022864c07b2c663373ae3958f0d120fb.ppt
- Количество слайдов: 73

DNA Computing Tutorial Biointelligence Lab. School of Compute Science and Engineering, Seoul National University 2002. 3. 13 (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 1

Outline Molecular Intelligence l DNA computing operators l Introduction of DNA computing l Examples l ¨ The first DNA computing approach ¨ Molecular theorem prover The difficulties of DNA computer l DNA computing applications l Conclusion l (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 2

Molecular Intelligence Solving artificial intelligence problems using molecular computing technology l Computing model: evolutionary computing l Computing devices: biomolecules (DNA) l Learning and inference using biomolecular computing l l l Biochemical reaction Massively parallel Micro or nanoscale Storage capacity Bio-compatible Molecular Computing Artificial Intelligence Molecular Intelligence 3
Biocomputing vs. Bioinformatics IT BT Biocomputing 4

What is DNA Computing? The use of biological molecules, primarily DNA, DNA analogs, and RNA, for computation purpose. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 5

DNA Computing 01101010001 ATGCTCGAAGCT 6

A different approach l What DNA computing is: ¨ a completely new method among a few others (e. g. , quantum computing) of general computation alternative to electronic/semiconductor technology ¨ uses biochemical processes based on DNA l What DNA computing isn’t: ¨ not to confuse with bio-computing which applies biological laws (evolution, selection) to computer algorithm design. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 7

Why try new stuff ? Will need a dramatically new technology to overcome some CMOS limitations and offer new opportunity. l Certain types of problems (learning, pattern recognition, fault-tolerant system, large set search algorithms) are intrinsically very difficult to solve even with fast evolution of CMOS. l Hope of achieving massive parallelism. l (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 8

Why try DNA Computing? 6. 022 1023 molecules / mole l Massively Parallel Search of All Possibilities l ¨ Desktop: 109 operations / sec ¨ Supercomputer: 1012 operations / sec ¨ 1 mmol of DNA: 1026 reactions l Favorable Energetics: Gibb’s Free Energy ¨ 1 J for 2 1019 operations l Storage Capacity: 1 bit per cubic nanometer (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 9

Why try DNA Computing? l The fastest supercomputer vs. DNA computer ¨ 106 op/sec vs. 1014 op/sec ¨ 109 op/J vs. 1019 op/J (in ligation step) ¨ 1 bit per 1012 nm 3 vs. 1 bit per 1 nm 3 (video tape vs. molecules) (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 10
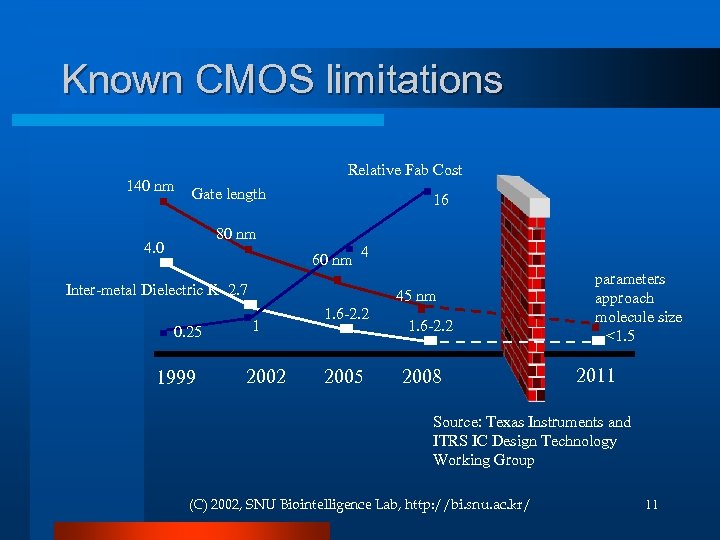
Known CMOS limitations 140 nm Relative Fab Cost Gate length 16 80 nm 4. 0 60 nm 4 Inter-metal Dielectric K 2. 7 0. 25 1999 45 nm 1 2002 1. 6 -2. 2 2005 1. 6 -2. 2 2008 parameters approach molecule size <1. 5 2011 Source: Texas Instruments and ITRS IC Design Technology Working Group (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 11
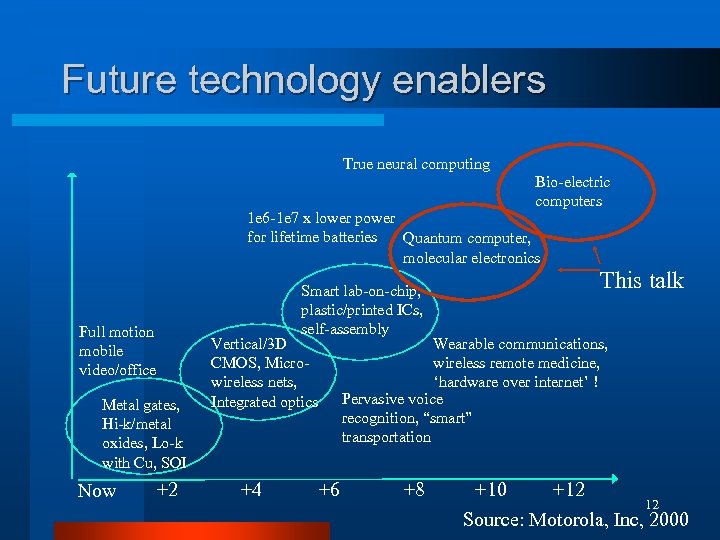

Future technology enablers True neural computing Bio-electric computers 1 e 6 -1 e 7 x lower power for lifetime batteries Quantum computer, molecular electronics Full motion mobile video/office Metal gates, Hi-k/metal oxides, Lo-k with Cu, SOI Now +2 Smart lab-on-chip, plastic/printed ICs, self-assembly Vertical/3 D CMOS, Microwireless nets, Integrated optics +4 This talk Wearable communications, wireless remote medicine, ‘hardware over internet’ ! Pervasive voice recognition, “smart” transportation +6 +8 +10 +12 12 Source: Motorola, Inc, 2000

Historical timeline Research 1950’s … 1994 R. Feynman’s paper on sub microscopic computers 1995 2000 2005 L. Adleman solves D. Boneh paper Lucent DNA computer Hamiltonian path on breaking DES builds DNA architecture ? problem using DNA. with DNA “motor” Field started Commercial 1970’s … DNA used in bio application 1996 Affymetrix sells Gene. Chip DNA analyzer 2000 Human Genome Sequenced (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 2015 Commercial computer ? 13

What’s up to date l l l Research still in very early stages but promise is big First groundbreaking work by Adleman at USC only 6 years ago No commercialization in sight within ~10 years Main work on settling for a logic abstraction model, ways of using DNA to compute Research sponsored by universities (Princeton, MIT, USC, Rutgers, etc) and NEC, Lucent Bell Labs, Telcordia, IBM One way or another, DNA computing will have a significant impact (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 14
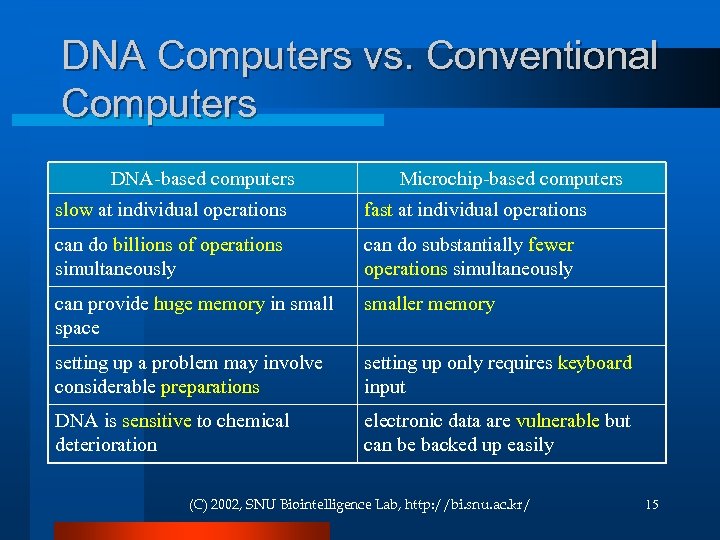

