0ba21210f7b7f7938212a9b67c60b7a1.ppt
- Количество слайдов: 70

Different Perspectives at Clustering: The “Number-of-Clusters” Case B. Mirkin School of Computer Science Birkbeck College, University of London IFCS 2006

Different Perspectives at Number of Clusters: Talk Outline Clustering and K-Means: A discussion Clustering goals and four perspectives Number of clusters in: - Classical statistics perspective - Machine learning perspective - Data Mining perspective (including a simulation study with 8 methods) - Knowledge discovery perspective (including a comparative genomics project)

WHAT IS CLUSTERING; WHAT IS DATA K-MEANS CLUSTERING: Conventional K-Means; Initialization of KMeans; Intelligent K-Means; Interpretation Aids WARD HIERARCHICAL CLUSTERING: Agglomeration; Divisive Clustering with Ward Criterion; Extensions of Ward Clustering DATA RECOVERY MODELS: Statistics Modelling as Data Recovery; Data Recovery Model for K-Means; for Ward; Extensions to Other Data Types; One-by-One Clustering DIFFERENT CLUSTERING APPROACHES: Extensions of K-Means; Graph -Theoretic Approaches; Conceptual Description of Clusters GENERAL ISSUES: Feature Selection and Extraction; Similarity on Subsets and Partitions; Validity and Reliability
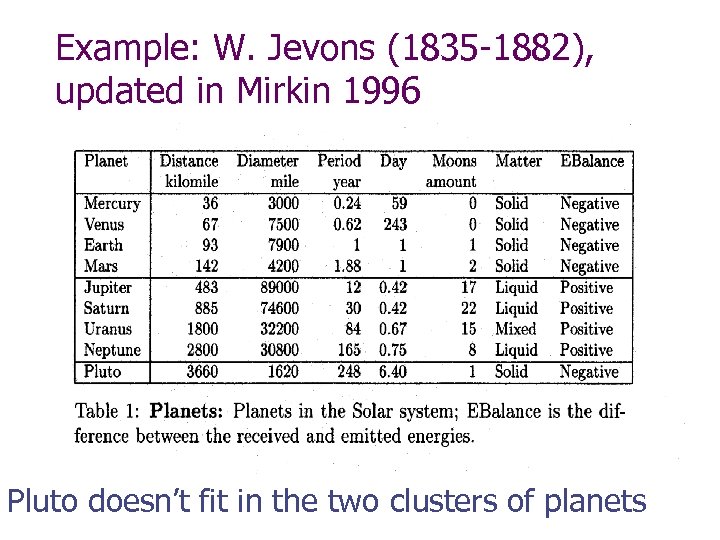
Example: W. Jevons (1835 -1882), updated in Mirkin 1996 Pluto doesn’t fit in the two clusters of planets
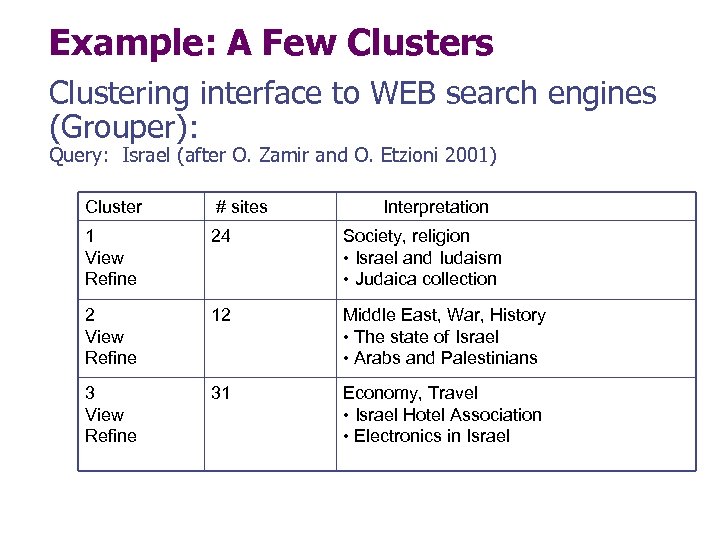
Example: A Few Clusters Clustering interface to WEB search engines (Grouper): Query: Israel (after O. Zamir and O. Etzioni 2001) Cluster # sites Interpretation 1 View Refine 24 Society, religion • Israel and Iudaism • Judaica collection 2 View Refine 12 Middle East, War, History • The state of Israel • Arabs and Palestinians 3 View Refine 31 Economy, Travel • Israel Hotel Association • Electronics in Israel

Clustering: Main Steps Data collecting Data pre-processing Finding clusters (the only step appreciated in conventional clustering) Interpretation Drawing conclusions

Conventional Clustering: Cluster Algorithms Single Linkage: Nearest Neighbour Ward Agglomeration Conceptual Clustering K-means Kohonen SOM ………………….
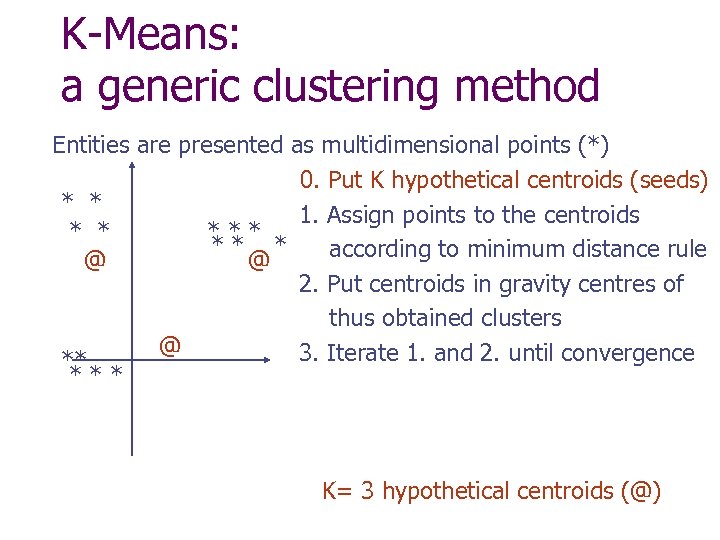

K-Means: a generic clustering method Entities are presented as multidimensional points (*) 0. Put K hypothetical centroids (seeds) * * 1. Assign points to the centroids * * ** * according to minimum distance rule @ @ 2. Put centroids in gravity centres of thus obtained clusters @ 3. Iterate 1. and 2. until convergence ** *** K= 3 hypothetical centroids (@)
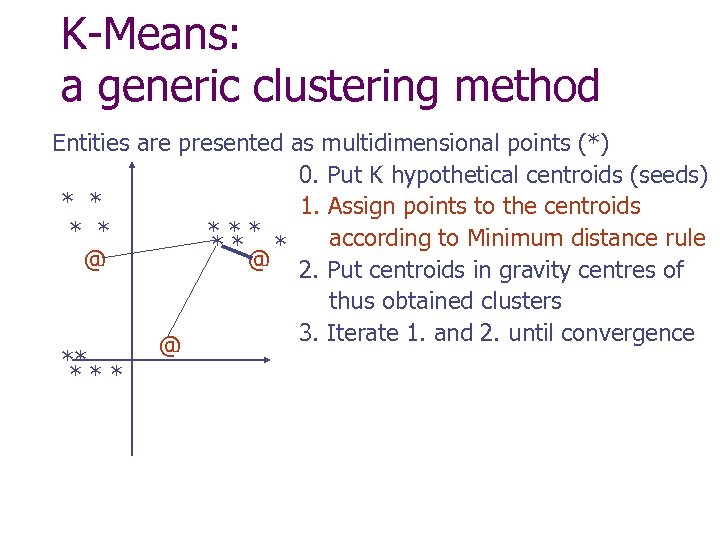

