f8aa4aee46631da31c15d1ba9f733c6e.ppt
- Количество слайдов: 45
 CONFIDENTIAL Meta. Tox: leveraging the power of systems biology for improving drug safety Andrej Bugrim Gene. Go, Inc. Copyright Gene. Go 2000 -2006
CONFIDENTIAL Meta. Tox: leveraging the power of systems biology for improving drug safety Andrej Bugrim Gene. Go, Inc. Copyright Gene. Go 2000 -2006
 Agenda CONFIDENTIAL • Understand what is missing today and why more drugs are failing human tox than ever before • How we plan to address this problems • Run through case studies to validate our ideas • Summary Copyright Gene. Go 2000 -2006
Agenda CONFIDENTIAL • Understand what is missing today and why more drugs are failing human tox than ever before • How we plan to address this problems • Run through case studies to validate our ideas • Summary Copyright Gene. Go 2000 -2006
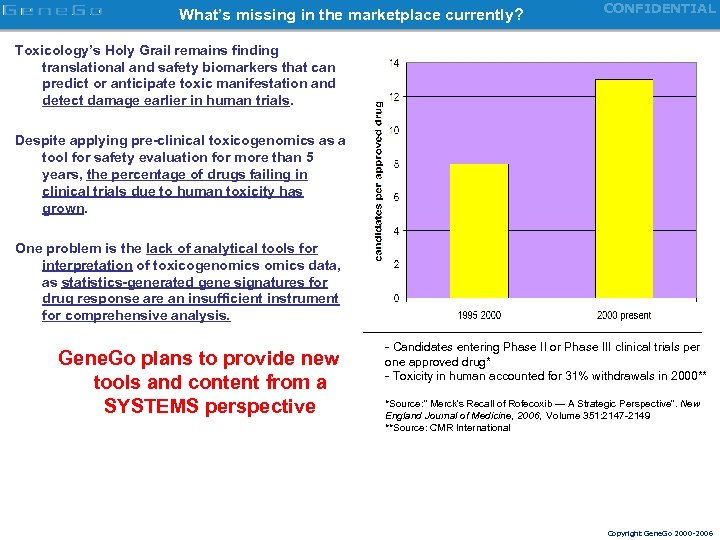 What’s missing in the marketplace currently? CONFIDENTIAL Toxicology’s Holy Grail remains finding translational and safety biomarkers that can predict or anticipate toxic manifestation and detect damage earlier in human trials. Despite applying pre-clinical toxicogenomics as a tool for safety evaluation for more than 5 years, the percentage of drugs failing in clinical trials due to human toxicity has grown. One problem is the lack of analytical tools for interpretation of toxicogenomics data, as statistics-generated gene signatures for drug response are an insufficient instrument for comprehensive analysis. Gene. Go plans to provide new tools and content from a SYSTEMS perspective - Candidates entering Phase II or Phase III clinical trials per one approved drug* - Toxicity in human accounted for 31% withdrawals in 2000** *Source: ” Merck's Recall of Rofecoxib — A Strategic Perspective”. New England Journal of Medicine, 2006, Volume 351: 2147 -2149 **Source: CMR International Copyright Gene. Go 2000 -2006
What’s missing in the marketplace currently? CONFIDENTIAL Toxicology’s Holy Grail remains finding translational and safety biomarkers that can predict or anticipate toxic manifestation and detect damage earlier in human trials. Despite applying pre-clinical toxicogenomics as a tool for safety evaluation for more than 5 years, the percentage of drugs failing in clinical trials due to human toxicity has grown. One problem is the lack of analytical tools for interpretation of toxicogenomics data, as statistics-generated gene signatures for drug response are an insufficient instrument for comprehensive analysis. Gene. Go plans to provide new tools and content from a SYSTEMS perspective - Candidates entering Phase II or Phase III clinical trials per one approved drug* - Toxicity in human accounted for 31% withdrawals in 2000** *Source: ” Merck's Recall of Rofecoxib — A Strategic Perspective”. New England Journal of Medicine, 2006, Volume 351: 2147 -2149 **Source: CMR International Copyright Gene. Go 2000 -2006
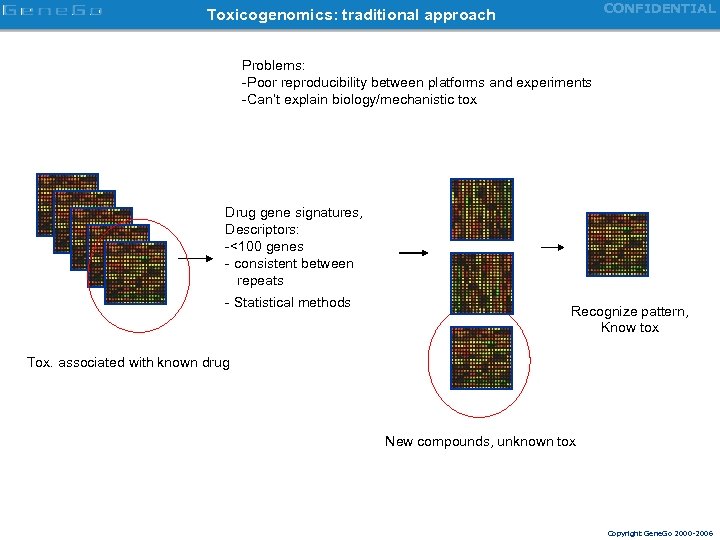 CONFIDENTIAL Toxicogenomics: traditional approach Problems: -Poor reproducibility between platforms and experiments -Can’t explain biology/mechanistic tox Drug gene signatures, Descriptors: -<100 genes - consistent between repeats - Statistical methods Recognize pattern, Know tox Tox. associated with known drug New compounds, unknown tox Copyright Gene. Go 2000 -2006
CONFIDENTIAL Toxicogenomics: traditional approach Problems: -Poor reproducibility between platforms and experiments -Can’t explain biology/mechanistic tox Drug gene signatures, Descriptors: -<100 genes - consistent between repeats - Statistical methods Recognize pattern, Know tox Tox. associated with known drug New compounds, unknown tox Copyright Gene. Go 2000 -2006
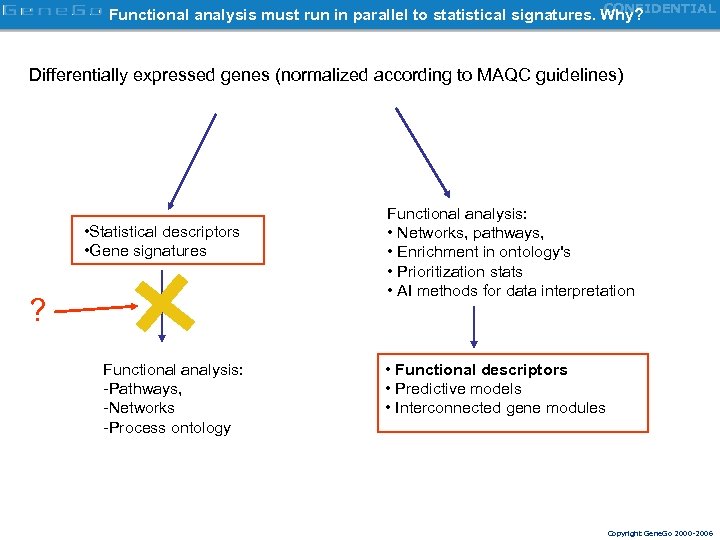 CONFIDENTIAL Functional analysis must run in parallel to statistical signatures. Why? Differentially expressed genes (normalized according to MAQC guidelines) • Statistical descriptors • Gene signatures ? Functional analysis: -Pathways, -Networks -Process ontology Functional analysis: • Networks, pathways, • Enrichment in ontology's • Prioritization stats • AI methods for data interpretation • Functional descriptors • Predictive models • Interconnected gene modules Copyright Gene. Go 2000 -2006
CONFIDENTIAL Functional analysis must run in parallel to statistical signatures. Why? Differentially expressed genes (normalized according to MAQC guidelines) • Statistical descriptors • Gene signatures ? Functional analysis: -Pathways, -Networks -Process ontology Functional analysis: • Networks, pathways, • Enrichment in ontology's • Prioritization stats • AI methods for data interpretation • Functional descriptors • Predictive models • Interconnected gene modules Copyright Gene. Go 2000 -2006
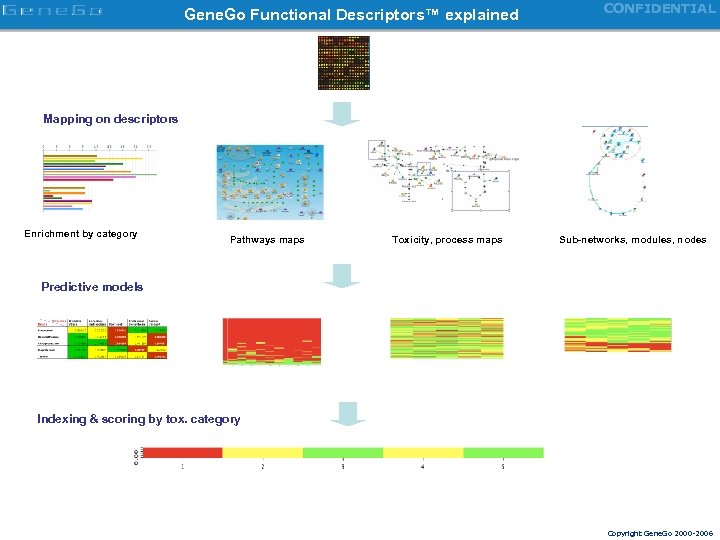 Gene. Go Functional Descriptors™ explained CONFIDENTIAL Mapping on descriptors Enrichment by category Pathways maps Toxicity, process maps Sub-networks, modules, nodes Predictive models Indexing & scoring by tox. category Copyright Gene. Go 2000 -2006
