7492bf1d2d8b1ce447300f4a92fe2b7a.ppt
- Количество слайдов: 19

Chassis George Church, Carlos Bustamante, Tom Knight Year Four Site Visit March 2, 2010

Chassis (&Bio. Fab) Goals Microbes, organelles, in vitro, plant, animal CAD-CAM: Agilent Chip Design / Assembly Opt. MAGE Genetic Design via Local Search (GDLS) Measurements: In situ sequencing Morphology & behavior Selection (evolution): Cell sorting Sensor set +/- Selector sets George Church, Carlos Bustamante, Tom Knight 2
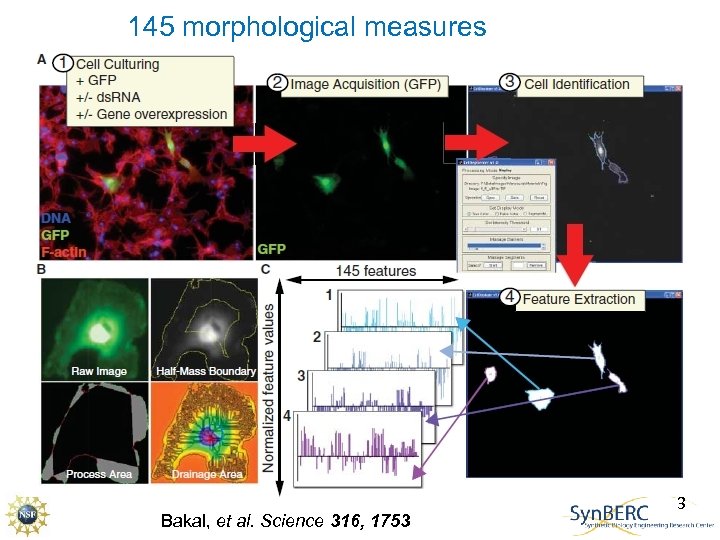
145 morphological measures Bakal, et al. Science 316, 1753 3
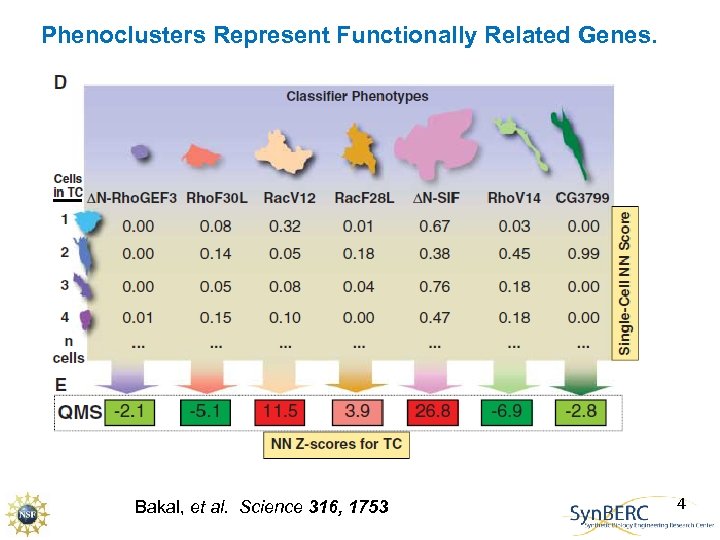
Phenoclusters Represent Functionally Related Genes. Bakal, et al. Science 316, 1753 4
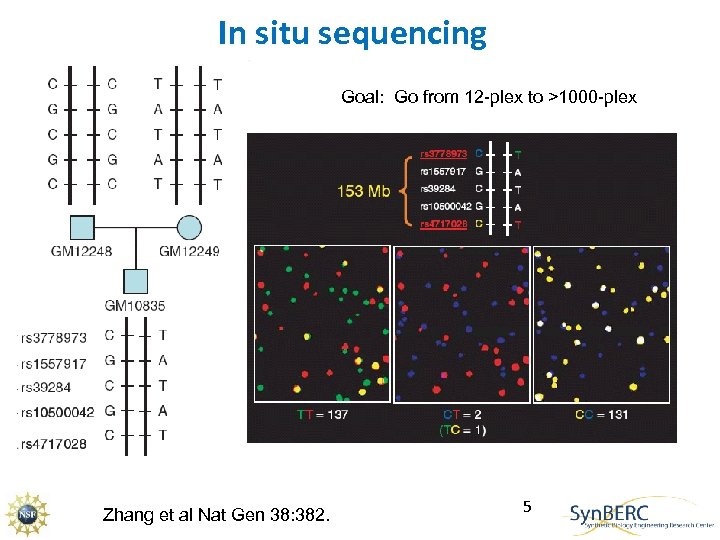
In situ sequencing Goal: Go from 12 -plex to >1000 -plex Zhang et al Nat Gen 38: 382. 5
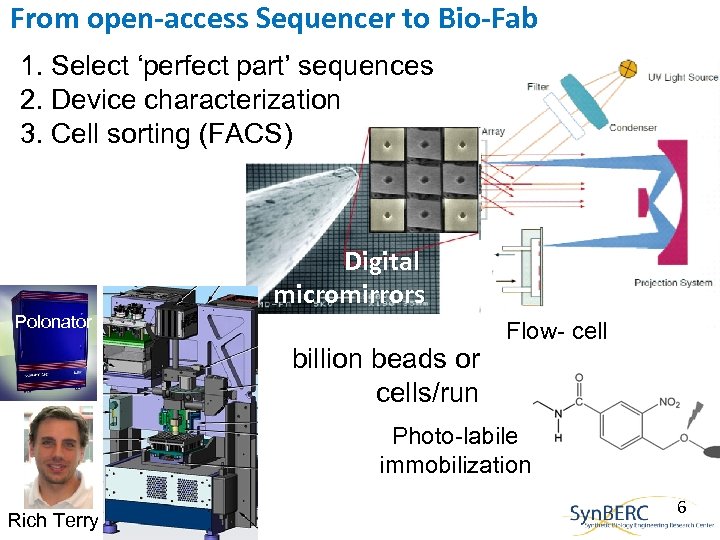