K-Means: a generic clustering method Entities are presented as multidimensional points (*) 0. Put K hypothetical centroids (seeds) * * 1. Assign points to the centroids * * *** according to Minimum distance rule ** * @ @ 2. Put centroids in gravity centres of thus obtained clusters 3. Iterate 1. and 2. until convergence @ ** ***
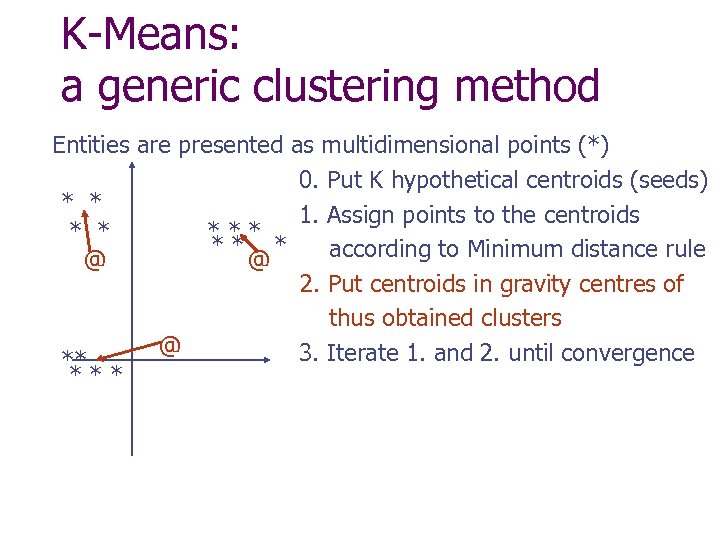

K-Means: a generic clustering method Entities are presented as multidimensional points (*) 0. Put K hypothetical centroids (seeds) * * 1. Assign points to the centroids * * ** * according to Minimum distance rule @ @ 2. Put centroids in gravity centres of thus obtained clusters @ 3. Iterate 1. and 2. until convergence ** ***
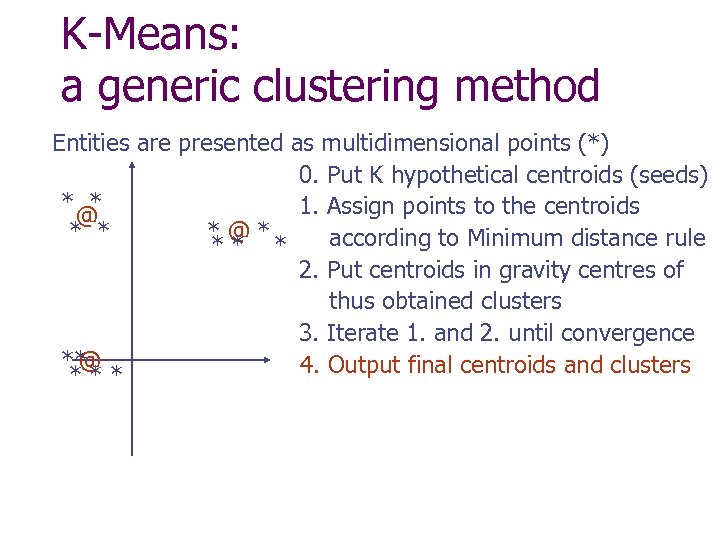

K-Means: a generic clustering method Entities are presented as multidimensional points (*) 0. Put K hypothetical centroids (seeds) * * 1. Assign points to the centroids @ * * *@* according to Minimum distance rule ** * 2. Put centroids in gravity centres of thus obtained clusters 3. Iterate 1. and 2. until convergence ** @ 4. Output final centroids and clusters ***

Advantages of K-Means Conventional: n Models typology building n Computationally effective n Can be incremental, `on-line’ Unconventional: n n Associates feature salience with feature scales and correlation/association Applicable to mixed scale data

Drawbacks of K-Means • No advice on: • Data pre-processing • Number of clusters • Initial setting • Instability of results • Criterion can be inadequate • Insufficient interpretation aids
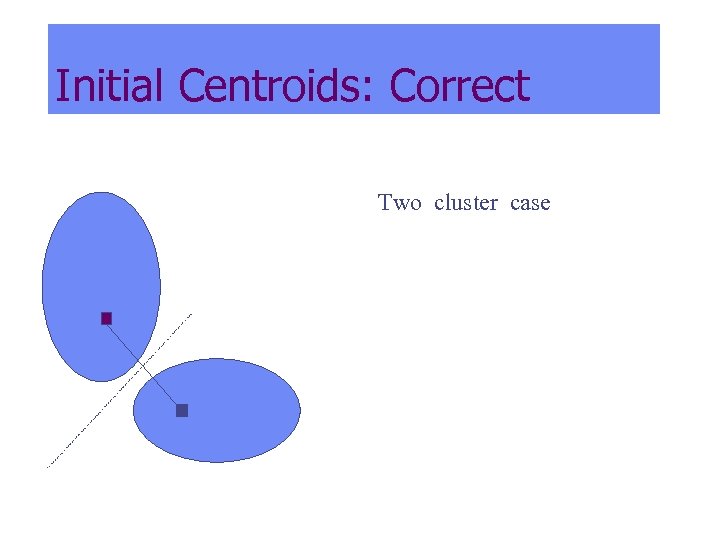
Initial Centroids: Correct Two cluster case
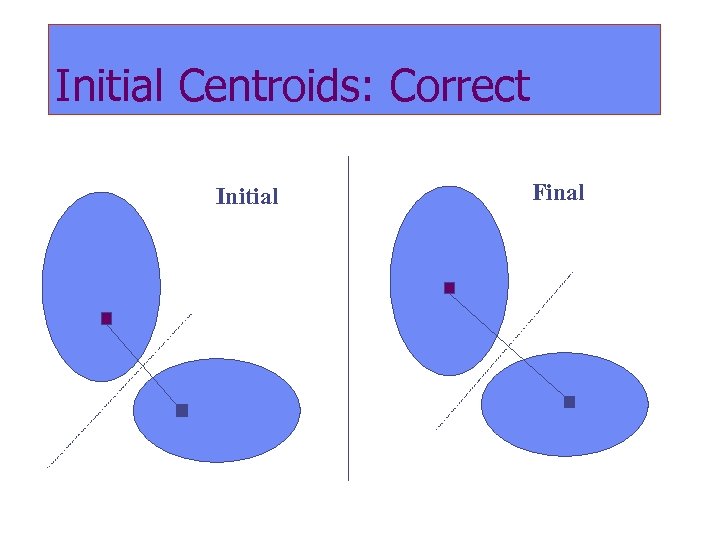
Initial Centroids: Correct Initial Final
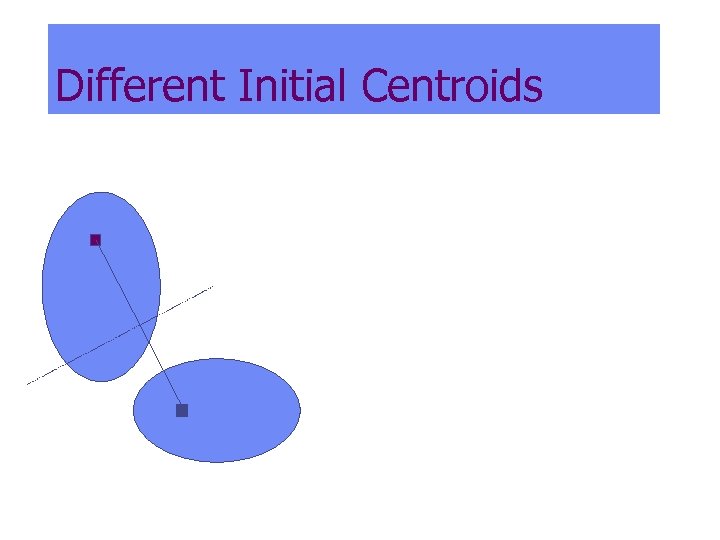
Different Initial Centroids
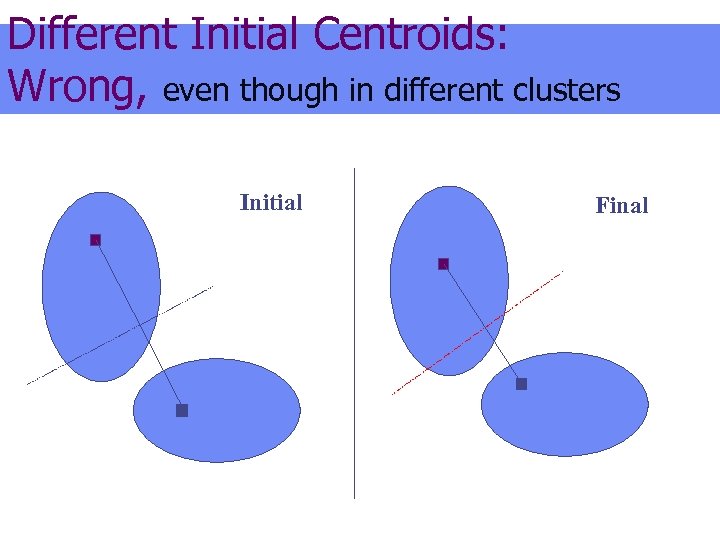
Different Initial Centroids: Wrong, even though in different clusters Initial Final