Gene. Go Functional Descriptors™ explained CONFIDENTIAL Mapping on descriptors Enrichment by category Pathways maps Toxicity, process maps Sub-networks, modules, nodes Predictive models Indexing & scoring by tox. category Copyright Gene. Go 2000 -2006
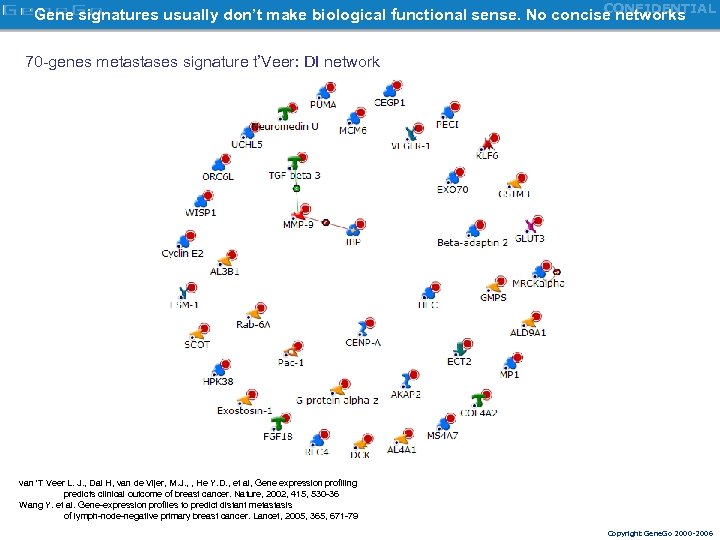 CONFIDENTIAL Gene signatures usually don’t make biological functional sense. No concise networks 70 -genes metastases signature t’Veer: DI network van 'T Veer L. J. , Dai H, van de Vijer, M. J. , , He Y. D. , et al, Gene expression profiling predicts clinical outcome of breast cancer. Nature, 2002, 415, 530 -36 Wang Y. et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet, 2005, 365, 671 -79 Copyright Gene. Go 2000 -2006
CONFIDENTIAL Gene signatures usually don’t make biological functional sense. No concise networks 70 -genes metastases signature t’Veer: DI network van 'T Veer L. J. , Dai H, van de Vijer, M. J. , , He Y. D. , et al, Gene expression profiling predicts clinical outcome of breast cancer. Nature, 2002, 415, 530 -36 Wang Y. et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet, 2005, 365, 671 -79 Copyright Gene. Go 2000 -2006
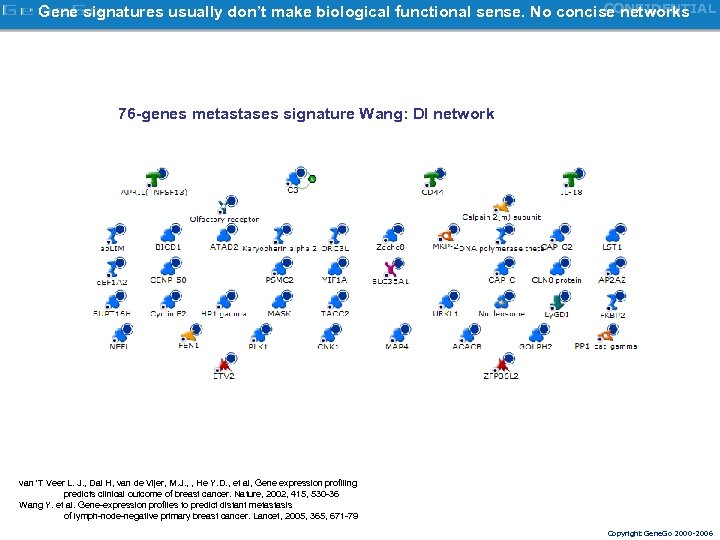 CONFIDENTIAL Gene signatures usually don’t make biological functional sense. No concise networks 76 -genes metastases signature Wang: DI network van 'T Veer L. J. , Dai H, van de Vijer, M. J. , , He Y. D. , et al, Gene expression profiling predicts clinical outcome of breast cancer. Nature, 2002, 415, 530 -36 Wang Y. et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet, 2005, 365, 671 -79 Copyright Gene. Go 2000 -2006
CONFIDENTIAL Gene signatures usually don’t make biological functional sense. No concise networks 76 -genes metastases signature Wang: DI network van 'T Veer L. J. , Dai H, van de Vijer, M. J. , , He Y. D. , et al, Gene expression profiling predicts clinical outcome of breast cancer. Nature, 2002, 415, 530 -36 Wang Y. et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet, 2005, 365, 671 -79 Copyright Gene. Go 2000 -2006
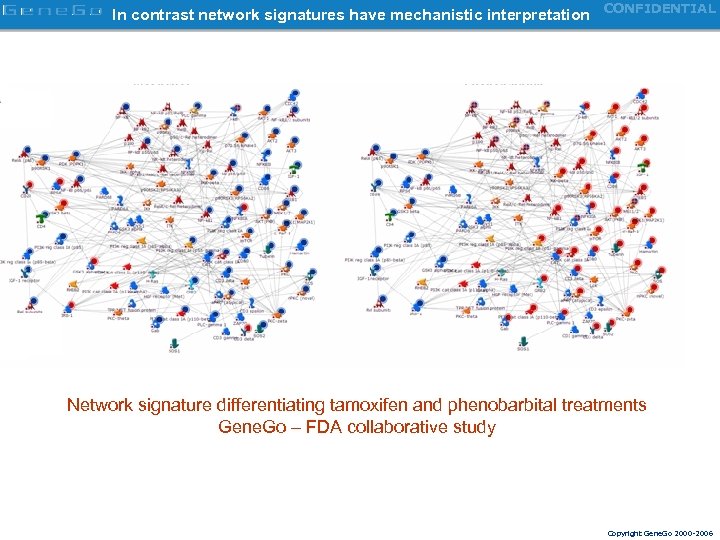 In contrast network signatures have mechanistic interpretation CONFIDENTIAL Network signature differentiating tamoxifen and phenobarbital treatments Gene. Go – FDA collaborative study Copyright Gene. Go 2000 -2006
In contrast network signatures have mechanistic interpretation CONFIDENTIAL Network signature differentiating tamoxifen and phenobarbital treatments Gene. Go – FDA collaborative study Copyright Gene. Go 2000 -2006
 Let’s take a closer look: nephrotoxicity case study 1 • • CONFIDENTIAL Training set: – 15 nephrotoxicants – 49 non-nephrotoxicants Test set: – 9 nephrotoxicants – 12 non-nephrotxicants • Microarray data profiling – Treatment: male Sprague-Dawley rats, 7 -8 weeks of age – Animal selection: 3 rats at day 5 for each treatment for microarray profiling – 10 rats at day 28 for histopathology profiling – Expression profiling: Amersham Codelink Uniset Rat 1 Bioarray • Nephrotoxicity is assessed by histopathology • Published models accuracy: – Training set error 16% – Test set error 25% Same lab, same platform, same animal group - why loss of accuracy? How would perform across platforms/labs/animal groups? 1 Source: Fielden MR, Eynon BP, Natsoulis G, Jarnagin K, Banas D, Kolaja KL. A gene expression signature that predicts the future onset of drug-induced renal tubular toxicity. Toxicol Pathol. 2005; 33(6): 675 -83. Copyright Gene. Go 2000 -2006
Let’s take a closer look: nephrotoxicity case study 1 • • CONFIDENTIAL Training set: – 15 nephrotoxicants – 49 non-nephrotoxicants Test set: – 9 nephrotoxicants – 12 non-nephrotxicants • Microarray data profiling – Treatment: male Sprague-Dawley rats, 7 -8 weeks of age – Animal selection: 3 rats at day 5 for each treatment for microarray profiling – 10 rats at day 28 for histopathology profiling – Expression profiling: Amersham Codelink Uniset Rat 1 Bioarray • Nephrotoxicity is assessed by histopathology • Published models accuracy: – Training set error 16% – Test set error 25% Same lab, same platform, same animal group - why loss of accuracy? How would perform across platforms/labs/animal groups? 1 Source: Fielden MR, Eynon BP, Natsoulis G, Jarnagin K, Banas D, Kolaja KL. A gene expression signature that predicts the future onset of drug-induced renal tubular toxicity. Toxicol Pathol. 2005; 33(6): 675 -83. Copyright Gene. Go 2000 -2006
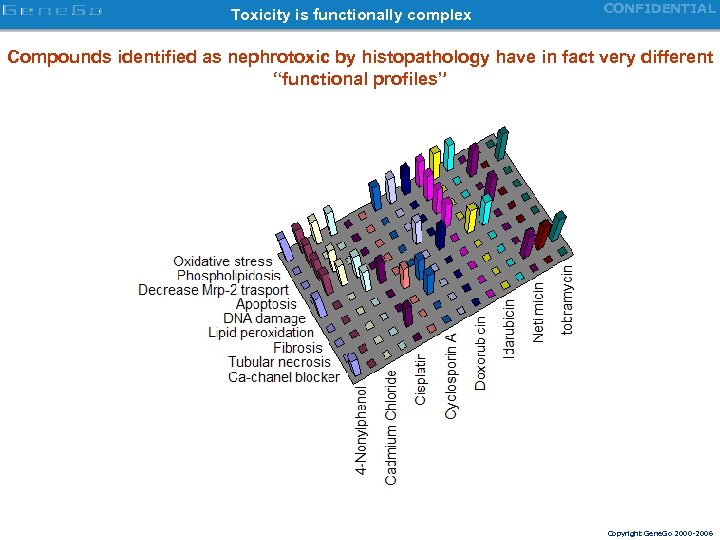 Toxicity is functionally complex CONFIDENTIAL Compounds identified as nephrotoxic by histopathology have in fact very different “functional profiles” Copyright Gene. Go 2000 -2006
Toxicity is functionally complex CONFIDENTIAL Compounds identified as nephrotoxic by histopathology have in fact very different “functional profiles” Copyright Gene. Go 2000 -2006
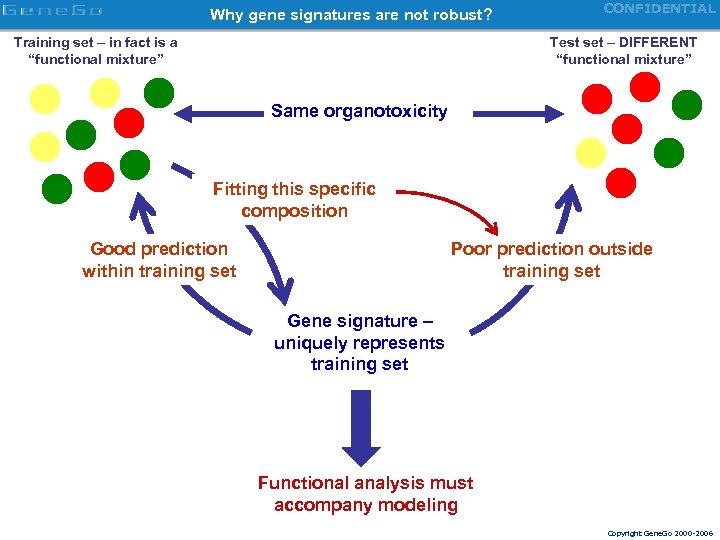 Why gene signatures are not robust? Training set – in fact is a “functional mixture” CONFIDENTIAL Test set – DIFFERENT “functional mixture” Same organotoxicity Fitting this specific composition Good prediction within training set Poor prediction outside training set Gene signature – uniquely represents training set Functional analysis must accompany modeling Copyright Gene. Go 2000 -2006
