39d44a0aad81942c575b38d5c8a0e745.ppt
- Количество слайдов: 71
Control of Eukaryotic Genes AP Biology 2007 -2008
The BIG Questions… How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions? AP Biology

Evolution of gene regulation Prokaryotes single-celled u evolved to grow & divide rapidly u must respond quickly to changes in external environment u exploit transient resources Gene regulation u turn genes on & off rapidly u AP Biology flexibility & reversibility adjust levels of enzymes for synthesis & digestion

Evolution of gene regulation Eukaryotes multicellular u evolved to maintain constant internal conditions while facing changing external conditions u u homeostasis regulate body as a whole growth & development w long term processes specialization w turn on & off large number of genes AP Biology must coordinate the body as a whole rather than serve the needs of individual cells
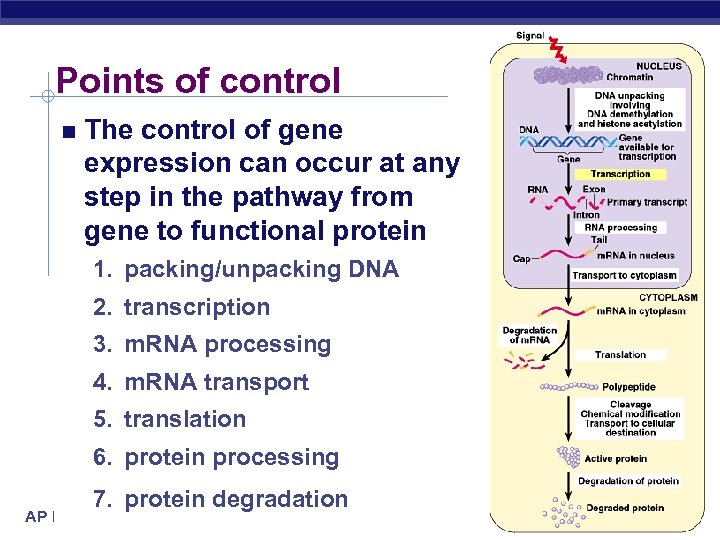
Points of control The control of gene expression can occur at any step in the pathway from gene to functional protein 1. packing/unpacking DNA 2. transcription 3. m. RNA processing 4. m. RNA transport 5. translation 6. protein processing 7. protein degradation AP Biology
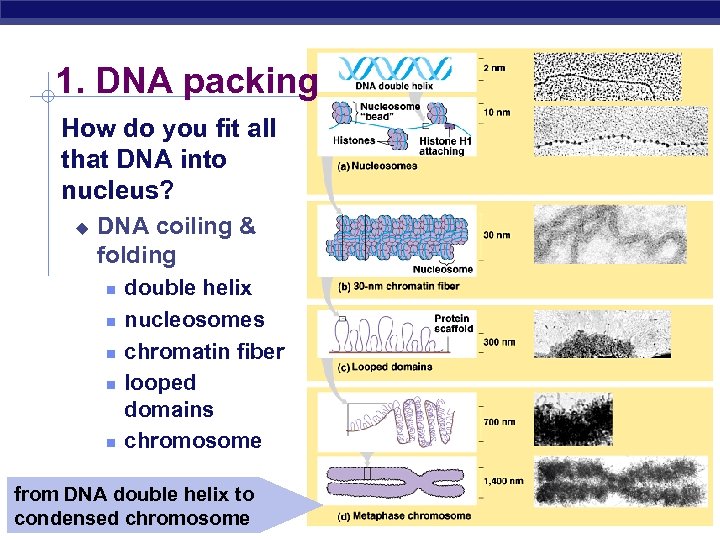
1. DNA packing How do you fit all that DNA into nucleus? u DNA coiling & folding double helix nucleosomes chromatin fiber looped domains chromosome from DNA double helix to AP Biology condensed chromosome
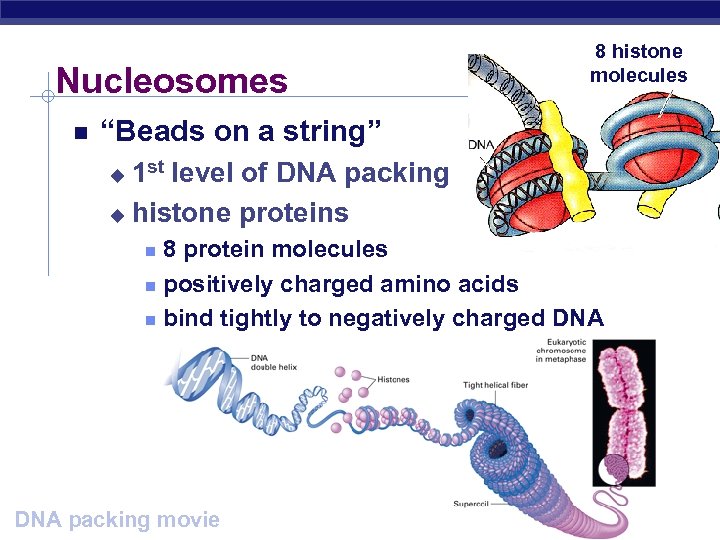
Nucleosomes 8 histone molecules “Beads on a string” 1 st level of DNA packing u histone proteins u 8 protein molecules positively charged amino acids bind tightly to negatively charged DNA AP Biology DNA packing movie
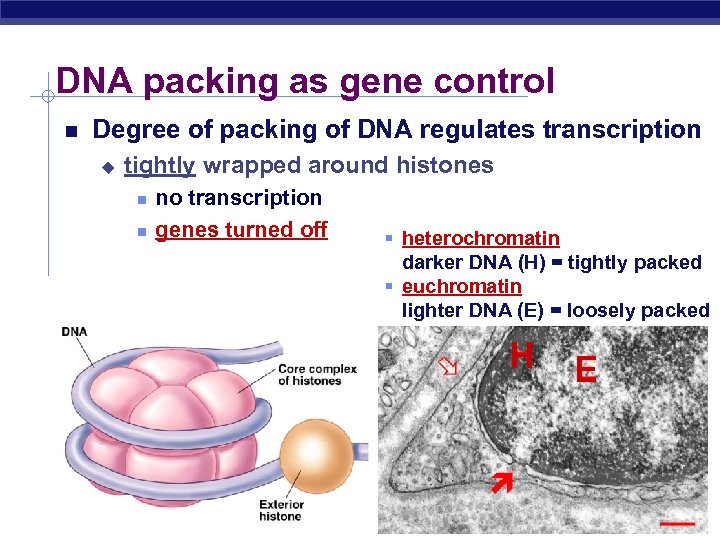
DNA packing as gene control Degree of packing of DNA regulates transcription u tightly wrapped around histones no transcription genes turned off § heterochromatin darker DNA (H) = tightly packed § euchromatin lighter DNA (E) = loosely packed H AP Biology E
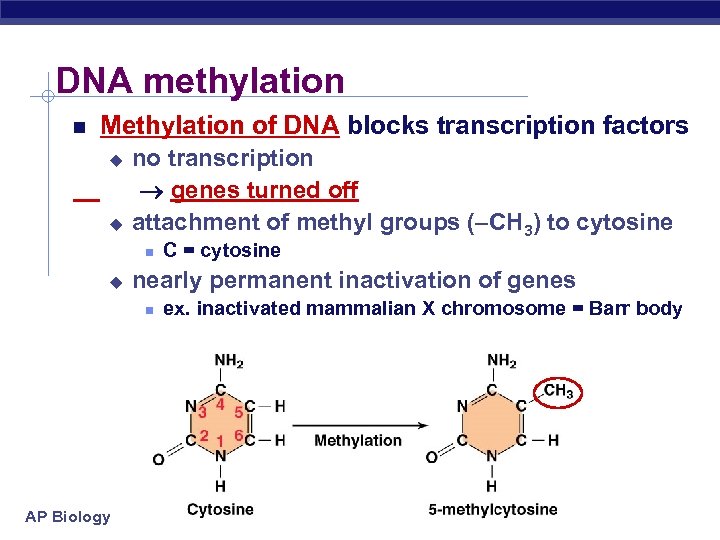
