b9fde4d3383f6712fc2f071a62e67e1e.ppt
- Количество слайдов: 31
 Computational prediction of mi. RNA and mi. RNA-disease relationship Quan Zou (邹权) PH. D. &Professor School of Computer Sci&Tech Tianjin University, China
Computational prediction of mi. RNA and mi. RNA-disease relationship Quan Zou (邹权) PH. D. &Professor School of Computer Sci&Tech Tianjin University, China
 Contents Ø background Ø micro. RNA identification Ø isomi. R Ø micro. RNA and disease Ø outlook 2
Contents Ø background Ø micro. RNA identification Ø isomi. R Ø micro. RNA and disease Ø outlook 2
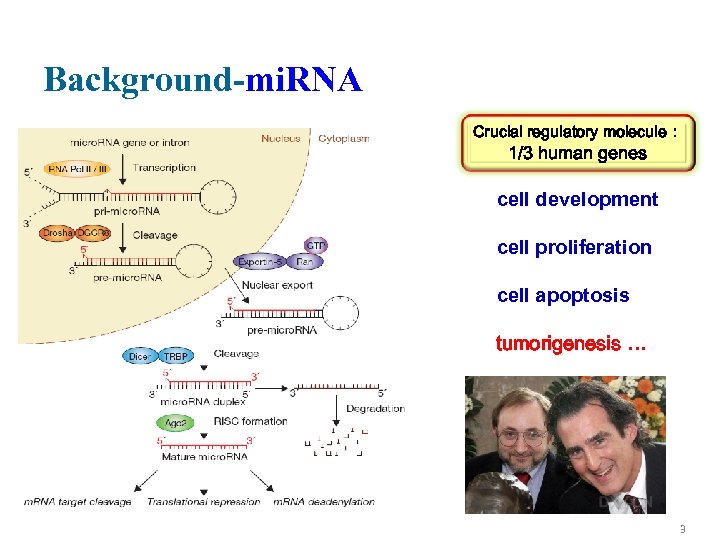 Background-mi. RNA Crucial regulatory molecule: 1/3 human genes cell development cell proliferation cell apoptosis tumorigenesis … 3
Background-mi. RNA Crucial regulatory molecule: 1/3 human genes cell development cell proliferation cell apoptosis tumorigenesis … 3
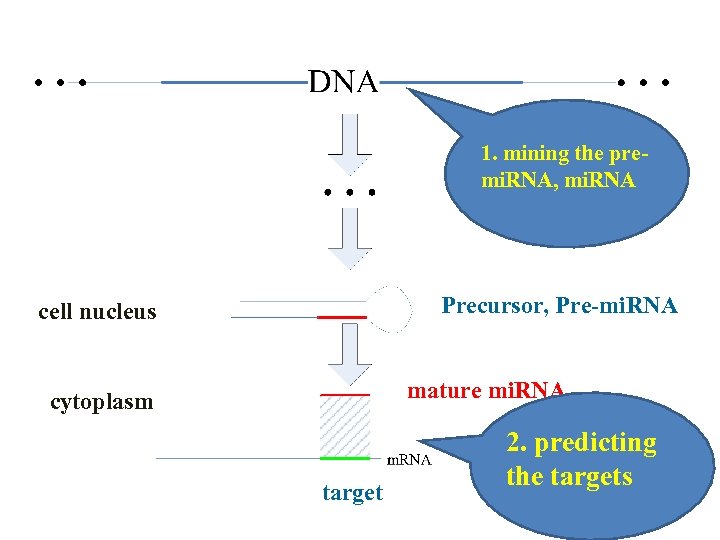 1. mining the premi. RNA, mi. RNA Precursor, Pre-mi. RNA cell nucleus mature mi. RNA cytoplasm target 2. predicting the targets
1. mining the premi. RNA, mi. RNA Precursor, Pre-mi. RNA cell nucleus mature mi. RNA cytoplasm target 2. predicting the targets
