5ce0c7c13457a3fed4af3e2484f6a9da.ppt
- Количество слайдов: 29

Discovering Gene-Disease Association using On-line Scientific Text Abstracts. Raj Adhikari Advisor: Javed Mostafa

Motivation n n 3/19/2018 A central problem in bioinformatics is how to capture information from the vast scientific literature and create an automated system for “knowledge discovery” that can be used in various areas. I address the special case of gene-disease interactions and show that using the frequencies/relevance of words in Pubmed abstracts can be used to find genes related to a disease. Bioinformatics capstone project 2

Goal n Use the combination of statistical methods and a database to: n n 3/19/2018 retrieve research abstracts from Pubmed. extract relevant information from the free texts using statistical methods. Measure the accuracy of the results and display the results using a Web based system. Complement and support existing knowledge base systems like Gene. Cards. Bioinformatics capstone project 3

Resources used in creating database n Pub. Med n n a database of human genes, their products and their involvement in diseases http: //bioinfo. weizmann. ac. il/cards/index. shtml HGNC n n http: //www. ncbi. nlm. nih. gov/entrez/query. fcgi Gene. Cards n n The US National Library of Medicine's database that contains more than 11 million references to journal articles in the health sciences. HUGO Gene Nomenclature Committee (approved over 19000 human gene symbols). consistent with OMIM and Locus. Link http: //www. gene. ucl. ac. uk/nomenclature Tools used: Perl, CGI, Java, My. SQL 3/19/2018 Bioinformatics capstone project 4

Creating the database n Data I used: n n A relatively small list of genes and diseases in humans An article set (around 8000) n For each Pubmed article: n n n 3/19/2018 PMID Article Title Abstract (filter with a list of stop words) The HUGO dataset. List of around 3500 related gene-disease pairs from Gene. Cards. Bioinformatics capstone project 5

Populating the database tables n n n Use the book Genes and Disease at OMIM to generate a list of around 60 diseases and 90 genes. Search Pubmed for each gene-disease pair on the Title/Abstract field. Use ESearch (tool that provides access to Pubmed database outside of the web interface) to retrieve data in XML file format. Use XML: : Simple Perl package to parse the XML file Filter the text using stop words and store each title and abstract along with the related PMID in a database table. Add more genes using HUGO OMIM: Database of genetic diseases with references to molecular medicine, cell biology, biochemistry and clinical details of the diseases. 3/19/2018 Bioinformatics capstone project 6

Populating the database tables n n Table structures: n. Term Derivative table Tfreq Dfreq Tfidf PMID LSI Parse the retrieved text files and create the following tables: HGNC genesymbol alias n HUGO table structure: n 3/19/2018 Gene. Cards table structure: Genesymbol disease Bioinformatics capstone project 7
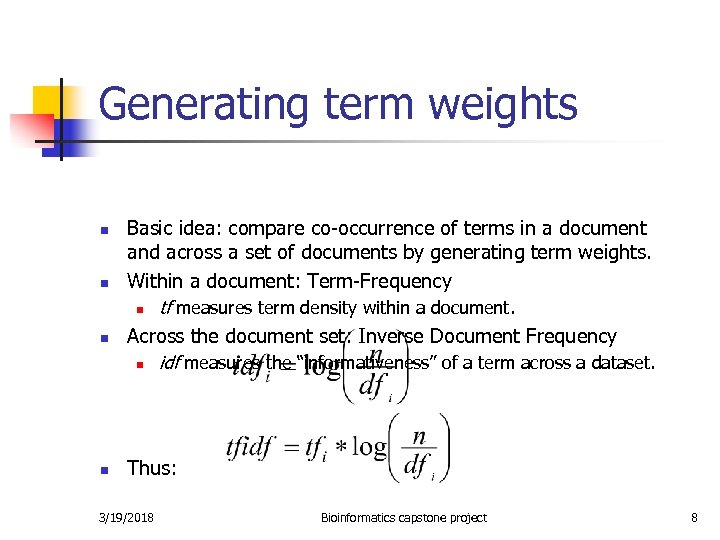

Generating term weights n n Basic idea: compare co-occurrence of terms in a document and across a set of documents by generating term weights. Within a document: Term-Frequency n tf measures term density within a document. Across the document set: Inverse Document Frequency n idf measures the “informativeness” of a term across a dataset. Thus: 3/19/2018 Bioinformatics capstone project 8