Two types of goals (with no clear-cut borderline) Engineering goals Data analysis goals

Engineering goals (examples) Devising a market segmentation to minimise the promotion and advertisement expenses Dividing a large scheme into modules to minimise the cost Organisation structure design

Data analysis goals (examples) Recovery of the distribution function Prediction Revealing patterns in data Enhancing knowledge with additional concepts and regularities Each of these is realised in a different perspective at clustering

Clustering Perspectives Classical statistics: Recovery of a multimodal distribution function Machine learning: Prediction Data mining: Revealing patterns in data Knowledge discovery: additional concepts and regularities

Clustering Perspectives at # Clusters Classical statistics: As many as meaningful modes (mixture items) Machine learning: As many as needed for acceptable prediction Data mining: As many as meaningful patterns in data (including incomplete clustering) Knowledge discovery: As many as needed to produce concepts and

Main Sources for Deriving # Clusters Classical statistics: Model of the world Machine learning: Cost & Accuracy Trade Off Data mining: Data Knowledge discovery: Domain knowledge

Classical Statistics Perspective There must be a model of data generation w E. g. , Mixture of Gaussians The task: identify all parameters of the model by using observed data w E. g. , The number of Gaussians and their probabilities, means and covariances
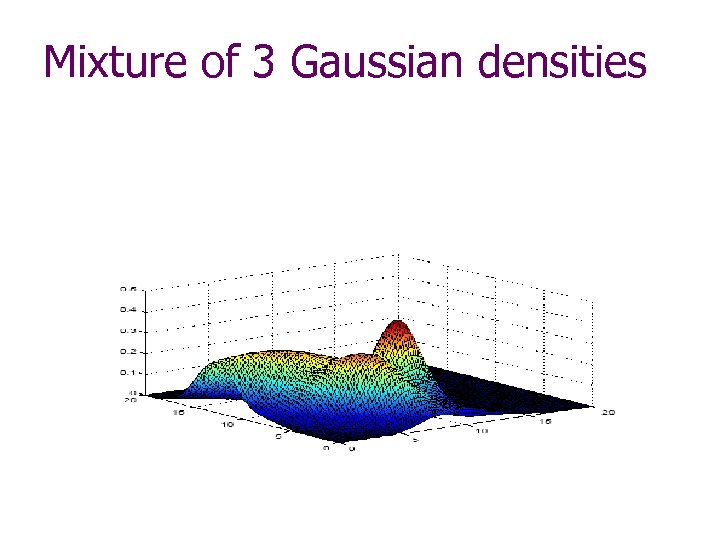
Mixture of 3 Gaussian densities

Classical statistics perspective on K-Means But a maximum likelihood method with spherical Gaussians of the same variance: - within a cluster, all variables are independent and Gaussian with the same cluster-independent variance (z - scoring is a must then); - the issue of the number of clusters can be approached with conventional approaches to hypothesis testing

Machine learning perspective Clusters should be of help in learning data incrementally generated The number should be specified by the trade-off between accuracy and cost A criterion should guarantee partitioning of the feature space with clearly separated high density areas; A method should be proven to be consistent with the criterion on the population

Machine learning on K-Means The number of clusters: to be specified according to prediction goals Pre-processing: no advice An incremental version of K-means converges to a minimum of the summary within-cluster variance, under conventional assumptions of data generation (Mc. Queen 1967 – major reference, though the method is traced a dozen or two years earlier)

Data mining perspective

Data recovery framework for data mining methods Type of Data n n Similarity Temporal Entity-to-feature Co-occurrence Type of Model n n n Regression Principal components Clusters Model: Data = Model_Derived_Data + Residual Pythagoras: Data 2 = Model_Derived_Data 2 + Residual 2 The better fit the better the model

K-Means as a data recovery method
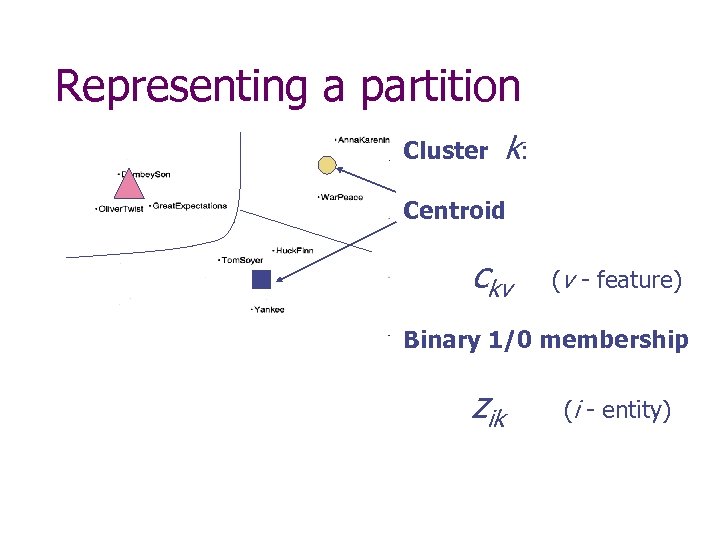
Representing a partition Cluster k: Centroid ckv (v - feature) Binary 1/0 membership zik (i - entity)
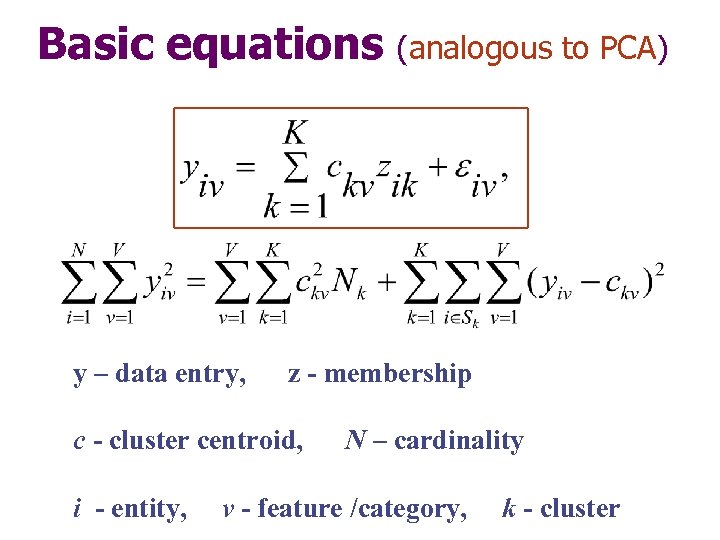

Basic equations y – data entry, z - membership c - cluster centroid, i - entity, (analogous to PCA) N – cardinality v - feature /category, k - cluster
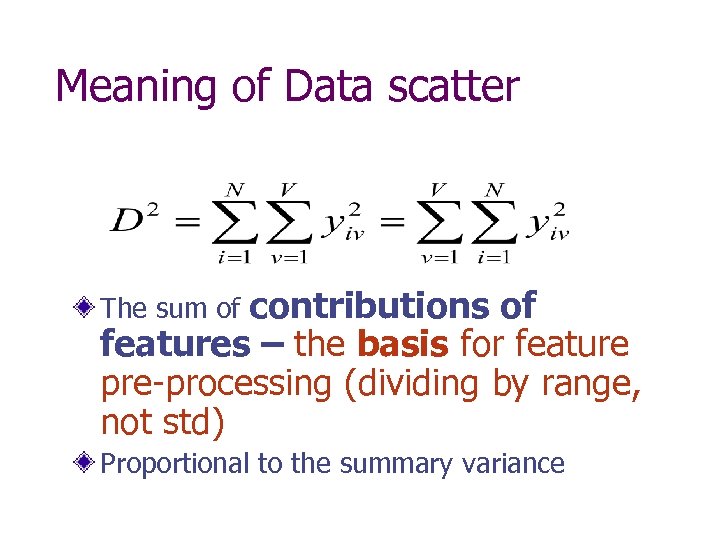