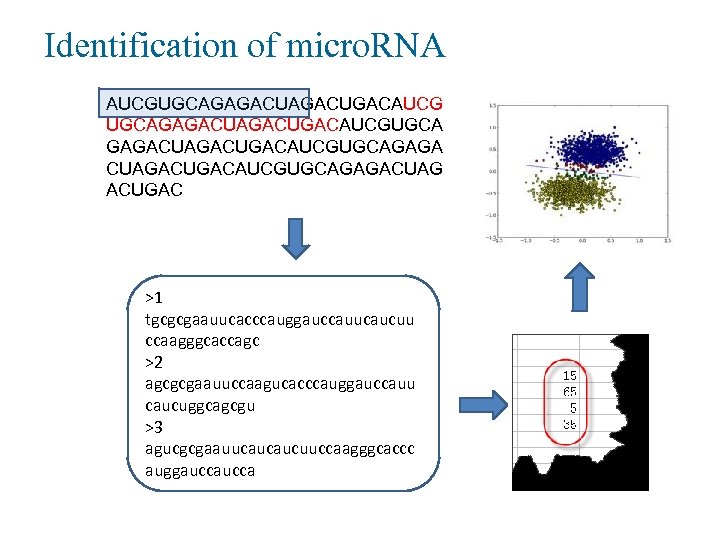 Identification of micro. RNA AUCGUGCAGAGACUGACAUCGUGCA GAGACUGACAUCGUGCAGAGA CUAGACUGACAUCGUGCAGAGACUAG ACUGAC >1 tgcgcgaauucacccauggauccauucaucuu ccaagggcaccagc >2 agcgcgaauuccaagucacccauggauccauu caucuggcagcgu >3 agucgcgaauucaucaucuuccaagggcaccc auggaucca
Identification of micro. RNA AUCGUGCAGAGACUGACAUCGUGCA GAGACUGACAUCGUGCAGAGA CUAGACUGACAUCGUGCAGAGACUAG ACUGAC >1 tgcgcgaauucacccauggauccauucaucuu ccaagggcaccagc >2 agcgcgaauuccaagucacccauggauccauu caucuggcagcgu >3 agucgcgaauucaucaucuuccaagggcaccc auggaucca
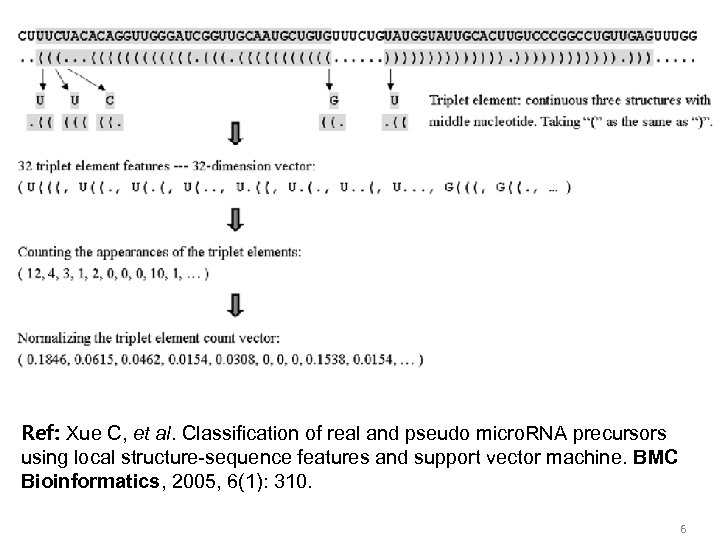 Ref: Xue C, et al. Classification of real and pseudo micro. RNA precursors using local structure-sequence features and support vector machine. BMC Bioinformatics, 2005, 6(1): 310. 6
Ref: Xue C, et al. Classification of real and pseudo micro. RNA precursors using local structure-sequence features and support vector machine. BMC Bioinformatics, 2005, 6(1): 310. 6
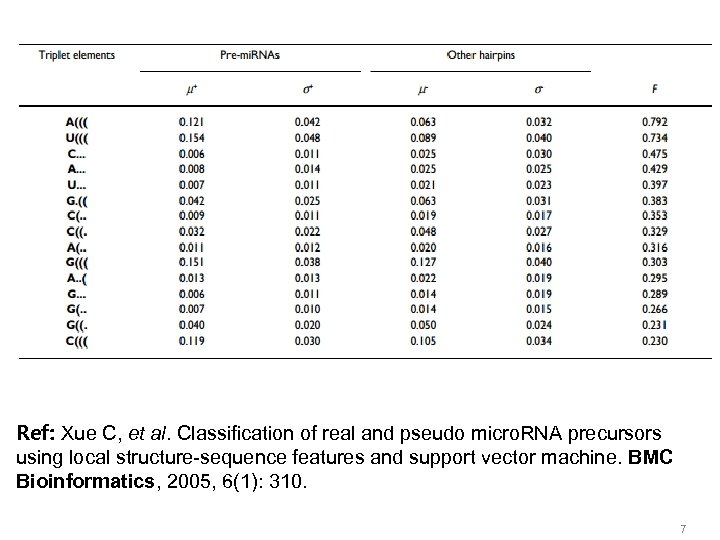 Ref: Xue C, et al. Classification of real and pseudo micro. RNA precursors using local structure-sequence features and support vector machine. BMC Bioinformatics, 2005, 6(1): 310. 7
Ref: Xue C, et al. Classification of real and pseudo micro. RNA precursors using local structure-sequence features and support vector machine. BMC Bioinformatics, 2005, 6(1): 310. 7