DNA methylation Methylation of DNA blocks transcription factors no transcription genes turned off u attachment of methyl groups (–CH 3) to cytosine u u nearly permanent inactivation of genes AP Biology C = cytosine ex. inactivated mammalian X chromosome = Barr body
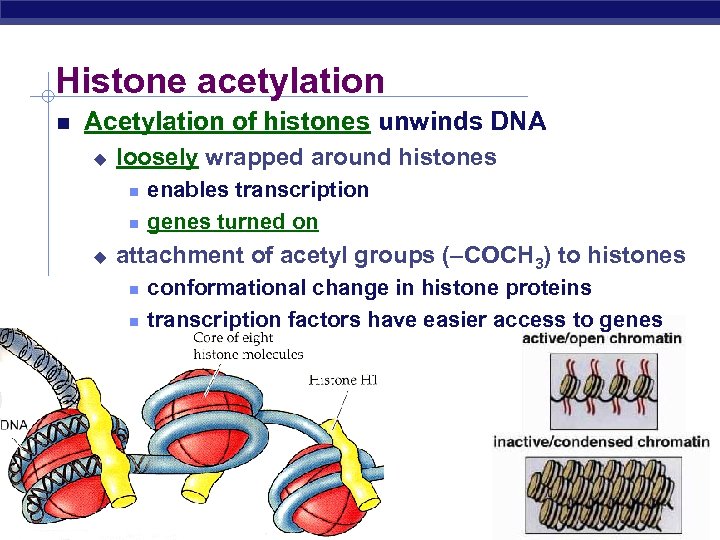
Histone acetylation Acetylation of histones unwinds DNA u loosely wrapped around histones u attachment of acetyl groups (–COCH 3) to histones AP Biology enables transcription genes turned on conformational change in histone proteins transcription factors have easier access to genes
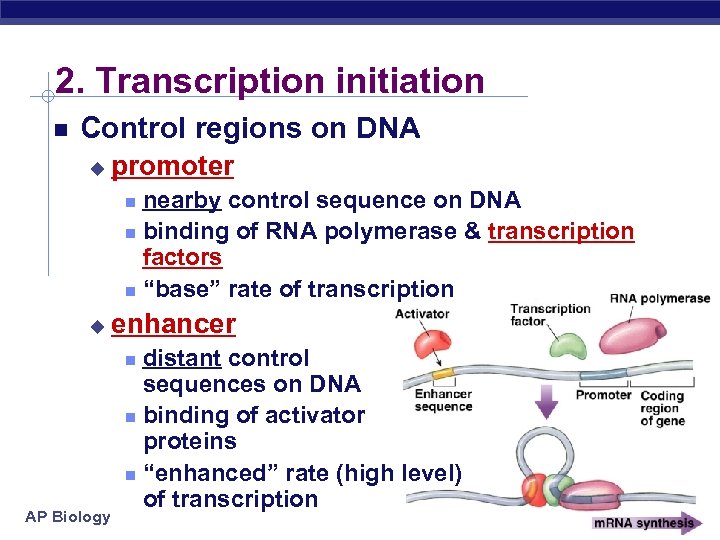
2. Transcription initiation Control regions on DNA u promoter nearby control sequence on DNA binding of RNA polymerase & transcription factors “base” rate of transcription u enhancer distant control sequences on DNA binding of activator proteins “enhanced” rate (high level) of transcription AP Biology
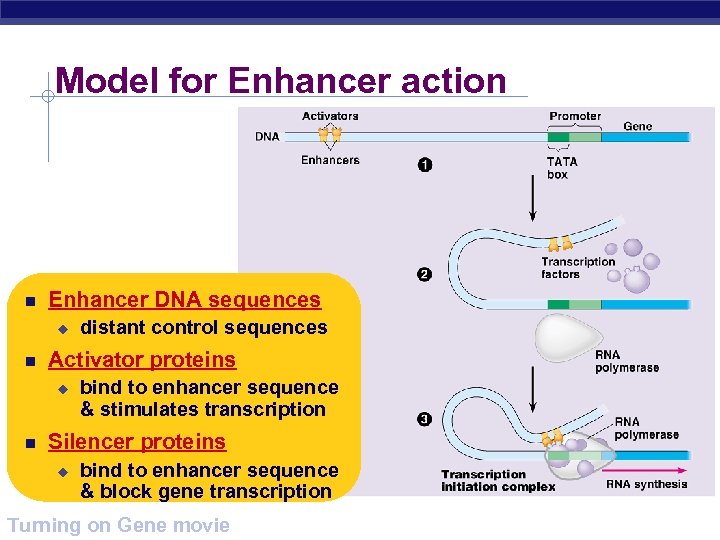
Model for Enhancer action Enhancer DNA sequences u Activator proteins u distant control sequences bind to enhancer sequence & stimulates transcription Silencer proteins u bind to enhancer sequence & block gene transcription AP Biology Turning on Gene movie
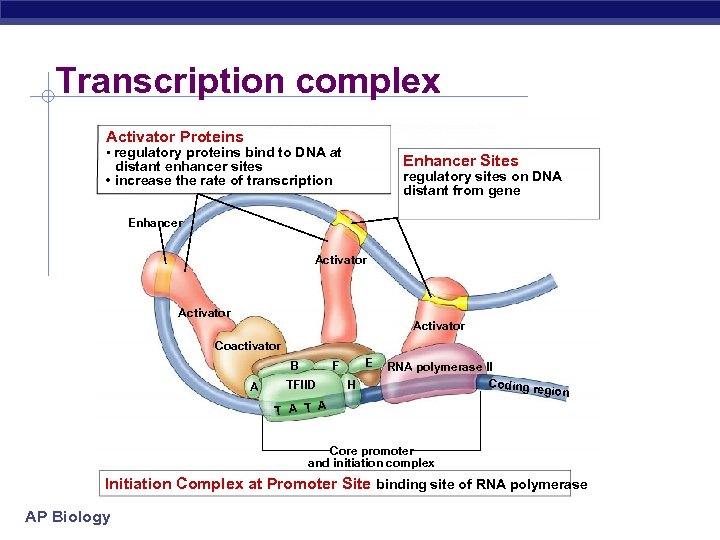
Transcription complex Activator Proteins • regulatory proteins bind to DNA at Enhancer Sites distant enhancer sites • increase the rate of transcription regulatory sites on DNA distant from gene Enhancer Activator Coactivator A E F B TFIID H RNA polymerase II Coding r egion T A Core promoter and initiation complex Initiation Complex at Promoter Site binding site of RNA polymerase AP Biology
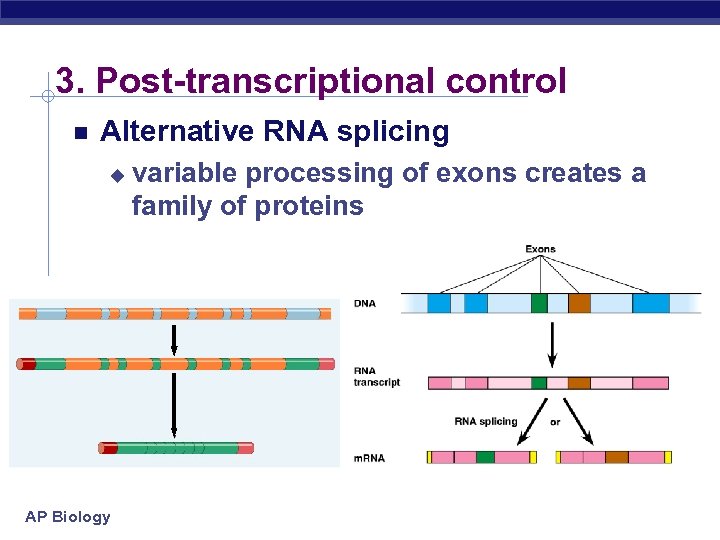
3. Post-transcriptional control Alternative RNA splicing u AP Biology variable processing of exons creates a family of proteins
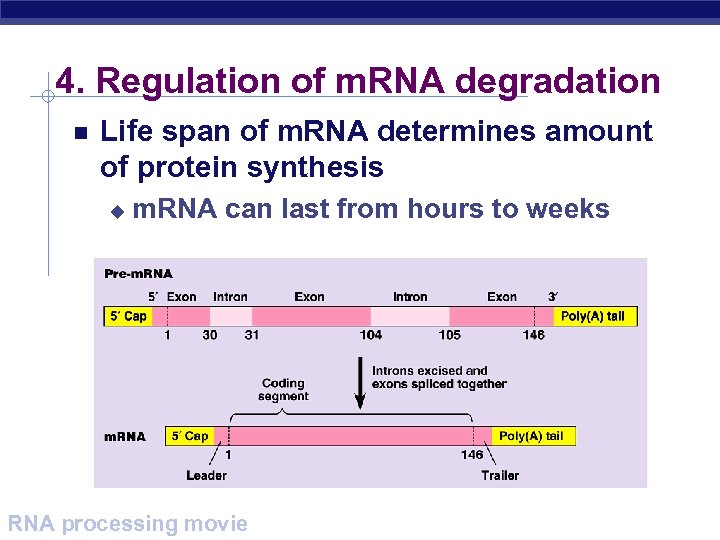
4. Regulation of m. RNA degradation Life span of m. RNA determines amount of protein synthesis u m. RNA can last from hours to weeks AP Biology RNA processing movie