Meaning of Data scatter contributions of features – the basis for feature pre-processing (dividing by range, not std) The sum of Proportional to the summary variance
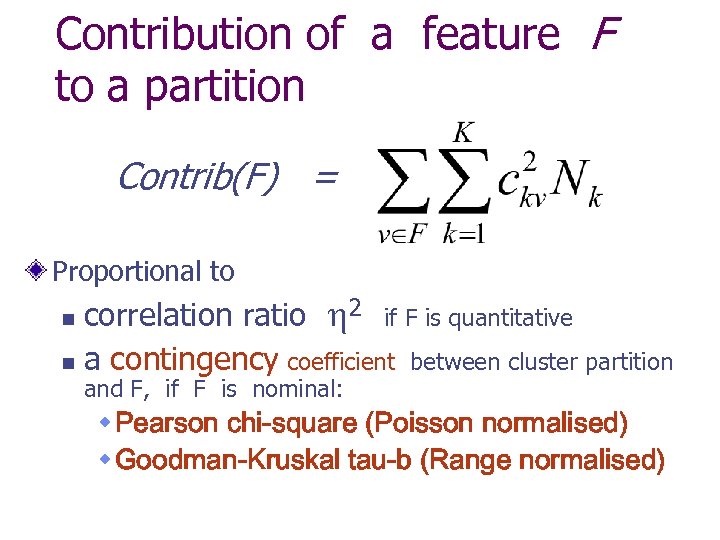

Contribution of a feature F to a partition Contrib(F) = Proportional to correlation ratio 2 if F is quantitative n a contingency coefficient between cluster partition n and F, if F is nominal: w Pearson chi-square (Poisson normalised) w Goodman-Kruskal tau-b (Range normalised)
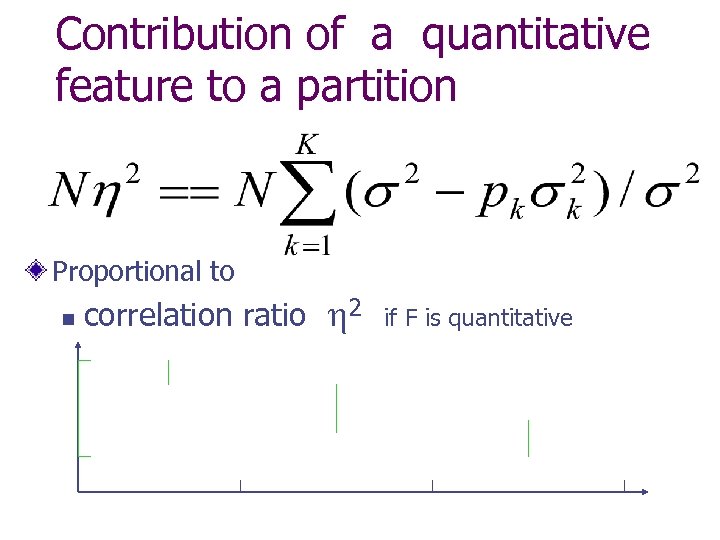

Contribution of a quantitative feature to a partition Proportional to n correlation ratio 2 if F is quantitative
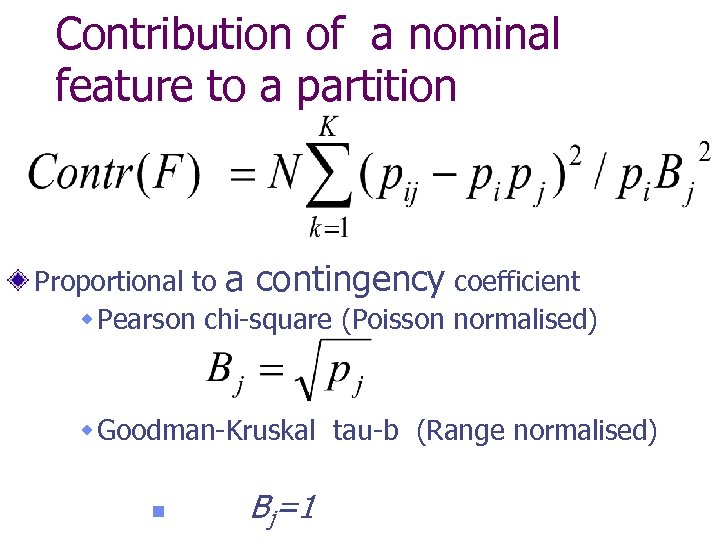

Contribution of a nominal feature to a partition Proportional to a contingency coefficient w Pearson chi-square (Poisson normalised) w Goodman-Kruskal tau-b (Range normalised) n Bj=1
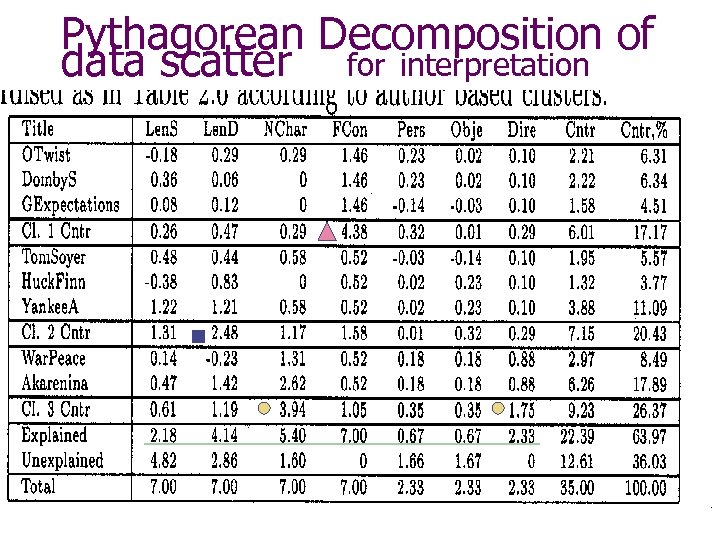
Pythagorean Decomposition of data scatter for interpretation
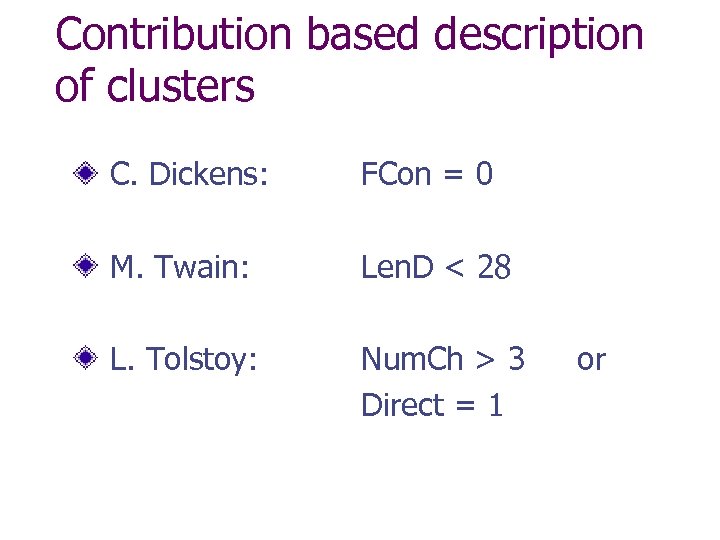

Contribution based description of clusters C. Dickens: FCon = 0 M. Twain: Len. D < 28 L. Tolstoy: Num. Ch > 3 Direct = 1 or
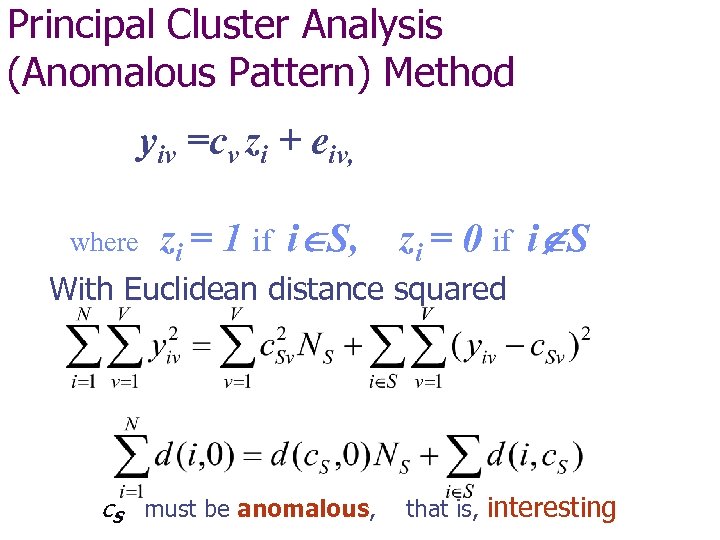