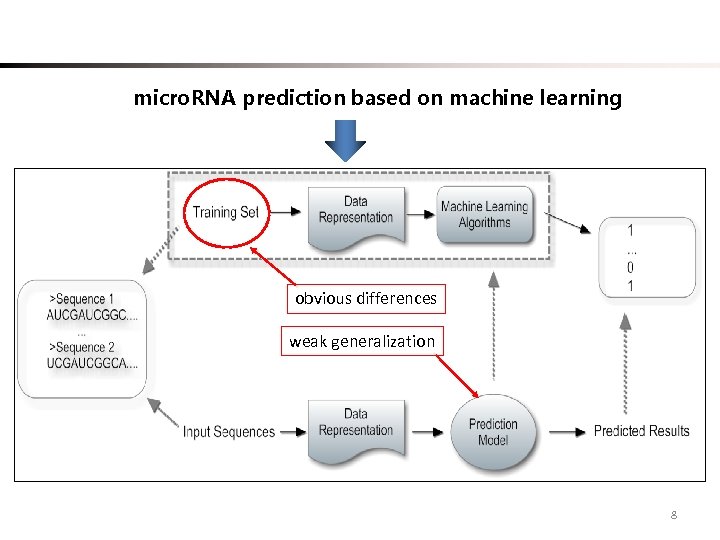 micro. RNA prediction based on machine learning obvious differences weak generalization 8
micro. RNA prediction based on machine learning obvious differences weak generalization 8
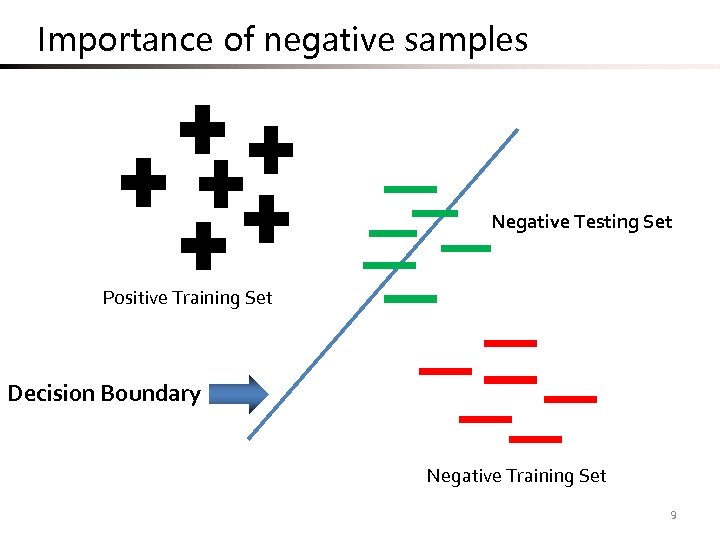 Importance of negative samples Negative Testing Set Positive Training Set Decision Boundary Negative Training Set 9
Importance of negative samples Negative Testing Set Positive Training Set Decision Boundary Negative Training Set 9
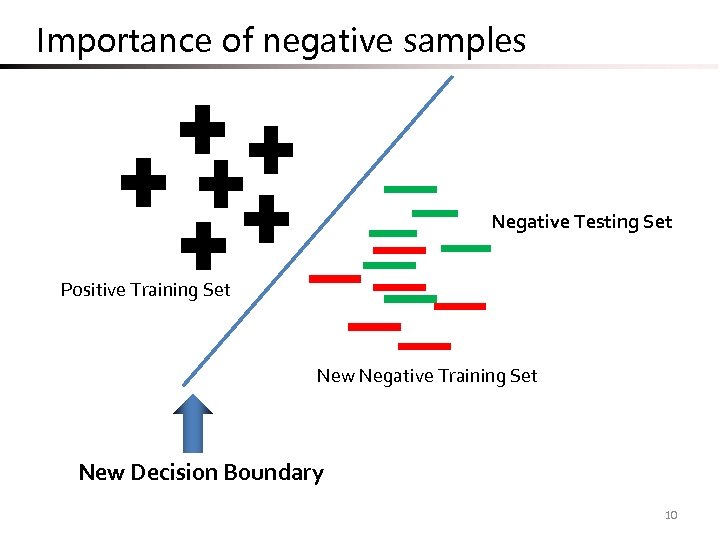 Importance of negative samples Negative Testing Set Positive Training Set New Negative Training Set New Decision Boundary 10
Importance of negative samples Negative Testing Set Positive Training Set New Negative Training Set New Decision Boundary 10
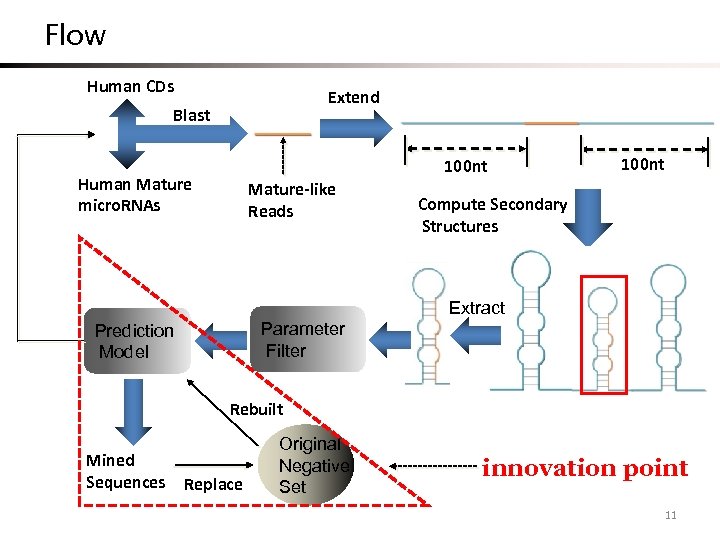 Flow Human CDs Extend Blast 100 nt Human Mature micro. RNAs Mature-like Reads 100 nt Compute Secondary Structures Extract Parameter Filter Prediction Model Rebuilt Mined Sequences Replace Original Negative Set innovation point 11