From open-access Sequencer to Bio-Fab 1. Select ‘perfect part’ sequences 2. Device characterization 3. Cell sorting (FACS) Digital micromirrors Polonator billion beads or cells/run Flow- cell Photo-labile immobilization Rich Terry 6
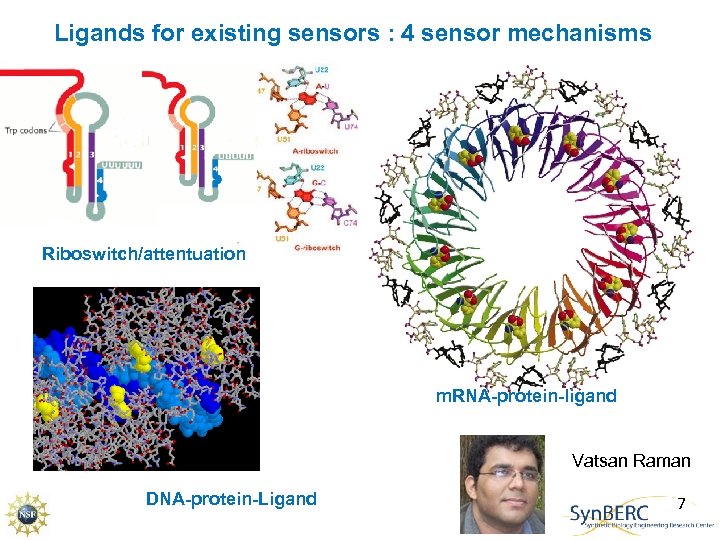
Ligands for existing sensors : 4 sensor mechanisms Riboswitch/attentuation m. RNA-protein-ligand Vatsan Raman DNA-protein-Ligand 7

Support Selections for new ligands for existing sensors 54 DNA binding proteins: ada ara. C arc. A arg. PR car. P cpx. R crp csp. A cyn. R cys. B cyt. R deo. R dna. A dgs. A fad. R far. R fhl. A flh. CD fnr fru. R fur gal. R gcv. A glp. R hip. B icl. R ilv. Y lac. I lex. A lrp mal. T mar. R mel. R met. J met. R mod. E nag. C nar. L nar. P ntr. C omp. R oxy. R pdh. R pho. B pur. R rha. S rpo. E rpo. H rpo. N rpo. S sox. S tor. R trp. R tyr. R 12 Riboswitches: Adenine B 12 