d85621a97272fad7ba2459c799825cd2.ppt
- Количество слайдов: 27

DMKPred: Specificity and Crossreactivity of Kinase Inhibitors G P S Raghava, Head Bioinformatics Centre Email: raghava@imtech. res. in; Web: http: //www. imtech. res. in/raghava/ Institute of Microbial Technology, Sector-39 A, Chandigarh, India

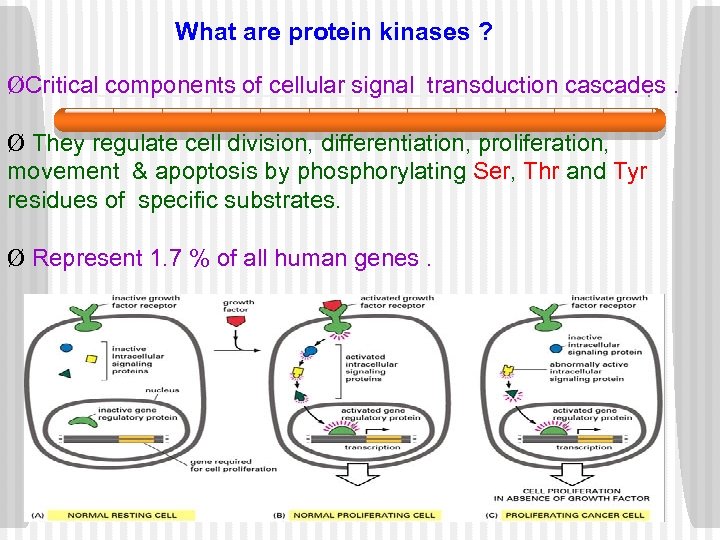
What are protein kinases ? ØCritical components of cellular signal transduction cascades. Ø They regulate cell division, differentiation, proliferation, movement & apoptosis by phosphorylating Ser, Thr and Tyr residues of specific substrates. Ø Represent 1. 7 % of all human genes.

Why kinases are so important? They are the key regulators of all aspects of neoplasia, including proliferation, invasion, angiogenesis and metastasis. A number of diseases, especially cancer involve unregulated kinase activity (overexpression / upregulation). This makes kinases as important targets for drug development. Kinase inhibitors are successfully used in cancer treatment.
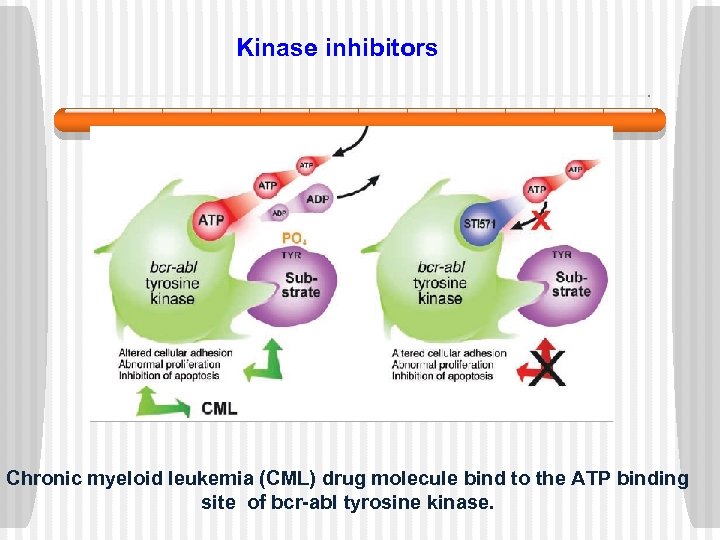
Kinase inhibitors Chronic myeloid leukemia (CML) drug molecule bind to the ATP binding site of bcr-abl tyrosine kinase.
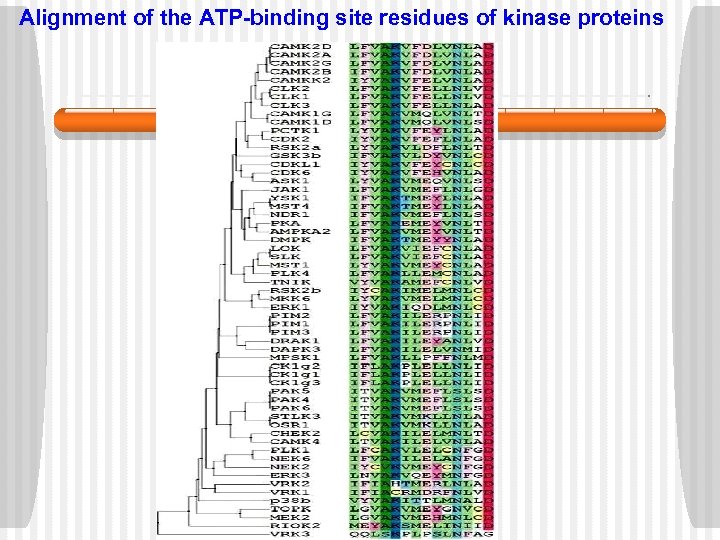
Alignment of the ATP-binding site residues of kinase proteins
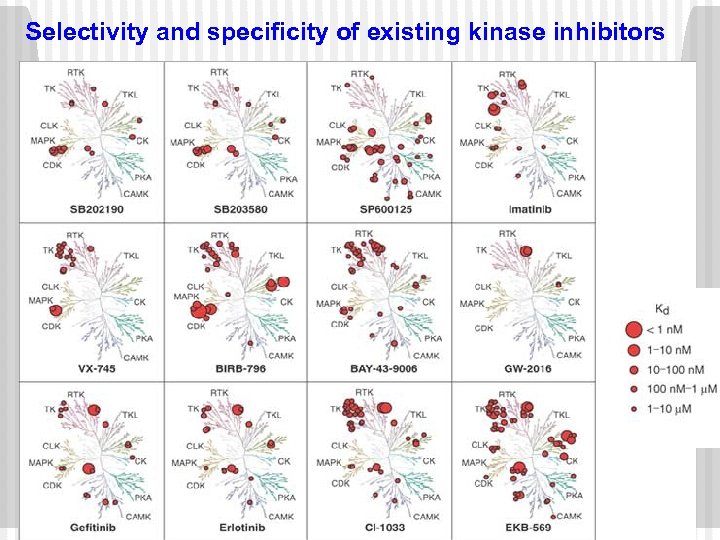
Selectivity and specificity of existing kinase inhibitors

