03c5b3d2d10c9acce839175a5e7691a4.ppt
- Количество слайдов: 42

EECS 800 Research Seminar Mining Biological Data Instructor: Luke Huan Fall, 2006 The UNIVERSITY of Kansas

Administrative Paper presentation schedule: Han, Bin, Kernel method in Analyzing Biological Data, Nov 6 th Barker, Brett, Data Mining in Systems Biology, Nov 8 th Leung, Daniel, High performance in Data Mining, Nov 13 th Ku, Matthew, Data Mining in Proteomics, Nov 15 th Lin, Cindy, Integrating Biological Data, Nov 20 th Jia, Yi, Analyzing Bionetworks, Nov 22 th 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 2

Sequential Pattern Mining Why sequential pattern mining? GSP algorithm Free. Span and Prefix. Span Boarder Collapsing Constraints and extensions 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 3

Sequence Databases and Sequential Pattern Analysis (Temporal) order is important in many situations Time-series databases and sequence databases Frequent patterns (frequent) sequential patterns Applications of sequential pattern mining Customer shopping sequences: First buy computer, then CD-ROM, and then digital camera, within 3 months. Medical treatment, natural disasters (e. g. , earthquakes), science & engineering processes, stocks and markets, telephone calling patterns, Weblog click streams, DNA sequences and gene structures 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 4
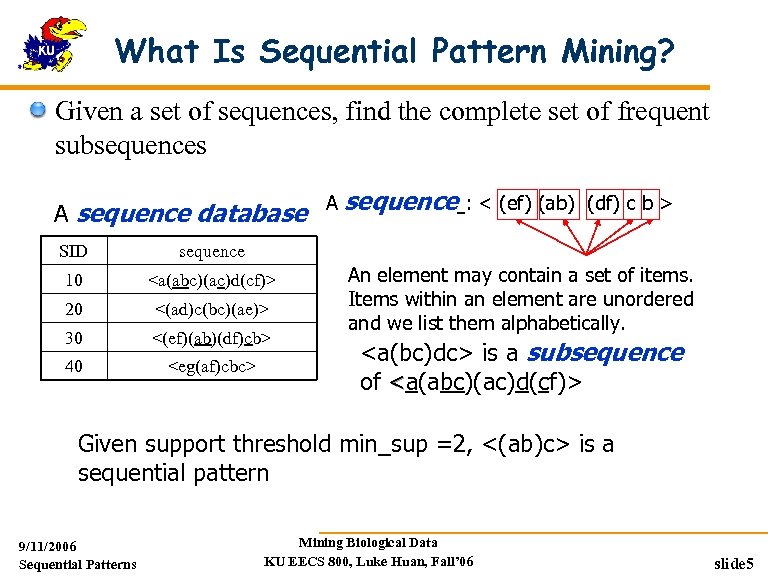
What Is Sequential Pattern Mining? Given a set of sequences, find the complete set of frequent subsequences A sequence database A sequence : < (ef) (ab) (df) c b > SID sequence 10 <a(abc)(ac)d(cf)> 20 <(ad)c(bc)(ae)> 30 <(ef)(ab)(df)cb> 40 <eg(af)cbc> An element may contain a set of items. Items within an element are unordered and we list them alphabetically. <a(bc)dc> is a subsequence of <a(abc)(ac)d(cf)> Given support threshold min_sup =2, <(ab)c> is a sequential pattern 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 5

Challenges on Sequential Pattern Mining A huge number of possible sequential patterns are hidden in databases A mining algorithm should Find the complete set of patterns satisfying the minimum support (frequency) threshold Be highly efficient, scalable, involving only a small number of database scans Be able to incorporate various kinds of user-specific constraints 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 6
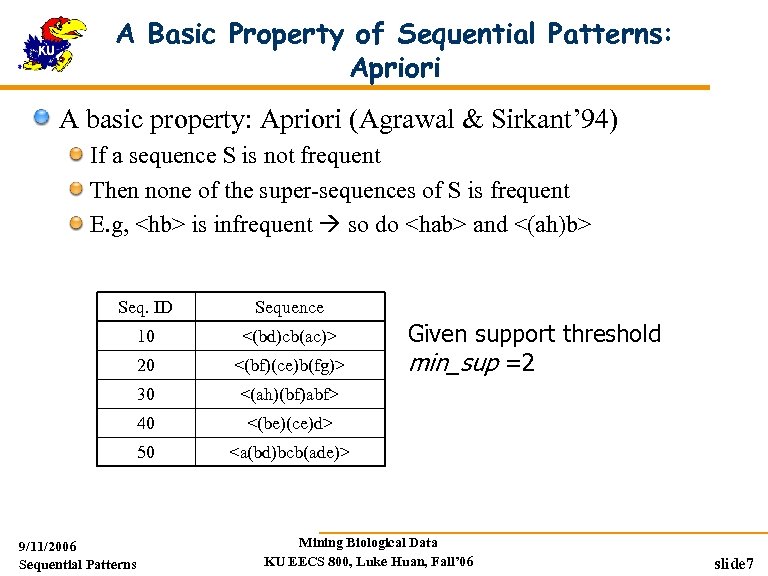
A Basic Property of Sequential Patterns: Apriori A basic property: Apriori (Agrawal & Sirkant’ 94) If a sequence S is not frequent Then none of the super-sequences of S is frequent E. g, <hb> is infrequent so do <hab> and <(ah)b> Seq. ID Sequence 10 <(bd)cb(ac)> 20 <(bf)(ce)b(fg)> 30 <(ah)(bf)abf> 40 <(be)(ce)d> 50 <a(bd)bcb(ade)> 9/11/2006 Sequential Patterns Given support threshold min_sup =2 Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 7
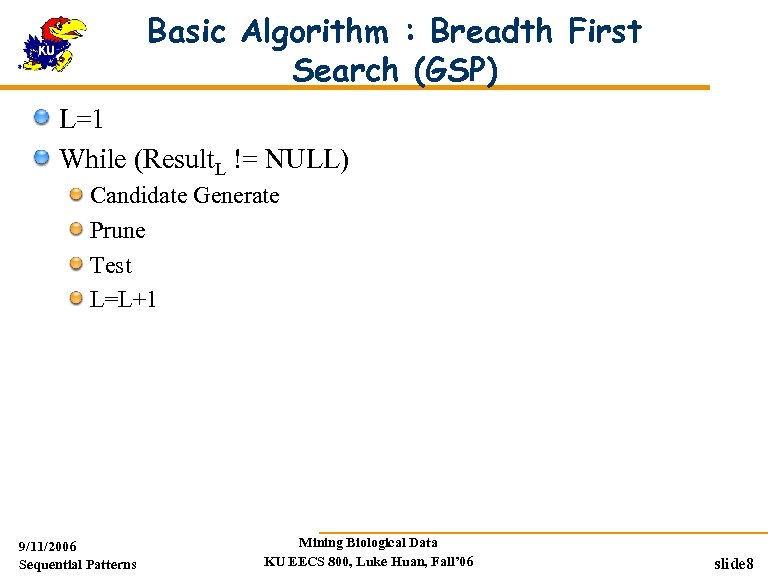