FMN Guanine Glucosamine-6 -phosphate Glycine di-GMP Lysine Molybdenum Pre. Q 1 SAM SAH TPP theophylline 3 methylxanthine 8 http: //pubs. acs. org/doi/abs/10. 1021/ja 048634 j
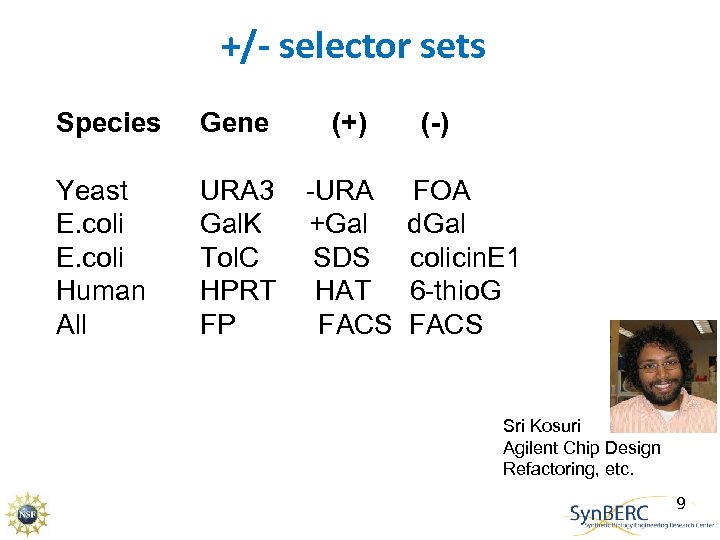

+/- selector sets Species Gene (+) Yeast E. coli Human All URA 3 Gal. K Tol. C HPRT FP -URA +Gal SDS HAT FACS (-) FOA d. Gal colicin. E 1 6 -thio. G FACS Sri Kosuri Agilent Chip Design Refactoring, etc. 9
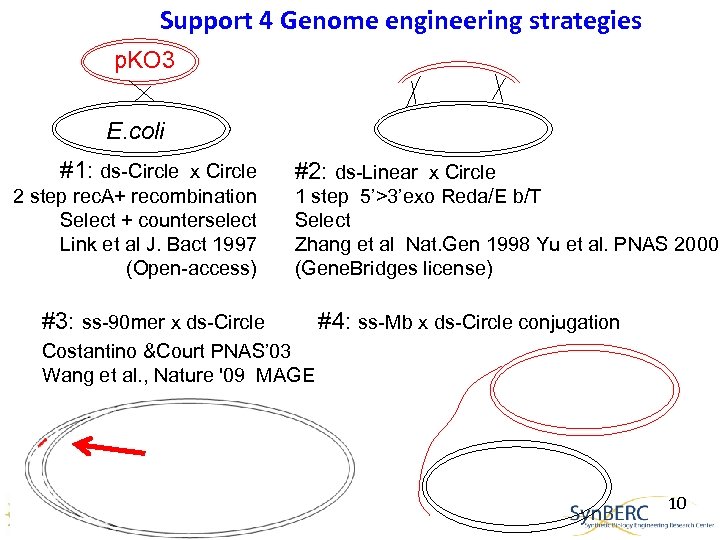
Support 4 Genome engineering strategies p. KO 3 E. coli #1: ds-Circle x Circle 2 step rec. A+ recombination Select + counterselect Link et al J. Bact 1997 (Open-access) #2: ds-Linear x Circle 1 step 5’>3’exo Reda/E b/T Select Zhang et al Nat. Gen 1998 Yu et al. PNAS 2000 (Gene. Bridges license) #3: ss-90 mer x ds-Circle #4: ss-Mb x ds-Circle conjugation Costantino &Court PNAS’ 03 Wang et al. , Nature '09 MAGE 10