Why gene signatures are not robust? Training set – in fact is a “functional mixture” CONFIDENTIAL Test set – DIFFERENT “functional mixture” Same organotoxicity Fitting this specific composition Good prediction within training set Poor prediction outside training set Gene signature – uniquely represents training set Functional analysis must accompany modeling Copyright Gene. Go 2000 -2006
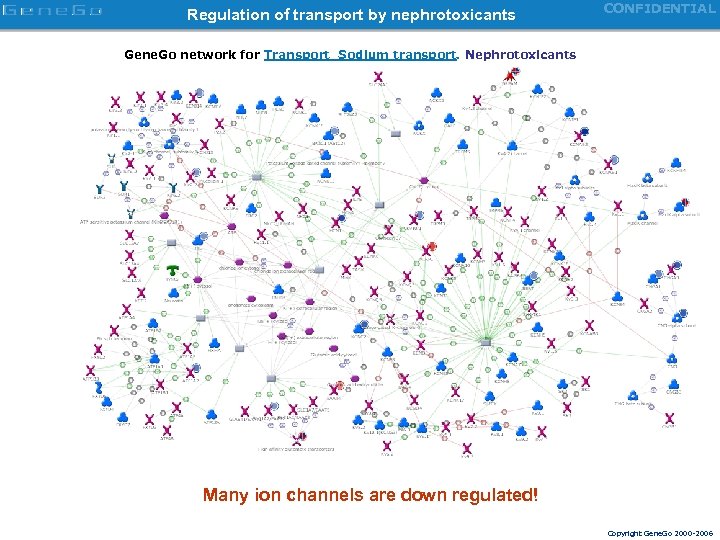 Regulation of transport by nephrotoxicants CONFIDENTIAL Gene. Go network for Transport_Sodium transport. Nephrotoxicants Many ion channels are down regulated! Copyright Gene. Go 2000 -2006
Regulation of transport by nephrotoxicants CONFIDENTIAL Gene. Go network for Transport_Sodium transport. Nephrotoxicants Many ion channels are down regulated! Copyright Gene. Go 2000 -2006
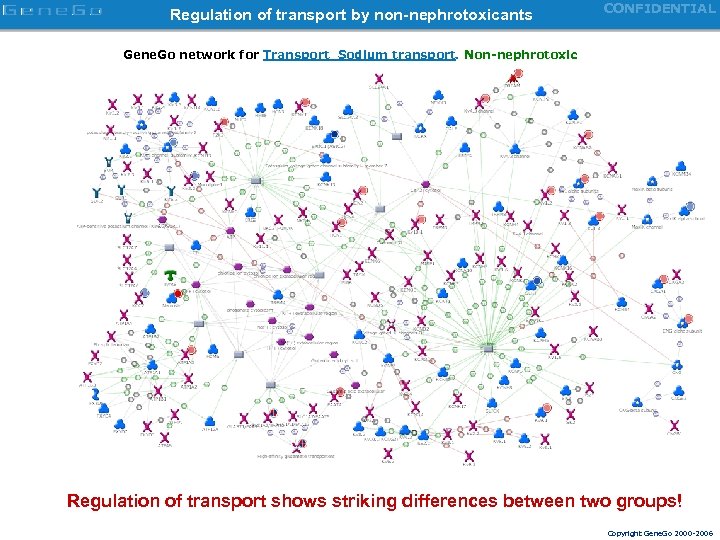 Regulation of transport by non-nephrotoxicants CONFIDENTIAL Gene. Go network for Transport_Sodium transport. Non-nephrotoxic Regulation of transport shows striking differences between two groups! Copyright Gene. Go 2000 -2006
Regulation of transport by non-nephrotoxicants CONFIDENTIAL Gene. Go network for Transport_Sodium transport. Non-nephrotoxic Regulation of transport shows striking differences between two groups! Copyright Gene. Go 2000 -2006
 Lessons learned: CONFIDENTIAL • Different mechanisms = same histopathology (nephrotoxicity) • Similar mechanisms could be shared by toxicants and non-toxicants alike • Dataset is too small to build a robust model What we need: Functional and mechanistic analysis must guide building predictive models Larger datasets are needed to address functional diversity Copyright Gene. Go 2000 -2006
Lessons learned: CONFIDENTIAL • Different mechanisms = same histopathology (nephrotoxicity) • Similar mechanisms could be shared by toxicants and non-toxicants alike • Dataset is too small to build a robust model What we need: Functional and mechanistic analysis must guide building predictive models Larger datasets are needed to address functional diversity Copyright Gene. Go 2000 -2006
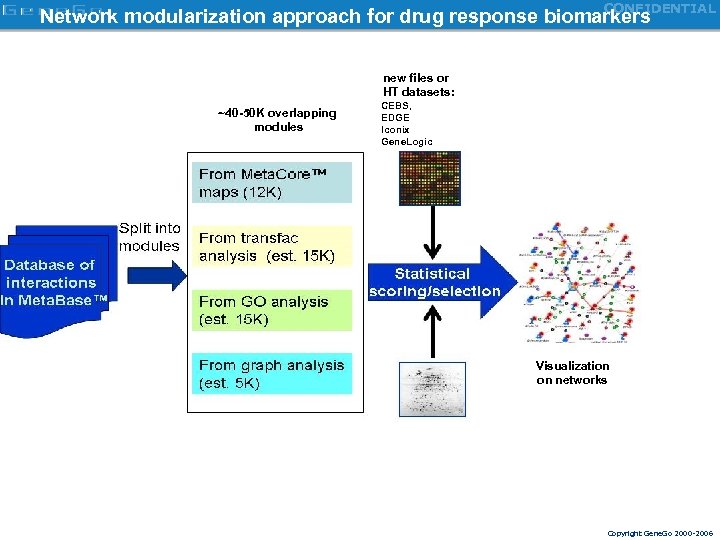 CONFIDENTIAL Network modularization approach for drug response biomarkers new files or HT datasets: ~40 -50 K overlapping modules CEBS, EDGE Iconix Gene. Logic Visualization on networks Copyright Gene. Go 2000 -2006
CONFIDENTIAL Network modularization approach for drug response biomarkers new files or HT datasets: ~40 -50 K overlapping modules CEBS, EDGE Iconix Gene. Logic Visualization on networks Copyright Gene. Go 2000 -2006
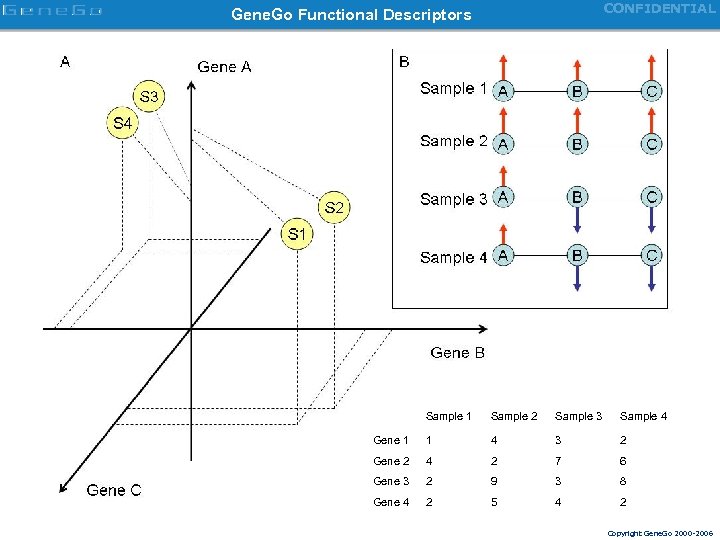 CONFIDENTIAL Gene. Go Functional Descriptors Sample 1 Sample 2 Sample 3 Sample 4 Gene 1 1 4 3 2 Gene 2 4 2 7 6 Gene 3 2 9 3 8 Gene 4 2 5 4 2 Copyright Gene. Go 2000 -2006
CONFIDENTIAL Gene. Go Functional Descriptors Sample 1 Sample 2 Sample 3 Sample 4 Gene 1 1 4 3 2 Gene 2 4 2 7 6 Gene 3 2 9 3 8 Gene 4 2 5 4 2 Copyright Gene. Go 2000 -2006
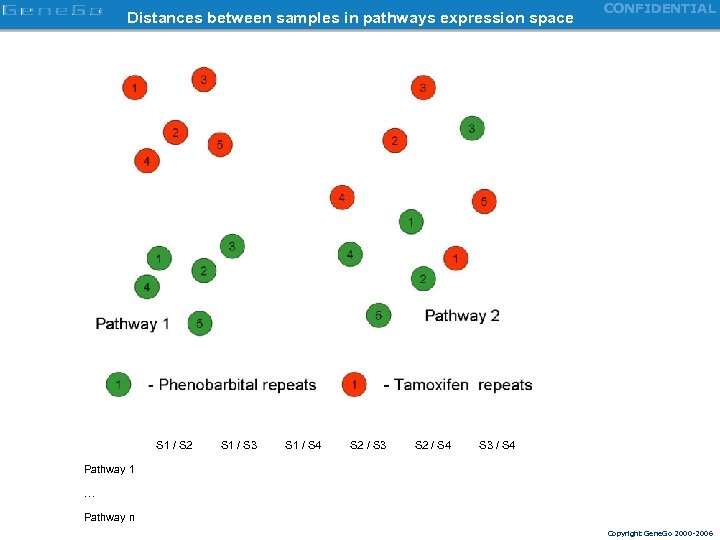 Distances between samples in pathways expression space S 1 / S 2 S 1 / S 3 S 1 / S 4 S 2 / S 3 S 2 / S 4 CONFIDENTIAL S 3 / S 4 Pathway 1 … Pathway n Copyright Gene. Go 2000 -2006
Distances between samples in pathways expression space S 1 / S 2 S 1 / S 3 S 1 / S 4 S 2 / S 3 S 2 / S 4 CONFIDENTIAL S 3 / S 4 Pathway 1 … Pathway n Copyright Gene. Go 2000 -2006
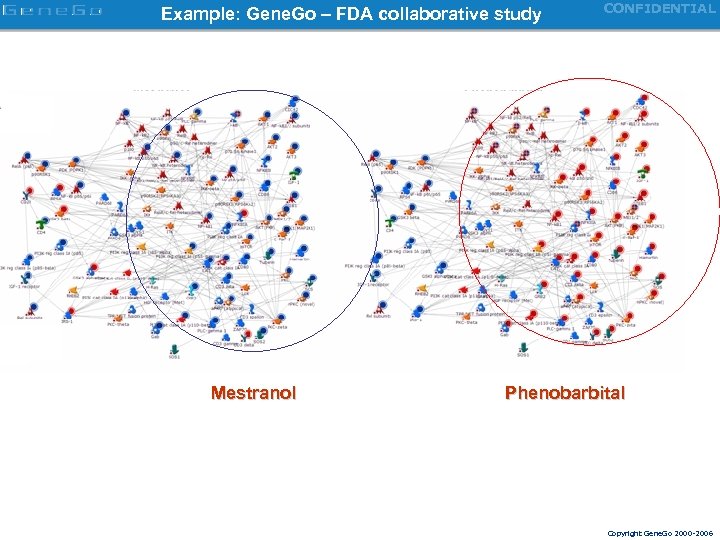 Example: Gene. Go – FDA collaborative study Mestranol CONFIDENTIAL Phenobarbital Copyright Gene. Go 2000 -2006
Example: Gene. Go – FDA collaborative study Mestranol CONFIDENTIAL Phenobarbital Copyright Gene. Go 2000 -2006
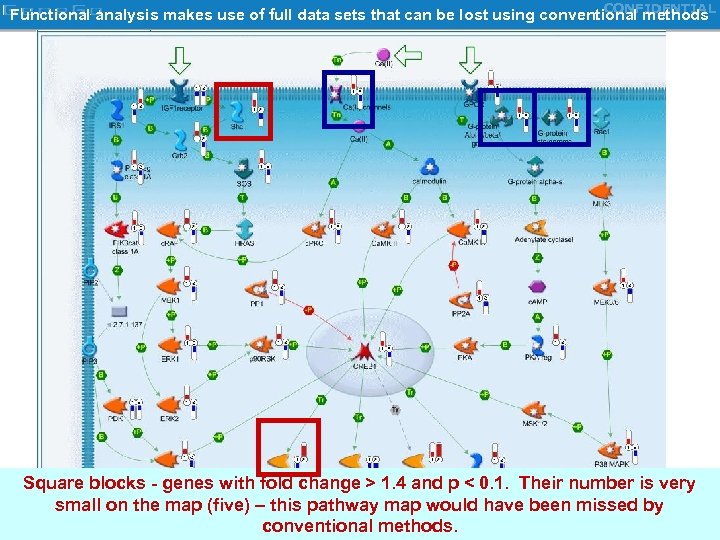 CONFIDENTIAL Functional analysis makes use of full data sets that can be lost using conventional methods Square blocks - genes with fold change > 1. 4 and p < 0. 1. Their number is very small on the map (five) – this pathway map would have been missed by conventional methods. Copyright Gene. Go 2000 -2006
CONFIDENTIAL Functional analysis makes use of full data sets that can be lost using conventional methods Square blocks - genes with fold change > 1. 4 and p < 0. 1. Their number is very small on the map (five) – this pathway map would have been missed by conventional methods. Copyright Gene. Go 2000 -2006
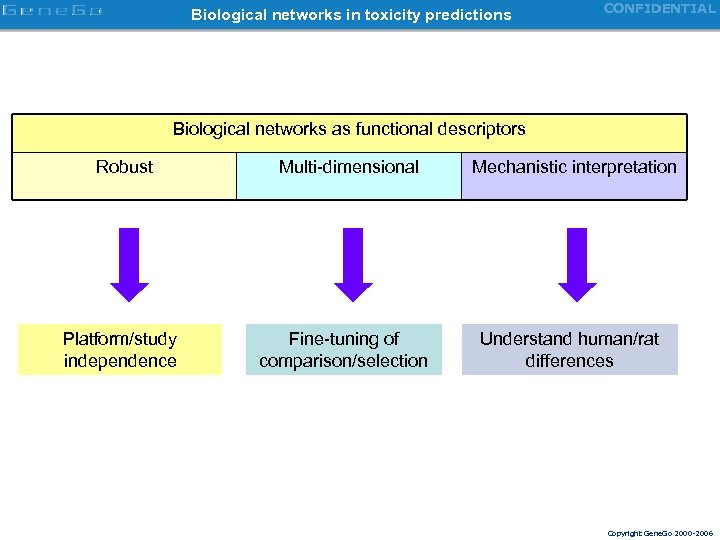 Biological networks in toxicity predictions CONFIDENTIAL Biological networks as functional descriptors Robust Platform/study independence Multi-dimensional Fine-tuning of comparison/selection Mechanistic interpretation Understand human/rat differences Copyright Gene. Go 2000 -2006
Biological networks in toxicity predictions CONFIDENTIAL Biological networks as functional descriptors Robust Platform/study independence Multi-dimensional Fine-tuning of comparison/selection Mechanistic interpretation Understand human/rat differences Copyright Gene. Go 2000 -2006