Basic Algorithm : Breadth First Search (GSP) L=1 While (Result. L != NULL) Candidate Generate Prune Test L=L+1 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 8
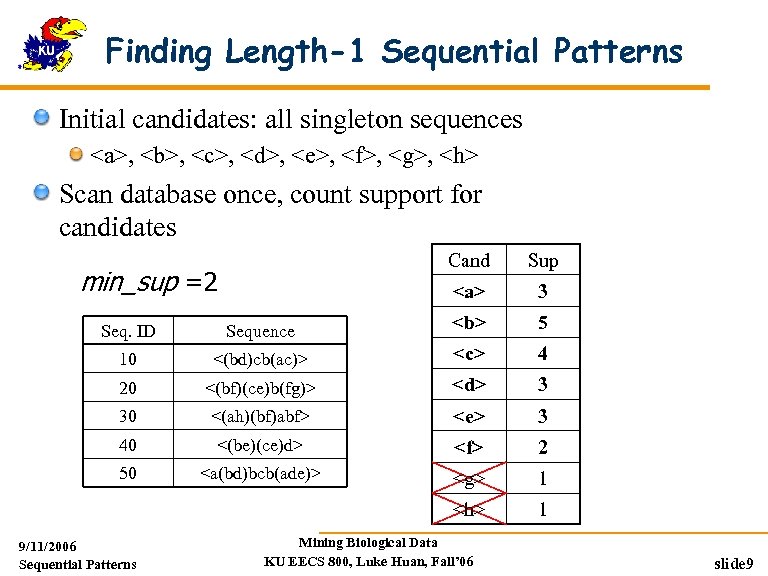

Finding Length-1 Sequential Patterns Initial candidates: all singleton sequences <a>, <b>, <c>, <d>, <e>, <f>, <g>, <h> Scan database once, count support for candidates Cand <a> min_sup =2 Sup 3 Seq. ID Sequence <b> 5 10 <(bd)cb(ac)> <c> 4 20 <(bf)(ce)b(fg)> <d> 3 30 <(ah)(bf)abf> <e> 3 40 <(be)(ce)d> <f> 2 50 <a(bd)bcb(ade)> <g> 1 <h> 1 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 9
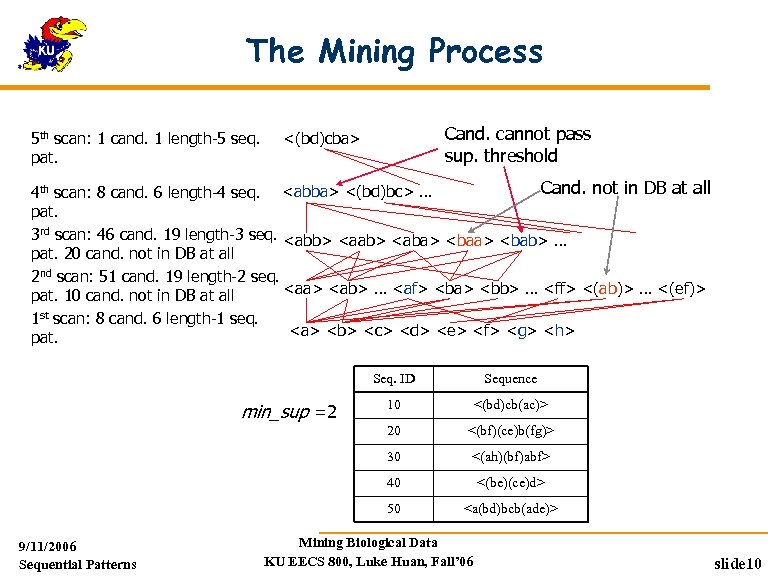
The Mining Process 5 th scan: 1 cand. 1 length-5 seq. pat. Cand. cannot pass sup. threshold <(bd)cba> Cand. not in DB at all 4 th scan: 8 cand. 6 length-4 seq. <abba> <(bd)bc> … pat. 3 rd scan: 46 cand. 19 length-3 seq. <abb> <aab> <aba> <bab> … pat. 20 cand. not in DB at all 2 nd scan: 51 cand. 19 length-2 seq. <aa> <ab> … <af> <ba> <bb> … <ff> <(ab)> … <(ef)> pat. 10 cand. not in DB at all 1 st scan: 8 cand. 6 length-1 seq. <a> <b> <c> <d> <e> <f> <g> <h> pat. Seq. ID <(bd)cb(ac)> <(bf)(ce)b(fg)> 30 <(ah)(bf)abf> 40 <(be)(ce)d> 50 9/11/2006 Sequential Patterns 10 20 min_sup =2 Sequence <a(bd)bcb(ade)> Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 10
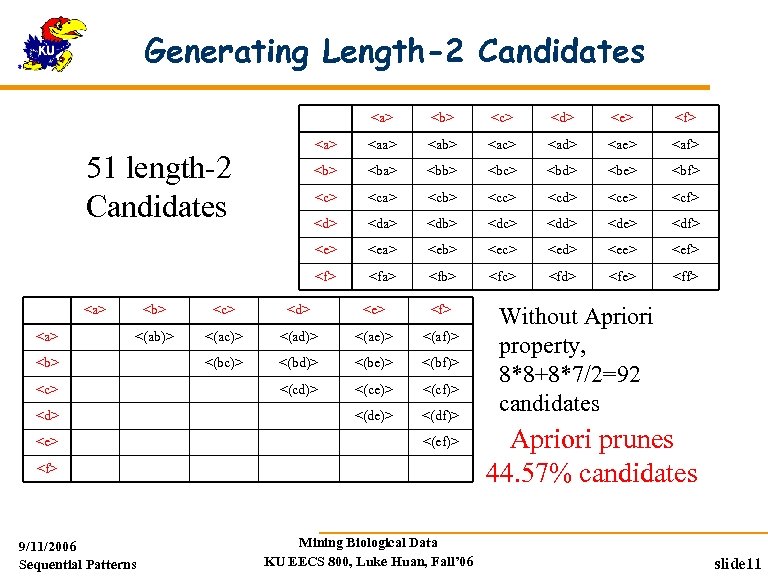
Generating Length-2 Candidates <a> <e> <f> <aa> <ab> <ac> <ad> <ae> <af> <ba> <bb> <bc> <bd> <be> <bf> <ca> <cb> <cc> <cd> <ce> <cf> <da> <db> <dc> <dd> <de> <df> <ea> <eb> <ec> <ed> <ee> <ef> <a> <d> <e> <a> <c> <a> 51 length-2 Candidates <b> <fa> <fb> <fc> <fd> <fe> <ff> <b> <c> <d> <e> <f> <(ab)> <(ac)> <(ad)> <(ae)> <(af)> <(bc)> <(bd)> <(be)> <(bf)> <(cd)> <(ce)> <(cf)> <(de)> <(df)> <b> <c> <d> <e> <(ef)> <f> 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 Without Apriori property, 8*8+8*7/2=92 candidates Apriori prunes 44. 57% candidates slide 11