Latent Symantec Indexing n n Calculating co-occurrence of terms might not suffice because of possible “noise” in the dataset. Use LSI, a statistical technique, to estimate a latent structure. Assume some underlying semantic structure in the dataset which could be partially obscured. Implementation n 3/19/2018 term by document matrix (tends to be sparse) convert matrix entries to weights, e. g. tfidf. Analyze the matrix by singular value decomposition (SVD) to derive latent semantic structure model. Bioinformatics capstone project 9

SVD n unique mathematical decomposition of a matrix into the product of three matrices: n n 3/19/2018 two with orthonormal columns one with singular values on the diagonal finds optimal projection into low-dimensional space tool for dimension reduction Bioinformatics capstone project 10
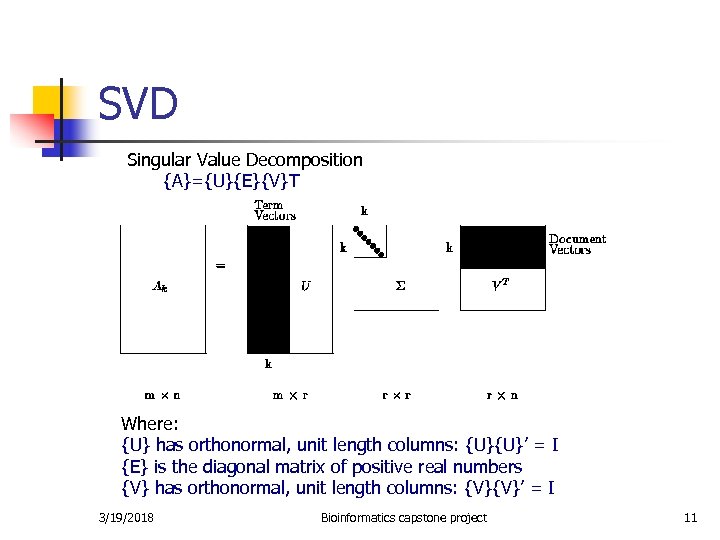
SVD Singular Value Decomposition {A}={U}{E}{V}T Where: {U} has orthonormal, unit length columns: {U}{U}’ = I {E} is the diagonal matrix of positive real numbers {V} has orthonormal, unit length columns: {V}{V}’ = I 3/19/2018 Bioinformatics capstone project 11

SVD n n Approximate Ak keeping only the first k singular values and the corresponding columns from U and V matrices. The new matrix Ak does not exactly match the original term by document matrix A. (It gets closer and closer as more singular values are kept). This is what we want: we don’t want perfect fit since we think some of the 0’s in A should be not be 0 and vice versa. Limitations of SVD – very memory intensive, cannot handle large datasets. 3/19/2018 Bioinformatics capstone project 12