 Case study#2. Networks analysis of J&J toxicogenomics dataset CONFIDENTIAL • Data* – CEBS J&J set of 137 compounds, 600 profiles – 1, 471 genes c. DNA array – Pros • The only large scale dataset available • High quality • Internally consistent – Cons • Few compounds in each tox category • Small array • Non-standard array • Classifications – Toxic vs. drugs with no severe toxicity – By molecular mechanism of toxicity – By system/organ affected – By carcinogenicity – By type of hepatology – By basic therapeutic effect • The Analysis – Collection of pre-built networks: general and toxicity-related biological processes – Whole array mapping (no filtering) – P-value on functional categories: Gene. Go process networks, canonical pathways maps, GO, tox. networks – Linear Discriminant Analysis (LDA) models are built to evaluate performance of networks as toxicity predictors. A. Y. Nie et al. Predictive toxicogenomics approaches reveal underlying toxicogenomics mechanisms of nongenotoxic cardiogenicity. Mol. Carcinogenesis, 2006 Copyright Gene. Go 2000 -2006
Case study#2. Networks analysis of J&J toxicogenomics dataset CONFIDENTIAL • Data* – CEBS J&J set of 137 compounds, 600 profiles – 1, 471 genes c. DNA array – Pros • The only large scale dataset available • High quality • Internally consistent – Cons • Few compounds in each tox category • Small array • Non-standard array • Classifications – Toxic vs. drugs with no severe toxicity – By molecular mechanism of toxicity – By system/organ affected – By carcinogenicity – By type of hepatology – By basic therapeutic effect • The Analysis – Collection of pre-built networks: general and toxicity-related biological processes – Whole array mapping (no filtering) – P-value on functional categories: Gene. Go process networks, canonical pathways maps, GO, tox. networks – Linear Discriminant Analysis (LDA) models are built to evaluate performance of networks as toxicity predictors. A. Y. Nie et al. Predictive toxicogenomics approaches reveal underlying toxicogenomics mechanisms of nongenotoxic cardiogenicity. Mol. Carcinogenesis, 2006 Copyright Gene. Go 2000 -2006
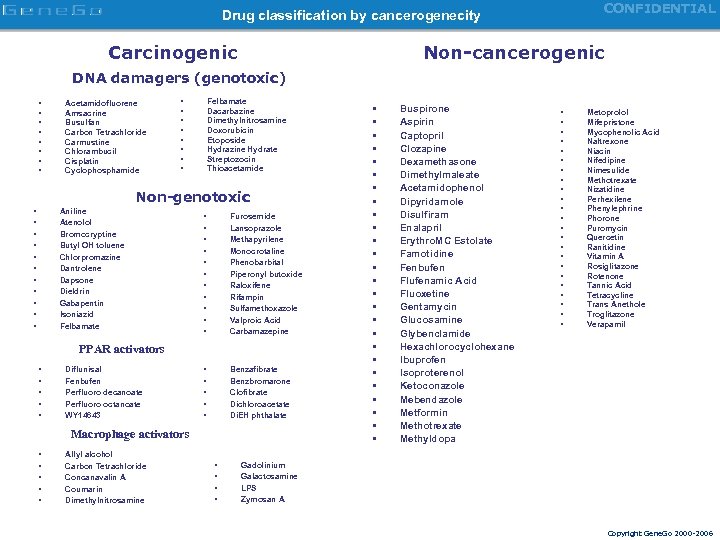 CONFIDENTIAL Drug classification by cancerogenecity Carcinogenic Non-cancerogenic DNA damagers (genotoxic) • • Acetamidofluorene Amsacrine Busulfan Carbon Tetrachloride Carmustine Chlorambucil Cisplatin Cyclophosphamide • • Felbamate Dacarbazine Dimethylnitrosamine Doxorubicin Etoposide Hydrazine Hydrate Streptozocin Thioacetamide Non-genotoxic • • • Aniline Atenolol Bromocryptine Butyl OH toluene Chlorpromazine Dantrolene Dapsone Dieldrin Gabapentin Isoniazid Felbamate • • • Furosemide Lansoprazole Methapyrilene Monocrotaline Phenobarbital Piperonyl butoxide Raloxifene Rifampin Sulfamethoxazole Valproic Acid Carbamazepine • • • Benzafibrate Benzbromarone Clofibrate Dichloroacetate Di. EH phthalate PPAR activators • • • Diflunisal Fenbufen Perfluoro decanoate Perfluoro octanoate WY 14643 Macrophage activators • • • Allyl alcohol Carbon Tetrachloride Concanavalin A Coumarin Dimethylnitrosamine • • • • • • • • Buspirone Aspirin Captopril Clozapine Dexamethasone Dimethylmaleate Acetamidophenol Dipyridamole Disulfiram Enalapril Erythro. MC Estolate Famotidine Fenbufen Flufenamic Acid Fluoxetine Gentamycin Glucosamine Glybenclamide Hexachlorocyclohexane Ibuprofen Isoproterenol Ketoconazole Mebendazole Metformin Methotrexate Methyldopa • • • • • • Metoprolol Mifepristone Mycophenolic Acid Naltrexone Niacin Nifedipine Nimesulide Methotrexate Nizatidine Perhexilene Phenylephrine Phorone Puromycin Quercetin Ranitidine Vitamin A Rosiglitazone Rotenone Tannic Acid Tetracycline Trans Anethole Troglitazone Verapamil Gadolinium Galactosamine LPS Zymosan A Copyright Gene. Go 2000 -2006
CONFIDENTIAL Drug classification by cancerogenecity Carcinogenic Non-cancerogenic DNA damagers (genotoxic) • • Acetamidofluorene Amsacrine Busulfan Carbon Tetrachloride Carmustine Chlorambucil Cisplatin Cyclophosphamide • • Felbamate Dacarbazine Dimethylnitrosamine Doxorubicin Etoposide Hydrazine Hydrate Streptozocin Thioacetamide Non-genotoxic • • • Aniline Atenolol Bromocryptine Butyl OH toluene Chlorpromazine Dantrolene Dapsone Dieldrin Gabapentin Isoniazid Felbamate • • • Furosemide Lansoprazole Methapyrilene Monocrotaline Phenobarbital Piperonyl butoxide Raloxifene Rifampin Sulfamethoxazole Valproic Acid Carbamazepine • • • Benzafibrate Benzbromarone Clofibrate Dichloroacetate Di. EH phthalate PPAR activators • • • Diflunisal Fenbufen Perfluoro decanoate Perfluoro octanoate WY 14643 Macrophage activators • • • Allyl alcohol Carbon Tetrachloride Concanavalin A Coumarin Dimethylnitrosamine • • • • • • • • Buspirone Aspirin Captopril Clozapine Dexamethasone Dimethylmaleate Acetamidophenol Dipyridamole Disulfiram Enalapril Erythro. MC Estolate Famotidine Fenbufen Flufenamic Acid Fluoxetine Gentamycin Glucosamine Glybenclamide Hexachlorocyclohexane Ibuprofen Isoproterenol Ketoconazole Mebendazole Metformin Methotrexate Methyldopa • • • • • • Metoprolol Mifepristone Mycophenolic Acid Naltrexone Niacin Nifedipine Nimesulide Methotrexate Nizatidine Perhexilene Phenylephrine Phorone Puromycin Quercetin Ranitidine Vitamin A Rosiglitazone Rotenone Tannic Acid Tetracycline Trans Anethole Troglitazone Verapamil Gadolinium Galactosamine LPS Zymosan A Copyright Gene. Go 2000 -2006