The SPADE Algorithm SPADE (Sequential PAttern Discovery using Equivalent Class) developed by Zaki 2001 A vertical format sequential pattern mining method A sequence database is mapped to a large set of Item: <SID, EID> Sequential pattern mining is performed by growing the subsequences (patterns) one item at a time by Apriori candidate generation 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 12

The SPADE Algorithm 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 13

Bottlenecks of GSP and SPADE A large set of candidates could be generated 1, 000 frequent length-1 sequences generate s huge number of length-2 candidates! Multiple scans of database in mining Breadth-first search Mining long sequential patterns 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 14
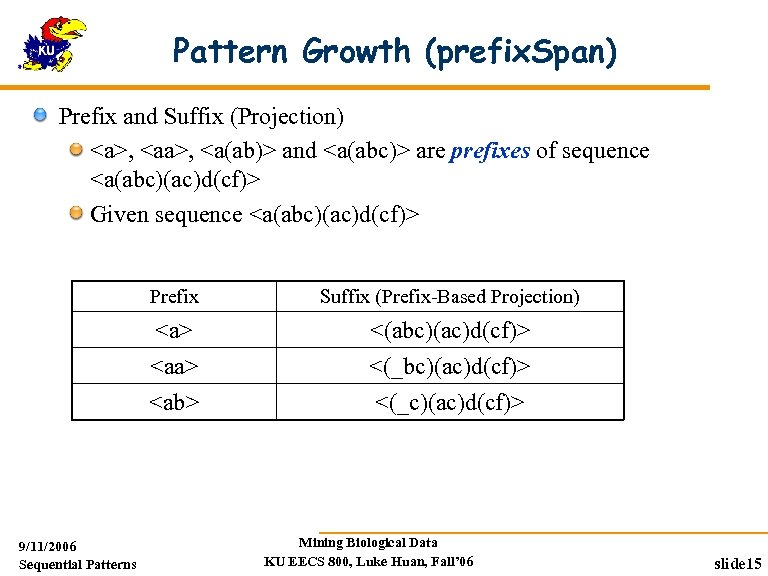
Pattern Growth (prefix. Span) Prefix and Suffix (Projection) <a>, <a(ab)> and <a(abc)> are prefixes of sequence <a(abc)(ac)d(cf)> Given sequence <a(abc)(ac)d(cf)> Prefix <a> <ab> 9/11/2006 Sequential Patterns Suffix (Prefix-Based Projection) <(abc)(ac)d(cf)> <(_c)(ac)d(cf)> Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 15
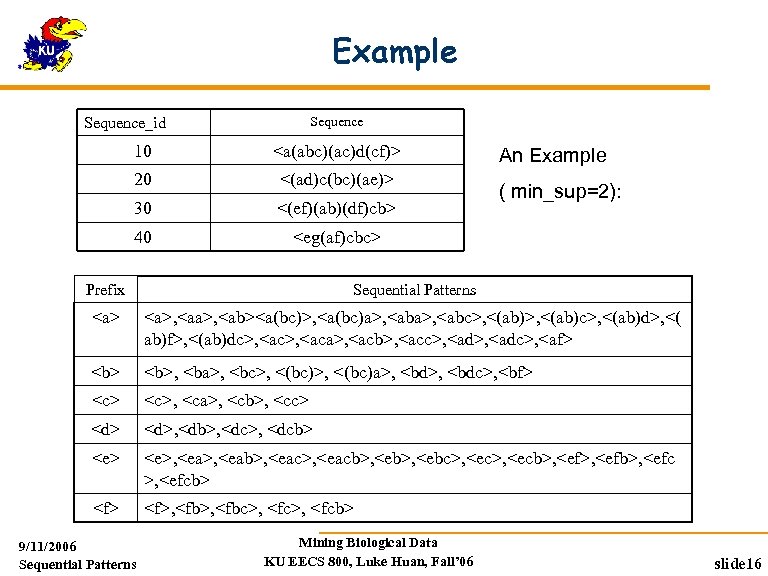