Scoring Matrix Generation n A scoring matrix is generated for each term weighting method using the data stored in the database. This matrix is used to find the relationships between genes and diseases. Relatively fast process since the weights are pre-computed and stored in a database. 3/19/2018 Bioinformatics capstone project 13
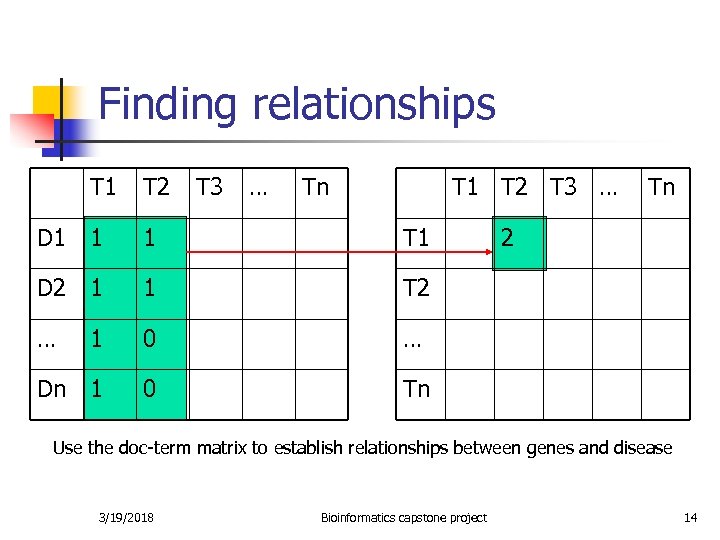
Finding relationships T 1 T 2 T 3 … Tn T 1 T 2 T 3 … D 1 1 1 T 1 D 2 1 1 T 2 … 1 0 … Dn 1 0 Tn 2 Tn Use the doc-term matrix to establish relationships between genes and disease 3/19/2018 Bioinformatics capstone project 14
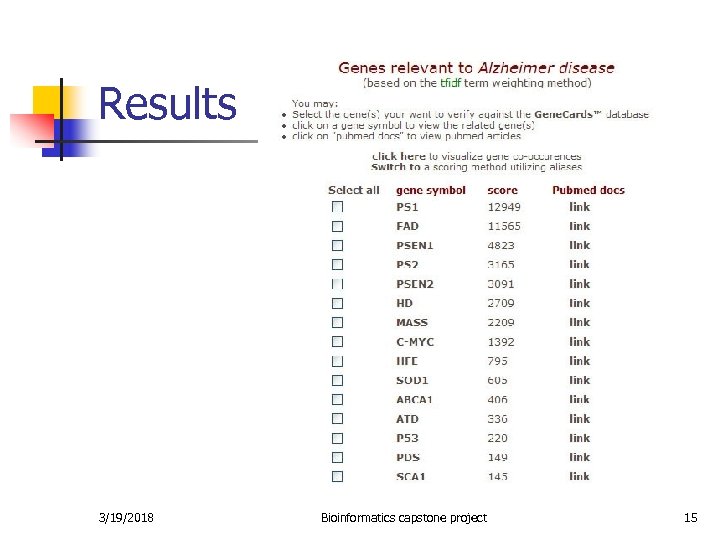
Results 3/19/2018 Bioinformatics capstone project 15

Verification of the relationship n n n Data from Gene. Cards and HUGO has been stored in a database. For each gene, if the symbol is an official genesymbol (according to HUGO), then search for the genesymbol in Gene. Cards and display the disease associated with it. Else (if the symbol is an alias), use HUGO to find the official genesymbol and search in Gene. Cards using this genesymbol and display the disease associated with the gene. 3/19/2018 Bioinformatics capstone project 16
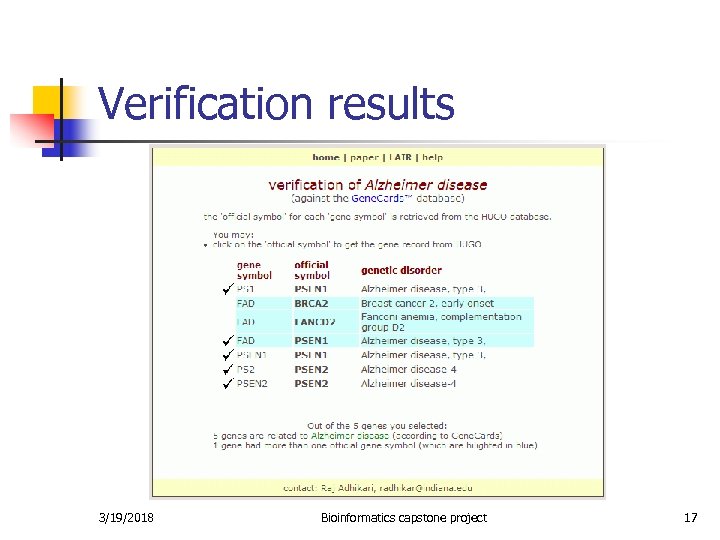

Verification results 3/19/2018 Bioinformatics capstone project 17

Using gene alias n n Make use of gene alias from HUGO to increase the chances of detecting correct genes for a given disease Method: n n 3/19/2018 Increment the weight of an official gene by adding the weight of the alias. Group the alias together with the official gene. Bioinformatics capstone project 18
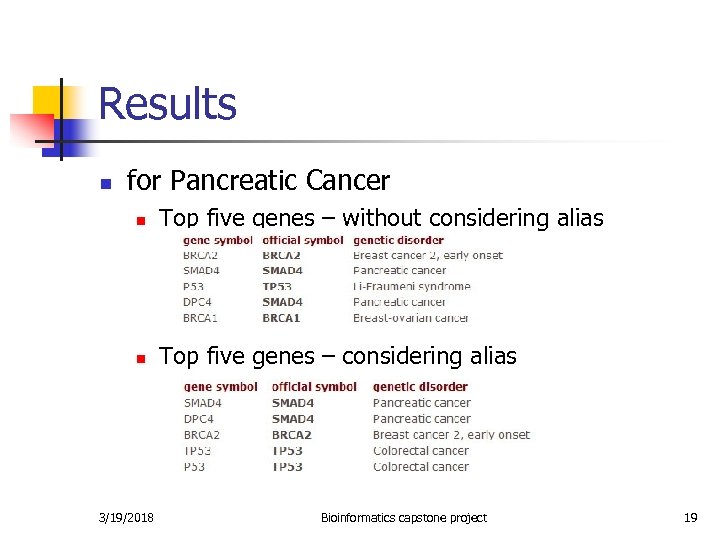