DNA Computers vs. Conventional Computers DNA-based computers Microchip-based computers slow at individual operations fast at individual operations can do billions of operations simultaneously can do substantially fewer operations simultaneously can provide huge memory in small space smaller memory setting up a problem may involve considerable preparations setting up only requires keyboard input DNA is sensitive to chemical deterioration electronic data are vulnerable but can be backed up easily (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 15

DNA Computing takes advantage of Our ability to produce massive numbers of DNA molecules with specific properties (size, sequence) l The natural proclivity of specific DNA molecules to chemically interact according to defined rules to produce new molecules l Laboratory techniques that allow the isolation/identification of product molecules with specific properties l ¨ PCR, Ligation, Gel Electrophoresis, etc. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 16

DNA Computing Operator (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 17
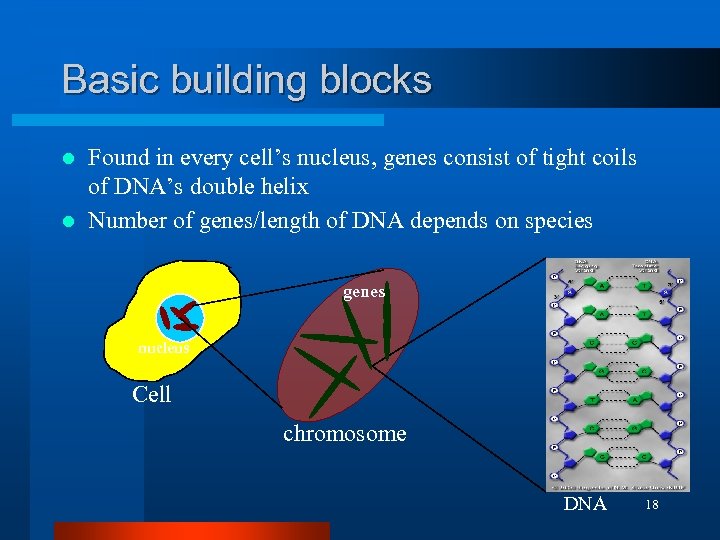
Basic building blocks Found in every cell’s nucleus, genes consist of tight coils of DNA’s double helix l Number of genes/length of DNA depends on species l genes nucleus Cell chromosome DNA 18
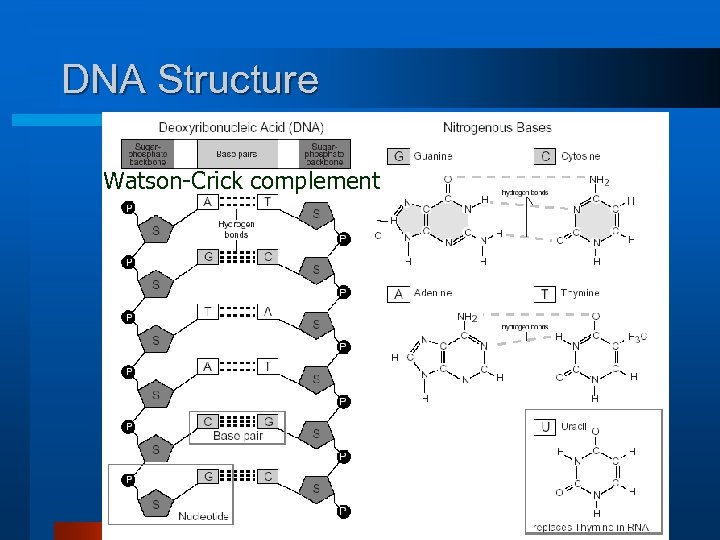
DNA Structure Watson-Crick complement 19
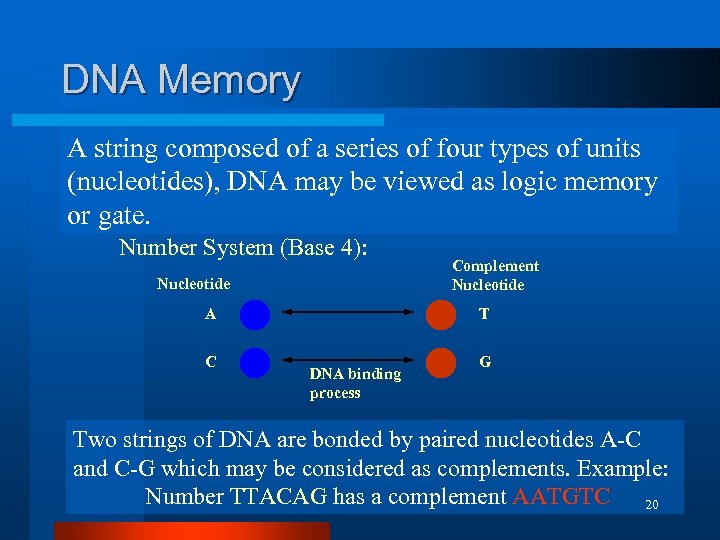

DNA Memory A string composed of a series of four types of units (nucleotides), DNA may be viewed as logic memory or gate. Number System (Base 4): Nucleotide A C Complement Nucleotide T DNA binding process G Two strings of DNA are bonded by paired nucleotides A-C and C-G which may be considered as complements. Example: Number TTACAG has a complement AATGTC 20
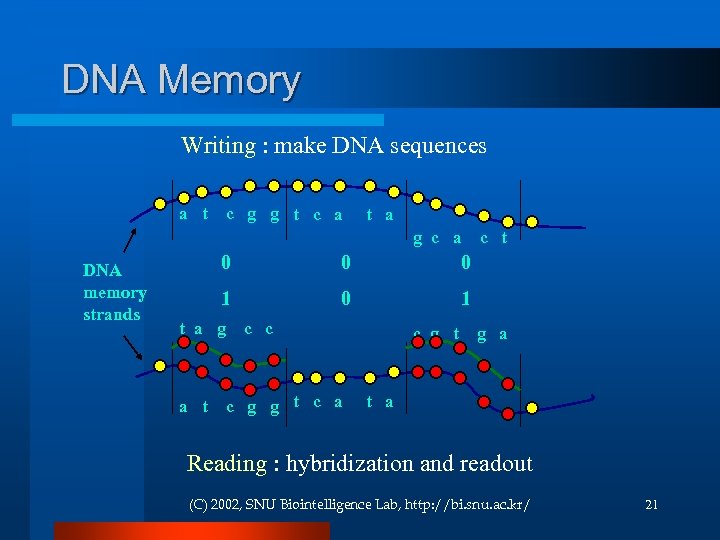
DNA Memory Writing : make DNA sequences a t c g g t c a t a g c a DNA memory strands 0 0 0 1 t a g a t c c c g g t c a c g t g a t a Reading : hybridization and readout (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 21