Principal Cluster Analysis (Anomalous Pattern) Method yiv =cv zi + eiv, where zi = 1 if i S, zi = 0 if i S With Euclidean distance squared c. S must be anomalous, that is, interesting
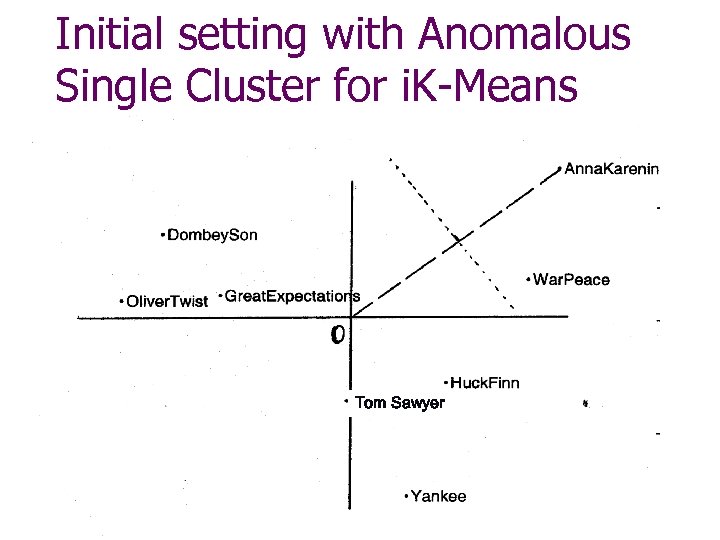
Initial setting with Anomalous Single Cluster for i. K-Means Tom Sawyer
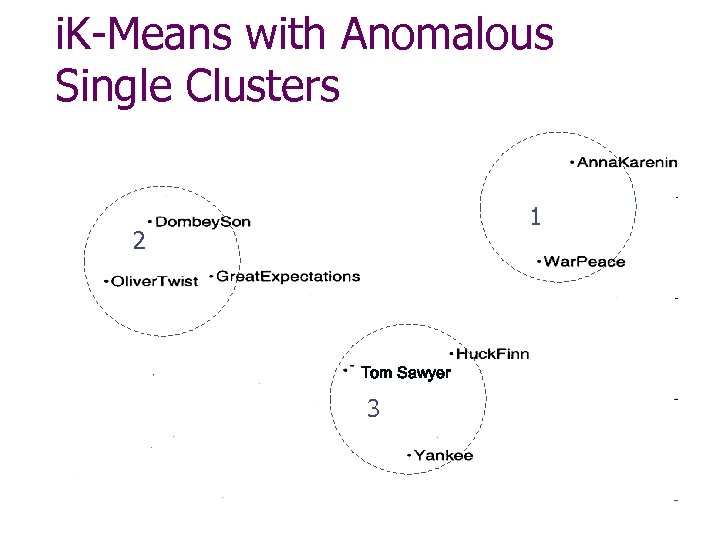
i. K-Means with Anomalous Single Clusters 1 2 0 Tom Sawyer 3
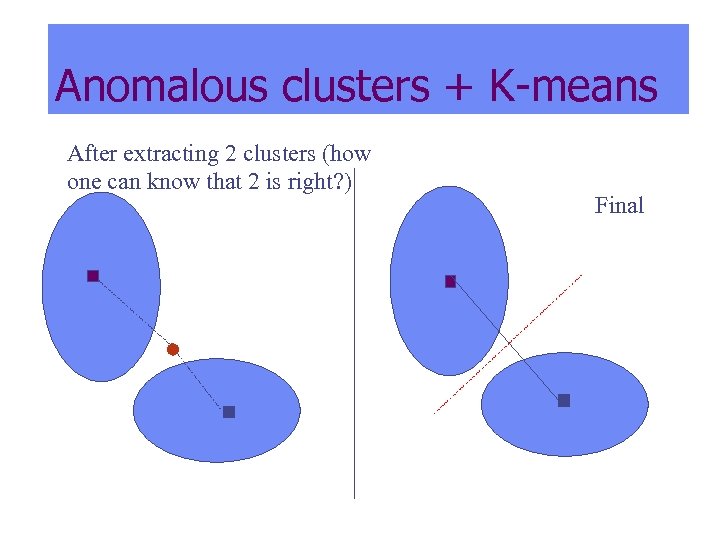
Anomalous clusters + K-means After extracting 2 clusters (how one can know that 2 is right? ) Final
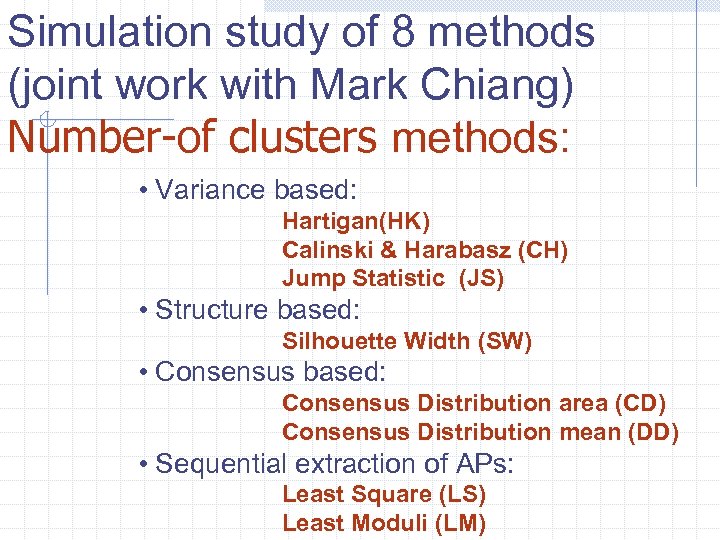

Simulation study of 8 methods (joint work with Mark Chiang) Number-of clusters methods: • Variance based: Hartigan(HK) Calinski & Harabasz (CH) Jump Statistic (JS) • Structure based: Silhouette Width (SW) • Consensus based: Consensus Distribution area (CD) Consensus Distribution mean (DD) • Sequential extraction of APs: Least Square (LS) Least Moduli (LM)
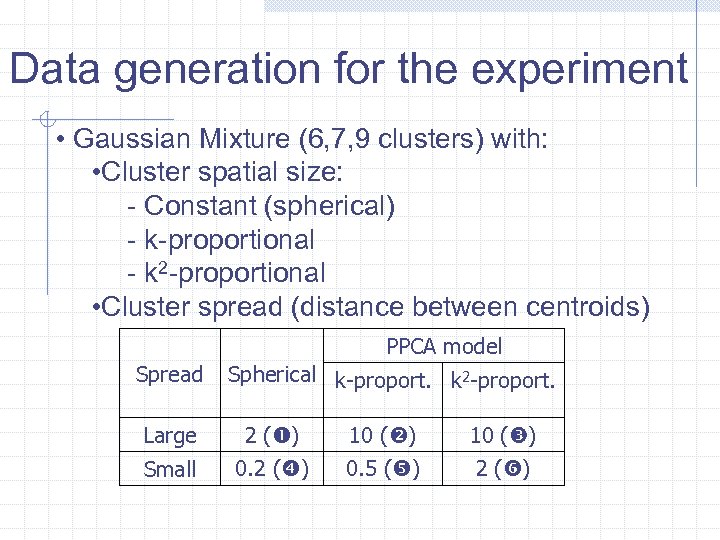

Data generation for the experiment • Gaussian Mixture (6, 7, 9 clusters) with: • Cluster spatial size: - Constant (spherical) - k-proportional - k 2 -proportional • Cluster spread (distance between centroids) PPCA model Spread Spherical k-proport. k 2 -proport. Large 2 ( ) 10 ( ) Small 0. 2 ( ) 0. 5 ( ) 2 ( )