Results n for Pancreatic Cancer n Top five genes – without considering alias n Top five genes – considering alias 3/19/2018 Bioinformatics capstone project 19
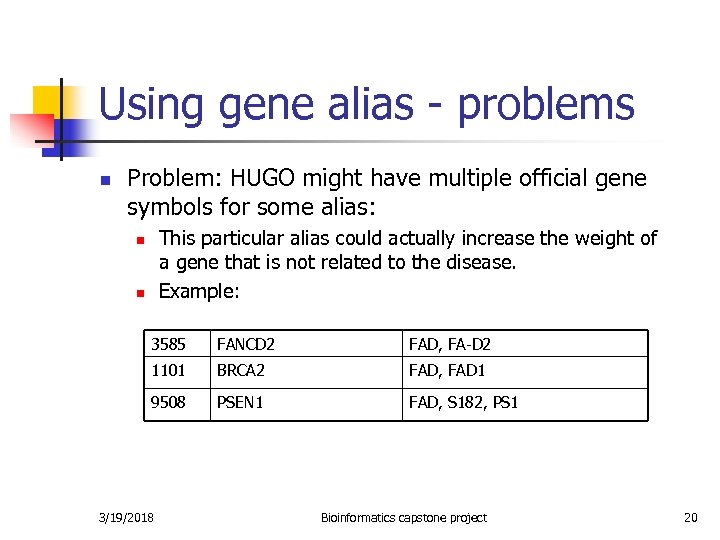

Using gene alias - problems n Problem: HUGO might have multiple official gene symbols for some alias: This particular alias could actually increase the weight of a gene that is not related to the disease. Example: n n 3585 FANCD 2 FAD, FA-D 2 1101 BRCA 2 FAD, FAD 1 9508 PSEN 1 FAD, S 182, PS 1 3/19/2018 Bioinformatics capstone project 20
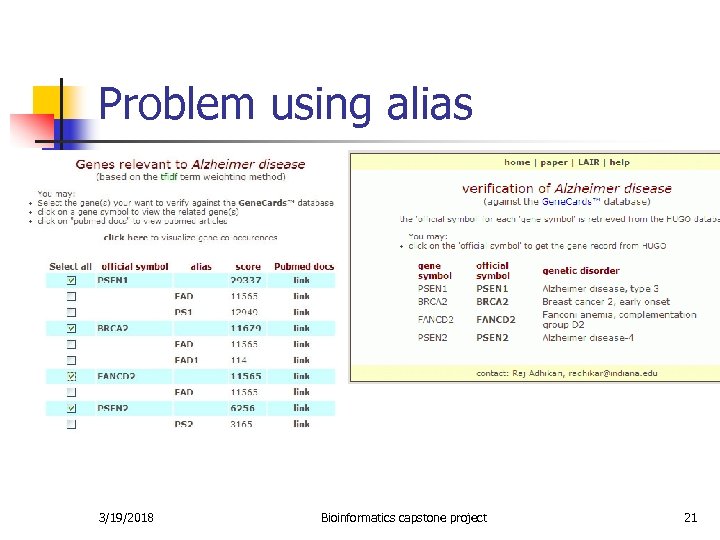
Problem using alias 3/19/2018 Bioinformatics capstone project 21