Flow Human CDs Extend Blast 100 nt Human Mature micro. RNAs Mature-like Reads 100 nt Compute Secondary Structures Extract Parameter Filter Prediction Model Rebuilt Mined Sequences Replace Original Negative Set innovation point 11
 Leyi Wei, Minghong Liao, Yue Gao, Rongrong Ji, Zengyou He*, Quan Zou(邹权)*. Improved and Promising Identification of Human Micro. RNAs by Incorporating a High-quality Negative Set. IEEE/ACM Transactions on Computational Biology and Bioinformatics. 2014, 11(1): 192 -201 (SCI, IF 2011=1. 543) 2018/3/18 12/30
Leyi Wei, Minghong Liao, Yue Gao, Rongrong Ji, Zengyou He*, Quan Zou(邹权)*. Improved and Promising Identification of Human Micro. RNAs by Incorporating a High-quality Negative Set. IEEE/ACM Transactions on Computational Biology and Bioinformatics. 2014, 11(1): 192 -201 (SCI, IF 2011=1. 543) 2018/3/18 12/30
 Novel mi. RNA found by our method 1 13/30
Novel mi. RNA found by our method 1 13/30
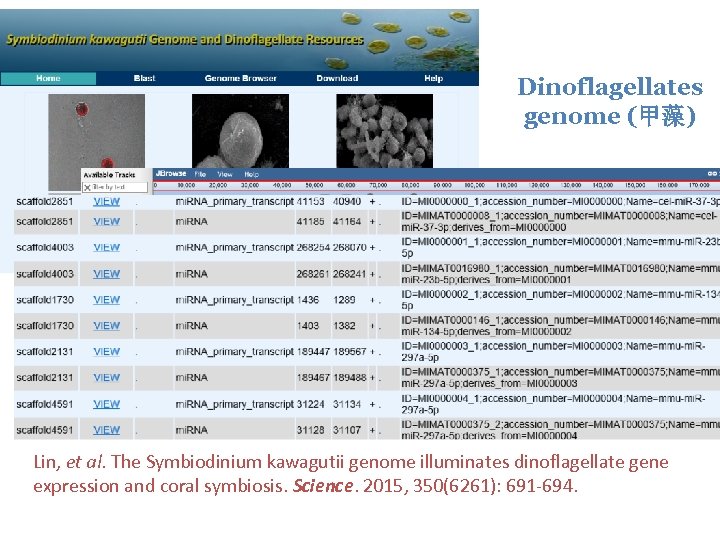 Dinoflagellates genome (甲藻) Lin, et al. The Symbiodinium kawagutii genome illuminates dinoflagellate gene expression and coral symbiosis. Science. 2015, 350(6261): 691 -694.
Dinoflagellates genome (甲藻) Lin, et al. The Symbiodinium kawagutii genome illuminates dinoflagellate gene expression and coral symbiosis. Science. 2015, 350(6261): 691 -694.