DNA Operators l The bio-lab technology. l Hybridization Ligation Polymerase Chain Reaction (PCR) Gel electrophoresis Affinity separation (Bead) Enzymes: restriction enzyme… l l l (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 22

Hybridization & Ligation l Hybridization ¨ base-pairing between two complementary single-strand molecules to form a double stranded DNA molecule l Ligation ¨ Joining DNA molecules together l Solution generation step! (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 23
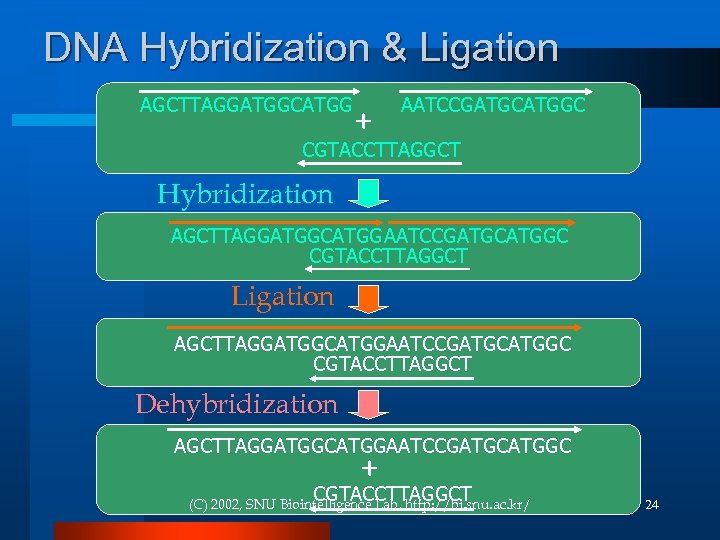
DNA Hybridization & Ligation AGCTTAGGATGGCATGG + AATCCGATGCATGGC CGTACCTTAGGCT Hybridization AGCTTAGGATGGCATGGAATCCGATGCATGGC CGTACCTTAGGCT Ligation AGCTTAGGATGGCATGGAATCCGATGCATGGC CGTACCTTAGGCT Dehybridization AGCTTAGGATGGCATGGAATCCGATGCATGGC + CGTACCTTAGGCT (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 24

PCR Polymerase Chain Reaction l Amplifies (produces identical copies of) selected ds. DNA molecules. l Make 2 n copies (n : number of iteration) l l Solution filtering or amplification step! (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 25
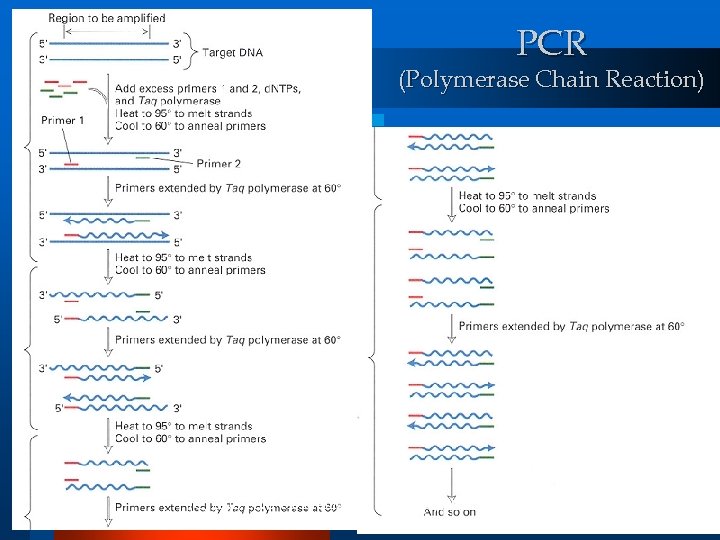
PCR (Polymerase Chain Reaction) (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 26

Gel electrophoresis l Molecular size fraction technique Bead Separation l Detect the specific DNA Solution detection or filtering step! (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 27
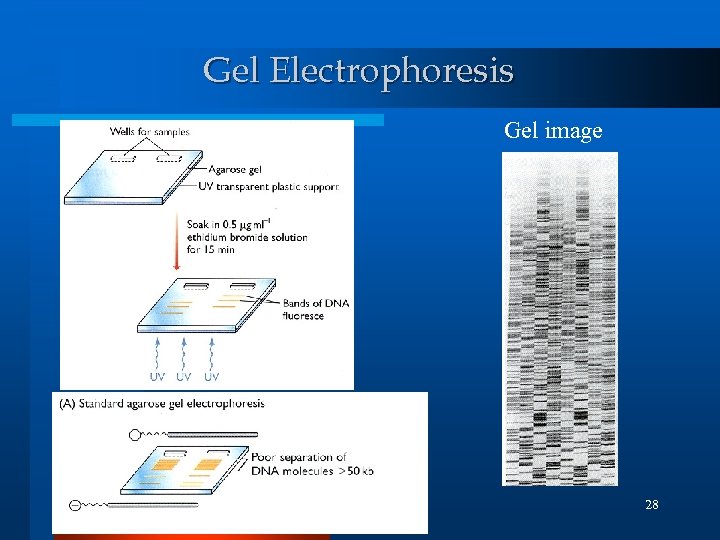
Gel Electrophoresis Gel image 28
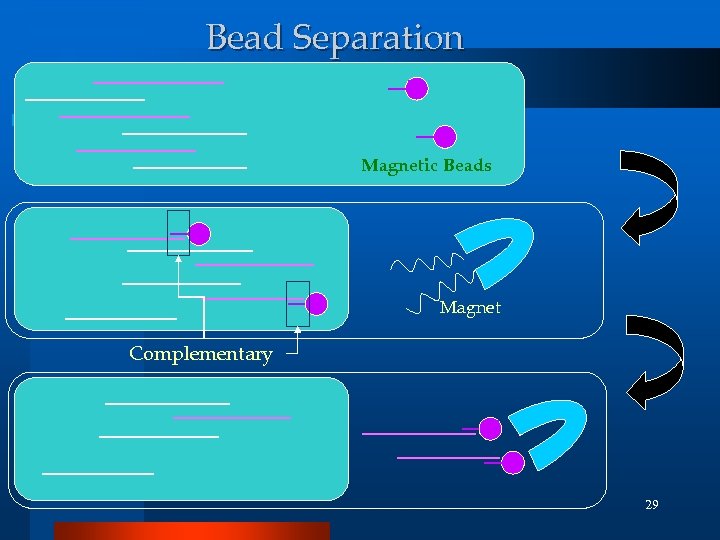
Bead Separation Magnetic Beads Magnet Complementary 29
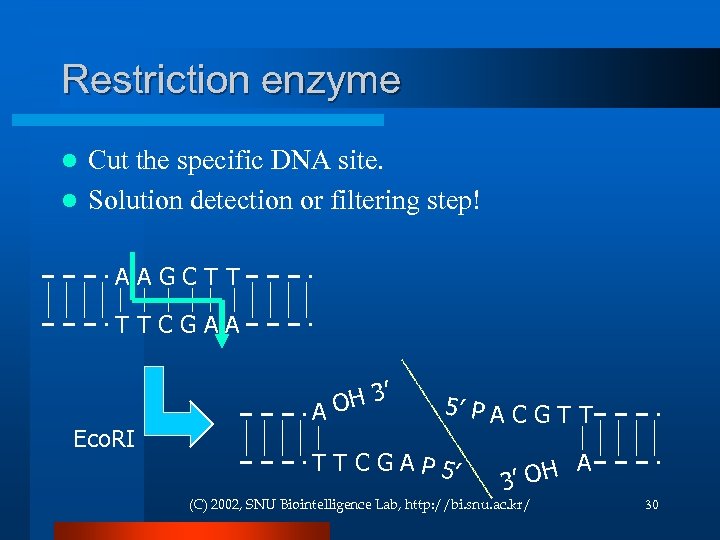
Restriction enzyme Cut the specific DNA site. l Solution detection or filtering step! l AAGCTT TTCGAA Eco. RI H 3’ AO 5’ P A C G T T C G A P 5’ H A 3’ O (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 30