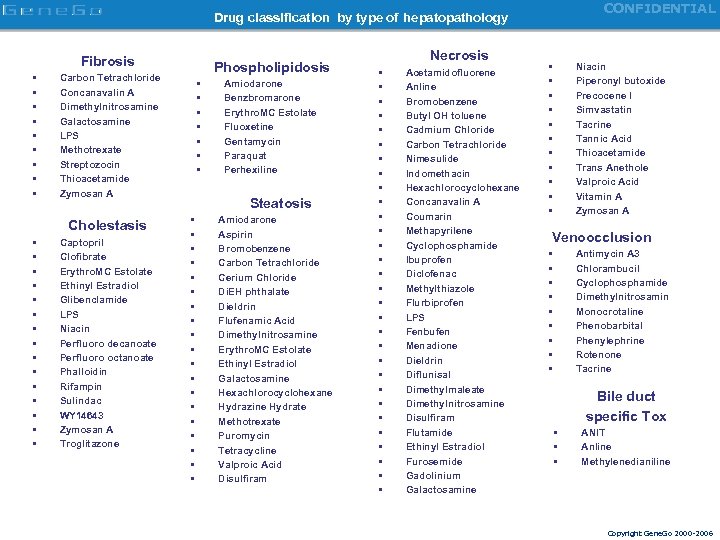 CONFIDENTIAL Drug classification by type of hepatopathology Fibrosis • • • Carbon Tetrachloride Concanavalin A Dimethylnitrosamine Galactosamine LPS Methotrexate Streptozocin Thioacetamide Zymosan A Cholestasis • • • • Phospholipidosis Captopril Clofibrate Erythro. MC Estolate Ethinyl Estradiol Glibenclamide LPS Niacin Perfluoro decanoate Perfluoro octanoate Phalloidin Rifampin Sulindac WY 14643 Zymosan A Troglitazone • • Amiodarone Benzbromarone Erythro. MC Estolate Fluoxetine Gentamycin Paraquat Perhexiline Steatosis • • • • • Amiodarone Aspirin Bromobenzene Carbon Tetrachloride Cerium Chloride Di. EH phthalate Dieldrin Flufenamic Acid Dimethylnitrosamine Erythro. MC Estolate Ethinyl Estradiol Galactosamine Hexachlorocyclohexane Hydrazine Hydrate Methotrexate Puromycin Tetracycline Valproic Acid Disulfiram Necrosis • • • • • • • • Acetamidofluorene Anline Bromobenzene Butyl OH toluene Cadmium Chloride Carbon Tetrachloride Nimesulide Indomethacin Hexachlorocyclohexane Concanavalin A Coumarin Methapyrilene Cyclophosphamide Ibuprofen Diclofenac Methylthiazole Flurbiprofen LPS Fenbufen Menadione Dieldrin Diflunisal Dimethylmaleate Dimethylnitrosamine Disulfiram Flutamide Ethinyl Estradiol Furosemide Gadolinium Galactosamine • • • Niacin Piperonyl butoxide Precocene I Simvastatin Tacrine Tannic Acid Thioacetamide Trans Anethole Valproic Acid Vitamin A Zymosan A Venoocclusion • • • Antimycin A 3 Chlorambucil Cyclophosphamide Dimethylnitrosamin Monocrotaline Phenobarbital Phenylephrine Rotenone Tacrine Bile duct specific Tox • • • ANIT Anline Methylenedianiline Copyright Gene. Go 2000 -2006
CONFIDENTIAL Drug classification by type of hepatopathology Fibrosis • • • Carbon Tetrachloride Concanavalin A Dimethylnitrosamine Galactosamine LPS Methotrexate Streptozocin Thioacetamide Zymosan A Cholestasis • • • • Phospholipidosis Captopril Clofibrate Erythro. MC Estolate Ethinyl Estradiol Glibenclamide LPS Niacin Perfluoro decanoate Perfluoro octanoate Phalloidin Rifampin Sulindac WY 14643 Zymosan A Troglitazone • • Amiodarone Benzbromarone Erythro. MC Estolate Fluoxetine Gentamycin Paraquat Perhexiline Steatosis • • • • • Amiodarone Aspirin Bromobenzene Carbon Tetrachloride Cerium Chloride Di. EH phthalate Dieldrin Flufenamic Acid Dimethylnitrosamine Erythro. MC Estolate Ethinyl Estradiol Galactosamine Hexachlorocyclohexane Hydrazine Hydrate Methotrexate Puromycin Tetracycline Valproic Acid Disulfiram Necrosis • • • • • • • • Acetamidofluorene Anline Bromobenzene Butyl OH toluene Cadmium Chloride Carbon Tetrachloride Nimesulide Indomethacin Hexachlorocyclohexane Concanavalin A Coumarin Methapyrilene Cyclophosphamide Ibuprofen Diclofenac Methylthiazole Flurbiprofen LPS Fenbufen Menadione Dieldrin Diflunisal Dimethylmaleate Dimethylnitrosamine Disulfiram Flutamide Ethinyl Estradiol Furosemide Gadolinium Galactosamine • • • Niacin Piperonyl butoxide Precocene I Simvastatin Tacrine Tannic Acid Thioacetamide Trans Anethole Valproic Acid Vitamin A Zymosan A Venoocclusion • • • Antimycin A 3 Chlorambucil Cyclophosphamide Dimethylnitrosamin Monocrotaline Phenobarbital Phenylephrine Rotenone Tacrine Bile duct specific Tox • • • ANIT Anline Methylenedianiline Copyright Gene. Go 2000 -2006
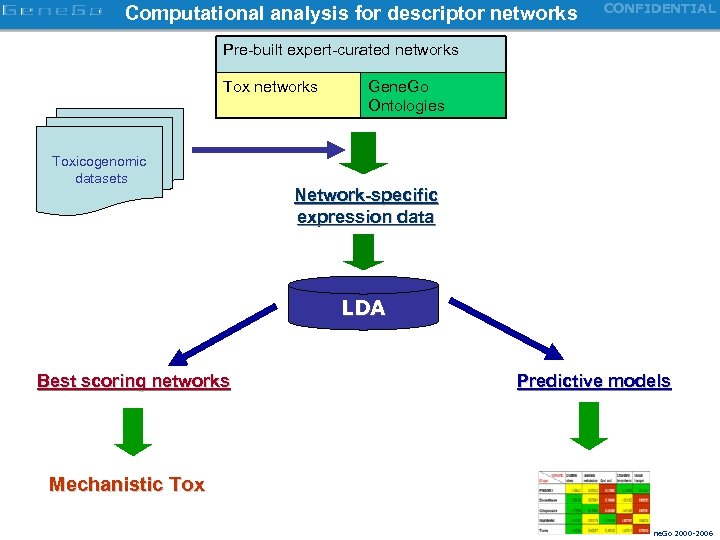 Computational analysis for descriptor networks CONFIDENTIAL Pre-built expert-curated networks Toxicogenomic datasets Gene. Go Ontologies Network-specific expression data LDA Best scoring networks Predictive models Mechanistic Tox Copyright Gene. Go 2000 -2006
Computational analysis for descriptor networks CONFIDENTIAL Pre-built expert-curated networks Toxicogenomic datasets Gene. Go Ontologies Network-specific expression data LDA Best scoring networks Predictive models Mechanistic Tox Copyright Gene. Go 2000 -2006
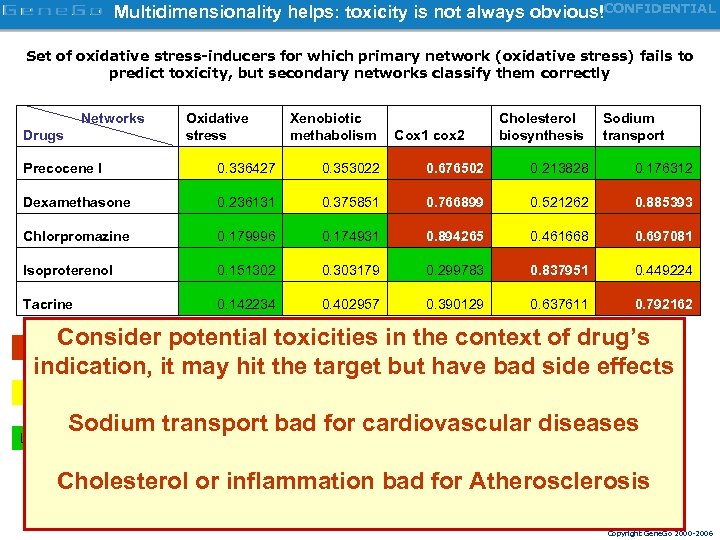 Multidimensionality helps: toxicity is not always obvious!CONFIDENTIAL Set of oxidative stress-inducers for which primary network (oxidative stress) fails to predict toxicity, but secondary networks classify them correctly Networks Drugs Oxidative stress Xenobiotic methabolism Cox 1 cox 2 Cholesterol biosynthesis Sodium transport Precocene I 0. 336427 0. 353022 0. 676502 0. 213828 0. 176312 Dexamethasone 0. 236131 0. 375851 0. 766899 0. 521262 0. 885393 Chlorpromazine 0. 179996 0. 174931 0. 894265 0. 461668 0. 697081 Isoproterenol 0. 151302 0. 303179 0. 299783 0. 837951 0. 449224 Tacrine 0. 142234 0. 402957 0. 390129 0. 637611 0. 792162 Consider potential toxicities in the context of drug’s indication, it may hit the target but have bad side effects Likely toxic May be toxic Sodium transport bad for cardiovascular diseases Likely nontoxic Cholesterol or inflammation bad for Atherosclerosis Copyright Gene. Go 2000 -2006