Verification n n In addition, the number of Pubmed articles containing a disease and a gene symbol can be an indication of how strong the association between a disease and a gene is. Same theory applies for a gene-gene relationship. 3/19/2018 Bioinformatics capstone project 22
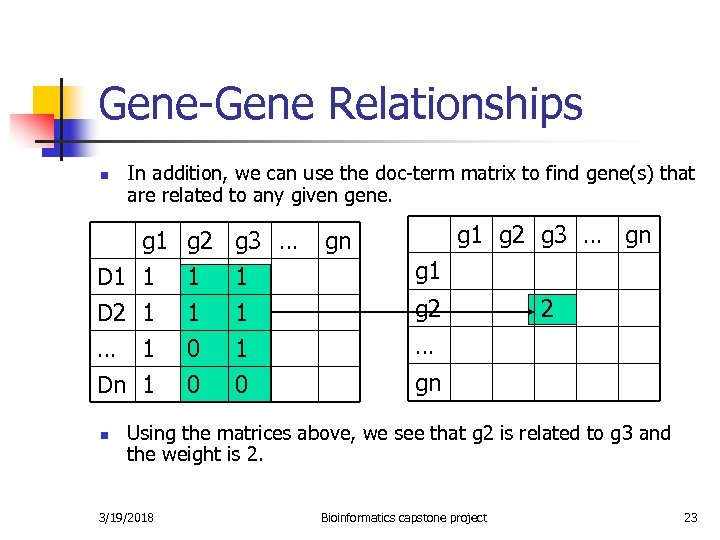
Gene-Gene Relationships n In addition, we can use the doc-term matrix to find gene(s) that are related to any given gene. g 1 D 1 1 D 2 1 … 1 g 2 1 1 0 g 3 … 1 1 1 Dn 1 0 0 n g 1 g 2 g 3 … gn gn g 1 g 2 2 … gn Using the matrices above, we see that g 2 is related to g 3 and the weight is 2. 3/19/2018 Bioinformatics capstone project 23

Discovering additional gene relationships n We can make use of the possibility that two genes might be related to each other via a disease as in: gene 1 -> disease 1 -> gene 2 gene 1 -> disease 2 -> gene 2 n to establish relationships between gene 1 and gene 2. In our case, the fact that gene 1 and gene 2 are related to each other via two different diseases makes the relationship between them even stronger. 3/19/2018 Bioinformatics capstone project 24
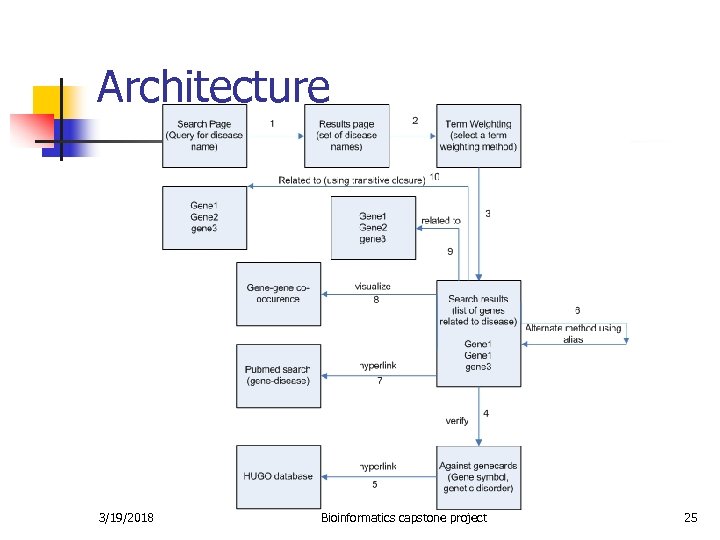
Architecture 3/19/2018 Bioinformatics capstone project 25

System Demonstration n n http: //biokdd. informatics. indiana. edu/radhikar /search. html Related URLs: n Genecards: n n HGNC: n 3/19/2018 http: //bioinfo. weizmann. ac. il/cards/index. shtml http: //www. gene. ucl. ac. uk/nomenclature/ Bioinformatics capstone project 26

Summary n n Using the combination of statistical methods and a database, the process of establishing gene-disease relationship using literature data is fast and efficient. With minimal changes, our system can be extended to discover other relationships like protein-protein interactions, etc. 3/19/2018 Bioinformatics capstone project 27

Future Work n n Extend our system to incorporate the entire Medline dataset. Incorporate full gene names. Find a better way to verify the gene-gene relationships. Incorporate other On-Line scientific literature databases. 3/19/2018 Bioinformatics capstone project 28

Acknowledgments n n Professor Javed Mostafa Professor Sun Kim Professor Memo Dalkilic Professor Haixu Tang 3/19/2018 Bioinformatics capstone project 29
5ce0c7c13457a3fed4af3e2484f6a9da.ppt