Can we solve specificity problem of kinase inhibitors ? ØMost of the drug molecules binds with other protein kinases and cause cross-reactivity. ØIt’s very difficult to design a specific kinase inhibitors against a protein kinase. ØIf Kd of inhibitors with primary intended target is ≤ 10 fold then chances of cross-reactivity is low. Fabin et al. , Nat. Biotechnol. , 2005, 23, 229236

Specificity and cross-reactivity of kinase inhibitors Molecules data: Data was taken from Nat. Biotechnol. , 2005, 23, 229 -236 (Fabin et al. , 119 × 20 Kd data) Select those kinases for which 6 chemical molecules have significant binding Finally we select 29 protein kinases for this study

Specificity and cross-reactivity of kinase inhibitors Structure Descriptors: We calculated 8 structure descriptors from Molinspiration (on line web server for descriptor calculation) We also calculated 600 structure descriptors using Pre. ADMET (on line web server for molecular descriptor calculation) Remove insignificant descriptors and select 62 molecular descriptors for further study
Specificity and cross-reactivity of kinase inhibitors Calculate molecular descriptors of kinase inhibitors Remove similar and insignificant descriptors Calculate correlation between Kd and descriptors Select highly significant molecular descriptors Developed model for prediction
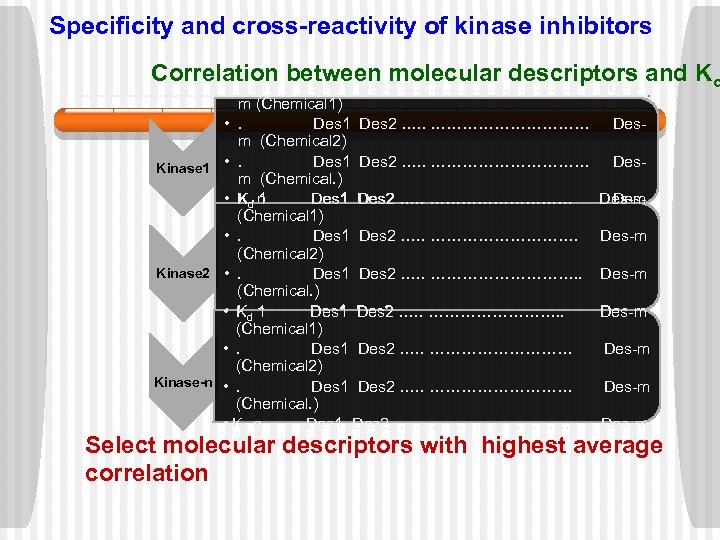