Evaluation of results: Estimated clustering versus that generated • Number of clusters • Distance between centroids • Similarity between partitions
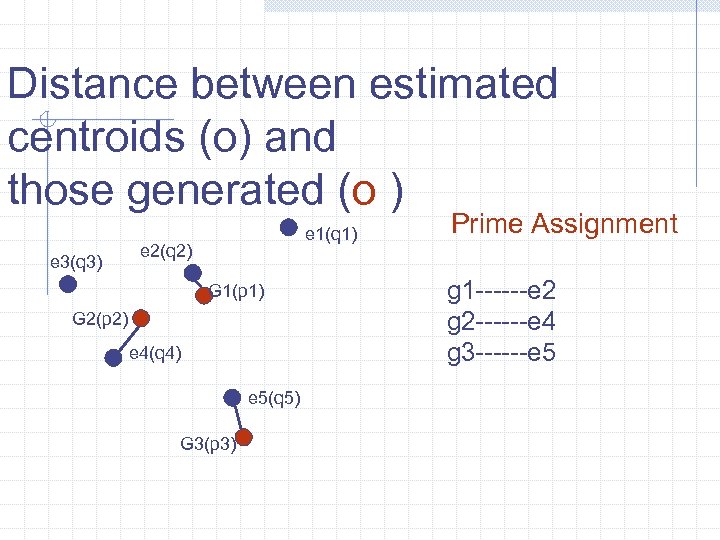

Distance between estimated centroids (o) and those generated (o ) e 3(q 3) e 1(q 1) e 2(q 2) G 1(p 1) G 2(p 2) e 4(q 4) e 5(q 5) G 3(p 3) Prime Assignment g 1 ------e 2 g 2 ------e 4 g 3 ------e 5
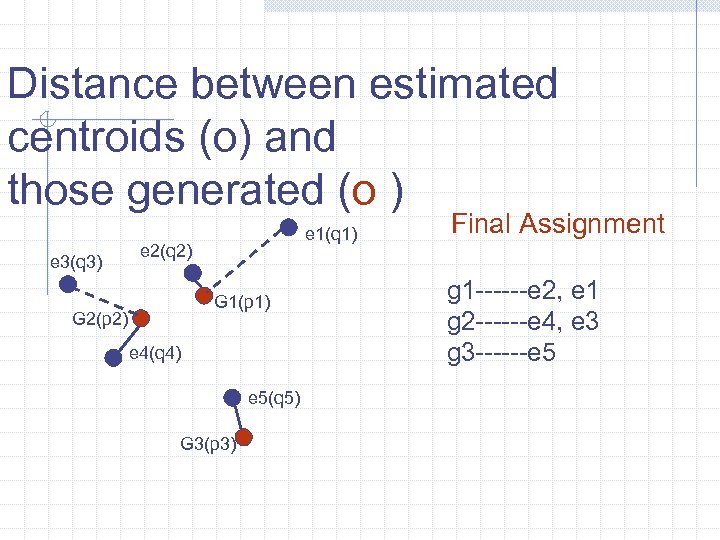

Distance between estimated centroids (o) and those generated (o ) e 3(q 3) e 1(q 1) e 2(q 2) G 1(p 1) G 2(p 2) e 4(q 4) e 5(q 5) G 3(p 3) Final Assignment g 1 ------e 2, e 1 g 2 ------e 4, e 3 g 3 ------e 5
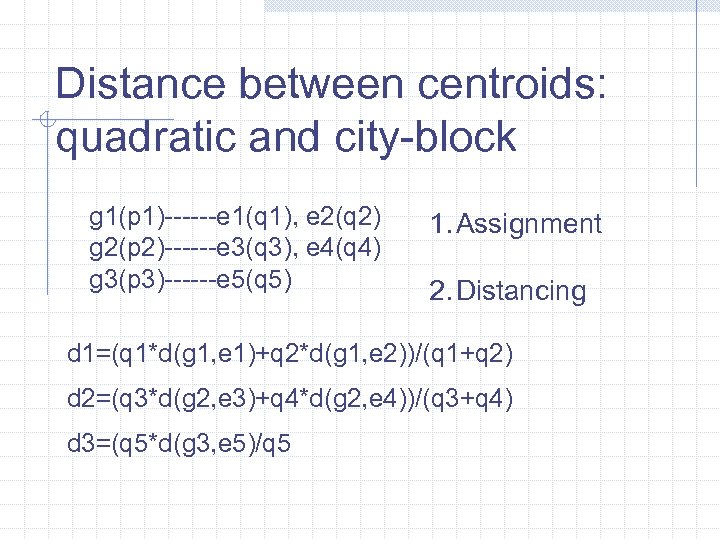

Distance between centroids: quadratic and city-block g 1(p 1)------e 1(q 1), e 2(q 2) g 2(p 2)------e 3(q 3), e 4(q 4) g 3(p 3)------e 5(q 5) 1. Assignment 2. Distancing d 1=(q 1*d(g 1, e 1)+q 2*d(g 1, e 2))/(q 1+q 2) d 2=(q 3*d(g 2, e 3)+q 4*d(g 2, e 4))/(q 3+q 4) d 3=(q 5*d(g 3, e 5)/q 5

Distance between centroids: quadratic and city-block p 1*d 1+p 2*d 2+p 3*d 3 1. Assignment 2. Distancing 3. Averaging

Similarity between partitions according to their confusion table • Relative distance (Mirkin-Cherny 1970) • Tchouprov coefficient (Cramer 1943) • Adjusted Rand Index (Arabie-Hubert, 1985) • Average Overlap (Mirkin 2005)
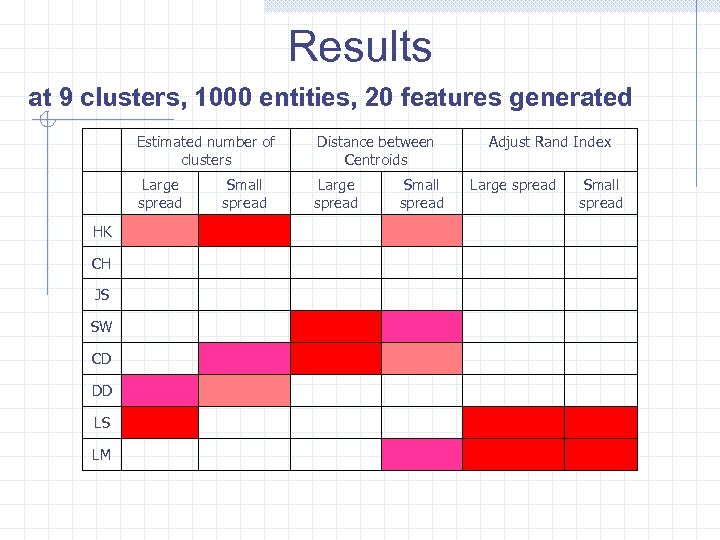
Results at 9 clusters, 1000 entities, 20 features generated Estimated number of clusters Large spread HK CH JS SW CD DD LS LM Distance between Centroids Large spread Small spread Adjust Rand Index Large spread Small spread

Knowledge discovery perspective on clustering Conforming to and enhancing domain knowledge: Informal considerations so far Relevant items: n n Decision trees External validation

A case to generalise Entities with a similarity measure Clustering interpretation tool developed Clustering method using a similarity threshold leading to a number of clusters Domain knowledge leading to constraints to the similarity threshold Best fitting interpretation provides for the best number of clusters
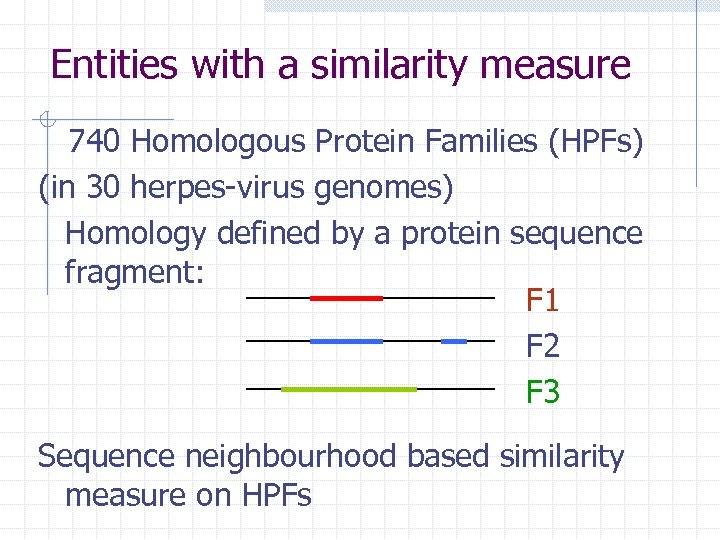