The First DNA Computing Method L. M. Adleman, Molecular Computation of Solutions to Combinatorial Problems, Science, 266: 1021 -1024, 1994 (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 31

First special DNA computer Special problem: given N points, find a path visiting each and every point only once, and starting and ending at a given locations. (Hamiltonian path problem) l Solved with a DNA computer by Leonard Adleman in 1994 for N=6 l Basic approach: code each point as an 8 unit DNA string, code each possible path, allow DNA bonding, suppress DNA with incorrect start/end points. l (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 32
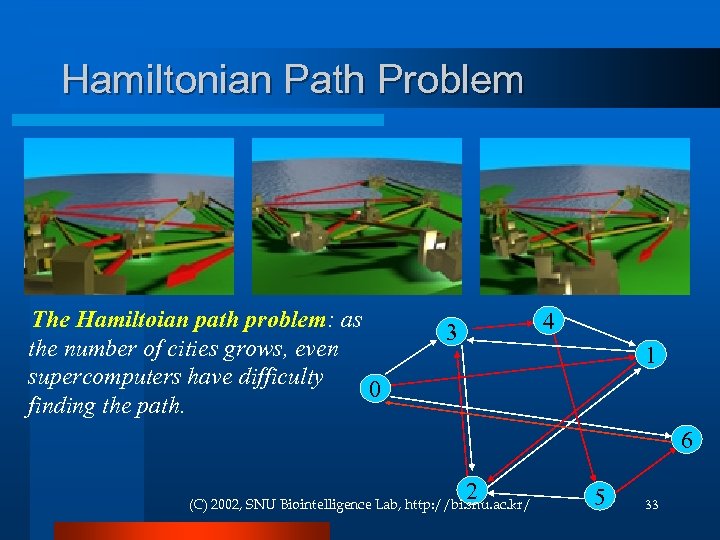
Hamiltonian Path Problem The Hamiltoian path problem: as the number of cities grows, even supercomputers have difficulty 0 finding the path. 4 3 1 6 2 (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 5 33
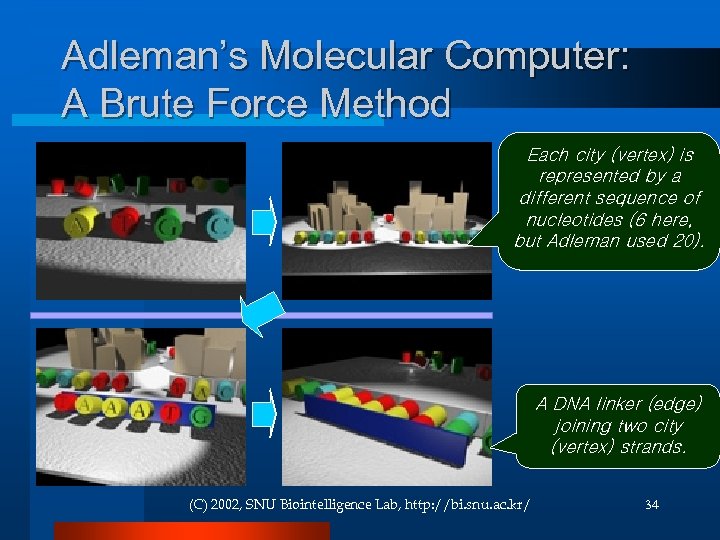
Adleman’s Molecular Computer: A Brute Force Method Each city (vertex) is represented by a different sequence of nucleotides (6 here, but Adleman used 20). A DNA linker (edge) joining two city (vertex) strands. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 34
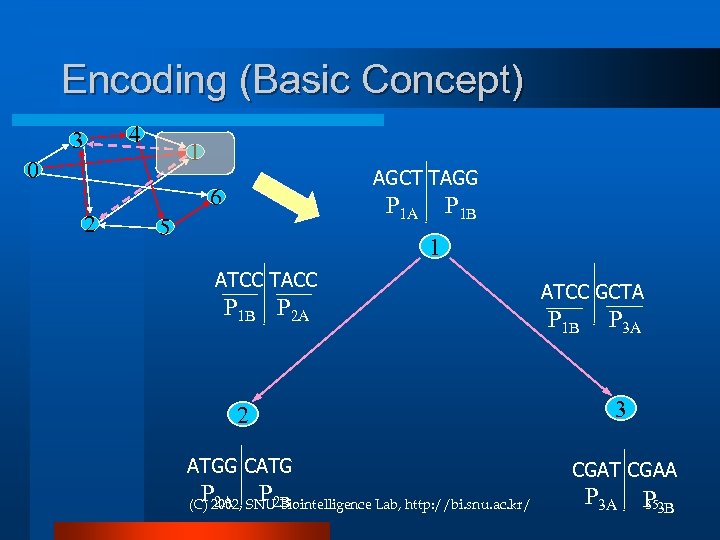
Encoding (Basic Concept) 4 3 1 0 AGCT TAGG 6 2 P 1 A 5 P 1 B 1 ATCC TACC P 1 B P 2 A ATCC GCTA P 1 B P 3 A 2 3 ATGG CATG CGAT CGAA P P 2 A 2 B (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ P 3 A P 3 B 35
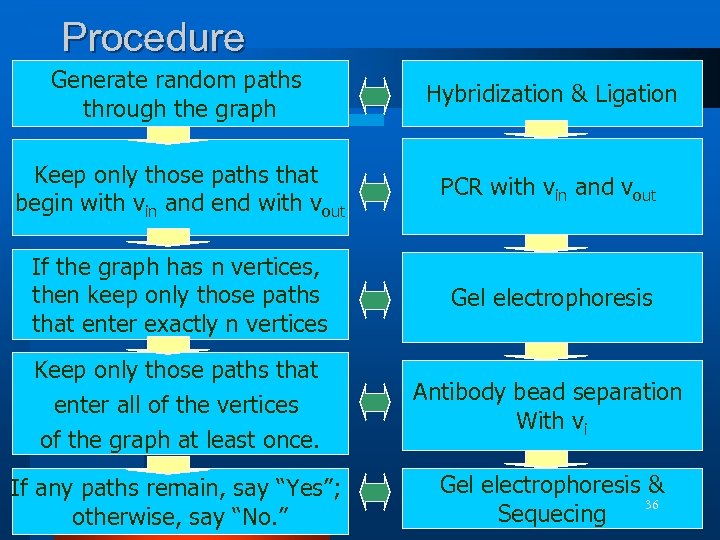

