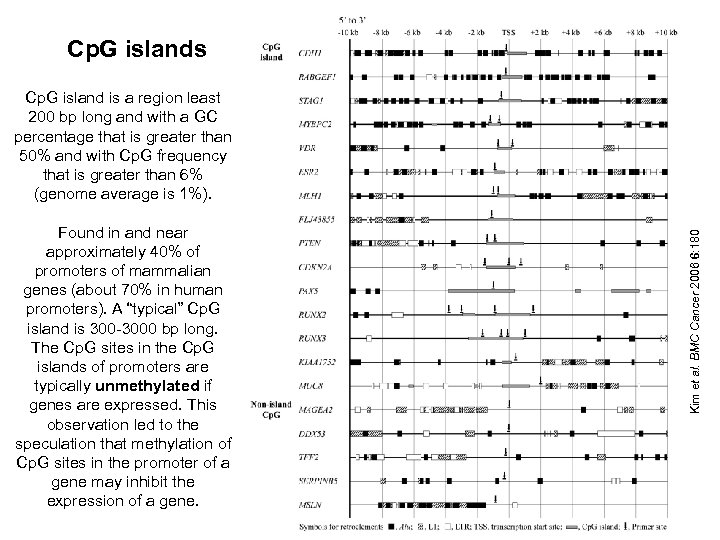
Cp. G islands Found in and near approximately 40% of promoters of mammalian genes (about 70% in human promoters). A “typical” Cp. G island is 300 -3000 bp long. The Cp. G sites in the Cp. G islands of promoters are typically unmethylated if genes are expressed. This observation led to the speculation that methylation of Cp. G sites in the promoter of a gene may inhibit the expression of a gene. Kim et al. BMC Cancer 2006 6: 180 Cp. G island is a region least 200 bp long and with a GC percentage that is greater than 50% and with Cp. G frequency that is greater than 6% (genome average is 1%).
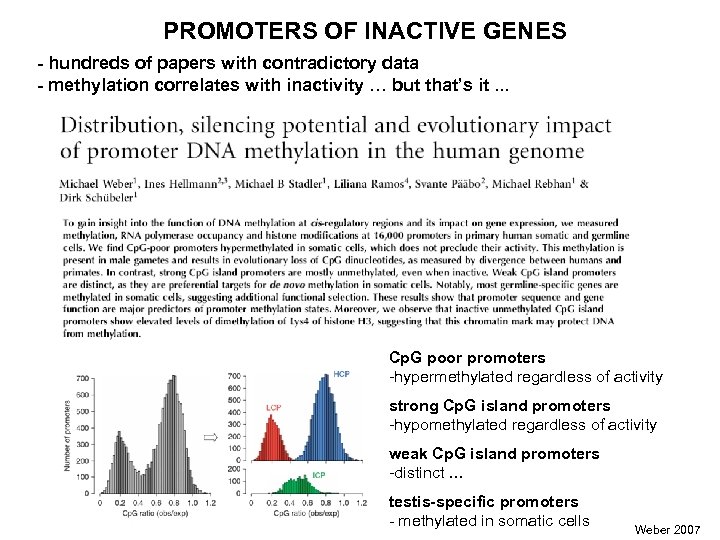
PROMOTERS OF INACTIVE GENES - hundreds of papers with contradictory data - methylation correlates with inactivity … but that’s it. . . Cp. G poor promoters -hypermethylated regardless of activity strong Cp. G island promoters -hypomethylated regardless of activity weak Cp. G island promoters -distinct … testis-specific promoters - methylated in somatic cells Weber 2007
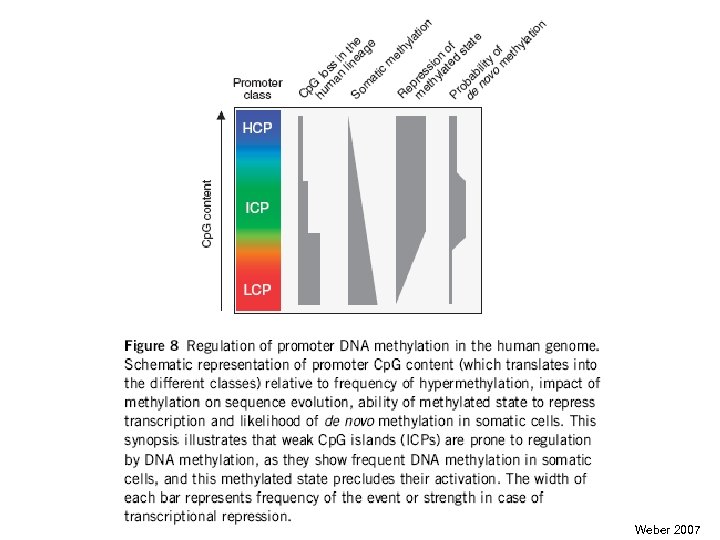
Weber 2007
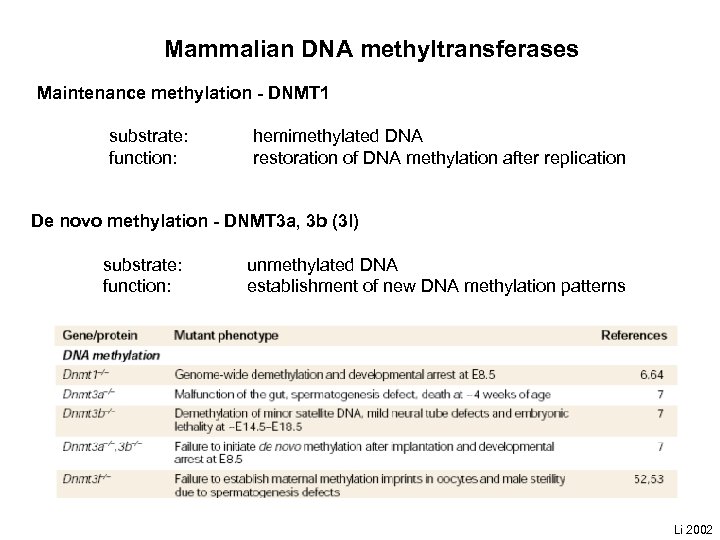

Mammalian DNA methyltransferases Maintenance methylation - DNMT 1 substrate: function: hemimethylated DNA restoration of DNA methylation after replication De novo methylation - DNMT 3 a, 3 b (3 l) substrate: function: unmethylated DNA establishment of new DNA methylation patterns Li 2002
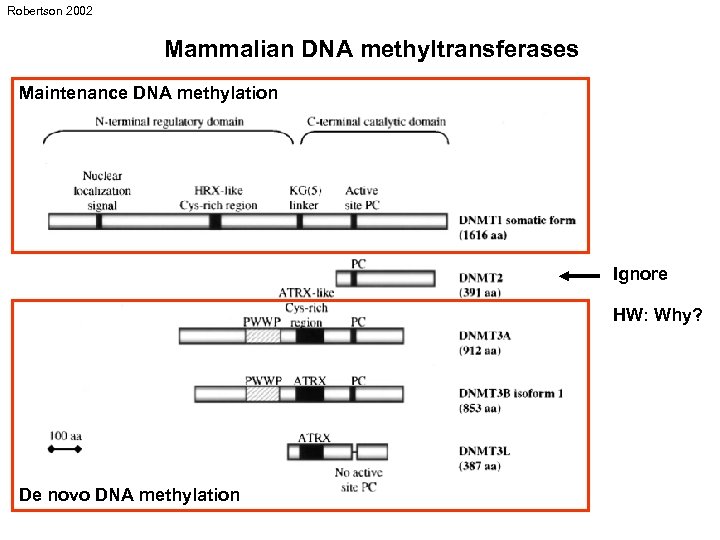
Robertson 2002 Mammalian DNA methyltransferases Maintenance DNA methylation Ignore HW: Why? De novo DNA methylation

Cooperativity between maintenance and de novo DNA methylation LINE-1 IAP, centromeres