Example Sequence_id 10 <a(abc)(ac)d(cf)> 20 <(ad)c(bc)(ae)> 30 <(ef)(ab)(df)cb> 40 <eg(af)cbc> An Example ( min_sup=2): Prefix Sequential Patterns <a>, <aa>, <ab><a(bc)>, <a(bc)a>, <abc>, <(ab)c>, <(ab)d>, <( ab)f>, <(ab)dc>, <aca>, <acb>, <acc>, <adc>, <af> <b>, <ba>, <bc>, <(bc)a>, <bdc>, <bf> <c>, <ca>, <cb>, <cc> <d>, <db>, <dcb> <e>, <eab>, <eacb>, <ebc>, <ecb>, <efb>, <efcb> <f>, <fbc>, <fcb> 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 16
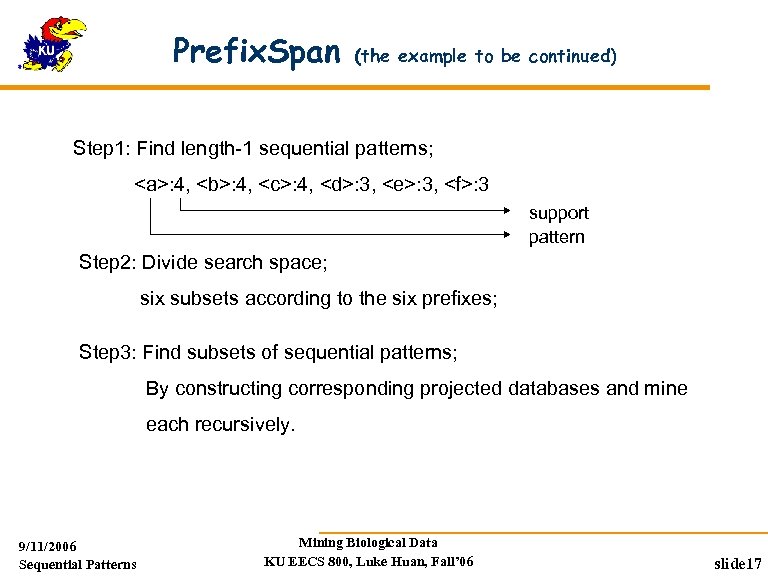

Prefix. Span (the example to be continued) Step 1: Find length-1 sequential patterns; <a>: 4, <b>: 4, <c>: 4, <d>: 3, <e>: 3, <f>: 3 support pattern Step 2: Divide search space; six subsets according to the six prefixes; Step 3: Find subsets of sequential patterns; By constructing corresponding projected databases and mine each recursively. 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 17

Example Find sequential patterns having prefix <a>: Scan sequence database S once. Sequences in S containing <a> are projected w. r. t <a> to form the <a>-projected database. Scan <a>-projected database once, get six length-2 sequential patterns having prefix <a> : <a>: 2 , <b>: 4, <(_b)>: 2, <c>: 4, <d>: 2, <f>: 2 <aa>: 2 , <ab>: 4, <(ab)>: 2, <ac>: 4, <ad>: 2, <af>: 2 Recursively, all sequential patterns having prefix <a> can be further partitioned into 6 subsets. Construct respective projected databases and mine each. e. g. <aa>-projected database has two sequences : <(_bc)(ac)d(cf)> and <(_e)>. 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 18
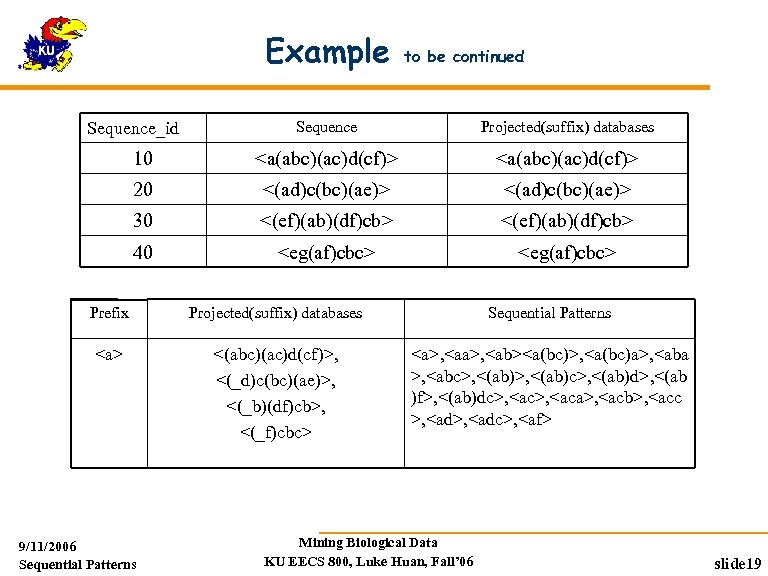
Example to be continued Sequence Projected(suffix) databases 10 <a(abc)(ac)d(cf)> 20 <(ad)c(bc)(ae)> 30 <(ef)(ab)(df)cb> 40 <eg(af)cbc> Sequence_id Prefix Projected(suffix) databases Sequential Patterns <a> <(abc)(ac)d(cf)>, <(_d)c(bc)(ae)>, <(_b)(df)cb>, <(_f)cbc> <a>, <ab><a(bc)>, <a(bc)a>, <aba >, <abc>, <(ab)c>, <(ab)d>, <(ab )f>, <(ab)dc>, <aca>, <acb>, <acc >, <adc>, <af> 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 19