RNA interference NEW Small interfering RNAs (si. RNA) u short segments of RNA (21 -28 bases) bind to m. RNA create sections of double-stranded m. RNA “death” tag for m. RNA w triggers degradation of m. RNA u cause gene “silencing” post-transcriptional control turns off gene = no protein produced si. RNA AP Biology !
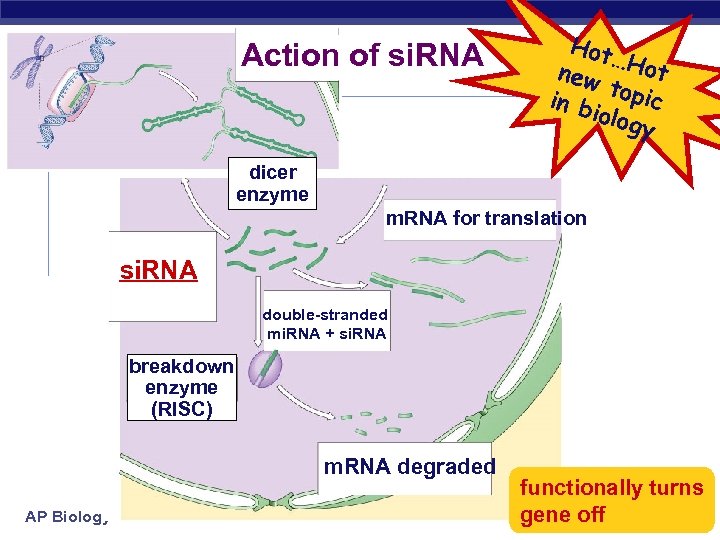
Action of si. RNA Hot … new Hot t in b opic iolog y dicer enzyme m. RNA for translation si. RNA double-stranded mi. RNA + si. RNA breakdown enzyme (RISC) m. RNA degraded AP Biology functionally turns gene off
si. RNA clip https: //www. youtube. com/watch? v=Fa 4 sk YBJHo. I AP Biology
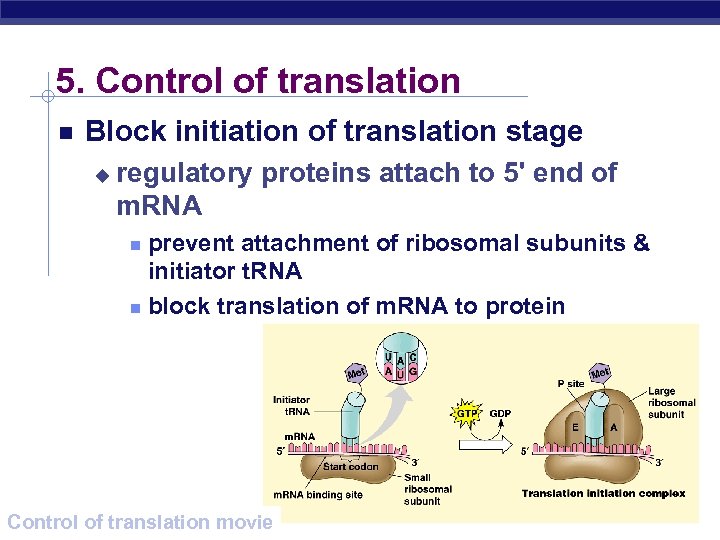
5. Control of translation Block initiation of translation stage u regulatory proteins attach to 5' end of m. RNA prevent attachment of ribosomal subunits & initiator t. RNA block translation of m. RNA to protein AP Biology Control of translation movie
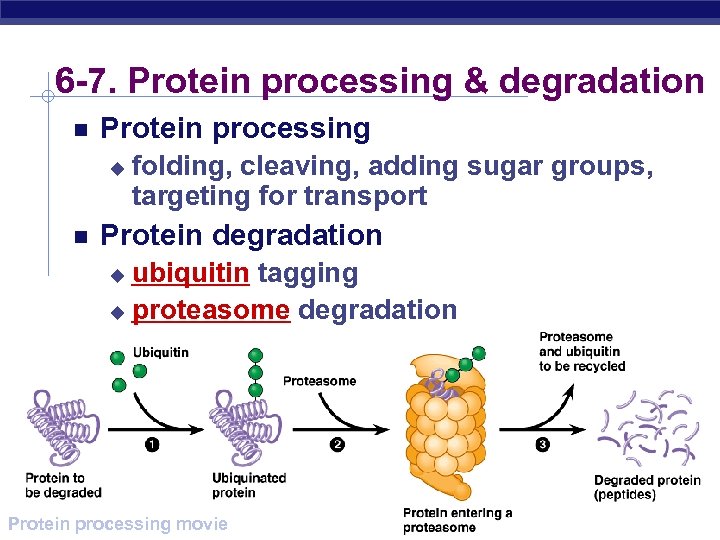
6 -7. Protein processing & degradation Protein processing u folding, cleaving, adding sugar groups, targeting for transport Protein degradation ubiquitin tagging u proteasome degradation u AP Biology Protein processing movie

1980 s | 2004 Ubiquitin “Death tag” mark unwanted proteins with a label u 76 amino acid polypeptide, ubiquitin u labeled proteins are broken down rapidly in "waste disposers" u proteasomes Aaron Ciechanover AP Biology Israel Avram Hershko Israel Irwin Rose UC Riverside
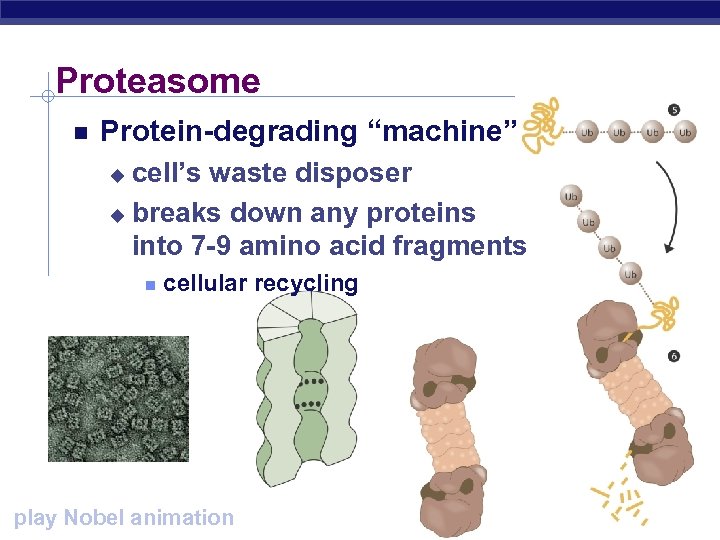
Proteasome Protein-degrading “machine” cell’s waste disposer u breaks down any proteins into 7 -9 amino acid fragments u cellular recycling AP Biology play Nobel animation

CENTRAL DOGMA Genetic information always goes from DNA to RNA to protein Gene regulation has been well studied in E. coli When a bacterial cell encounters a potential food source it will manufacture the enzymes necessary to metabolize that food AP Biology

CENTRAL DOGMA Genetic information always goes from DNA to RNA to protein Gene regulation has been well studied in E. coli When a bacterial cell encounters a potential food source it will manufacture the enzymes necessary to metabolize that food AP Biology

Gene Regulation In addition to sugars like glucose and lactose E. coli cells also require amino acids One essential aa is tryptophan. When E. coli is swimming in tryptophan (milk & poultry) it will absorb the amino acids from the media When tryptophan is not present in the media then the cell must manufacture its’ own amino acids AP Biology

Trp Operon E. coli uses several proteins encoded by a cluster of 5 genes to manufacture the amino acid tryptophan All 5 genes are transcribed together as a unit called an operon, which produces a single long piece of m. RNA for all the genes RNA polymerase binds to a promoter located at the beginning of the first gene and proceeds down the DNA transcribing the genes in sequence AP Biology
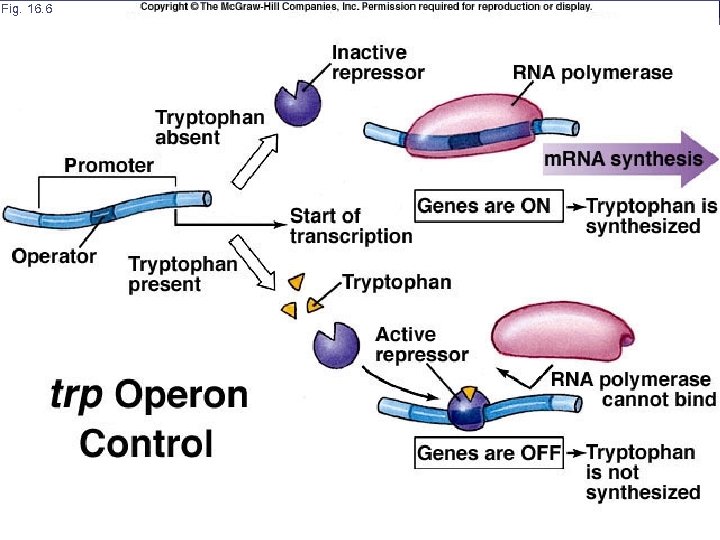
Fig. 16. 6 AP Biology