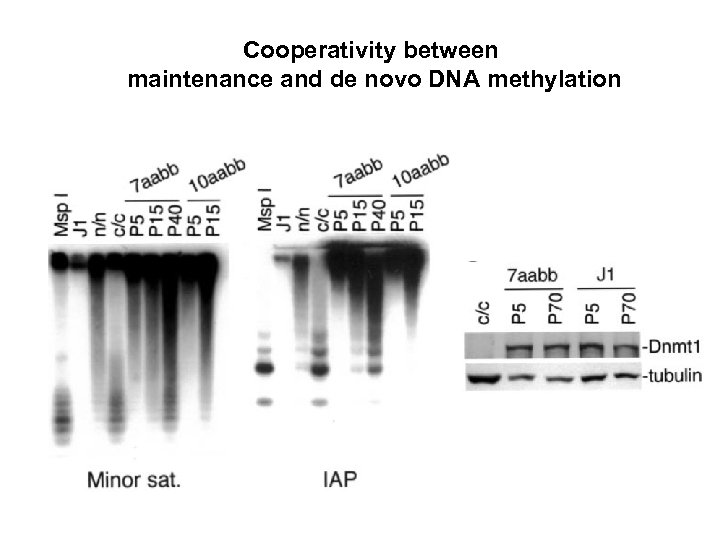
Cooperativity between maintenance and de novo DNA methylation
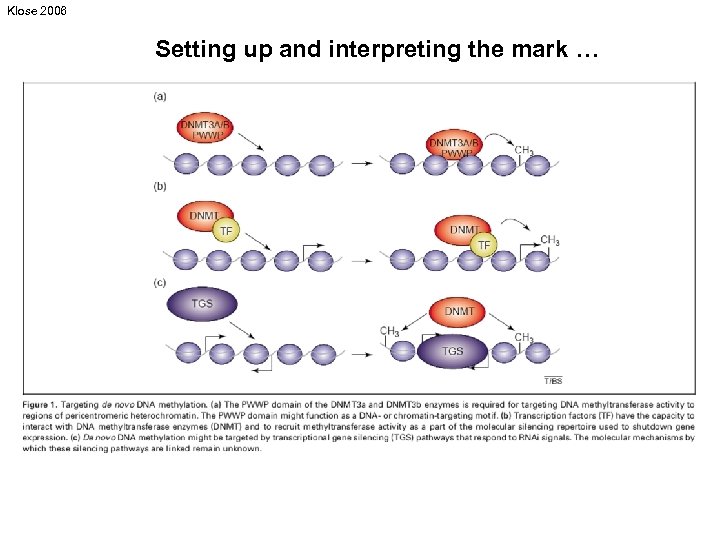
Klose 2006 Setting up and interpreting the mark …
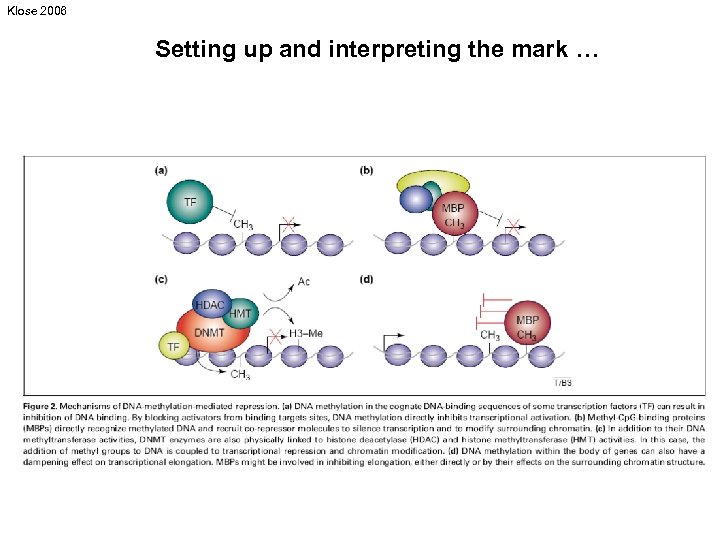
Klose 2006 Setting up and interpreting the mark …
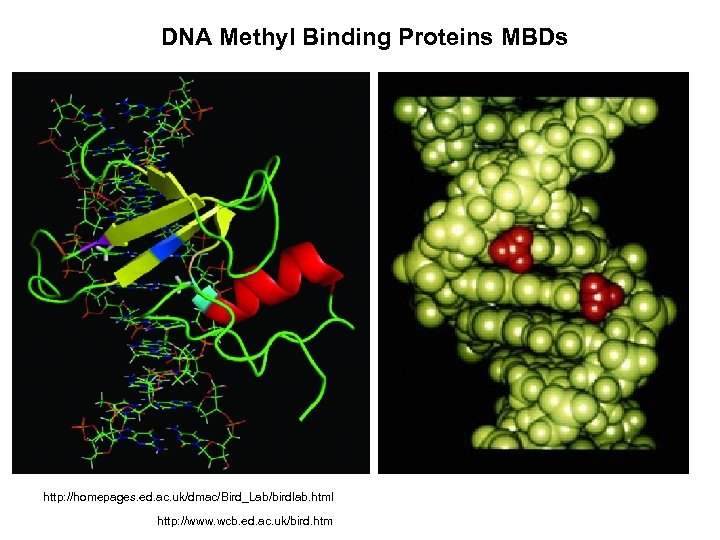
DNA Methyl Binding Proteins MBDs http: //homepages. ed. ac. uk/dmac/Bird_Lab/birdlab. html http: //www. wcb. ed. ac. uk/bird. htm
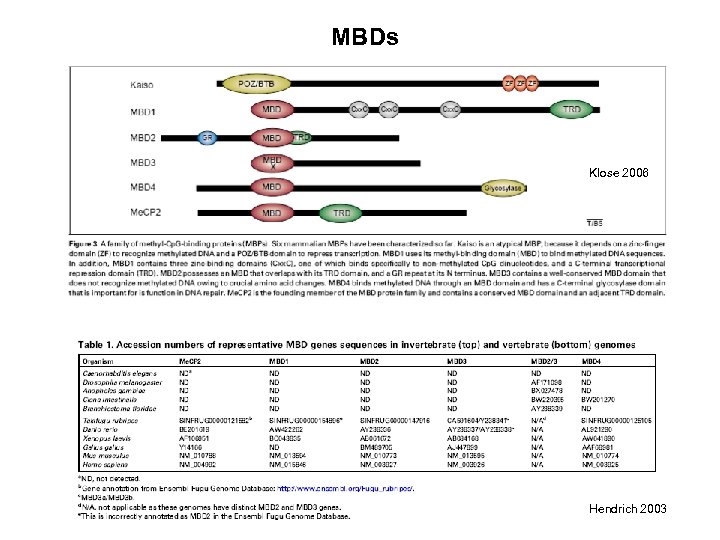
MBDs Klose 2006 Hendrich 2003
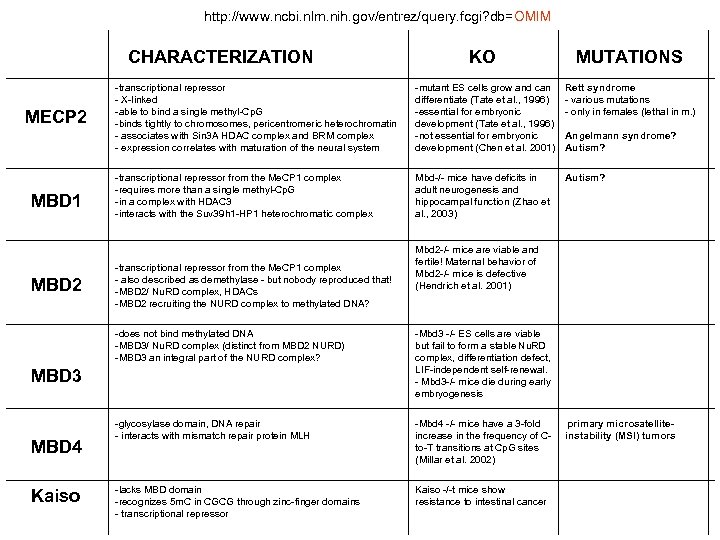

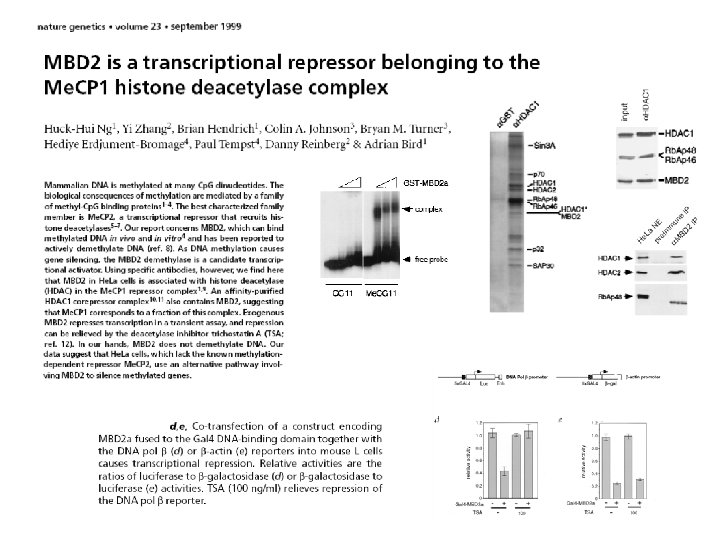
http: //www. ncbi. nlm. nih. gov/entrez/query. fcgi? db=OMIM CHARACTERIZATION MECP 2 MBD 1 MBD 2 KO MUTATIONS -transcriptional repressor - X-linked -able to bind a single methyl-Cp. G -binds tightly to chromosomes, pericentromeric