Procedure Generate random paths through the graph Hybridization & Ligation Keep only those paths that begin with vin and end with vout PCR with vin and vout If the graph has n vertices, then keep only those paths that enter exactly n vertices Gel electrophoresis Keep only those paths that enter all of the vertices of the graph at least once. Antibody bead separation With vi If any paths remain, say “Yes”; otherwise, say “No. ” Gel electrophoresis & 36 Sequecing
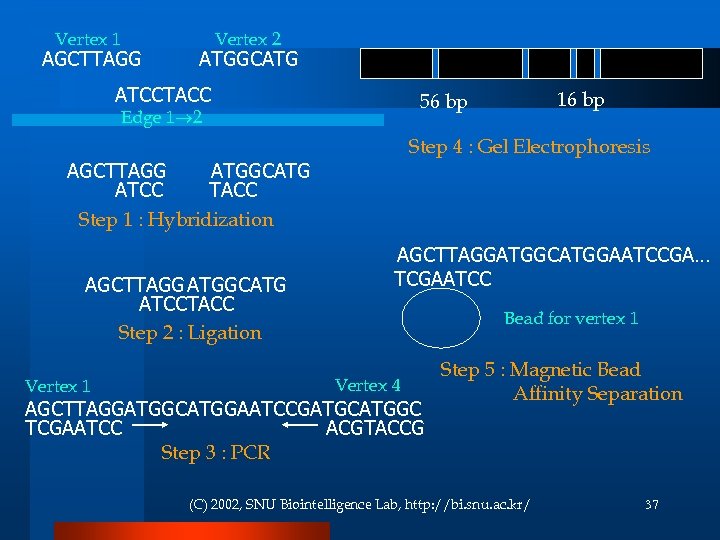
Vertex 1 AGCTTAGG Vertex 2 ATGGCATG ATCCTACC Edge 1 2 Step 4 : Gel Electrophoresis AGCTTAGG ATGGCATG ATCC TACC Step 1 : Hybridization AGCTTAGG ATGGCATG ATCCTACC Step 2 : Ligation Vertex 1 16 bp 56 bp AGCTTAGGATGGCATGGAATCCGA… TCGAATCC Bead for vertex 1 Vertex 4 AGCTTAGGATGGCATGGAATCCGATGCATGGC TCGAATCC ACGTACCG Step 3 : PCR Step 5 : Magnetic Bead Affinity Separation (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 37
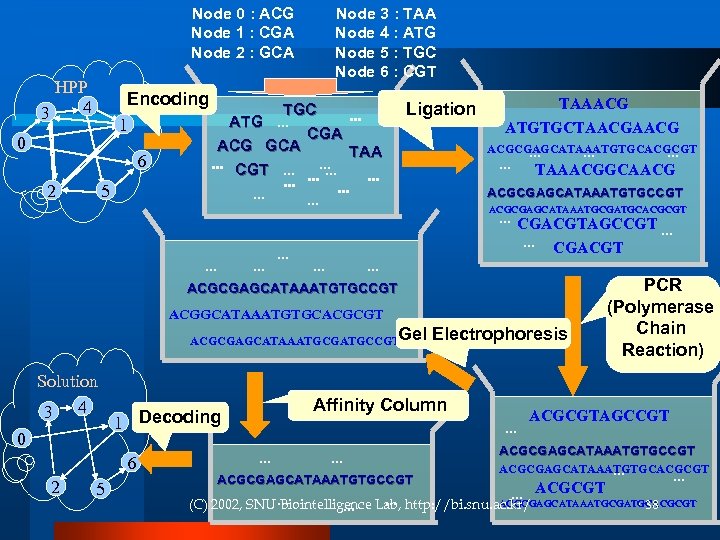
Node 0 : ACG Node 1 : CGA Node 2 : GCA HPP 4 3 Encoding 1 0 6 2 5 Node 3 : TAA Node 4 : ATG Node 5 : TGC Node 6 : CGT TGC. . . ATG. . . CGA ACG GCA TAA. . . CGT. . . . . Ligation TAAACG ATGTGCTAACG ACGCGAGCATAAATGTGCACGCGT. . . TAAACGGCAACG ACGCGAGCATAAATGTGCCGT ACGCGAGCATAAATGCGATGCACGCGT . . . CGACGTAGCCGT. . . CGACGT . . . ACGCGAGCATAAATGTGCCGT ACGGCATAAATGTGCACGCGT ACGCGAGCATAAATGCGATGCCGTGel Solution 3 4 0 2 5 Affinity Column 1 Decoding 6 Electrophoresis . . ACGCGAGCATAAATGTGCCGT PCR (Polymerase Chain Reaction) ACGCGTAGCCGT ACGCGAGCATAAATGTGCACGCGT. . . ACGCGT . . ACGCGAGCATAAATGCGATGCACGCGT (C) 2002, SNU. . . Biointelligence Lab, http: //bi. snu. ac. kr/ 38. . .

DNA finds a solution! A Hamiltonian path with all vertices included is isolated and recovered (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 39

Another Practical Example Molecular theorem prover (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 40
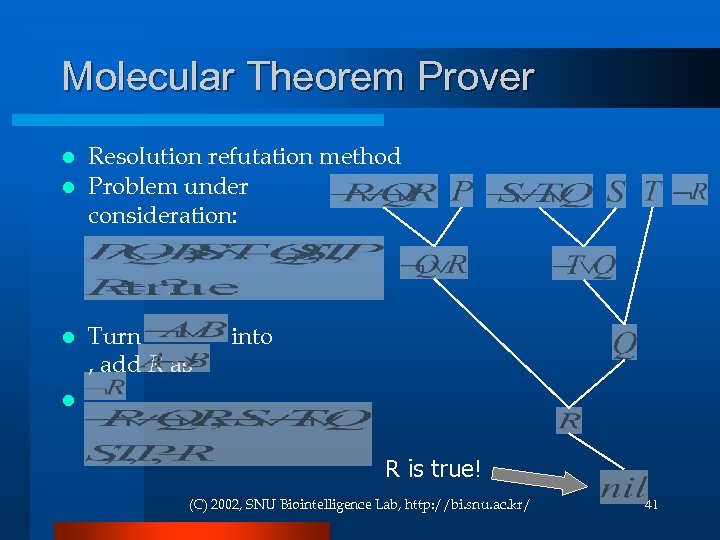
Molecular Theorem Prover Resolution refutation method l Problem under consideration: l l Turn , add R as into l R is true! (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 41
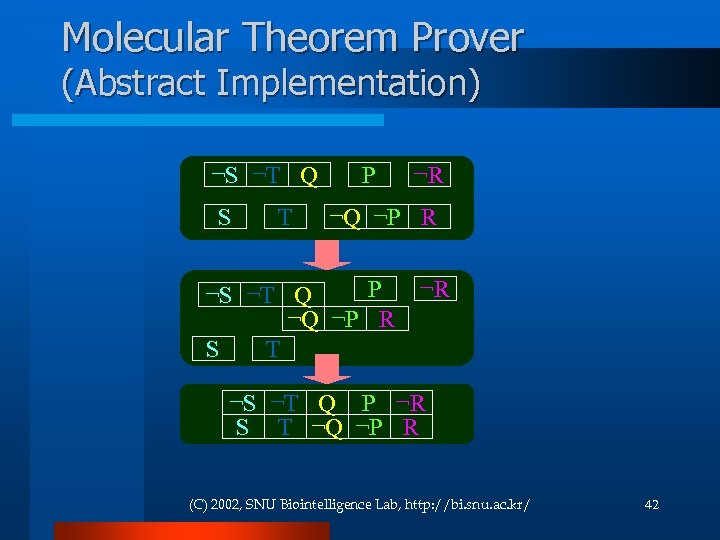