Prefix. Span Algorithm Main Idea: Use frequent prefixes to divide the search space and to project sequence databases. only search the relevant sequences. Prefix. Span( , i, S| ) 1. Initially is a single frequent element in S 2. Scan S| once, find the set of frequent items b such that • b can be assembled to the last element of to form a sequential pattern; or • <b> can be appended to form a sequential pattern. 3. For each frequent item b, appended it to form a sequential pattern ’, and output ’; 4. For each ’, construct ’-projected database S| ’, and call Prefix. Span( ’, i+1, S| ’). 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 20
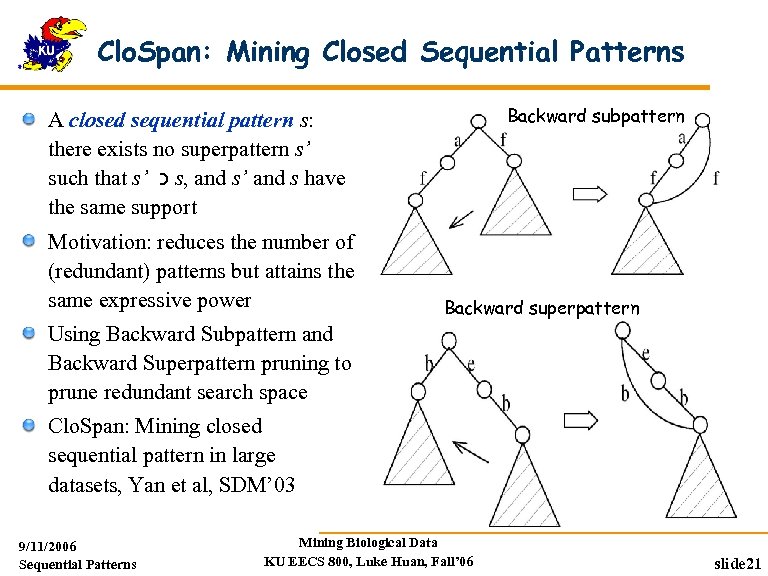
Clo. Span: Mining Closed Sequential Patterns Backward subpattern A closed sequential pattern s: there exists no superpattern s’ such that s’ כ s, and s’ and s have the same support Motivation: reduces the number of (redundant) patterns but attains the same expressive power Backward superpattern Using Backward Subpattern and Backward Superpattern pruning to prune redundant search space Clo. Span: Mining closed sequential pattern in large datasets, Yan et al, SDM’ 03 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 21
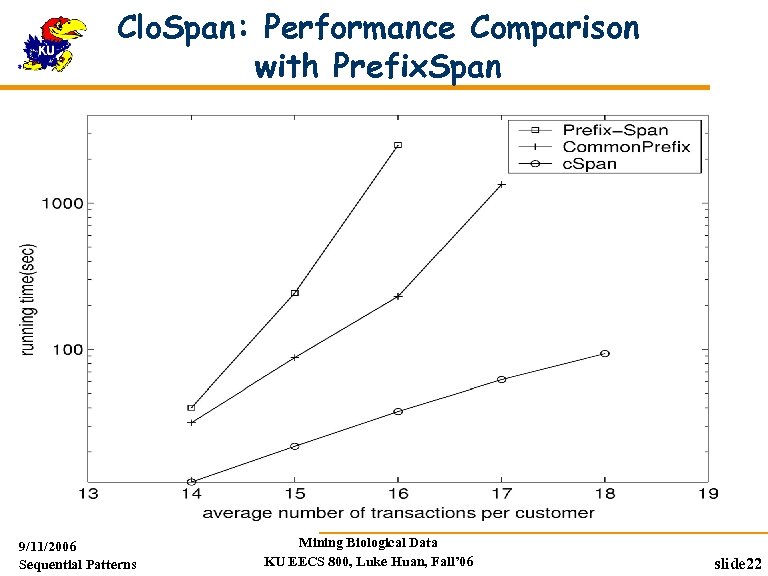
Clo. Span: Performance Comparison with Prefix. Span 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 22