heterochromatin - associates with Sin 3 A HDAC complex and BRM complex - expression correlates with maturation of the neural system -mutant ES cells grow and can differentiate (Tate et al. , 1996) -essential for embryonic development (Tate et al. , 1996) -not essential for embryonic development (Chen et al. 2001) Rett syndrome - various mutations - only in females (lethal in m. ) -transcriptional repressor from the Me. CP 1 complex -requires more than a single methyl-Cp. G -in a complex with HDAC 3 -interacts with the Suv 39 h 1 -HP 1 heterochromatic complex Mbd-/- mice have deficits in adult neurogenesis and hippocampal function (Zhao et al. , 2003) Autism? -transcriptional repressor from the Me. CP 1 complex - also described as demethylase - but nobody reproduced that! -MBD 2/ Nu. RD complex, HDACs -MBD 2 recruiting the NURD complex to methylated DNA? Mbd 2 -/- mice are viable and fertile! Maternal behavior of Mbd 2 -/- mice is defective (Hendrich et al. 2001) -does not bind methylated DNA -MBD 3/ Nu. RD complex (distinct from MBD 2 NURD) -MBD 3 an integral part of the NURD complex? -Mbd 3 -/- ES cells are viable but fail to form a stable Nu. RD complex, differentiation defect, LIF-independent self-renewal. - Mbd 3 -/- mice die during early embryogenesis -glycosylase domain, DNA repair - interacts with mismatch repair protein MLH -Mbd 4 -/- mice have a 3 -fold increase in the frequency of Cto-T transitions at Cp. G sites (Millar et al. 2002) -lacks MBD domain -recognizes 5 m. C in CGCG through zinc-finger domains - transcriptional repressor Kaiso -/-t mice show resistance to intestinal cancer MBD 3 MBD 4 Kaiso Angelmann syndrome? Autism? primary microsatelliteinstability (MSI) tumors