GENE REGULATION In addition to amino acids, E. coli cells also metabolize sugars in their environment In 1959 Jacques Monod and Fracois Jacob looked at the ability of E. coli cells to digest the sugar lactose AP Biology

GENE REGULATION In the presence of the sugar lactose, E. coli makes an enzyme called beta galactosidase Beta galactosidase breaks down the sugar lactose so the E. coli can digest it for food It is the LAC Z gene in E coli that codes for the enzyme beta galactosidase AP Biology

Lac Z Gene The tryptophan gene is turned on when there is no tryptophan in the media That is when the cell wants to make its’ own tryptophan E. coli cells can not make the sugar lactose They can only have lactose when it is present in their environment Then they turn on genes to beak down lactose AP Biology

GENE REGULATION The E. coli bacteria only needs beta galactosidase if there is lactose in the environment to digest There is no point in making the enzyme if there is no lactose sugar to break down It is the combination of the promoter and the DNA that regulate when a gene will be transcribed AP Biology
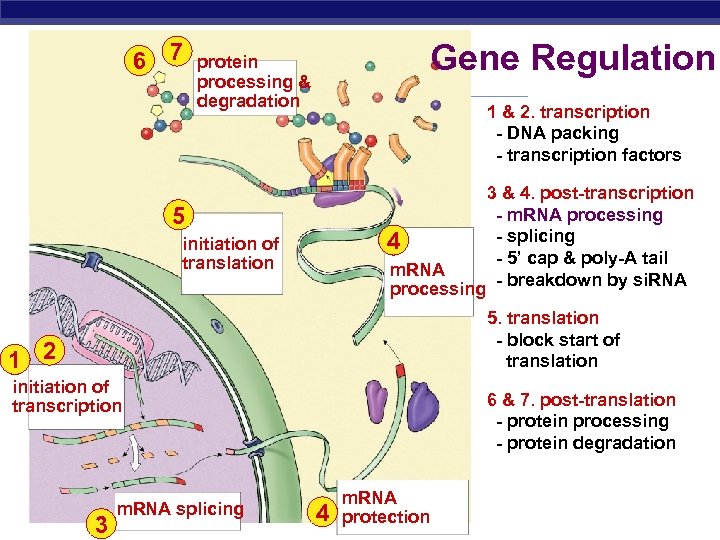
6 7 Gene Regulation protein processing & degradation 1 & 2. transcription - DNA packing - transcription factors 5 4 initiation of translation m. RNA processing 3 & 4. post-transcription - m. RNA processing - splicing - 5’ cap & poly-A tail - breakdown by si. RNA 5. translation - block start of translation 1 2 initiation of transcription AP Biology m. RNA splicing 3 6 & 7. post-translation - protein processing - protein degradation 4 m. RNA protection

GENE REGULATION This combination of a promoter and a gene is called an OPERON Operon is a cluster of genes encoding related enzymes that are regulated together AP Biology

GENE REGULATION Operon consists of A promoter site where RNA polyerase binds and begins transcribing the message u A region that makes a repressor u Repressor sits on the DNA at a spot between the promoter and the gene to be transcribed This site is called the operator AP Biology
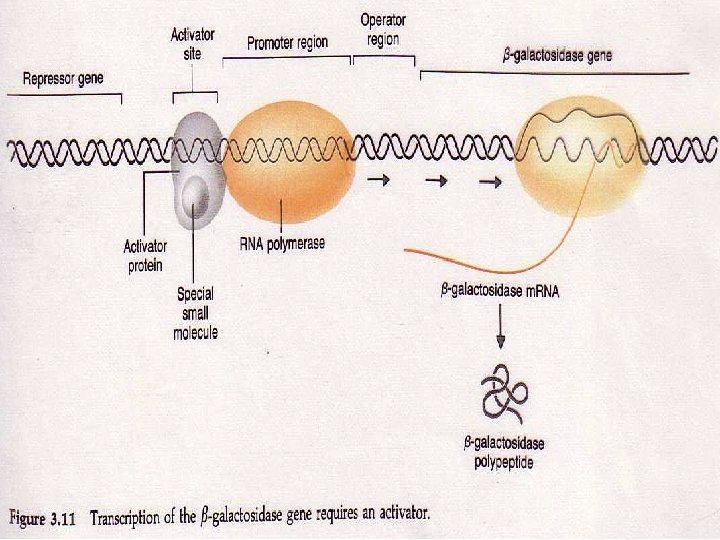
AP Biology

LAC Z GENE EE. coli regulate the production of Beta Galactocidase by using a regulatory protein called a repressor The repressor binds to the lac Z gene at a site between the promotor and the start of the coding sequence The site the repressor binds to is called the operator. AP Biology
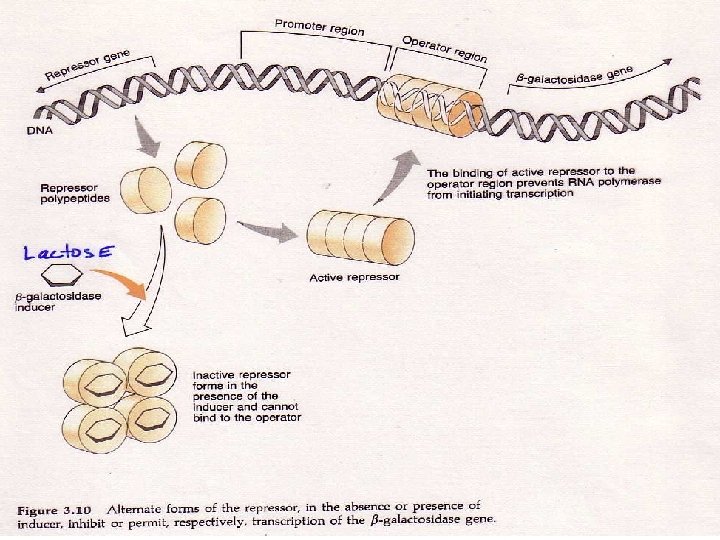
AP Biology

LAC Z GENE Normally the repressor sits on the operator repressing transcription of the lac Z gene In the presence of lactose the repressor binds to the sugar and this allows the polymerase to move down the lac Z gene AP Biology

LAC Z GENE This results in the production of beta galactosidase which breaks down the sugar When there is no sugar left the repressor will return to its spot on the chromosome and stop the transcription of the lac Z gene AP Biology
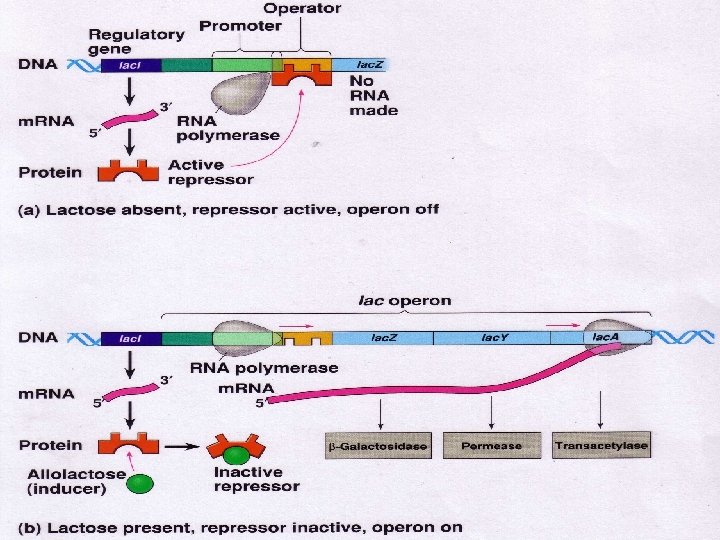
AP Biology
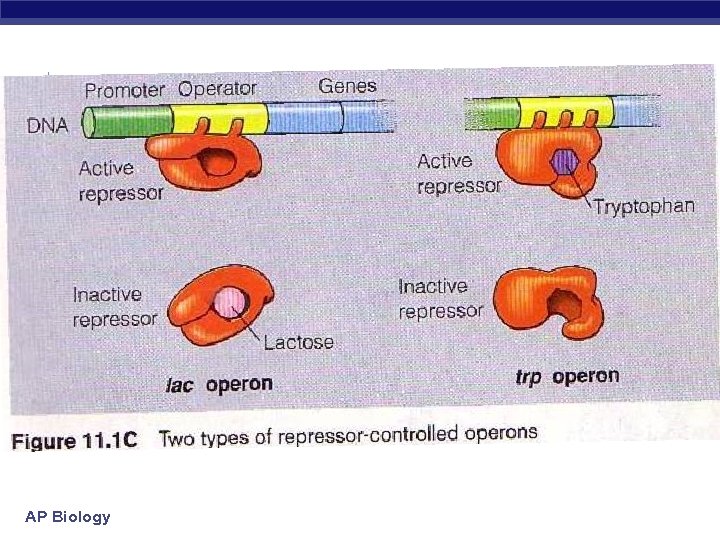
AP Biology

Concept 18. 5: Cancer results from genetic changes that affect cell cycle control The gene regulation systems that go wrong during cancer are the very same systems involved in embryonic development AP Biology © 2011 Pearson Education, Inc.