Specificity and cross-reactivity of kinase inhibitors Correlation between molecular descriptors and Kd • K 1 Des 2 …. . ……………. . Desd m (Chemical 1) • . Des 1 Des 2 …. . …………… Desm (Chemical 2) Des 1 Des 2 …. . …………… Des. Kinase 1 • . m (Chemical. ) • Kd n 1 Des 2…… …………… Des 2 …. . …………… Des-m Des(Chemical 1) m (Chemical-n) • . Des 1 Des 2 …. . ……………. Des-m (Chemical 2) Kinase 2 • . Des 1 Des 2 …. . ……………. . Des-m (Chemical. ) Des 1 Des 2…… …………. . • Kd n Des 2 …. . ……………. Des-m d 1 (Chemical-n) (Chemical 1) • . Des 1 Des 2 …. . …………… Des-m (Chemical 2) Kinase-n • . Des 1 Des 2 …. . …………… Des-m (Chemical. ) • Kd n Des 1 Des 2…… …………… Des-m (Chemical-n) Select molecular descriptors with highest average correlation
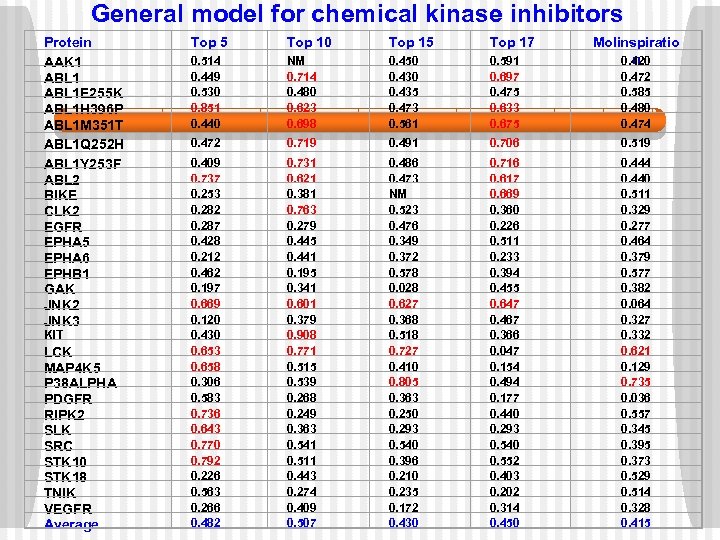
General model for chemical kinase inhibitors Protein AAK 1 ABL 1 E 255 K ABL 1 H 396 P ABL 1 M 351 T ABL 1 Q 252 H ABL 1 Y 253 F ABL 2 BIKE CLK 2 EGFR EPHA 5 EPHA 6 EPHB 1 GAK JNK 2 JNK 3 KIT LCK MAP 4 K 5 P 38 ALPHA PDGFR RIPK 2 SLK SRC STK 10 STK 18 TNIK VEGFR Average Top 5 Top 10 Top 15 Top 17 Molinspiratio n 0. 420 0. 514 0. 449 0. 530 0. 851 0. 440 NM 0. 714 0. 480 0. 623 0. 698 0. 450 0. 435 0. 473 0. 561 0. 591 0. 697 0. 475 0. 633 0. 675 0. 472 0. 719 0. 491 0. 706 0. 519 0. 409 0. 737 0. 253 0. 282 0. 287 0. 428 0. 212 0. 462 0. 197 0. 669 0. 120 0. 430 0. 653 0. 658 0. 306 0. 583 0. 736 0. 643 0. 770 0. 792 0. 226 0. 563 0. 266 0. 482 0. 731 0. 621 0. 381 0. 763 0. 279 0. 445 0. 441 0. 195 0. 341 0. 601 0. 379 0. 908 0. 771 0. 515 0. 539 0. 268 0. 249 0. 363 0. 541 0. 511 0. 443 0. 274 0. 409 0. 507 0. 486 0. 473 NM 0. 523 0. 476 0. 349 0. 372 0. 578 0. 028 0. 627 0. 368 0. 518 0. 727 0. 410 0. 805 0. 363 0. 250 0. 293 0. 540 0. 396 0. 210 0. 235 0. 