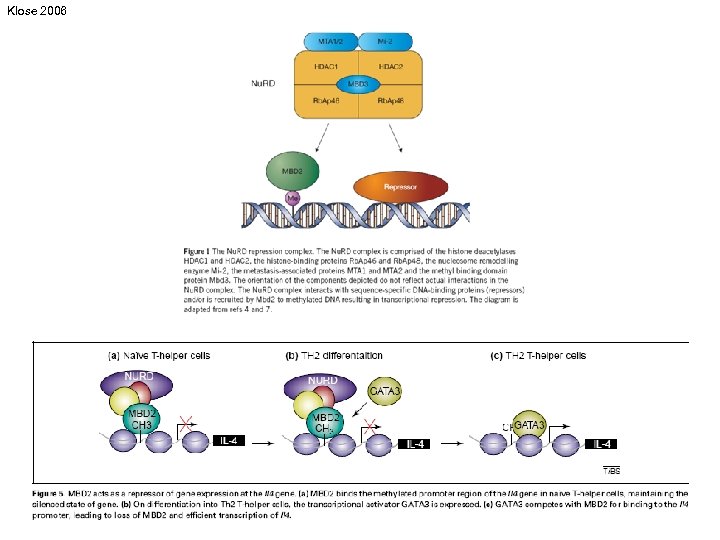
Klose 2006
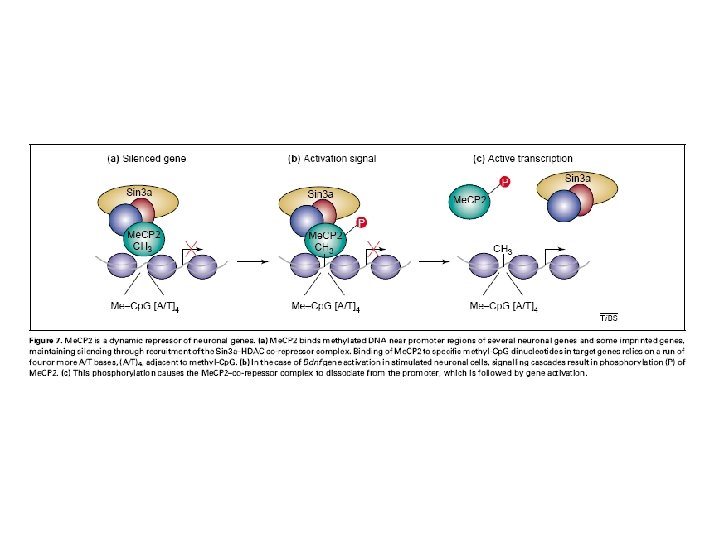
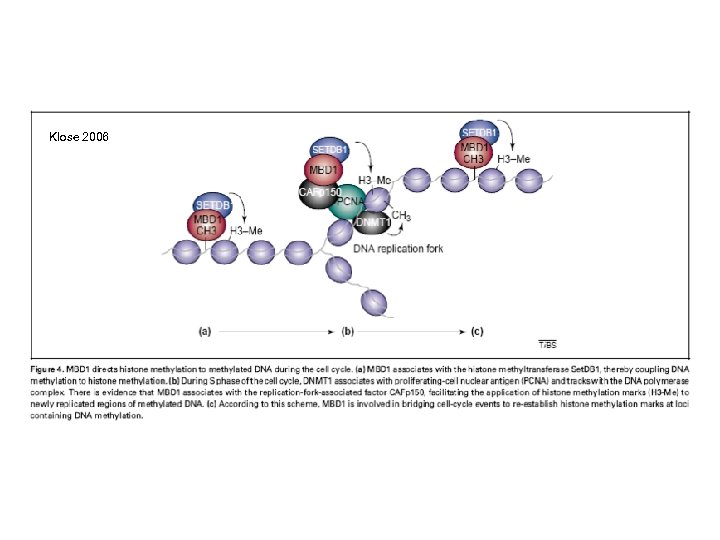
Klose 2006

A few final notes regarding DNA methylation - when it is associated with transcriptional repression, there are usually also chromatin modifications consistent with the pattern - information about DNA sequence methylation without information about location (promoter, intergenic, intronic, exonic …) and expression has low information content - methylated promoter does not automatically belong to an inactive gene while unmethylated promoter does not automatically belong to an active gene - it’s OK to say that DNA methylation is typically associated with transcriptional repression - be open minded and expect the unexpected. . .