Types of Genes Associated with Cancer can be caused by mutations to genes that regulate cell growth and division Tumor viruses can cause cancer in animals including humans AP Biology © 2011 Pearson Education, Inc.

Oncogenes are cancer-causing genes Proto-oncogenes are the corresponding normal cellular genes that are responsible for normal cell growth and division Conversion of a proto-oncogene to an oncogene can lead to abnormal stimulation of the cell cycle AP Biology © 2011 Pearson Education, Inc.
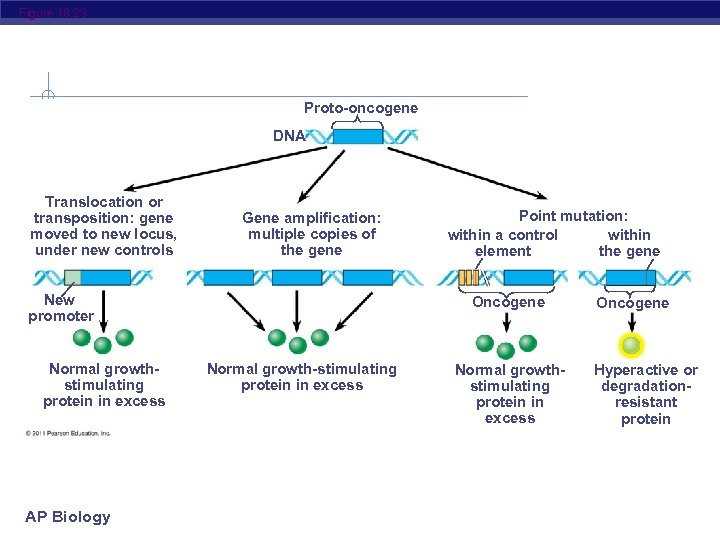
Figure 18. 23 Proto-oncogene DNA Translocation or transposition: gene moved to new locus, under new controls Gene amplification: multiple copies of the gene New promoter Normal growthstimulating protein in excess AP Biology Point mutation: within a control within element the gene Oncogene Normal growth-stimulating protein in excess Normal growthstimulating protein in excess Oncogene Hyperactive or degradationresistant protein

Proto-oncogenes can be converted to oncogenes by Movement of DNA within the genome: if it ends up near an active promoter, transcription may increase u Amplification of a proto-oncogene: increases the number of copies of the gene u Point mutations in the proto-oncogene or its control elements: cause an increase in gene expression u AP Biology © 2011 Pearson Education, Inc.

Tumor-Suppressor Genes Tumor-suppressor genes help prevent uncontrolled cell growth Mutations that decrease protein products of tumor-suppressor genes may contribute to cancer onset Tumor-suppressor proteins u Repair damaged DNA u Control cell adhesion u AP Biology Inhibit the cell cycle in the cell-signaling pathway © 2011 Pearson Education, Inc.
Interference with Normal Cell. Signaling Pathways Mutations in the ras proto-oncogene and p 53 tumor-suppressor gene are common in human cancers Mutations in the ras gene can lead to production of a hyperactive Ras protein and increased cell division AP Biology © 2011 Pearson Education, Inc.
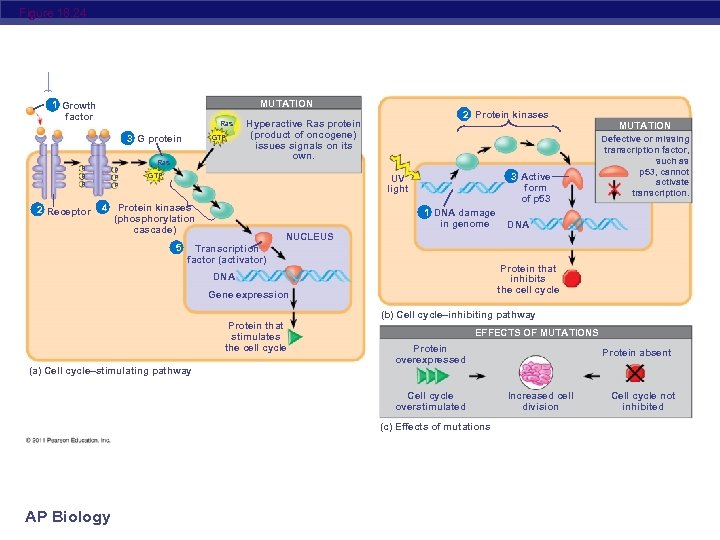
Figure 18. 24 MUTATION 1 Growth factor Ras 3 G protein GTP Ras P P P 2 Receptor 4 2 Protein kinases Hyperactive Ras protein (product of oncogene) issues signals on its own. GTP 3 Active form of p 53 UV light Protein kinases (phosphorylation cascade) 5 1 DNA damage in genome MUTATION Defective or missing transcription factor, such as p 53, cannot activate transcription. DNA NUCLEUS Transcription factor (activator) Protein that inhibits the cell cycle DNA Gene expression (b) Cell cycle–inhibiting pathway Protein that stimulates the cell cycle (a) Cell cycle–stimulating pathway EFFECTS OF MUTATIONS Protein overexpressed Cell cycle overstimulated (c) Effects of mutations AP Biology Protein absent Increased cell division Cell cycle not inhibited
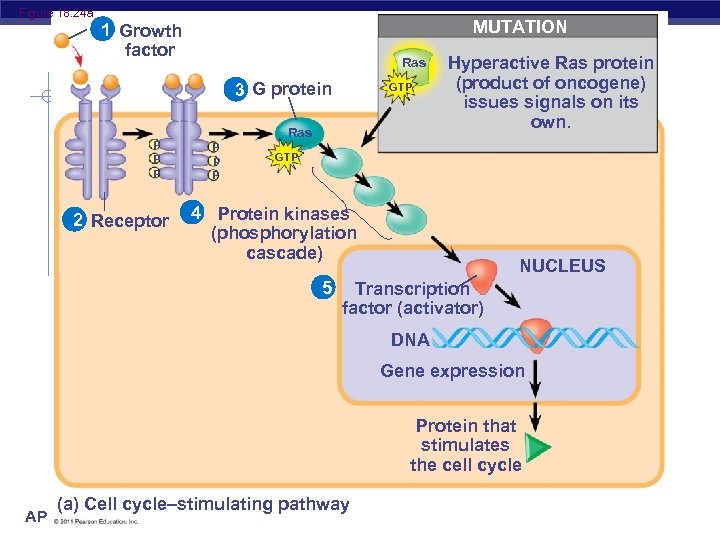
Figure 18. 24 a MUTATION 1 Growth factor Ras 3 G protein P P P 2 Receptor GTP Ras P P P Hyperactive Ras protein (product of oncogene) issues signals on its own. GTP 4 Protein kinases (phosphorylation cascade) 5 NUCLEUS Transcription factor (activator) DNA Gene expression Protein that stimulates the cell cycle (a) Cell cycle–stimulating pathway AP Biology
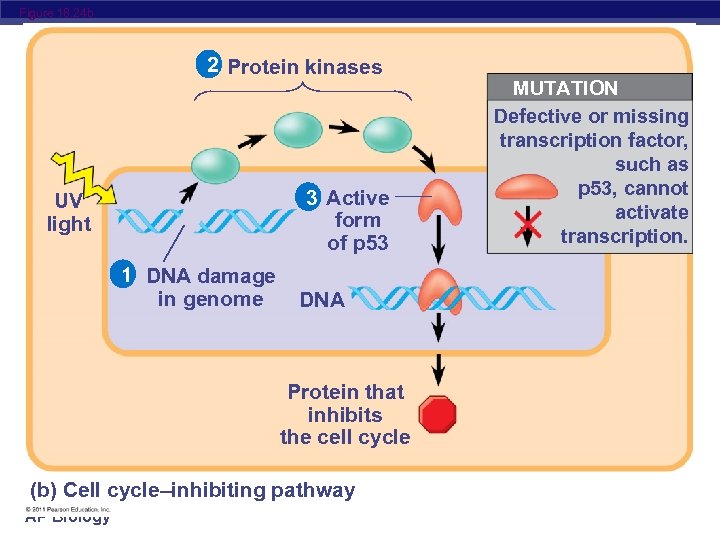
Figure 18. 24 b 2 Protein kinases 3 Active form of p 53 UV light 1 DNA damage in genome DNA Protein that inhibits the cell cycle (b) Cell cycle–inhibiting pathway AP Biology MUTATION Defective or missing transcription factor, such as p 53, cannot activate transcription.