Entities with a similarity measure 740 Homologous Protein Families (HPFs) (in 30 herpes-virus genomes) Homology defined by a protein sequence fragment: F 1 F 2 F 3 Sequence neighbourhood based similarity measure on HPFs
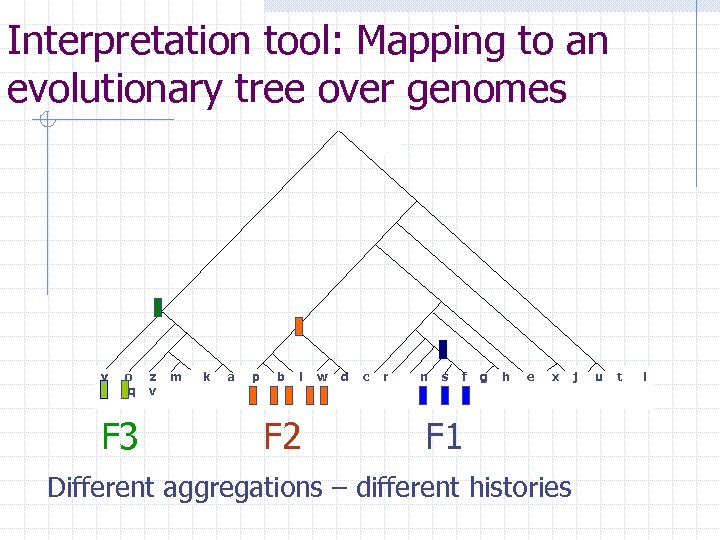
Interpretation tool: Mapping to an evolutionary tree over genomes y o q F 3 z v m k a p b l F 2 w d c r n s f g h e x F 1 Different aggregations – different histories j u t i

Algorithm: ADDI-S (Mirkin Jo. C 1987), a data approximation technique To maximize Contribution to Data Scatter, Average within-cluster similarity c multiplied by the cluster’s size #S Algorithm ADDI-S: n n n Take S={ j } for arbitrary j Given S, find c=c(S) and similarities b(i, S) to S for all entities i in and out of S; Check the differences b(i, S)-c/2. If they are consistent, change the state of a most contributing entity. Else, stop and output S. Resulting S: a tightness property. Holzinger (1941) B-coefficient, Arkadiev&Braverman (1964, 1967) Specter, Mirkin (1976, 1987) ADDI family, Ben-Dor, Shamir, Yakhini (1999) CAST
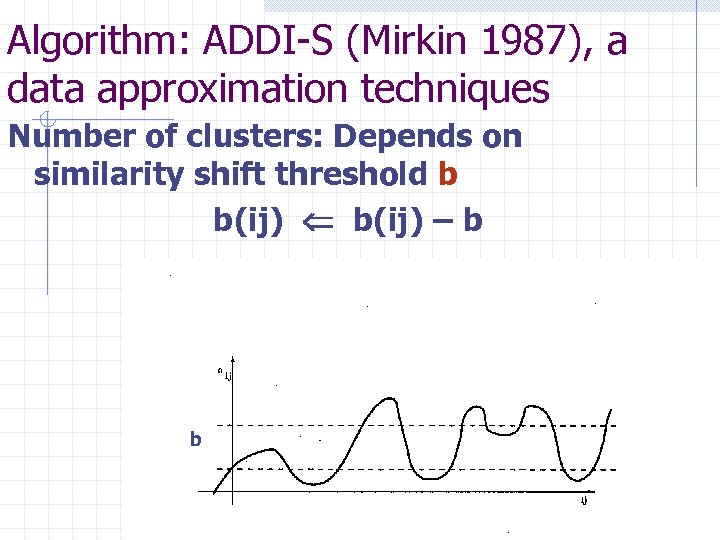
Algorithm: ADDI-S (Mirkin 1987), a data approximation techniques Number of clusters: Depends on similarity shift threshold b b(ij) – b b
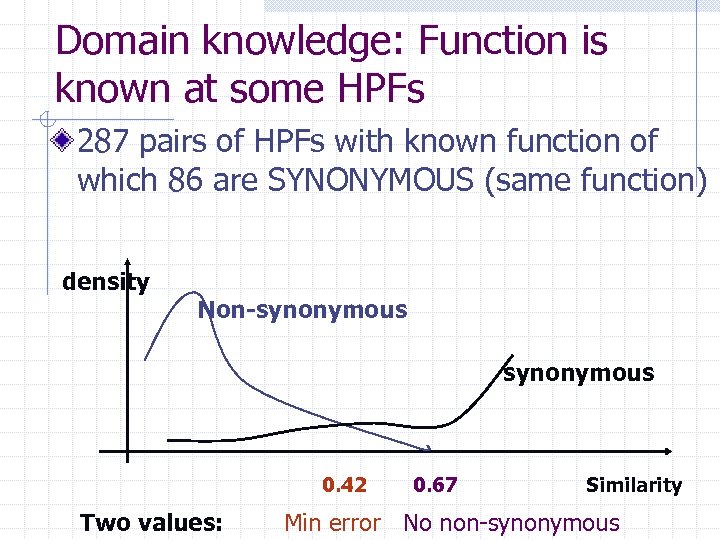
Domain knowledge: Function is known at some HPFs 287 pairs of HPFs with known function of which 86 are SYNONYMOUS (same function) density Non-synonymous 0. 42 Two values: 0. 67 Similarity Min error No non-synonymous
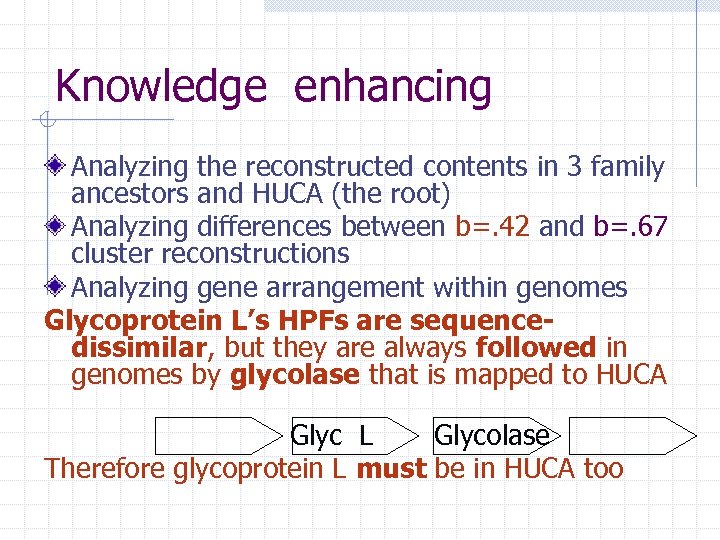

Knowledge enhancing Analyzing the reconstructed contents in 3 family ancestors and HUCA (the root) Analyzing differences between b=. 42 and b=. 67 cluster reconstructions Analyzing gene arrangement within genomes Glycoprotein L’s HPFs are sequencedissimilar, but they are always followed in genomes by glycolase that is mapped to HUCA Glyc L Glycolase Therefore glycoprotein L must be in HUCA too

Final HPFs and APFs HPFs with a sequence-based similarity measure Interpretation: parsimonious histories Clustering ADDI-S using a similarity threshold leading to a number of clusters Domain knowledge: 86 pairs should be in same clusters, and 201 in different clusters 2 suggested similarity thresholds Best fitting: 102 APF (aggregating 249 HPFs) and 491 singleton HPFs
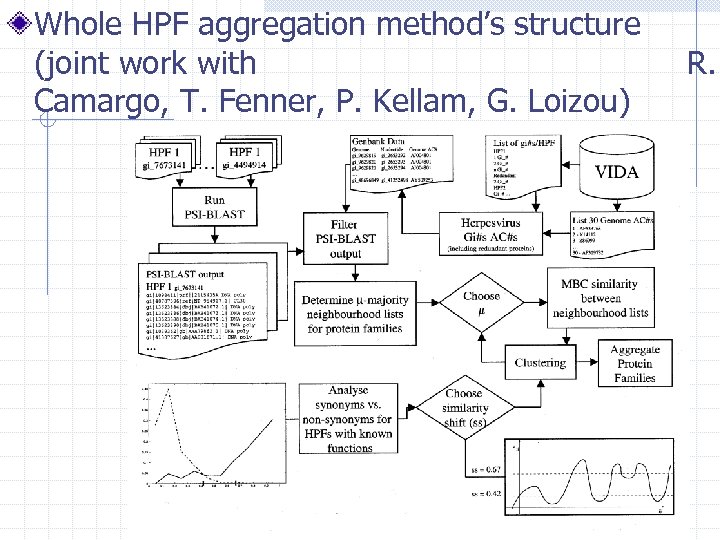

Whole HPF aggregation method’s structure (joint work with Camargo, T. Fenner, P. Kellam, G. Loizou) R.