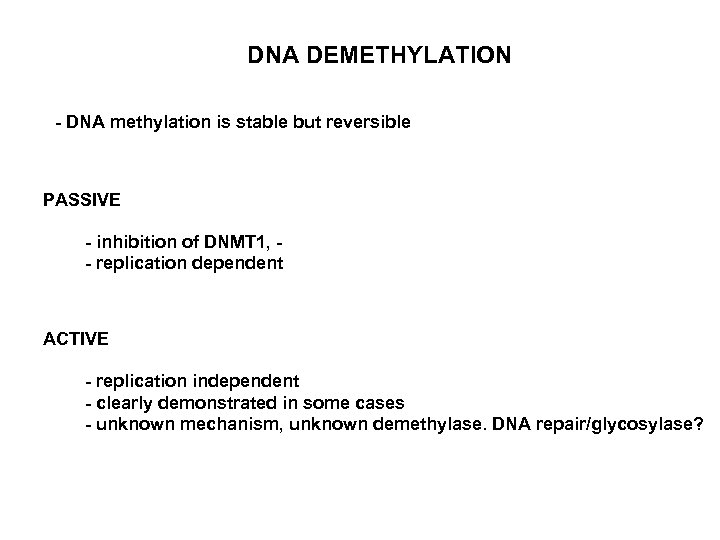


DNA DEMETHYLATION - DNA methylation is stable but reversible PASSIVE - inhibition of DNMT 1, - replication dependent ACTIVE - replication independent - clearly demonstrated in some cases - unknown mechanism, unknown demethylase. DNA repair/glycosylase?
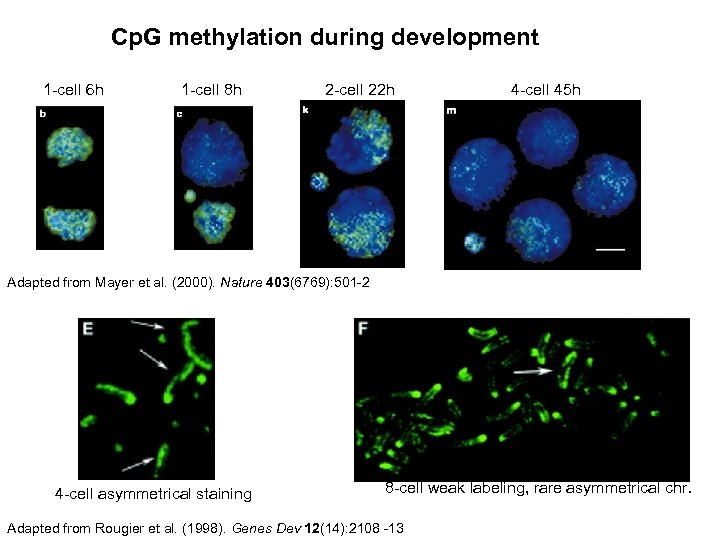
Cp. G methylation during development 1 -cell 6 h 1 -cell 8 h 2 -cell 22 h 4 -cell 45 h Adapted from Mayer et al. (2000). Nature 403(6769): 501 -2 4 -cell asymmetrical staining 8 -cell weak labeling, rare asymmetrical chr. Adapted from Rougier et al. (1998). Genes Dev 12(14): 2108 -13
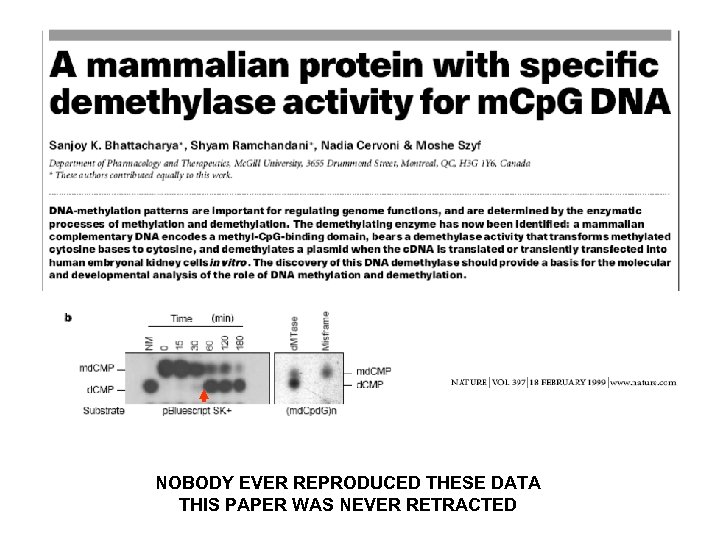

NOBODY EVER REPRODUCED THESE DATA THIS PAPER WAS NEVER RETRACTED

Reading the original literature. . . Introduction and Background: What hypotheses are being tested in this paper? What information made the authors to perform the experiments? What