Suppression of the cell cycle can be important in the case of damage to a cell’s DNA; p 53 prevents a cell from passing on mutations due to DNA damage Mutations in the p 53 gene prevent suppression of the cell cycle AP Biology © 2011 Pearson Education, Inc.
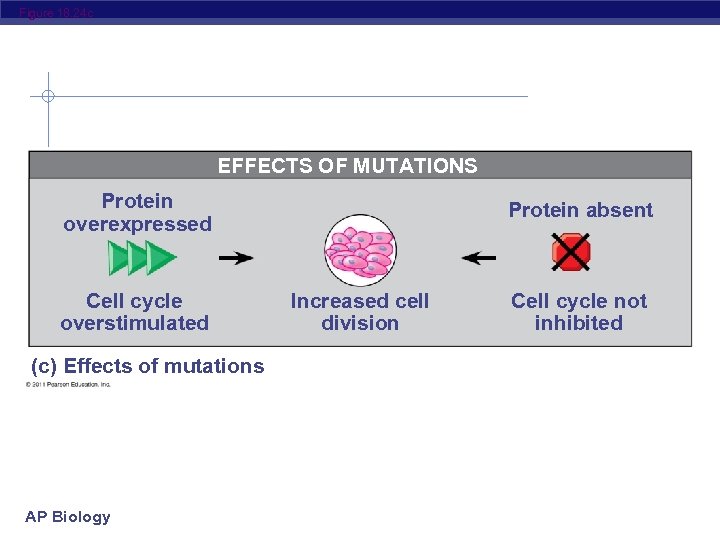
Figure 18. 24 c EFFECTS OF MUTATIONS Protein overexpressed Cell cycle overstimulated (c) Effects of mutations AP Biology Protein absent Increased cell division Cell cycle not inhibited

The Multistep Model of Cancer Development Multiple mutations are generally needed for full -fledged cancer; thus the incidence increases with age At the DNA level, a cancerous cell is usually characterized by at least one active oncogene and the mutation of several tumor-suppressor genes AP Biology © 2011 Pearson Education, Inc.
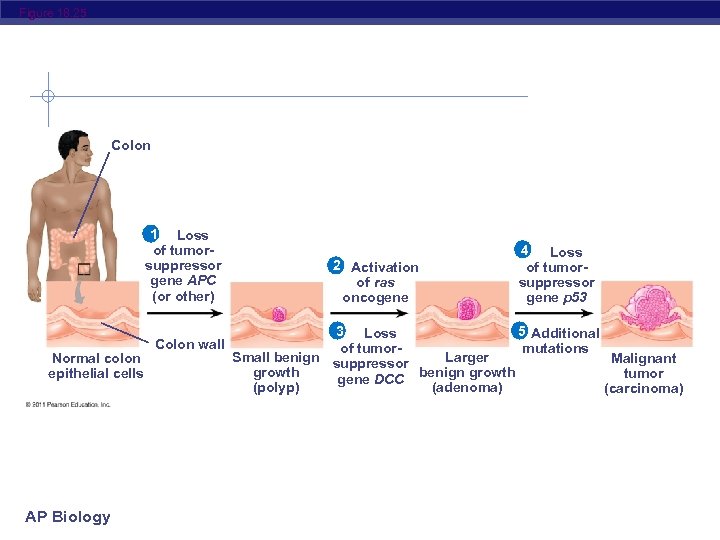
Figure 18. 25 Colon 1 Loss of tumorsuppressor gene APC (or other) Normal colon epithelial cells AP Biology Colon wall 2 Activation of ras oncogene 4 Loss of tumorsuppressor gene p 53 3 Loss 5 Additional mutations of tumor. Small benign suppressor Larger Malignant growth tumor gene DCC benign growth (polyp) (adenoma) (carcinoma)
Figure 18. 25 a Colon wall Normal colon epithelial cells AP Biology
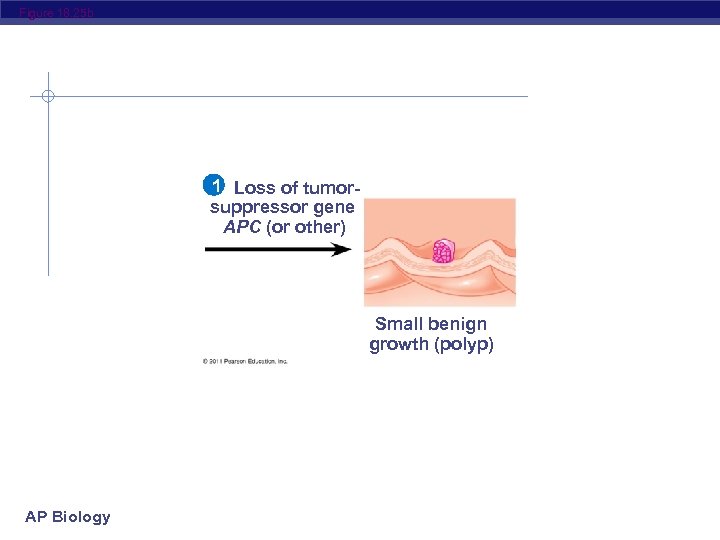
Figure 18. 25 b 1 Loss of tumorsuppressor gene APC (or other) Small benign growth (polyp) AP Biology
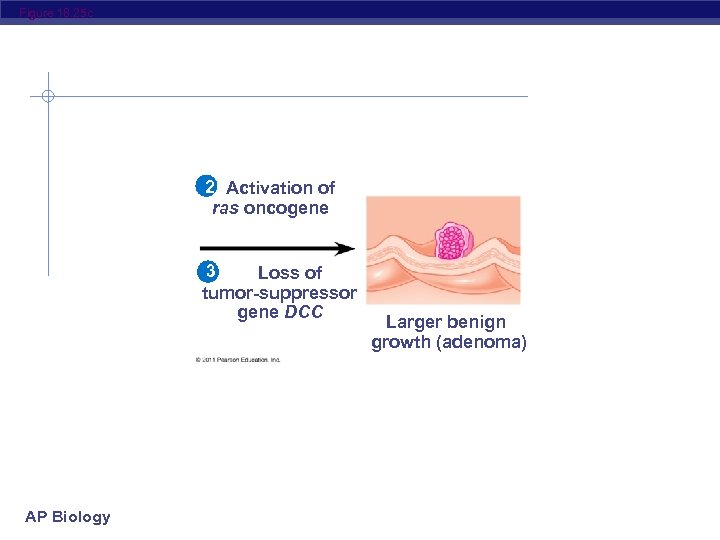
Figure 18. 25 c 2 Activation of ras oncogene 3 Loss of tumor-suppressor gene DCC AP Biology Larger benign growth (adenoma)
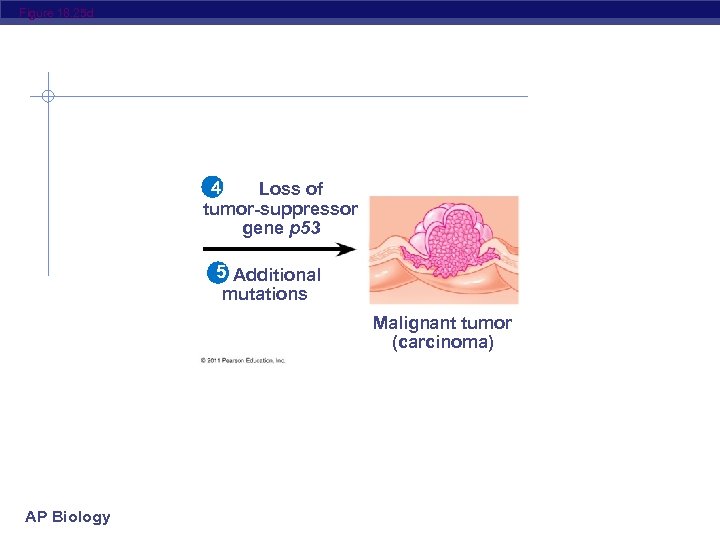
Figure 18. 25 d 4 Loss of tumor-suppressor gene p 53 5 Additional mutations Malignant tumor (carcinoma) AP Biology

Inherited Predisposition and Other Factors Contributing to Cancer Individuals can inherit oncogenes or mutant alleles of tumor-suppressor genes Inherited mutations in the tumor-suppressor gene adenomatous polyposis coli are common in individuals with colorectal cancer Mutations in the BRCA 1 or BRCA 2 gene are found in at least half of inherited breast cancers, and tests using DNA sequencing can detect these mutations AP Biology © 2011 Pearson Education, Inc.