Molecular Theorem Prover (Abstract Implementation) ¬S ¬T Q S T P ¬R ¬Q ¬P R P ¬S ¬T Q ¬Q ¬P R S T ¬R ¬S ¬T Q P ¬R S T ¬Q ¬P R (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 42

Experimental Procedure Step 1. DNA Sequence Design Step 2. DNA Synthesis Step 3. Chemical Reaction in a Test Tube ¨ Hybridization ¨ Ligation Step 4. Detection of Solutions ¨ PCR ¨ Gel Electrophresis (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 43
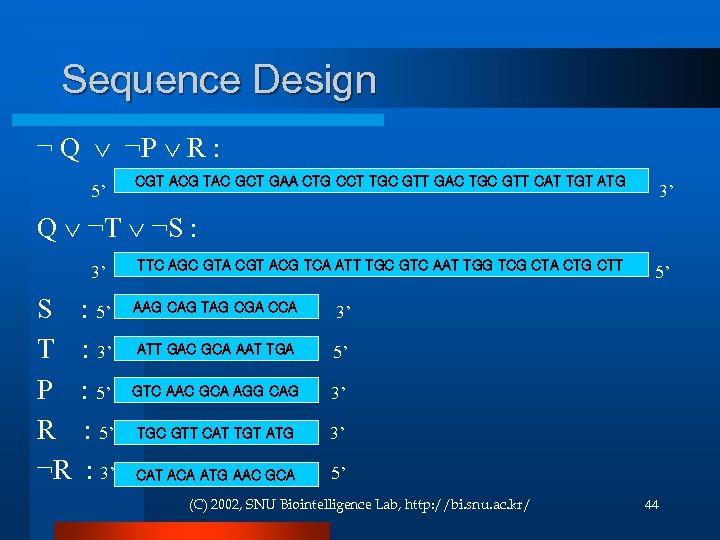

Sequence Design ¬ Q ¬P R : CGT ACG TAC GCT GAA CTG CCT TGC GTT GAC TGC GTT CAT TGT ATG 5’ 3’ Q ¬T ¬S : TTC AGC GTA CGT ACG TCA ATT TGC GTC AAT TGG TCG CTA CTG CTT 3’ 5’ AAG CAG TAG CGA CCA S : 5’ 3’ ATT GAC GCA AAT TGA T : 3’ 5’ GTC AAC GCA AGG CAG P : 5’ 3’ TGC GTT CAT TGT ATG R : 5’ 3’ ¬R : 3’ 5’ CAT ACA ATG AAC GCA (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 44
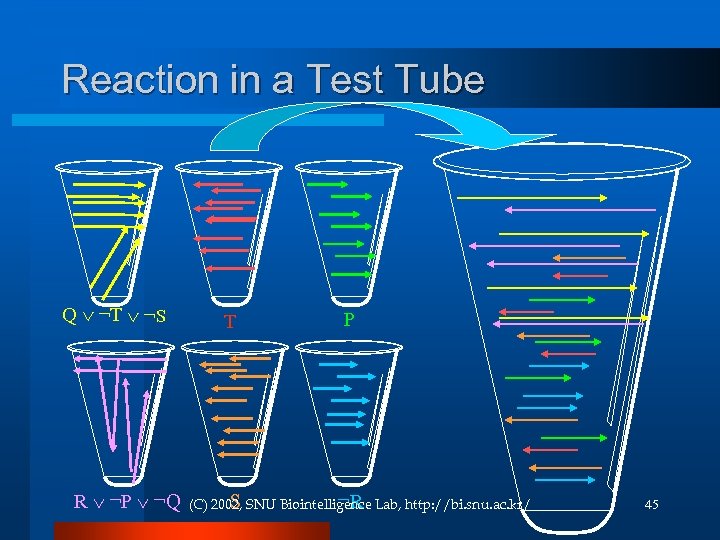
Reaction in a Test Tube Q ¬T ¬S R ¬P ¬Q T P S ¬R (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 45
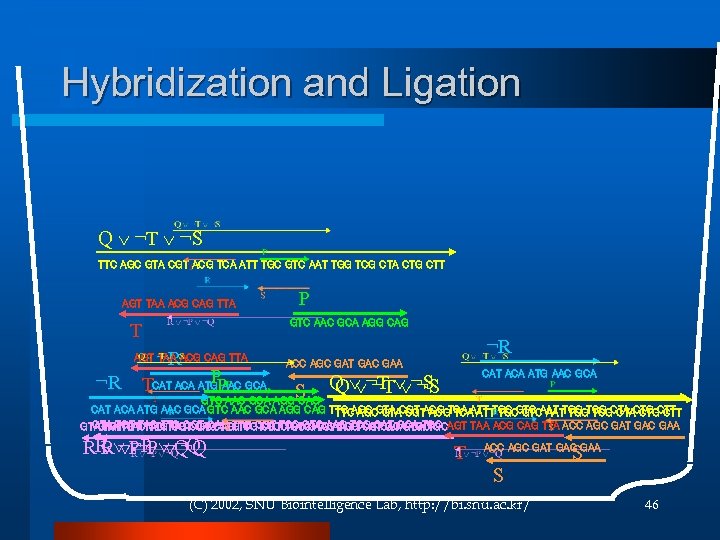
Hybridization and Ligation Q ¬T ¬S TTC AGC GTA CGT ACG TCA ATT TGC GTC AAT TGG TCG CTA CTG CTT AGT TAA ACG CAG TTA GTC AAC GCA AGG CAG T ¬R AGT TAA ACG CAG TTA ¬R P PAAC TCAT ACA ATGP GCA ¬R ACC AGC GAT GAC GAA S CAT ACA ATG AAC GCA Q ¬T ¬S GTC AAC GCA AGG CAT ACA ATG AAC GCA GTC AAC GCA AGG CAG TTC AGC GTA CGT ACG TCA ATT TGC GTC AAT TGG TCG CTA CTG CTT GTA TGT TAC TTG CGT CAG TTG CGT TCC GTC AAG TCG CAT GCA TGC GTAGTA TGT TAC TTG CGT CAG TTG CGT TCC GTC AAG TCG CAT GCA TGCAGT TAA ACG CAG TTA ACC AGC GAT GAC GAA TGT TAC TTG CGT CAG TTG CGT TCC GTC AAG TCG CAT GCA TGC R ¬P ¬Q T S ACC AGC GAT GAC GAA S (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 46
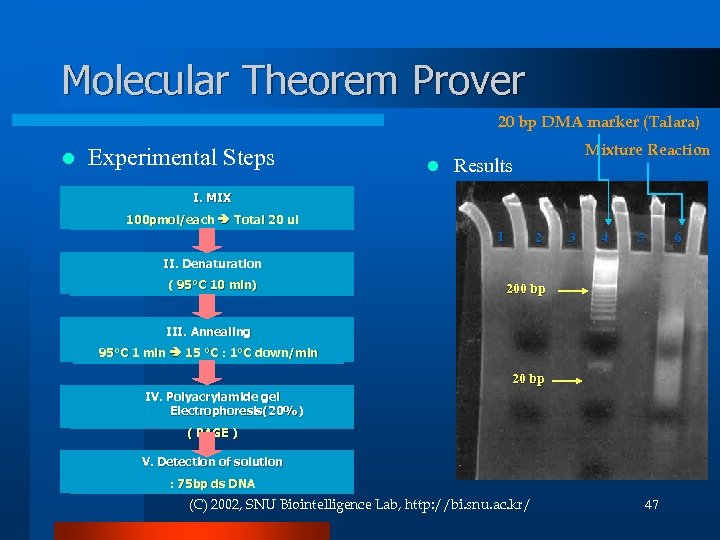
Molecular Theorem Prover 20 bp DMA marker (Talara) l Experimental Steps l Mixture Reaction Results I. MIX 100 pmol/each Total 20 ul 1 2 3 4 6 5 II. Denaturation ( 95°C 10 min) 200 bp III. Annealing 95°C 1 min 15 °C : 1°C down/min 20 bp IV. Polyacrylamide gel Electrophoresis(20%) ( PAGE ) V. Detection of solution : 75 bp ds DNA (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 47