172 0. 430 0. 716 0. 617 0. 669 0. 360 0. 226 0. 511 0. 233 0. 394 0. 455 0. 647 0. 467 0. 366 0. 047 0. 154 0. 494 0. 177 0. 440 0. 293 0. 540 0. 552 0. 403 0. 202 0. 314 0. 450 0. 444 0. 440 0. 511 0. 329 0. 277 0. 464 0. 379 0. 577 0. 382 0. 064 0. 327 0. 332 0. 621 0. 129 0. 735 0. 036 0. 557 0. 345 0. 395 0. 373 0. 529 0. 514 0. 328 0. 415 0. 472 0. 585 0. 480 0. 474
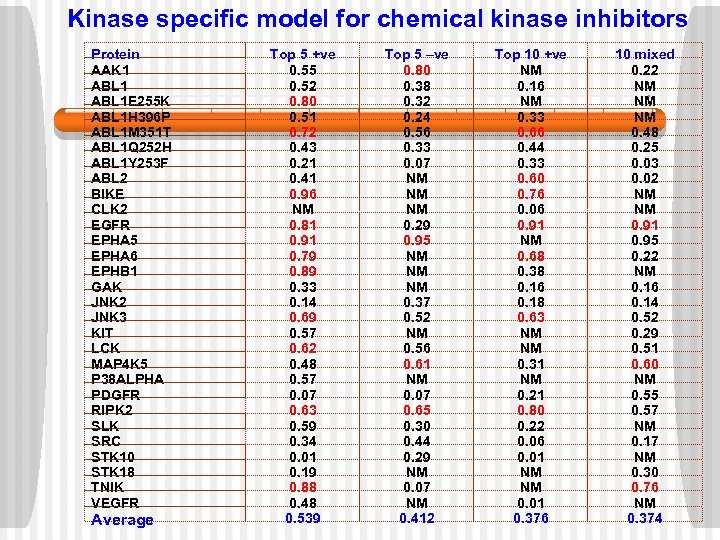
Kinase specific model for chemical kinase inhibitors Protein AAK 1 ABL 1 E 255 K ABL 1 H 396 P ABL 1 M 351 T ABL 1 Q 252 H ABL 1 Y 253 F ABL 2 BIKE CLK 2 EGFR EPHA 5 EPHA 6 EPHB 1 GAK JNK 2 JNK 3 KIT LCK MAP 4 K 5 P 38 ALPHA PDGFR RIPK 2 SLK SRC STK 10 STK 18 TNIK VEGFR Average Top 5 +ve 0. 55 0. 52 0. 80 0. 51 0. 72 0. 43 0. 21 0. 41 0. 96 NM 0. 81 0. 91 0. 79 0. 89 0. 33 0. 14 0. 69 0. 57 0. 62 0. 48 0. 57 0. 07 0. 63 0. 59 0. 34 0. 01 0. 19 0. 88 0. 48 0. 539 Top 5 –ve 0. 80 0. 38 0. 32 0. 24 0. 56 0. 33 0. 07 NM NM NM 0. 29 0. 95 NM NM NM 0. 37 0. 52 NM 0. 56 0. 61 NM 0. 07 0. 65 0. 30 0. 44 0. 29 NM 0. 07 NM 0. 412 Top 10 +ve NM 0. 16 NM 0. 33 0. 66 0. 44 0. 33 0. 60 0. 76 0. 06 0. 91 NM 0. 68 0. 38 0. 16 0. 18 0. 63 NM NM 0. 31 NM 0. 21 0. 80 0. 22 0. 06 0. 01 NM NM 0. 01 0. 376 10 mixed 0. 22 NM NM NM 0. 48 0. 25 0. 03 0. 02 NM NM 0. 91 0. 95 0. 22 NM 0. 16 0. 14 0. 52 0. 29 0. 51 0. 60 NM 0. 55 0. 57 NM 0. 17 NM 0. 30 0. 76 NM 0. 374
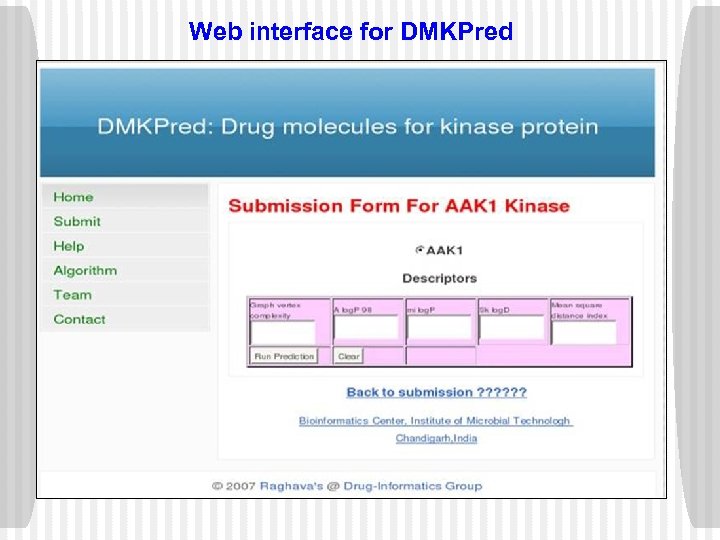

Web interface for DMKPred

Computational Resources for Drug Discovery An Insilico Module of Open Source Drug Discovery

Meta-Server: Prediction of subcellular localization of proteins using various server

URLs: http: //www. imtech. res. in/raghava/ & http: //crdd. osdd. net/ & bic. uams. edu/
d85621a97272fad7ba2459c799825cd2.ppt