Figure 18. 26 AP Biology
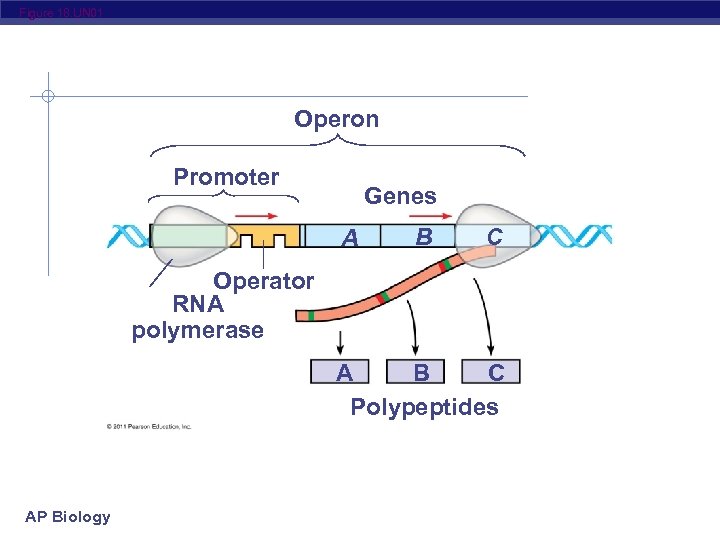
Figure 18. UN 01 Operon Promoter Genes A B C Operator RNA polymerase A B C Polypeptides AP Biology
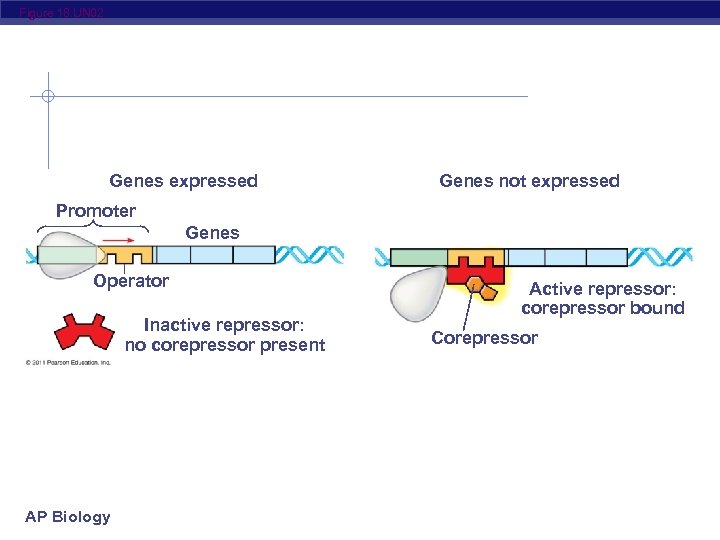
Figure 18. UN 02 Genes expressed Genes not expressed Promoter Genes Operator Inactive repressor: no corepressor present AP Biology Active repressor: corepressor bound Corepressor
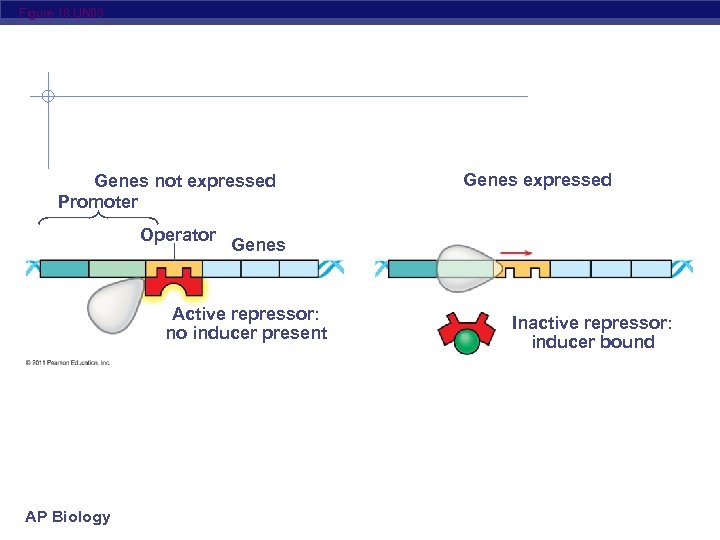
Figure 18. UN 03 Genes not expressed Promoter Operator Genes Active repressor: no inducer present AP Biology Genes expressed Inactive repressor: inducer bound
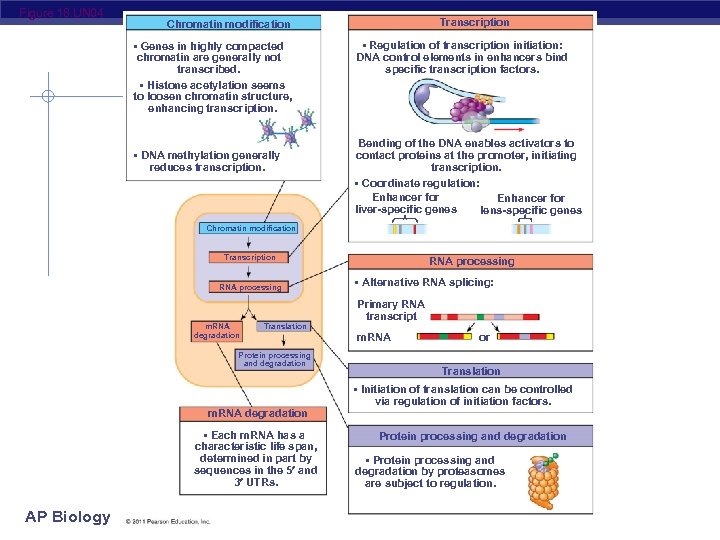
Figure 18. UN 04 Transcription Chromatin modification • Genes in highly compacted chromatin are generally not transcribed. • Histone acetylation seems to loosen chromatin structure, enhancing transcription. • DNA methylation generally reduces transcription. • Regulation of transcription initiation: DNA control elements in enhancers bind specific transcription factors. Bending of the DNA enables activators to contact proteins at the promoter, initiating transcription. • Coordinate regulation: Enhancer for liver-specific genes lens-specific genes Chromatin modification Transcription RNA processing m. RNA degradation Translation Protein processing and degradation m. RNA degradation • Each m. RNA has a characteristic life span, determined in part by sequences in the 5 and 3 UTRs. AP Biology RNA processing • Alternative RNA splicing: Primary RNA transcript m. RNA or Translation • Initiation of translation can be controlled via regulation of initiation factors. Protein processing and degradation • Protein processing and degradation by proteasomes are subject to regulation.
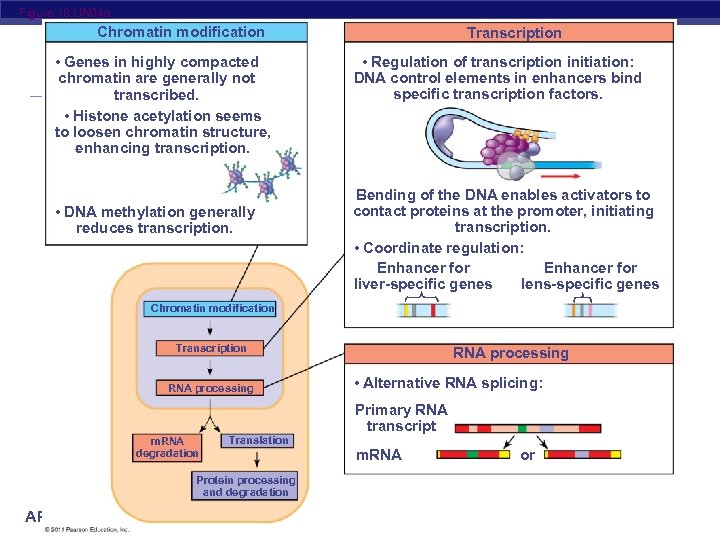