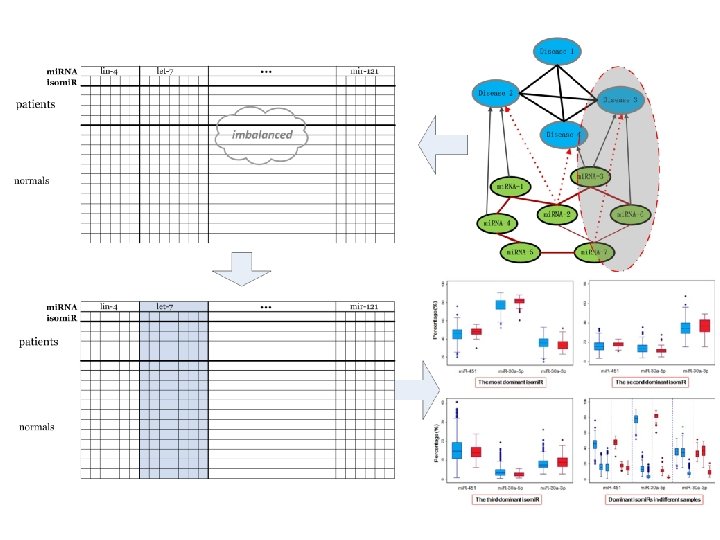
 mi. RNA family classification • PFAM(~2000) VS mi. RNA family(~2000) • Troubles – Multiple classes – Few samples – imbalaned 1 15/30
mi. RNA family classification • PFAM(~2000) VS mi. RNA family(~2000) • Troubles – Multiple classes – Few samples – imbalaned 1 15/30
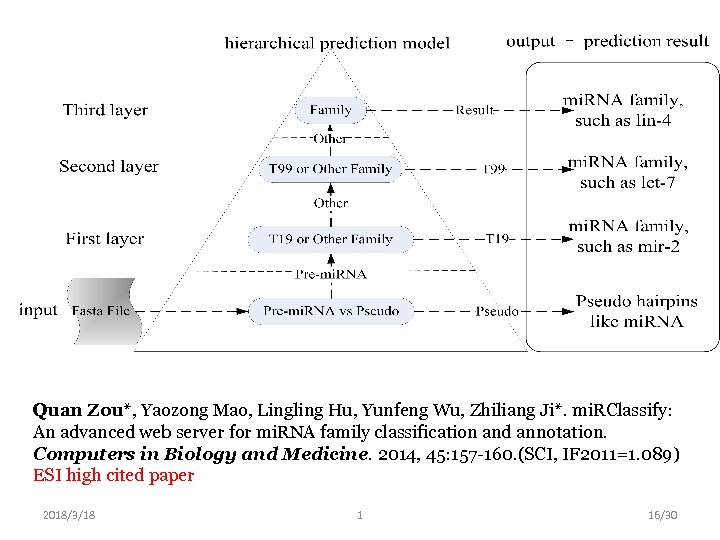 Quan Zou*, Yaozong Mao, Lingling Hu, Yunfeng Wu, Zhiliang Ji*. mi. RClassify: An advanced web server for mi. RNA family classification and annotation. Computers in Biology and Medicine. 2014, 45: 157 -160. (SCI, IF 2011=1. 089) ESI high cited paper 2018/3/18 1 16/30
Quan Zou*, Yaozong Mao, Lingling Hu, Yunfeng Wu, Zhiliang Ji*. mi. RClassify: An advanced web server for mi. RNA family classification and annotation. Computers in Biology and Medicine. 2014, 45: 157 -160. (SCI, IF 2011=1. 089) ESI high cited paper 2018/3/18 1 16/30
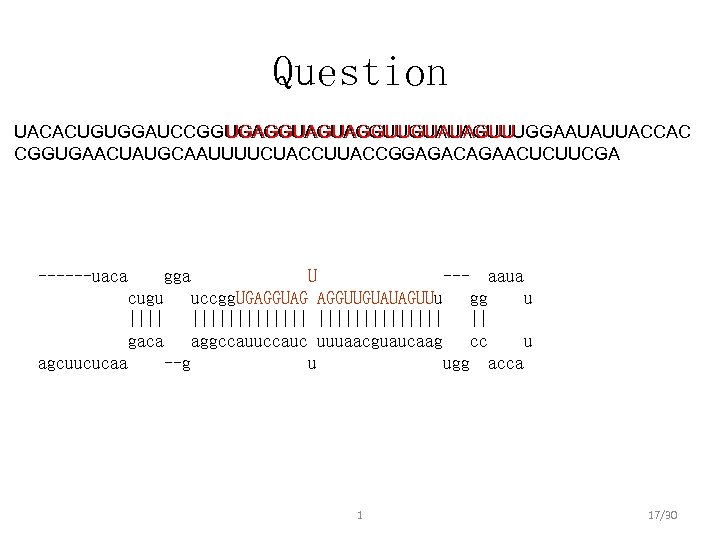 Question UGAGGUAGUAGGUUGUAUAGUU UACACUGUGGAUCCGGUGAGGUAGUAGGUUGUAUAGUUUGGAAUAUUACCAC CGGUGAACUAUGCAAUUUUCUACCUUACCGGAGACAGAACUCUUCGA ------uaca gga U --- aaua cugu uccgg. UGAGGUAG AGGUUGUAUAGUUu gg u ||||||||| || gaca aggccauuccauc uuuaacguaucaag cc u agcuucucaa --g u ugg acca 1 17/30
Question UGAGGUAGUAGGUUGUAUAGUU UACACUGUGGAUCCGGUGAGGUAGUAGGUUGUAUAGUUUGGAAUAUUACCAC CGGUGAACUAUGCAAUUUUCUACCUUACCGGAGACAGAACUCUUCGA ------uaca gga U --- aaua cugu uccgg. UGAGGUAG AGGUUGUAUAGUUu gg u ||||||||| || gaca aggccauuccauc uuuaacguaucaag cc u agcuucucaa --g u ugg acca 1 17/30
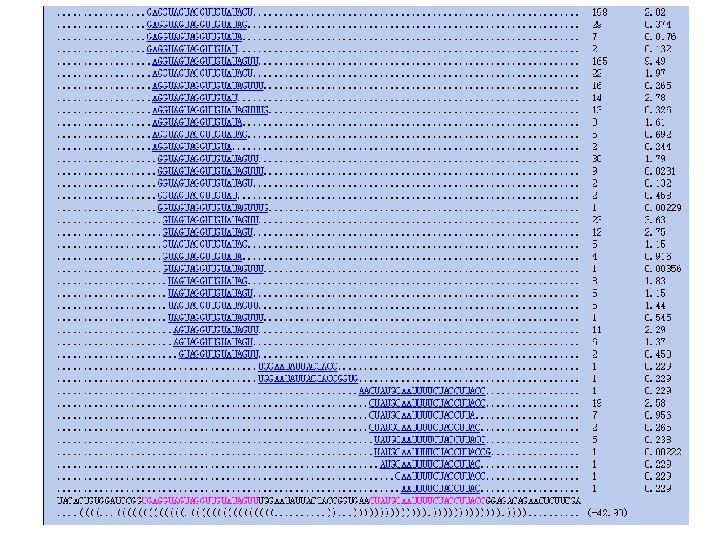 1 18/30