new methods or insights brought to bear on the problem? Methods: What are the critical methods of the paper? What enabling technologies are used? What are the weaknesses of the methods used? Are there other or better approaches that could be used? If this is a genetics approach, what would be a biochemical or molecular approach? If this is a biochemical or molecular approach, what genetics methods could be used? Results and Discussion What are the primary conclusions of the paper? Did the authors test their hypotheses properly? What novel information or directions come from this work? What control experiments were performed? What assumptions still remain in the work? How could these assumptions be tested? What other explanations for the observations are still possible?
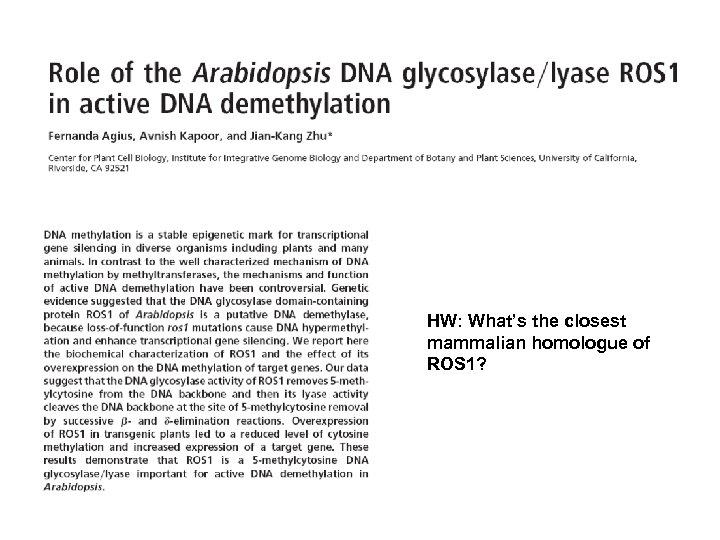

HW: What’s the closest mammalian homologue of ROS 1?
ce04f7de4f9128ab013952812e5169c4.ppt