The difficulties of DNA computer (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 48

The Reality of DNA based computation l l l Symmetry makes control difficult. Importance of Non-Watson-Crick Pairing. Importance of hybridization protocols. Importance of concentration and environment. Fidelity of ligation is high, but NOT perfect. DNA molecules breath. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 49

The Reality of DNA based computation l l l Affinity binding is a filtration process. Complex ligation produces complex products. Restriction enzymes produces partial digestion products. Base stacking is often determining interaction. DNA is a physical chemical system. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 50

Encoding Problems l Encoding Problems ¨ encoding problem is mapping the problem instance onto a set of DNA molecules and molecular biology protocols so that the resulting products contain an answer to instance of the problem ¨ prevent errors ¨ enable extraction (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 51

Operational Problems l It takes TOO long times ¨ hybridization/ligation operation over 4 hours ¨ In Adleman’s experiments : 7 days! (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 52

Operational Problems l Not Perfect Operation ¨ Hybridization Mismatches ¨ Extraction Errors ¨ Volume and Mass to solve a problem False Negatives l False Positives l (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 53
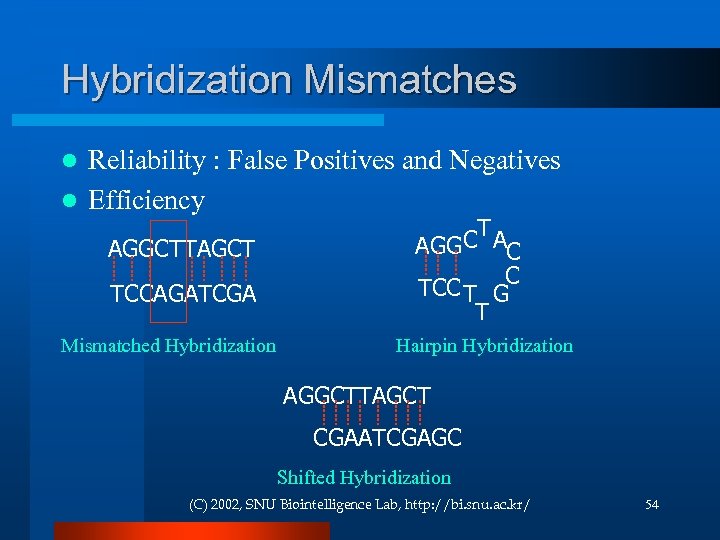

Hybridization Mismatches Reliability : False Positives and Negatives l Efficiency l AGGCTTAGCT TCCAGATCGA Mismatched Hybridization CT AC AGG TCC T GC T Hairpin Hybridization AGGCTTAGCT CGAATCGAGC Shifted Hybridization (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 54

Detection problems NOT exact detection l Gel electrophoresis is widely used detection technique. l ¨ We can not separate the exact solution! (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 55
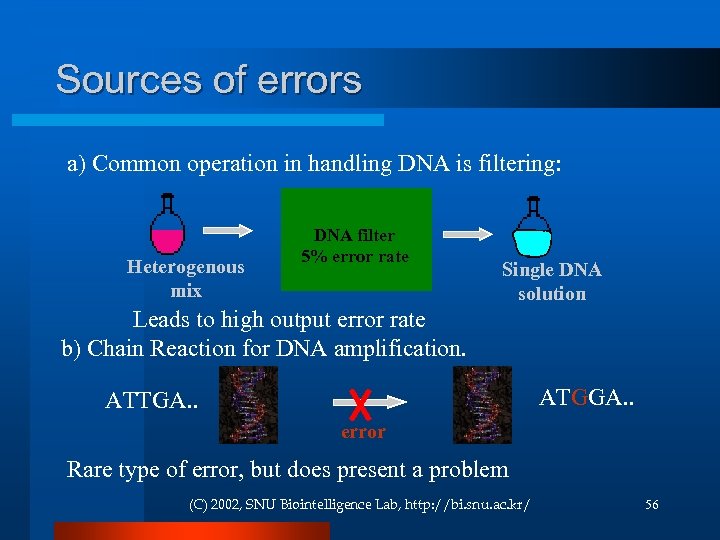

Sources of errors a) Common operation in handling DNA is filtering: Heterogenous mix DNA filter 5% error rate Single DNA solution Leads to high output error rate b) Chain Reaction for DNA amplification. ATGGA. . ATTGA. . error Rare type of error, but does present a problem (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 56
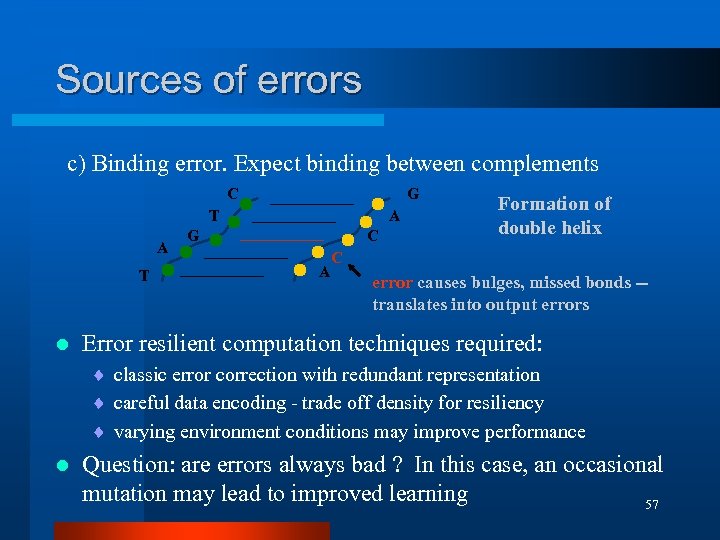

Sources of errors c) Binding error. Expect binding between complements C G T A T l A G C A Formation of double helix C error causes bulges, missed bonds -translates into output errors Error resilient computation techniques required: ¨ classic error correction with redundant representation ¨ careful data encoding - trade off density for resiliency ¨ varying environment conditions may improve performance l Question: are errors always bad ? In this case, an occasional mutation may lead to improved learning 57

DNA Computing Applications (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 58