Multidimensionality helps: toxicity is not always obvious!CONFIDENTIAL Set of oxidative stress-inducers for which primary network (oxidative stress) fails to predict toxicity, but secondary networks classify them correctly Networks Drugs Oxidative stress Xenobiotic methabolism Cox 1 cox 2 Cholesterol biosynthesis Sodium transport Precocene I 0. 336427 0. 353022 0. 676502 0. 213828 0. 176312 Dexamethasone 0. 236131 0. 375851 0. 766899 0. 521262 0. 885393 Chlorpromazine 0. 179996 0. 174931 0. 894265 0. 461668 0. 697081 Isoproterenol 0. 151302 0. 303179 0. 299783 0. 837951 0. 449224 Tacrine 0. 142234 0. 402957 0. 390129 0. 637611 0. 792162 Consider potential toxicities in the context of drug’s indication, it may hit the target but have bad side effects Likely toxic May be toxic Sodium transport bad for cardiovascular diseases Likely nontoxic Cholesterol or inflammation bad for Atherosclerosis Copyright Gene. Go 2000 -2006
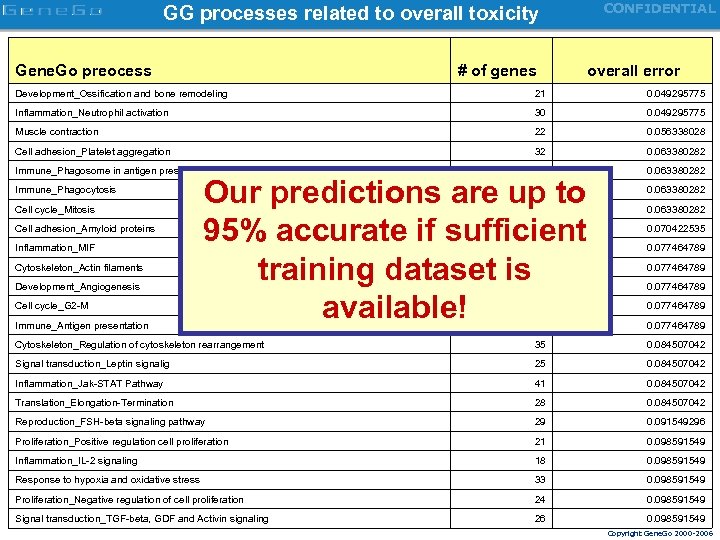 CONFIDENTIAL GG processes related to overall toxicity Gene. Go preocess # of genes overall error Development_Ossification and bone remodeling 21 0. 049295775 Inflammation_Neutrophil activation 30 0. 049295775 Muscle contraction 22 0. 056338028 Cell adhesion_Platelet aggregation 32 0. 063380282 Immune_Phagosome in antigen presentation 61 0. 063380282 Immune_Phagocytosis 54 0. 063380282 20 0. 063380282 30 0. 070422535 46 0. 077464789 22 0. 077464789 26 0. 077464789 23 0. 077464789 44 0. 077464789 Cytoskeleton_Regulation of cytoskeleton rearrangement 35 0. 084507042 Signal transduction_Leptin signalig 25 0. 084507042 Inflammation_Jak-STAT Pathway 41 0. 084507042 Translation_Elongation-Termination 28 0. 084507042 Reproduction_FSH-beta signaling pathway 29 0. 091549296 Proliferation_Positive regulation cell proliferation 21 0. 098591549 Inflammation_IL-2 signaling 18 0. 098591549 Response to hypoxia and oxidative stress 33 0. 098591549 Proliferation_Negative regulation of cell proliferation 24 0. 098591549 Signal transduction_TGF-beta, GDF and Activin signaling 26 0. 098591549 Cell cycle_Mitosis Cell adhesion_Amyloid proteins Inflammation_MIF Cytoskeleton_Actin filaments Development_Angiogenesis Cell cycle_G 2 -M Immune_Antigen presentation Our predictions are up to 95% accurate if sufficient training dataset is available! Copyright Gene. Go 2000 -2006
CONFIDENTIAL GG processes related to overall toxicity Gene. Go preocess # of genes overall error Development_Ossification and bone remodeling 21 0. 049295775 Inflammation_Neutrophil activation 30 0. 049295775 Muscle contraction 22 0. 056338028 Cell adhesion_Platelet aggregation 32 0. 063380282 Immune_Phagosome in antigen presentation 61 0. 063380282 Immune_Phagocytosis 54 0. 063380282 20 0. 063380282 30 0. 070422535 46 0. 077464789 22 0. 077464789 26 0. 077464789 23 0. 077464789 44 0. 077464789 Cytoskeleton_Regulation of cytoskeleton rearrangement 35 0. 084507042 Signal transduction_Leptin signalig 25 0. 084507042 Inflammation_Jak-STAT Pathway 41 0. 084507042 Translation_Elongation-Termination 28 0. 084507042 Reproduction_FSH-beta signaling pathway 29 0. 091549296 Proliferation_Positive regulation cell proliferation 21 0. 098591549 Inflammation_IL-2 signaling 18 0. 098591549 Response to hypoxia and oxidative stress 33 0. 098591549 Proliferation_Negative regulation of cell proliferation 24 0. 098591549 Signal transduction_TGF-beta, GDF and Activin signaling 26 0. 098591549 Cell cycle_Mitosis Cell adhesion_Amyloid proteins Inflammation_MIF Cytoskeleton_Actin filaments Development_Angiogenesis Cell cycle_G 2 -M Immune_Antigen presentation Our predictions are up to 95% accurate if sufficient training dataset is available! Copyright Gene. Go 2000 -2006
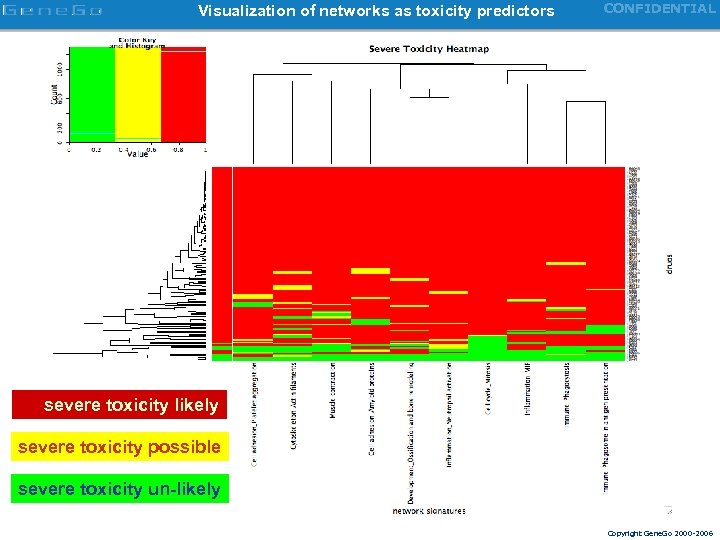 Visualization of networks as toxicity predictors CONFIDENTIAL severe toxicity likely severe toxicity possible severe toxicity un-likely Copyright Gene. Go 2000 -2006
Visualization of networks as toxicity predictors CONFIDENTIAL severe toxicity likely severe toxicity possible severe toxicity un-likely Copyright Gene. Go 2000 -2006
 Conclusions for case study #2 CONFIDENTIAL • Drugs with subtle manifestations of toxicity could be detected by multidimensional network signatures • Network descriptors allow to evaluate toxicity in the context of drug’s indications • For datasets of sufficient size, networks descriptors feature very high prediction accuracy (4% error) Copyright Gene. Go 2000 -2006
Conclusions for case study #2 CONFIDENTIAL • Drugs with subtle manifestations of toxicity could be detected by multidimensional network signatures • Network descriptors allow to evaluate toxicity in the context of drug’s indications • For datasets of sufficient size, networks descriptors feature very high prediction accuracy (4% error) Copyright Gene. Go 2000 -2006
 Annotation projects for Meta. Tox • Toxicity ontology and processes • Toxicity networks and maps • Human/mouse/rat differences – differential drug toxicity • Drug action/drug targets maps • CONFIDENTIAL Drug response from structure/ligand-receptor interactions Copyright Gene. Go 2000 -2006