Noise-tolerant Sequence Patterns There are noises in real-world sequences data Biological sequences Gene expression profiles Web-log collection Compatibility matrix is introduced to tolerate certain level of noise Yang et al. Mining Long Sequential Patterns in a Noisy Environment, SIGMOD’ 01 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 23
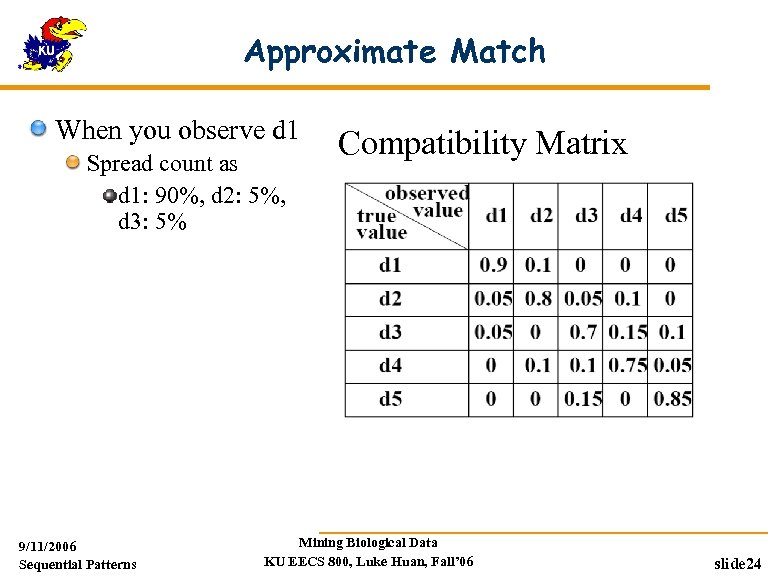
Approximate Match When you observe d 1 Spread count as d 1: 90%, d 2: 5%, d 3: 5% 9/11/2006 Sequential Patterns Compatibility Matrix Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 24
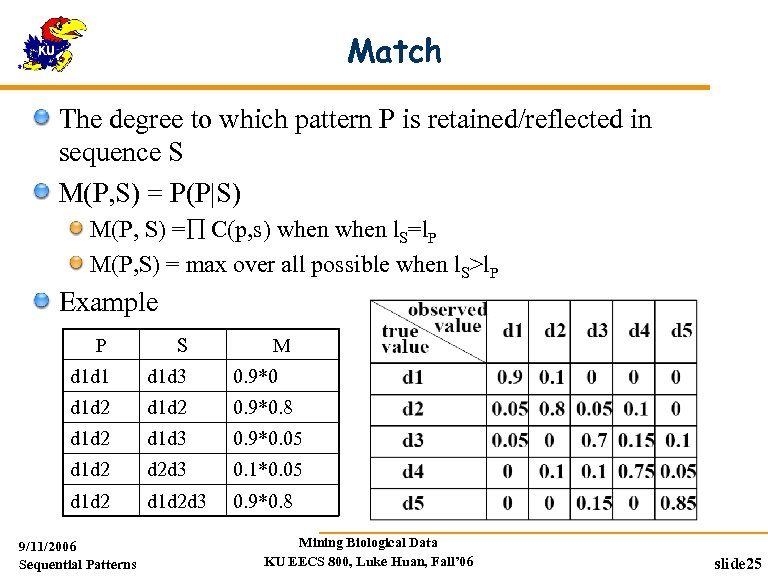
Match The degree to which pattern P is retained/reflected in sequence S M(P, S) = P(P|S) M(P, S) = C(p, s) when l. S=l. P M(P, S) = max over all possible when l. S>l. P Example P S d 1 d 1 d 1 d 3 0. 9*0 d 1 d 2 0. 9*0. 8 d 1 d 2 d 1 d 3 0. 9*0. 05 d 1 d 2 d 2 d 3 0. 1*0. 05 d 1 d 2 d 3 0. 9*0. 8 9/11/2006 Sequential Patterns M Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 25
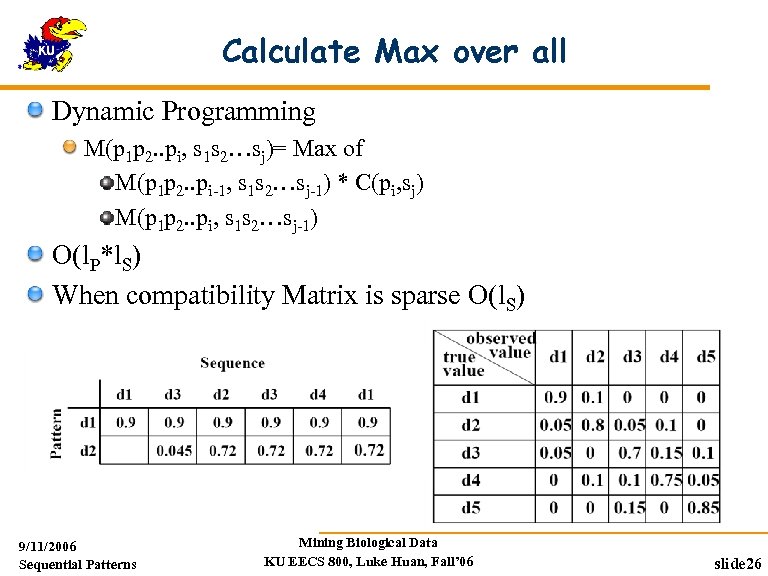

Calculate Max over all Dynamic Programming M(p 1 p 2. . pi, s 1 s 2…sj)= Max of M(p 1 p 2. . pi-1, s 1 s 2…sj-1) * C(pi, sj) M(p 1 p 2. . pi, s 1 s 2…sj-1) O(l. P*l. S) When compatibility Matrix is sparse O(l. S) 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 26
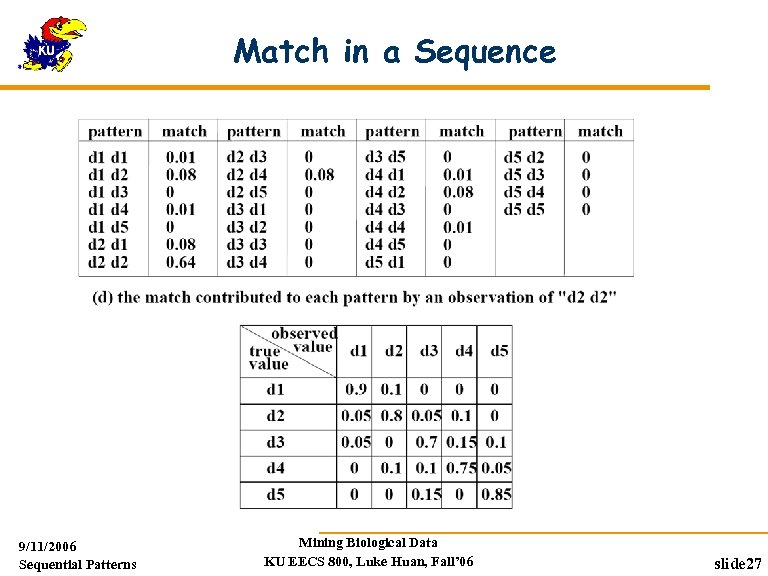
Match in a Sequence 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 27
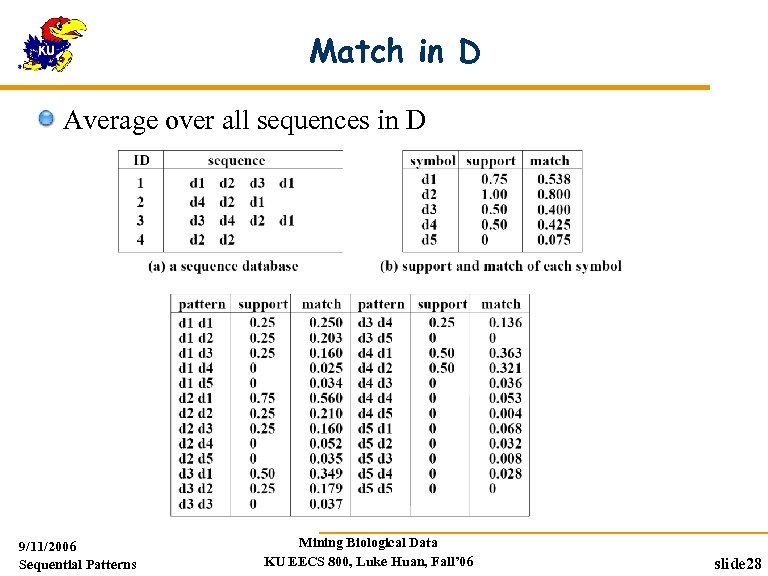
Match in D Average over all sequences in D 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 28