1 18/30
 Contents Ø background Ø micro. RNA identification Ø isomi. R Ø micro. RNA and disease Ø outlook 19
Contents Ø background Ø micro. RNA identification Ø isomi. R Ø micro. RNA and disease Ø outlook 19
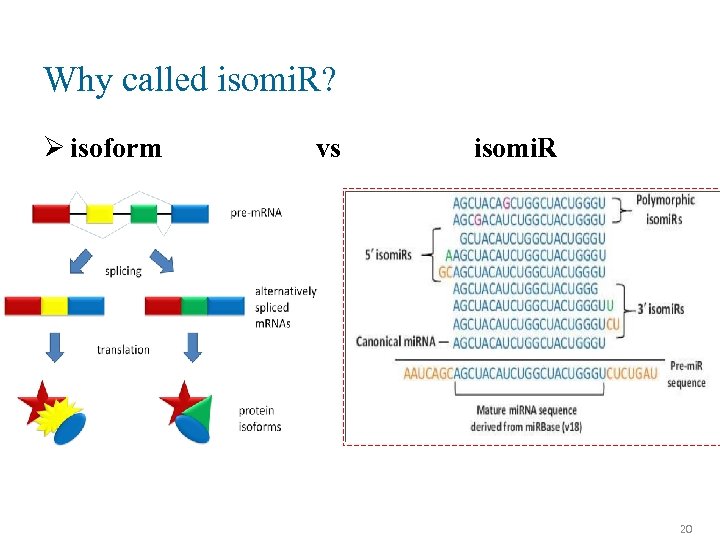 Why called isomi. R? Ø isoform vs isomi. R 20
Why called isomi. R? Ø isoform vs isomi. R 20
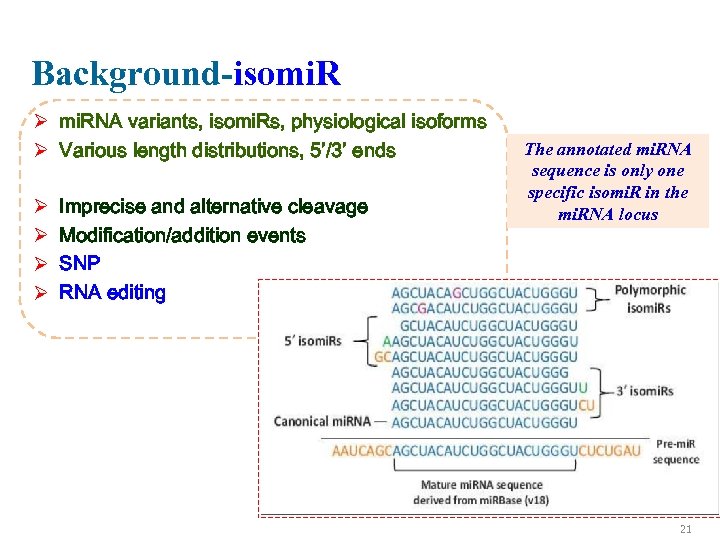 Background-isomi. R Ø mi. RNA variants, isomi. Rs, physiological isoforms Ø Various length distributions, 5’/3’ ends Ø Ø Imprecise and alternative cleavage Modification/addition events SNP RNA editing The annotated mi. RNA sequence is only one specific isomi. R in the mi. RNA locus 21
Background-isomi. R Ø mi. RNA variants, isomi. Rs, physiological isoforms Ø Various length distributions, 5’/3’ ends Ø Ø Imprecise and alternative cleavage Modification/addition events SNP RNA editing The annotated mi. RNA sequence is only one specific isomi. R in the mi. RNA locus 21
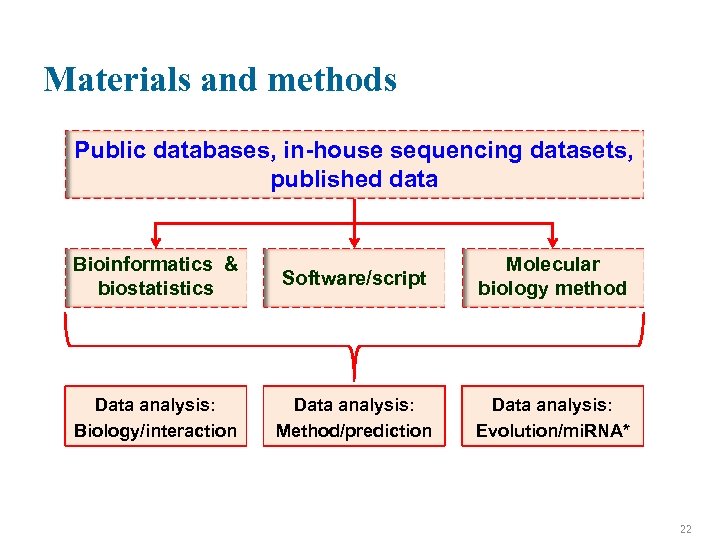 Materials and methods Public databases, in-house sequencing datasets, published data Bioinformatics & biostatistics Software/script Molecular biology method Data analysis: Biology/interaction Data analysis: Method/prediction Data analysis: Evolution/mi. RNA* 22
Materials and methods Public databases, in-house sequencing datasets, published data Bioinformatics & biostatistics Software/script Molecular biology method Data analysis: Biology/interaction Data analysis: Method/prediction Data analysis: Evolution/mi. RNA* 22