Annotation projects for Meta. Tox • Toxicity ontology and processes • Toxicity networks and maps • Human/mouse/rat differences – differential drug toxicity • Drug action/drug targets maps • CONFIDENTIAL Drug response from structure/ligand-receptor interactions Copyright Gene. Go 2000 -2006
 CONFIDENTIAL Drug-induced pathological processes (Gene. Go annotations) • • • Drug-induced pathological processes Drug-induced pathology classification has a hierarchical structure and is based on literature data Genes associated with drug-induced pathologies Compounds associated with drug-induced pathologies Cited articles (unique) 38 600 30 250 (!) Public domain DOES NOT have structured information (which can be queried for miming and download) about drug-induced pathological processes connections and genes. Copyright Gene. Go 2000 -2006
CONFIDENTIAL Drug-induced pathological processes (Gene. Go annotations) • • • Drug-induced pathological processes Drug-induced pathology classification has a hierarchical structure and is based on literature data Genes associated with drug-induced pathologies Compounds associated with drug-induced pathologies Cited articles (unique) 38 600 30 250 (!) Public domain DOES NOT have structured information (which can be queried for miming and download) about drug-induced pathological processes connections and genes. Copyright Gene. Go 2000 -2006
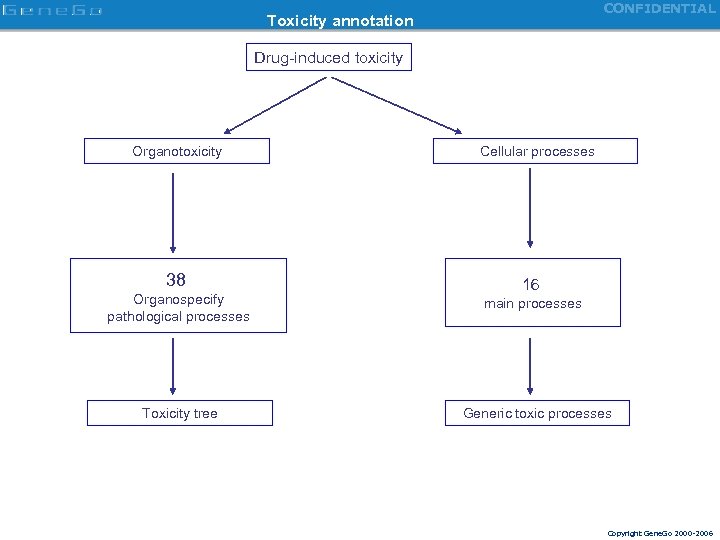 CONFIDENTIAL Toxicity annotation Drug-induced toxicity Organotoxicity Cellular processes 38 16 Organospecify pathological processes Toxicity tree main processes Generic toxic processes Copyright Gene. Go 2000 -2006
CONFIDENTIAL Toxicity annotation Drug-induced toxicity Organotoxicity Cellular processes 38 16 Organospecify pathological processes Toxicity tree main processes Generic toxic processes Copyright Gene. Go 2000 -2006
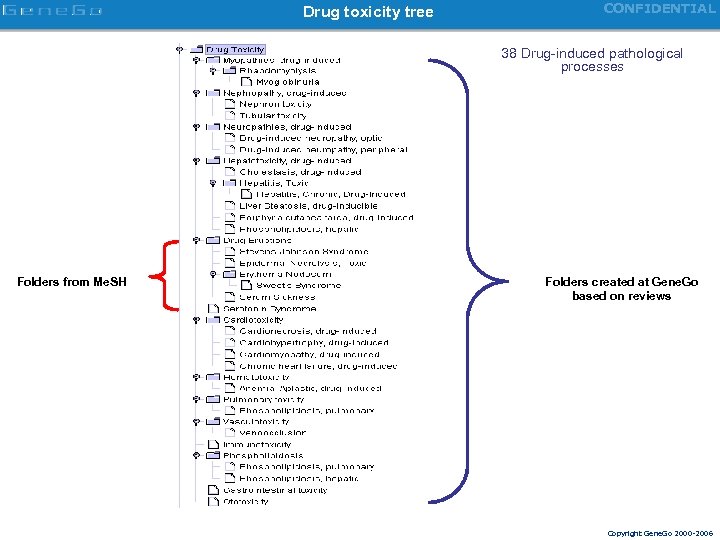 Drug toxicity tree CONFIDENTIAL 38 Drug-induced pathological processes Folders from Me. SH Folders created at Gene. Go based on reviews Copyright Gene. Go 2000 -2006
Drug toxicity tree CONFIDENTIAL 38 Drug-induced pathological processes Folders from Me. SH Folders created at Gene. Go based on reviews Copyright Gene. Go 2000 -2006
 Generic tox-response processes CONFIDENTIAL Process Cell cycle Apoptosis DNA damage and repair Adhesion, cytoskeleton, ECM Proliferation Inflammation Immune response Blood coagulation Oxidative stress Gene expression (transcription, translation) Chemotaxis Development Signal Transduction Transport Metabolism Protein folding Copyright Gene. Go 2000 -2006
Generic tox-response processes CONFIDENTIAL Process Cell cycle Apoptosis DNA damage and repair Adhesion, cytoskeleton, ECM Proliferation Inflammation Immune response Blood coagulation Oxidative stress Gene expression (transcription, translation) Chemotaxis Development Signal Transduction Transport Metabolism Protein folding Copyright Gene. Go 2000 -2006
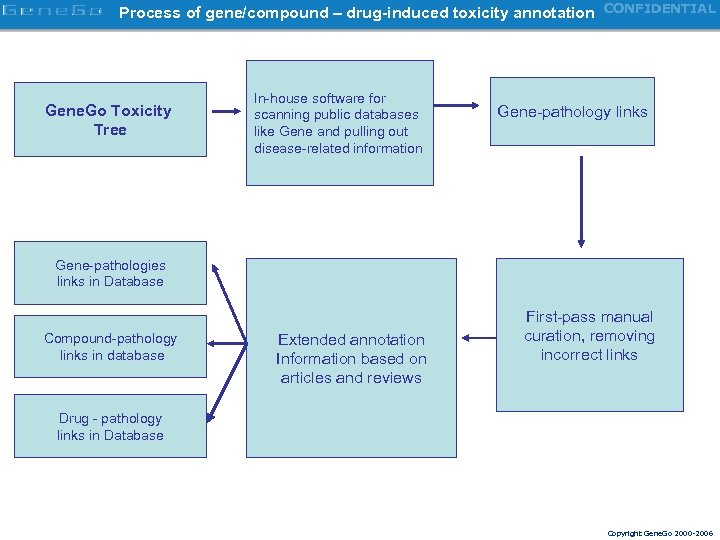 Process of gene/compound – drug-induced toxicity annotation CONFIDENTIAL Gene. Go Toxicity Tree In-house software for scanning public databases like Gene and pulling out disease-related information Gene-pathology links Gene-pathologies links in Database Compound-pathology links in database Extended annotation Information based on articles and reviews First-pass manual curation, removing incorrect links Drug - pathology links in Database Copyright Gene. Go 2000 -2006
Process of gene/compound – drug-induced toxicity annotation CONFIDENTIAL Gene. Go Toxicity Tree In-house software for scanning public databases like Gene and pulling out disease-related information Gene-pathology links Gene-pathologies links in Database Compound-pathology links in database Extended annotation Information based on articles and reviews First-pass manual curation, removing incorrect links Drug - pathology links in Database Copyright Gene. Go 2000 -2006
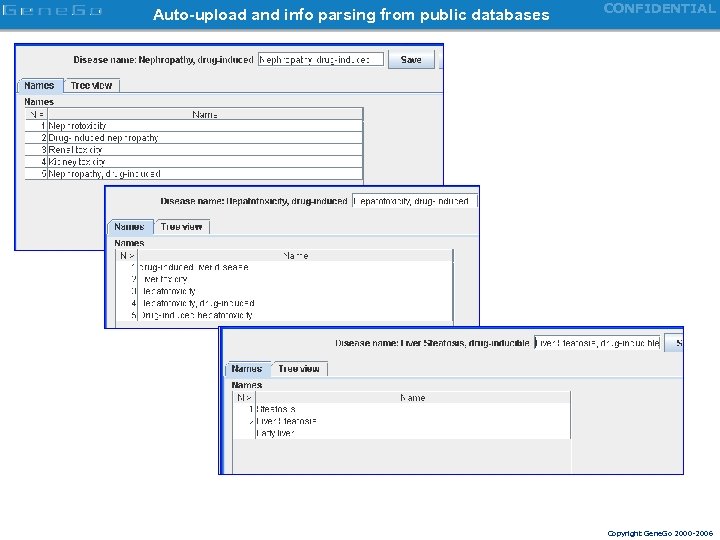 Auto-upload and info parsing from public databases CONFIDENTIAL Copyright Gene. Go 2000 -2006
Auto-upload and info parsing from public databases CONFIDENTIAL Copyright Gene. Go 2000 -2006
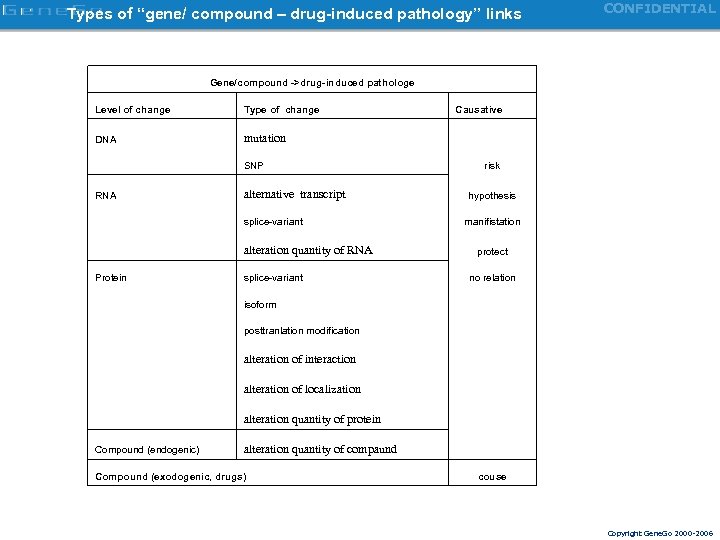 Types of “gene/ compound – drug-induced pathology” links CONFIDENTIAL Gene/compound ->drug-induced pathologe Level of change Type of change Causative DNA mutation SNP RNA alternative transcript splice-variant alteration quantity of RNA Protein splice-variant isoform posttranlation modification alteration of interaction alteration of localization alteration quantity of protein Compound (endogenic) alteration quantity of compaund Compound (exodogenic, drugs) risk hypothesis manifistation protect no relation couse Copyright Gene. Go 2000 -2006
Types of “gene/ compound – drug-induced pathology” links CONFIDENTIAL Gene/compound ->drug-induced pathologe Level of change Type of change Causative DNA mutation SNP RNA alternative transcript splice-variant alteration quantity of RNA Protein splice-variant isoform posttranlation modification alteration of interaction alteration of localization alteration quantity of protein Compound (endogenic) alteration quantity of compaund Compound (exodogenic, drugs) risk hypothesis manifistation protect no relation couse Copyright Gene. Go 2000 -2006