Anti-Monotone If compatibility matrix is identity matrix, match = support Theorem: the match of a pattern P in a symbol sequence S is less than or equal to the match of any subpattern of P in S Corollary: the match of a pattern P in a sequence database D is less than or equal to the match of any subpattern of P in D Can use any support based algorithm More patterns match so require efficient solution Sample based algorithms Border collapsing of ambiguous patterns 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 29

Chernoff Bound Given sample size=n, sample mean = μ, and we know that the range of the data is R, then we have: population mean is μ = sqrt([R 2 ln(1/ )]/2 n) with probability 1 - (almost certain) Can the estimation be replaced by normal due to the law of large number? Distribution free More conservative Sample size: fit in memory Restricted spread : Frequent Patterns min_match + For pattern P= p 1 p 2. . p. L R=min (match[pi]) for all 1 i L min_match - Infrequent patterns 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 30

Algorithm Scan DB: O(N*Ls*m) Find the match of each individual symbol Take a random sample of sequences N, # of sequence, Ls, average sequence length, m: # of symbols Identify borders that embrace the set of ambiguous patterns O(m. Lp * |S| * Lp * n) Min_match existing methods for association rule mining Lp is the length of the largest patter, S, average length in sample sequence, n # of samples Locate the border of frequent patterns via border collapsing 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 31

Border Collapsing If memory can not hold the counters for all ambiguous counters Probe-and-collapse : binary search Probe patterns with highest collapsing power until memory is filled If memory can hold all patterns up to the 1/x layer the space of of ambiguous patterns can be narrowed to at least 1/x of the original one where x is a power of 2 If it takes a level-wise search y scans of the DB, only O(logxy) scans are necessary when the border collapsing technique is employed 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 32
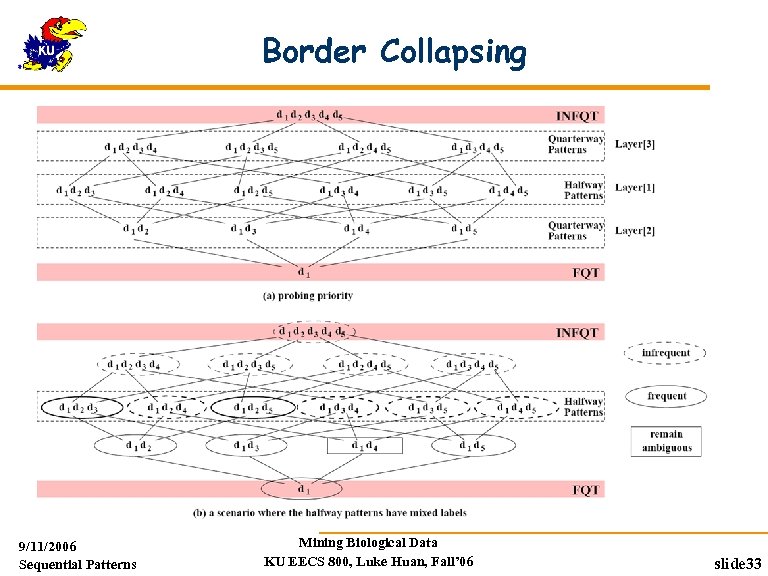
Border Collapsing 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 33
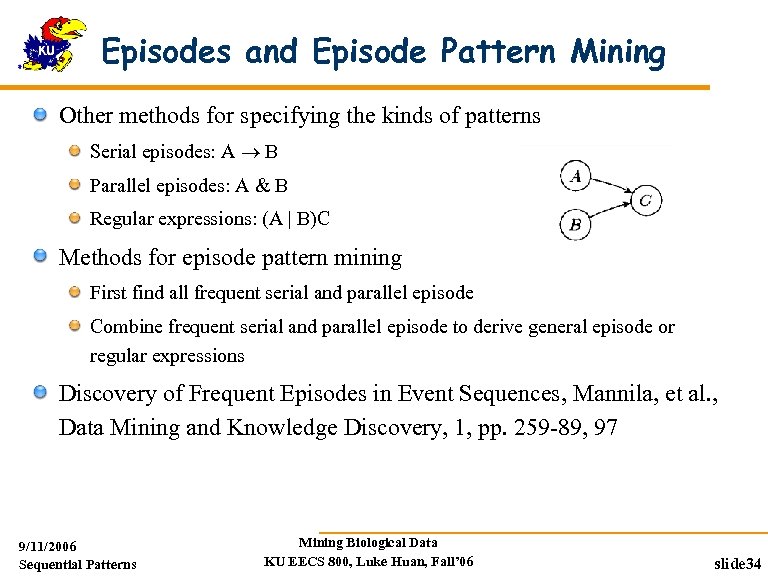