 Where does isomi. R happen? Ø across different species Ø normal vs cancer n isomi. R data - TCGA 23
Where does isomi. R happen? Ø across different species Ø normal vs cancer n isomi. R data - TCGA 23
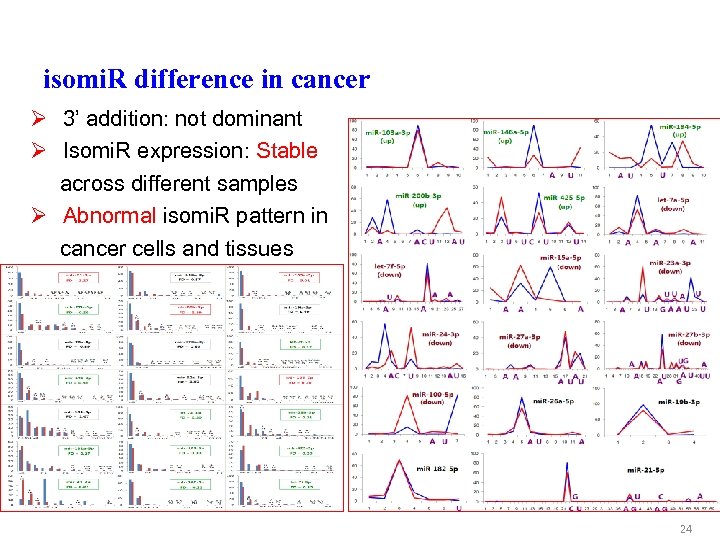 isomi. R difference in cancer Ø 3’ addition: not dominant Ø Isomi. R expression: Stable across different samples Ø Abnormal isomi. R pattern in cancer cells and tissues 24
isomi. R difference in cancer Ø 3’ addition: not dominant Ø Isomi. R expression: Stable across different samples Ø Abnormal isomi. R pattern in cancer cells and tissues 24
 Contents Ø background Ø micro. RNA identification Ø isomi. R Ø micro. RNA and disease Ø outlook 25
Contents Ø background Ø micro. RNA identification Ø isomi. R Ø micro. RNA and disease Ø outlook 25
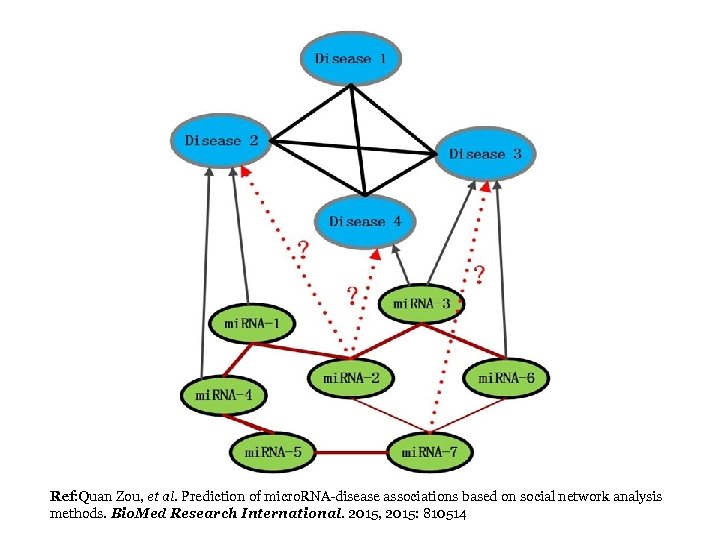 Ref: Quan Zou, et al. Prediction of micro. RNA-disease associations based on social network analysis methods. Bio. Med Research International. 2015, 2015: 810514
Ref: Quan Zou, et al. Prediction of micro. RNA-disease associations based on social network analysis methods. Bio. Med Research International. 2015, 2015: 810514
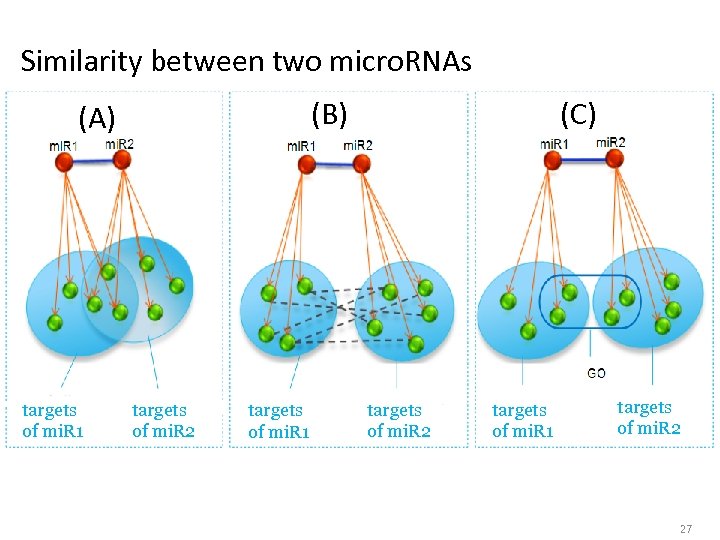 Similarity between two micro. RNAs (B) (A) targets of mi. R 1 targets of mi. R 2 targets of mi. R 1 (C) targets of mi. R 2 targets of mi. R 1 targets of mi. R 2 27
Similarity between two micro. RNAs (B) (A) targets of mi. R 1 targets of mi. R 2 targets of mi. R 1 (C) targets of mi. R 2 targets of mi. R 1 targets of mi. R 2 27