 Gene – toxicity annotation. Example 1 CONFIDENTIAL Copyright Gene. Go 2000 -2006
Gene – toxicity annotation. Example 1 CONFIDENTIAL Copyright Gene. Go 2000 -2006
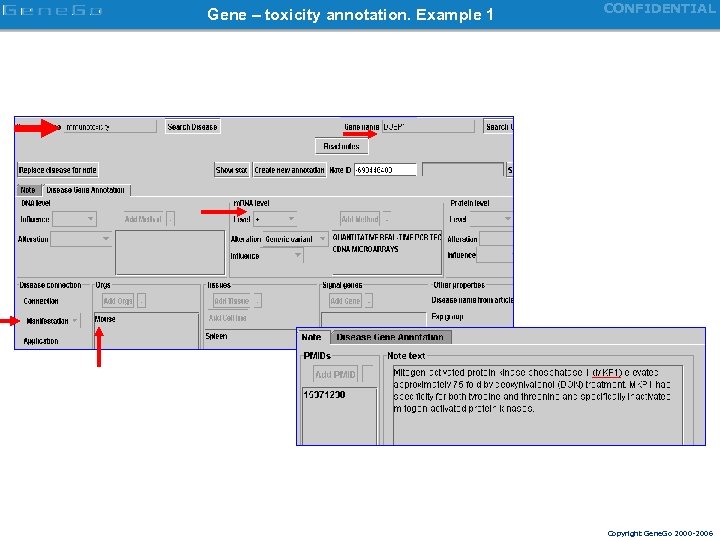 Gene – toxicity annotation. Example 1 CONFIDENTIAL Copyright Gene. Go 2000 -2006
Gene – toxicity annotation. Example 1 CONFIDENTIAL Copyright Gene. Go 2000 -2006
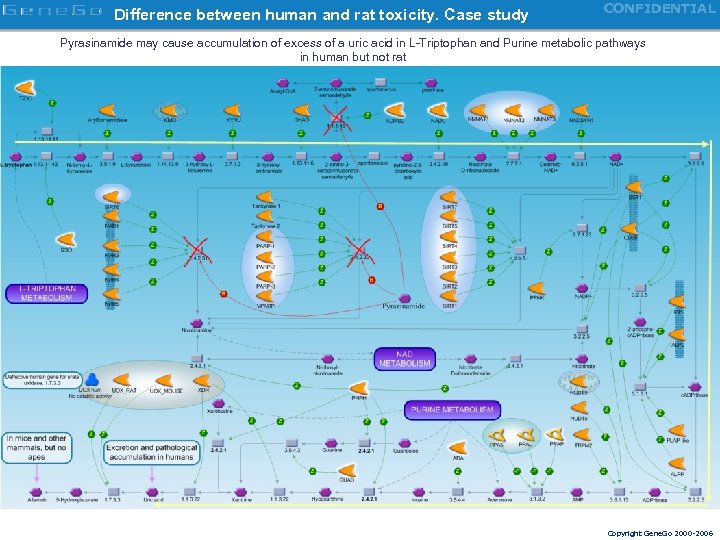 Difference between human and rat toxicity. Case study CONFIDENTIAL Pyrasinamide may cause accumulation of excess of a uric acid in L-Triptophan and Purine metabolic pathways in human but not rat Copyright Gene. Go 2000 -2006
Difference between human and rat toxicity. Case study CONFIDENTIAL Pyrasinamide may cause accumulation of excess of a uric acid in L-Triptophan and Purine metabolic pathways in human but not rat Copyright Gene. Go 2000 -2006
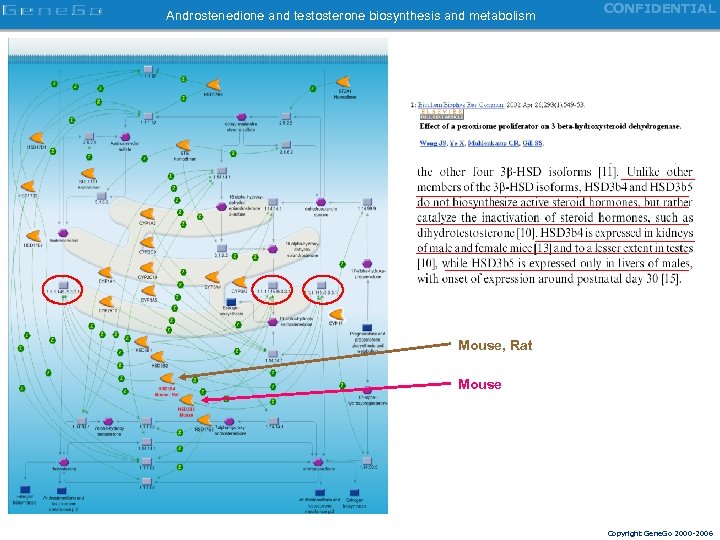 Androstenedione and testosterone biosynthesis and metabolism CONFIDENTIAL Mouse, Rat Mouse Copyright Gene. Go 2000 -2006
Androstenedione and testosterone biosynthesis and metabolism CONFIDENTIAL Mouse, Rat Mouse Copyright Gene. Go 2000 -2006
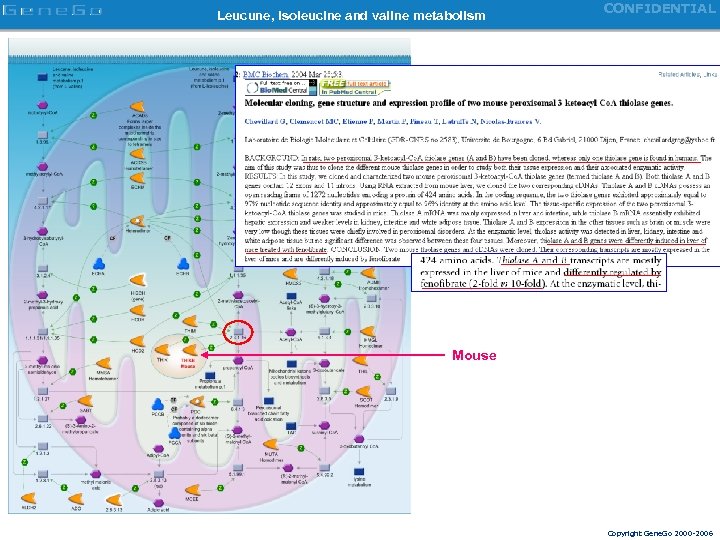 Leucune, isoleucine and valine metabolism CONFIDENTIAL Mouse Copyright Gene. Go 2000 -2006
Leucune, isoleucine and valine metabolism CONFIDENTIAL Mouse Copyright Gene. Go 2000 -2006
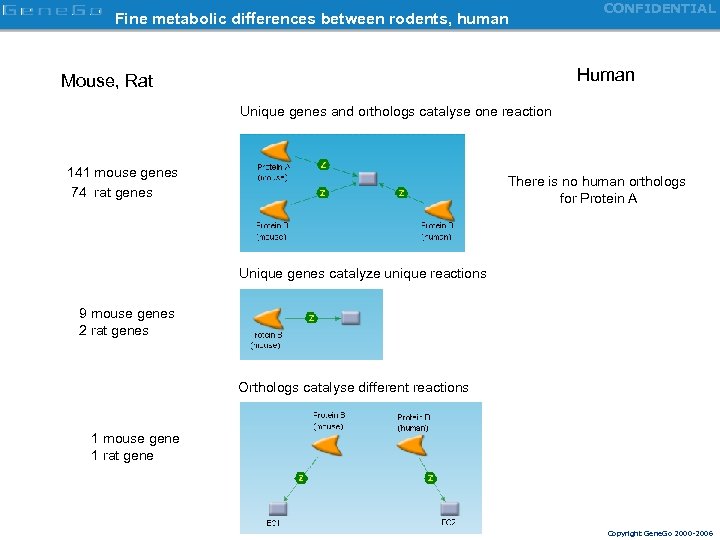 Fine metabolic differences between rodents, human CONFIDENTIAL Unique genes Human Mouse, Rat Unique genes and orthologs catalyse one reaction 141 mouse genes 74 rat genes There is no human orthologs for Protein A Unique genes catalyze unique reactions 9 mouse genes 2 rat genes Orthologs catalyse different reactions 1 mouse gene 1 rat gene Copyright Gene. Go 2000 -2006
Fine metabolic differences between rodents, human CONFIDENTIAL Unique genes Human Mouse, Rat Unique genes and orthologs catalyse one reaction 141 mouse genes 74 rat genes There is no human orthologs for Protein A Unique genes catalyze unique reactions 9 mouse genes 2 rat genes Orthologs catalyse different reactions 1 mouse gene 1 rat gene Copyright Gene. Go 2000 -2006
 Annotation of protein complexes common, different for three organisms CONFIDENTIAL Object contains protein-complex for three organisms Copyright Gene. Go 2000 -2006
Annotation of protein complexes common, different for three organisms CONFIDENTIAL Object contains protein-complex for three organisms Copyright Gene. Go 2000 -2006
 Meta. Tox Consortium Business Model • • CONFIDENTIAL Potential members: Boehringer Ingelheim, JNJ, Merck KGa. A, Pfizer, Novartis, GSK, Wyeth, Roche, Vertex, Serono, Altana, BMS, Eli Lilly…. Charter member: FDA Director: Richard Brennan 3 years: annual meetings including training Early access to content, workflows and analysis tools – Includes unlimited access to all Meta. Tox products for the term of the consortium and one perpetual named user license thereafter Immediate deliverables – Two additional Meta. Drug seats, One named user Map. Editor and One named user Meta. Link ($144 k value) – 400 Pre built Tox networks as part of Meta. Drug – Two-ways connectivity with Array. Track (FDA) – An uploaded normalized set of 137 compound-response signatures from CEBS JNJ • Gene. Go experts have made the data ready for analysis and comparison • Compound profile grouped by type of toxicity and mode of action Consortium member option: – Each member may provide 100 compounds with expression data using whole genome arrays under CDA. This information will not be shared with other members. The data will be used to develop tox signature networks that will be available to all members. The original data will be returned to the member Copyright Gene. Go 2000 -2006
Meta. Tox Consortium Business Model • • CONFIDENTIAL Potential members: Boehringer Ingelheim, JNJ, Merck KGa. A, Pfizer, Novartis, GSK, Wyeth, Roche, Vertex, Serono, Altana, BMS, Eli Lilly…. Charter member: FDA Director: Richard Brennan 3 years: annual meetings including training Early access to content, workflows and analysis tools – Includes unlimited access to all Meta. Tox products for the term of the consortium and one perpetual named user license thereafter Immediate deliverables – Two additional Meta. Drug seats, One named user Map. Editor and One named user Meta. Link ($144 k value) – 400 Pre built Tox networks as part of Meta. Drug – Two-ways connectivity with Array. Track (FDA) – An uploaded normalized set of 137 compound-response signatures from CEBS JNJ • Gene. Go experts have made the data ready for analysis and comparison • Compound profile grouped by type of toxicity and mode of action Consortium member option: – Each member may provide 100 compounds with expression data using whole genome arrays under CDA. This information will not be shared with other members. The data will be used to develop tox signature networks that will be available to all members. The original data will be returned to the member Copyright Gene. Go 2000 -2006