Episodes and Episode Pattern Mining Other methods for specifying the kinds of patterns Serial episodes: A B Parallel episodes: A & B Regular expressions: (A | B)C Methods for episode pattern mining First find all frequent serial and parallel episode Combine frequent serial and parallel episode to derive general episode or regular expressions Discovery of Frequent Episodes in Event Sequences, Mannila, et al. , Data Mining and Knowledge Discovery, 1, pp. 259 -89, 97 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 34

Periodicity Analysis Periodicity is everywhere: tides, seasons, daily power consumption, etc. Full periodicity Every point in time contributes (precisely or approximately) to the periodicity Partial periodicit: A more general notion Only some segments contribute to the periodicity Jim reads NY Times 7: 00 -7: 30 am every week day Cyclic association rules Associations which form cycles Methods Full periodicity: FFT, other statistical analysis methods Partial and cyclic periodicity: Variations of Apriori-like mining methods 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 35

Periodic Pattern Full periodic pattern ABC ABC Partial periodic pattern ABC ADC ACC ABC Pattern hierarchy ABC ABC ABC DE DE ABC ABC DE DE 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 36

Periodic Pattern Recent Achievements Partial Periodic Pattern Asynchronous Periodic Pattern Meta Pattern Info. Miner/Info. Miner+/STAMP 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 37

Constraint-Based Seq. Pattern Mining Constraint-based sequential pattern mining Constraints: User-specified, for focused mining of desired patterns How to explore efficient mining with constraints? — Optimization Classification of constraints Anti-monotone: E. g. , value_sum(S) < 150, min(S) > 10 Monotone: E. g. , count (S) > 5, S {PC, digital_camera} Succinct: E. g. , length(S) 10, S {Pentium, MS/Office, MS/Money} Convertible: E. g. , value_avg(S) < 25, profit_sum (S) > 160, max(S)/avg(S) < 2, median(S) – min(S) > 5 Inconvertible: E. g. , avg(S) – median(S) = 0 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 38

From Sequential Patterns to Structured Patterns Sets, sequences, trees, graphs, and other structures Transaction DB: Sets of items {{i 1, i 2, …, im}, …} Sets of Sequences: {{<i 1, i 2>, …, <im, in, ik>}, …} Sets of trees: {t 1, t 2, …, tn} Sets of graphs (mining for frequent subgraphs): {g 1, g 2, …, gn} Mining structured patterns in XML documents, bio-molecule structures, etc. 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 39

References: Sequential Pattern Mining Methods R. Agrawal and R. Srikant. Mining sequential patterns. ICDE'95, 3 -14, Taipei, Taiwan. R. Srikant and R. Agrawal. Mining sequential patterns: Generalizations and performance improvements. EDBT’ 96. J. Han, J. Pei, B. Mortazavi-Asl, Q. Chen, U. Dayal, M. -C. Hsu, "Free. Span: Frequent Pattern-Projected Sequential Pattern Mining", Proc. 2000 Int. Conf. on Knowledge Discovery and Data Mining (KDD'00), Boston, MA, August 2000. H. Mannila, H Toivonen, and A. I. Verkamo. Discovery of frequent episodes in event sequences. Data Mining and Knowledge Discovery, 1: 259 -289, 1997. J. Pei, J. Han, H. Pinto, Q. Chen, U. Dayal, and M. -C. Hsu, "Prefix. Span: Mining Sequential Patterns Efficiently by Prefix-Projected Pattern Growth", Proc. 2001 Int. Conf. on Data Engineering (ICDE'01), Heidelberg, Germany, April 2001. 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 40

References: Sequential Pattern Mining Methods B. Ozden, S. Ramaswamy, and A. Silberschatz. Cyclic association rules. ICDE'98, 412 -421, Orlando, FL. S. Ramaswamy, S. Mahajan, and A. Silberschatz. On the discovery of interesting patterns in association rules. VLDB'98, 368 -379, New York, NY. M. J. Zaki. Efficient enumeration of frequent sequences. CIKM’ 98. Novermber 1998. M. N. Garofalakis, R. Rastogi, K. Shim: SPIRIT: Sequential Pattern Mining with Regular Expression Constraints. VLDB 1999: 223 -234, Edinburgh, Scotland. Wei Wang, Jiong Yang, Philip S. Yu: Mining Patterns in Long Sequential Data with Noise. SIGKDD Explorations 2(2): 28 -33 (2000) Jiong Yang, Wei Wang, Philip S. Yu, Jiawei Han: Mining Long Sequential Patterns in a Noisy Environment. SIGMOD Conference 2002 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 41

References: Periodic Pattern Mining Methods Jiawei Han, Wan Gong, Yiwen Yin: Mining Segment-Wise Periodic Patterns in Time-Related Databases. KDD 1998: 214 -218 Jiawei Han, Guozhu Dong, Yiwen Yin: Efficient Mining of Partial Periodic Patterns in Time Series Database. ICDE 1999: 106 -115 Jiong Yang, Wei Wang, Philip S. Yu: Mining asynchronous periodic patterns in time series data. KDD 2000: 275 -279 Wei Wang, Jiong Yang, Philip S. Yu: Meta-patterns: Revealing Hidden Periodic Patterns. ICDM 2001: 550 -557 Jiong Yang, Wei Wang, Philip S. Yu: Infominer: mining surprising periodic patterns. KDD 2001: 395 -400 9/11/2006 Sequential Patterns Mining Biological Data KU EECS 800, Luke Huan, Fall’ 06 slide 42
03c5b3d2d10c9acce839175a5e7691a4.ppt