Applications of Biomolecular Computing l l l l Massively parallel problem solving Combinatorial optimization Molecular nano-memory with fast associative search AI problem solving Medical diagnosis, drug discovery Cryptography Biocomputer Further impact in biology and medicine: ¨ ¨ Wet biological data bases Processing of DNA labeled with digital data Sequence comparison Fingerprinting (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 59
![DNA Adder l [UC Berkely, USA] The Addition Reaction: Primer Extension 5’ 3’ Template DNA Adder l [UC Berkely, USA] The Addition Reaction: Primer Extension 5’ 3’ Template](https://present5.com/presentation/022864c07b2c663373ae3958f0d120fb/image-60.jpg)
DNA Adder l [UC Berkely, USA] The Addition Reaction: Primer Extension 5’ 3’ Template DNA Primer Extension Polymerase 5’ Primer DNA (= 2 nd digit) (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 60
![Massively Parallel Theorem Proving l [Warsaw Univ. of Technology, Poland] IF Premise THEN Conclusion Massively Parallel Theorem Proving l [Warsaw Univ. of Technology, Poland] IF Premise THEN Conclusion](https://present5.com/presentation/022864c07b2c663373ae3958f0d120fb/image-61.jpg)
Massively Parallel Theorem Proving l [Warsaw Univ. of Technology, Poland] IF Premise THEN Conclusion Premise A A A Conclusion C A D B C C B B B D D 61
![Breaking DES l [Princeton, USA] B 2(0) B 1(0) S 1 S 0 B Breaking DES l [Princeton, USA] B 2(0) B 1(0) S 1 S 0 B](https://present5.com/presentation/022864c07b2c663373ae3958f0d120fb/image-62.jpg)
Breaking DES l [Princeton, USA] B 2(0) B 1(0) S 1 S 0 B 1(1) BN(0) SN …. . B 2(1) BN(1) DES circuit initial graph S 0 l S 0 r B 1 l(0) B 1 r(0) S 1 l S 1 r B 2 l(1) B 2 r(1) S 2 l A path in the graph SN l SNr BNr(1) SNl SNr 62

Future Applications (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 63
Interesting possibilities a) Self-replication: Two for one Based on DNA self-replication b) Self-repair: Based on regeneration c) DNA computer mutation/evolution d) New meaning of a Learning. computer virus ? or biohazard May be malignant 64

Evolvable Biomolecular Hardware l Sequence programmable and evolvable molecular systems have been constructed as cell-free chemical systems using biomolecules such as DNA and proteins. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 65
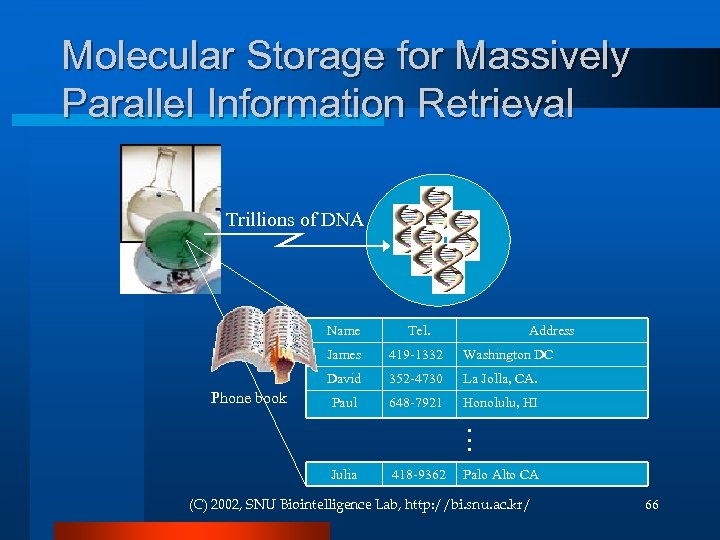
Molecular Storage for Massively Parallel Information Retrieval Trillions of DNA Name James Address Washington DC 352 -4730 La Jolla, CA. Paul 648 -7921 Honolulu, HI Julia 418 -9362 … 419 -1332 David Phone book Tel. Palo Alto CA (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 66
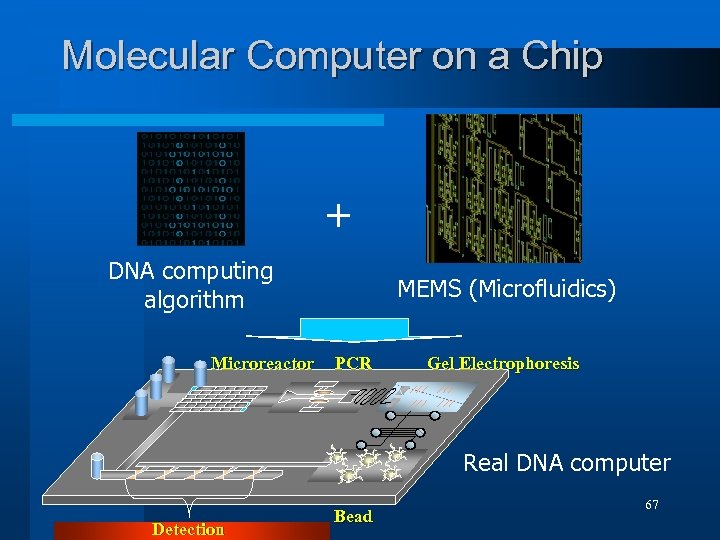
Molecular Computer on a Chip + DNA computing algorithm Microreactor MEMS (Microfluidics) PCR Gel Electrophoresis Real DNA computer Detection Bead 67

Bio. MEMS – the Next Transistors (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 68

Lab-on-a-chip Technology Integrates sample handling, separation and detection and data analysis for: DNA, RNA and protein solutions using Lab. Chip technology. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 69

Conclusion DNA Computing uses DNA molecules to computing or storage materials. l DNA computing technology has many interesting properties, including l ¨ ¨ l Massively parallel, solution-based, biochemical Nano-scale, biocompatible high energy efficiency high memory storage density DNA computing is in very early stage of development. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 70

Research Groups l l l MIT, Caltech, Princeton University, Bell Labs EMCC (European Molecular Computing Consortium) is composed of national groups from 11 European countries Bio. MIP Institute (Bio. Molecular Information Processing) at the German National Research Center for Information Technology (GMD) Molecular Computer Project (MCP) in Japan Leiden Center for Natural Computation (LCNC) (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 71

Web Resources European Molecular Computing Consortium (EMCC): http: //www. csc. liv. ac. uk/~emcc/ l Bio. Molecular Information Processing (Bio. Mip): http: //www. gmd. de/BIOMIP l Leiden Center for Natural Computation (LCNC): http: //www. wi. leidenuniv. nl/~lcnc/ l Biomolecular Computation (BMC): http: //bmc. cs. duke. edu/ l DNA Computing and Informatics at Surfaces: http: //www. corninfo. chem. wisc. edu/writings/DNAcomputi ng. html l SNU Molecular Evolutionary Computing (MEC) Project: http: //scai. snu. ac. kr/Research/ 72 l

Books l l l Cristian S, Calude and Gheorghe Paun, Computing with Cells and Atoms: An introduction to quantum, DNA and membrane computing, Taylor & Francis, 2001. Pâun, G. , Ed. , Computing With Bio-Molecules: Theory and Experiments, Springer, 1999. Gheorghe Paun, Grzegorz Rozenberg and Arto Salomaa, DNA Computing, New Computing Paradigms, Springer, 1998. C. S. Calude, J. Casti and M. J. Dinneen, Unconventional Models of Computation, Springer, 1998. Tono Gramss, Stefan Bornholdt, Michael Gross, Melanie Mitchell and thomas Pellizzari, Non-Standard Computation: Molecular Computation-Cellular Automata-Evolutionary Algorithms-Quantum Computers, Wiley-Vch, 1997. (C) 2002, SNU Biointelligence Lab, http: //bi. snu. ac. kr/ 73
022864c07b2c663373ae3958f0d120fb.ppt