Conclusion I: Number of clusters? Engineering perspective: defined by cost/effect Classical statistics perspective: can and should be determined from data with a model Machine learning perspective: can be specified according to the prediction accuracy to achieve Data mining perspective: not to pre-specify; only those are of interest that bear interesting patterns Knowledge discovery perspective: not to prespecify; those that are best in knowledge enhancing

Conclusion II: Each other data analysis concept Classical statistics perspective: can be determined from data with a model Machine learning perspective: prediction accuracy to achieve Data mining perspective: data approximation Knowledge discovery perspective: knowledge enhancing
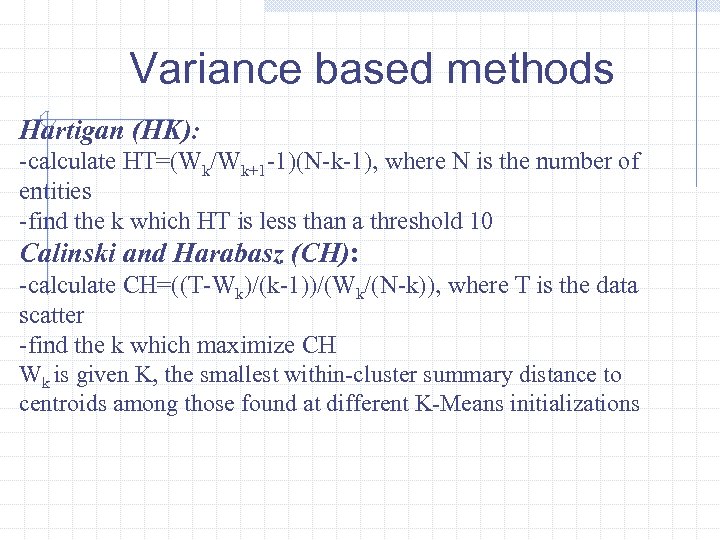

Variance based methods Hartigan (HK): -calculate HT=(Wk/Wk+1 -1)(N-k-1), where N is the number of entities -find the k which HT is less than a threshold 10 Calinski and Harabasz (CH): -calculate CH=((T-Wk)/(k-1))/(Wk/(N-k)), where T is the data scatter -find the k which maximize CH Wk is given K, the smallest within-cluster summary distance to centroids among those found at different K-Means initializations
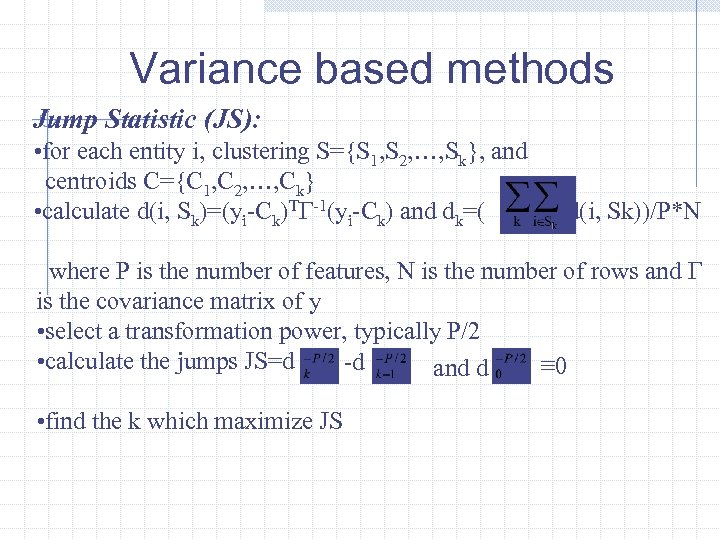

Variance based methods Jump Statistic (JS): • for each entity i, clustering S={S 1, S 2, …, Sk}, and centroids C={C 1, C 2, …, Ck} • calculate d(i, Sk)=(yi-Ck)TΓ-1(yi-Ck) and dk=( d(i, Sk))/P*N where P is the number of features, N is the number of rows and Γ is the covariance matrix of y • select a transformation power, typically P/2 • calculate the jumps JS=d -d ≡ 0 and d • find the k which maximize JS
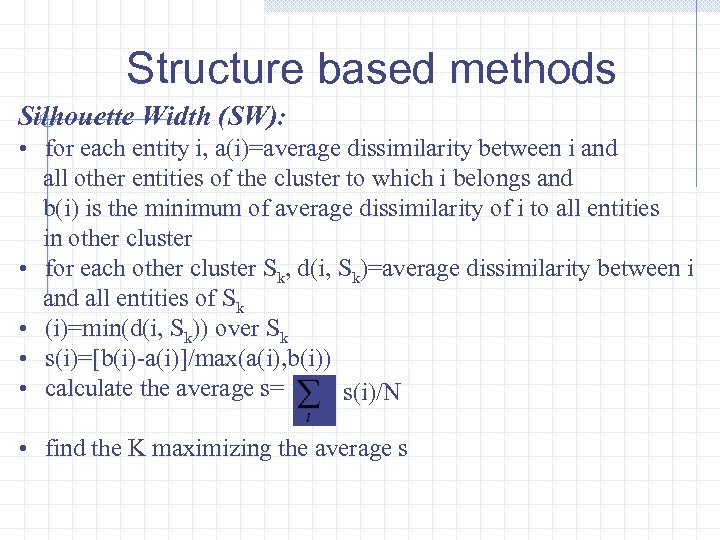

Structure based methods Silhouette Width (SW): • for each entity i, a(i)=average dissimilarity between i and all other entities of the cluster to which i belongs and b(i) is the minimum of average dissimilarity of i to all entities in other cluster • for each other cluster Sk, d(i, Sk)=average dissimilarity between i and all entities of Sk • (i)=min(d(i, Sk)) over Sk • s(i)=[b(i)-a(i)]/max(a(i), b(i)) • calculate the average s= s(i)/N • find the K maximizing the average s
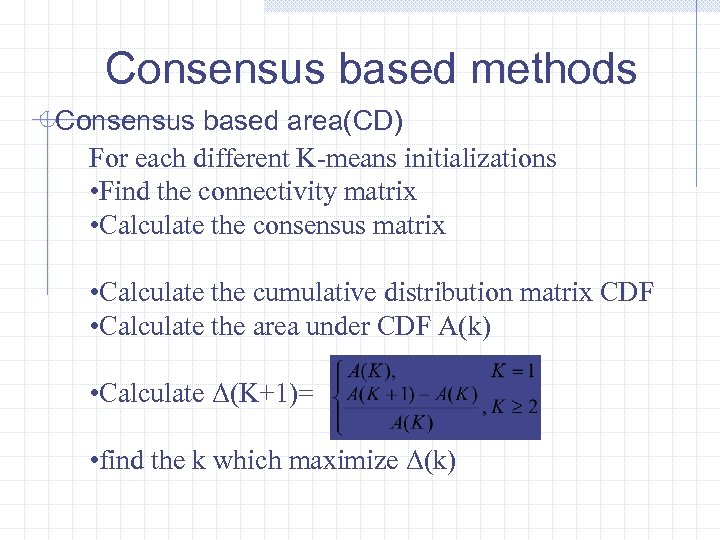

Consensus based methods Consensus based area(CD) For each different K-means initializations • Find the connectivity matrix • Calculate the consensus matrix • Calculate the cumulative distribution matrix CDF • Calculate the area under CDF A(k) • Calculate Δ(K+1)= • find the k which maximize Δ(k)

Consensus based methods μK is the mean of the consensus matrix σK is the variance of the consensus matrix avdis(K)= μK*(1 - μK)- σK 2 davdis(K)=(avdis(K)-avdis(K+1))/avdis(K+1) Find the K which maximize davdis(DD)

Sequential cluster extraction Intelligent K-Means: • Anomalous Pattern (Initial clusters) • Removal of singletons • K-Means • Euclidean Distance + The within-cluster mean Least Square Criterion (LS) • Manhattan Distance + The within-cluster median Least Modules Criterion (LM)
0ba21210f7b7f7938212a9b67c60b7a1.ppt