Figure 18. UN 04 a Chromatin modification • Genes in highly compacted chromatin are generally not transcribed. • Histone acetylation seems to loosen chromatin structure, enhancing transcription. • DNA methylation generally reduces transcription. Transcription • Regulation of transcription initiation: DNA control elements in enhancers bind specific transcription factors. Bending of the DNA enables activators to contact proteins at the promoter, initiating transcription. • Coordinate regulation: Enhancer for lens-specific genes liver-specific genes Chromatin modification Transcription RNA processing m. RNA degradation Translation Protein processing and degradation AP Biology RNA processing • Alternative RNA splicing: Primary RNA transcript m. RNA or
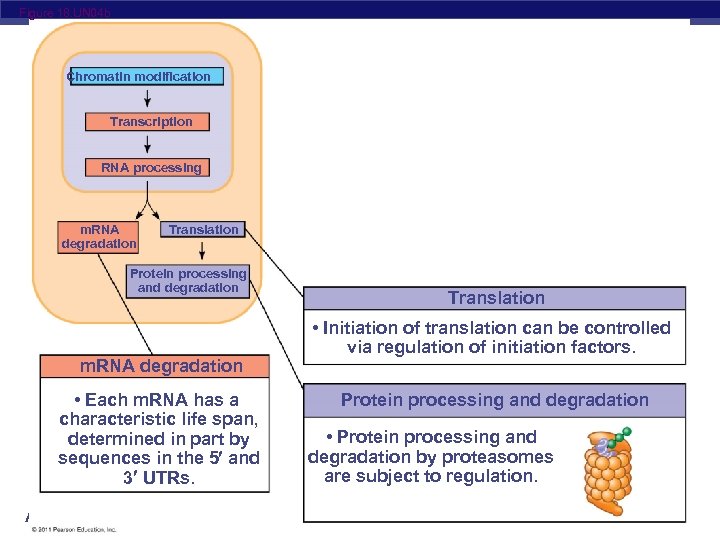
Figure 18. UN 04 b Chromatin modification Transcription RNA processing m. RNA degradation Translation Protein processing and degradation m. RNA degradation • Each m. RNA has a characteristic life span, determined in part by sequences in the 5 and 3 UTRs. AP Biology Translation • Initiation of translation can be controlled via regulation of initiation factors. Protein processing and degradation • Protein processing and degradation by proteasomes are subject to regulation.
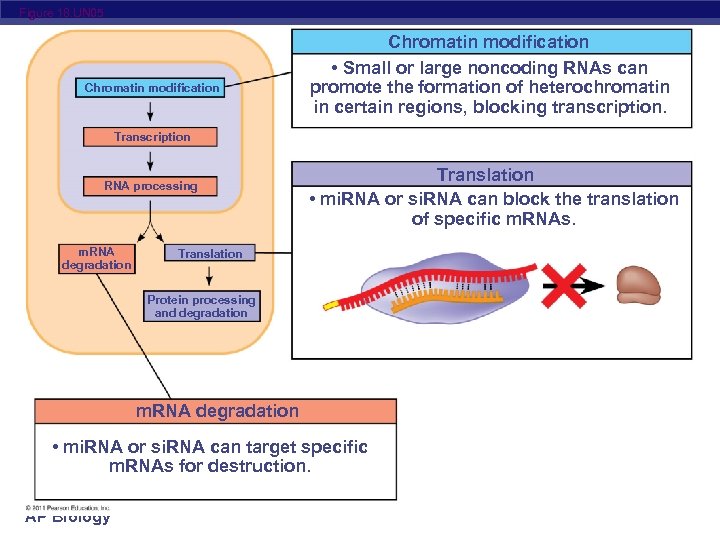
Figure 18. UN 05 Chromatin modification • Small or large noncoding RNAs can promote the formation of heterochromatin in certain regions, blocking transcription. Transcription RNA processing m. RNA degradation Translation • mi. RNA or si. RNA can block the translation of specific m. RNAs. Translation Protein processing and degradation m. RNA degradation • mi. RNA or si. RNA can target specific m. RNAs for destruction. AP Biology
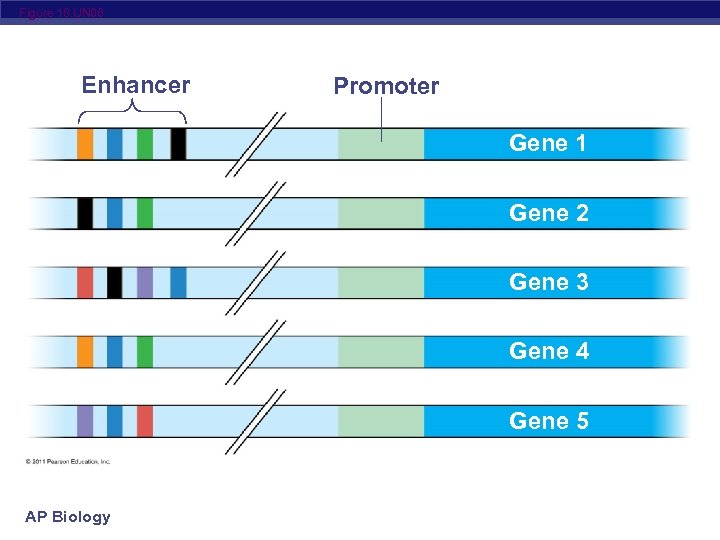
Figure 18. UN 06 Enhancer Promoter Gene 1 Gene 2 Gene 3 Gene 4 Gene 5 AP Biology
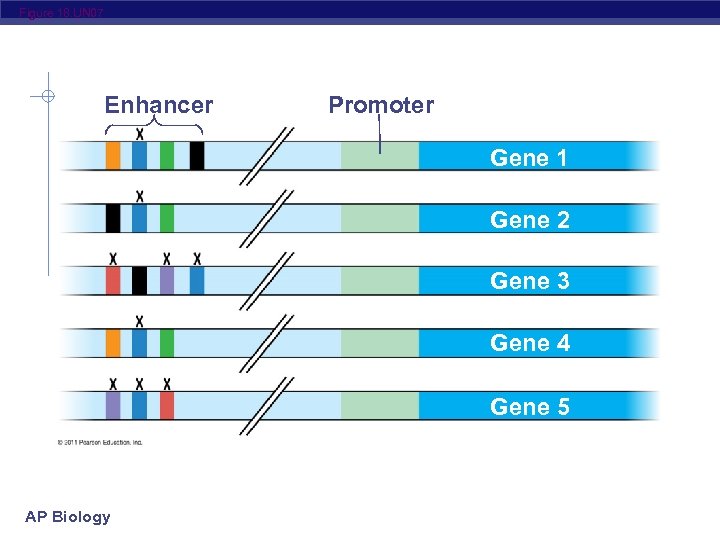
Figure 18. UN 07 Enhancer Promoter Gene 1 Gene 2 Gene 3 Gene 4 Gene 5 AP Biology
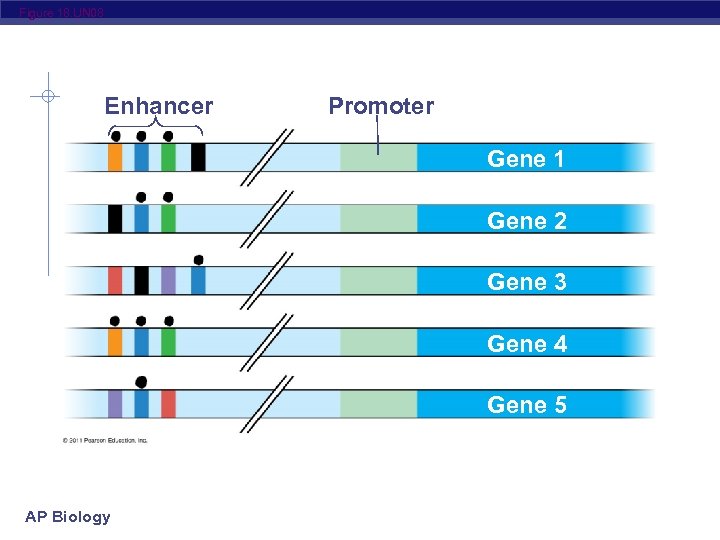
Figure 18. UN 08 Enhancer Promoter Gene 1 Gene 2 Gene 3 Gene 4 Gene 5 AP Biology
39d44a0aad81942c575b38d5c8a0e745.ppt