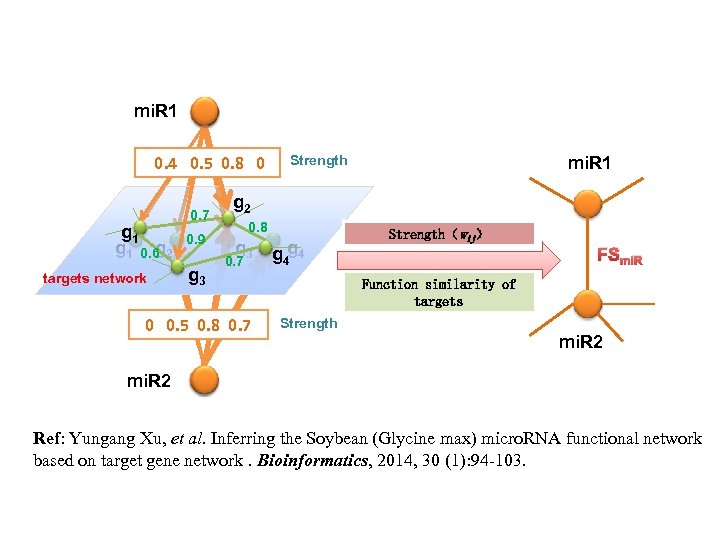 mi. R 1 0. 4 0. 5 0. 8 0 g 1 0. 6 2 g targets network 0. 7 0. 9 g 3 Strength mi. R 1 g 2 0. 8 g 3 0. 7 0 0. 5 0. 8 0. 7 g 4 g 4 Strength(wij) FSmi. R Function similarity of targets Strength mi. R 2 Ref: Yungang Xu, et al. Inferring the Soybean (Glycine max) micro. RNA functional network based on target gene network. Bioinformatics, 2014, 30 (1): 94 -103.
mi. R 1 0. 4 0. 5 0. 8 0 g 1 0. 6 2 g targets network 0. 7 0. 9 g 3 Strength mi. R 1 g 2 0. 8 g 3 0. 7 0 0. 5 0. 8 0. 7 g 4 g 4 Strength(wij) FSmi. R Function similarity of targets Strength mi. R 2 Ref: Yungang Xu, et al. Inferring the Soybean (Glycine max) micro. RNA functional network based on target gene network. Bioinformatics, 2014, 30 (1): 94 -103.
 Outlook How many novel micro. RNAs are still left? All the micro. RNA research methods can be extended to nc. RNA and lnc. RNA isomi. R would be the next hot topic in micro. RNA research Diseases would be the hot spots for ever!
Outlook How many novel micro. RNAs are still left? All the micro. RNA research methods can be extended to nc. RNA and lnc. RNA isomi. R would be the next hot topic in micro. RNA research Diseases would be the hot spots for ever!
 Quan Zou, Ph. D&Professor School of Computer Science and Technology Tianjin University Email: zouquan@nclab. net http: //lab. malab. cn/~zq/ 31
Quan Zou, Ph. D&Professor School of Computer Science and Technology Tianjin University Email: zouquan@nclab. net http: //lab. malab. cn/~zq/ 31