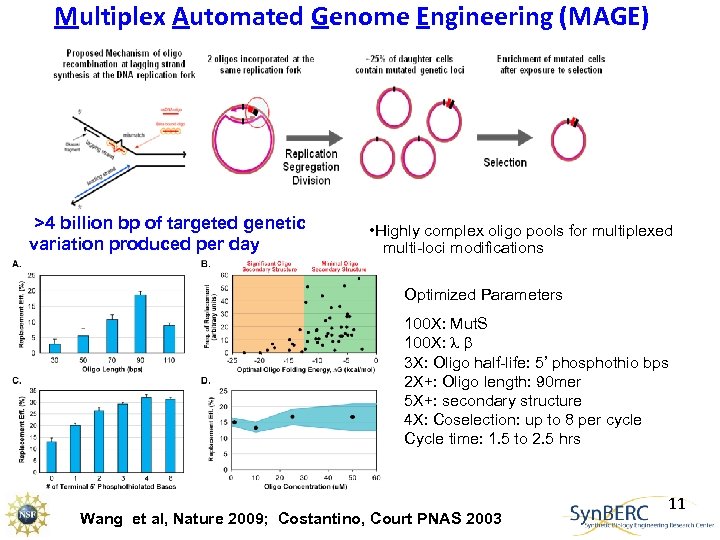
Multiplex Automated Genome Engineering (MAGE) >4 billion bp of targeted genetic variation produced per day • Highly complex oligo pools for multiplexed multi-loci modifications Optimized Parameters 100 X: Mut. S 100 X: b 3 X: Oligo half-life: 5’ phosphothio bps 2 X+: Oligo length: 90 mer 5 X+: secondary structure 4 X: Coselection: up to 8 per cycle Cycle time: 1. 5 to 2. 5 hrs Wang et al, Nature 2009; Costantino, Court PNAS 2003 11

MAGE Instrumentation Sep 2009 12
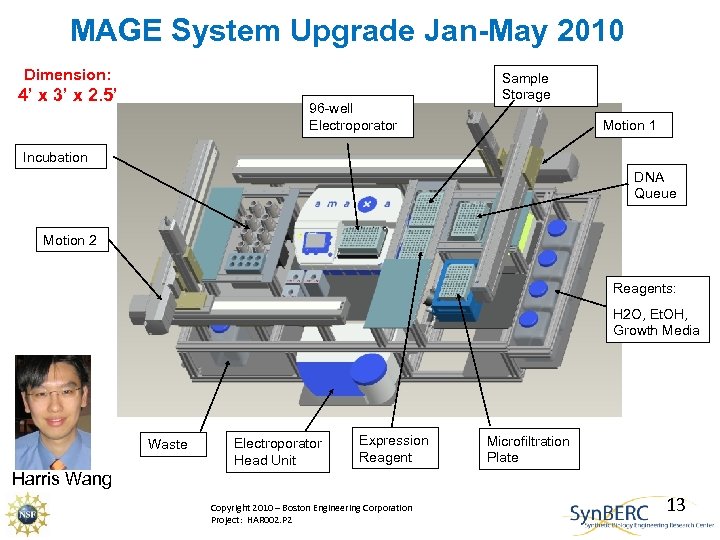
MAGE System Upgrade Jan-May 2010 Dimension: 4’ x 3’ x 2. 5’ 96 -well Electroporator Sample Storage Motion 1 Incubation DNA Queue Motion 2 Reagents: H 2 O, Et. OH, Growth Media Waste Electroporator Head Unit Expression Reagent Microfiltration Plate Harris Wang Copyright 2010 – Boston Engineering Corporation Project: HAR 002. P 2 13
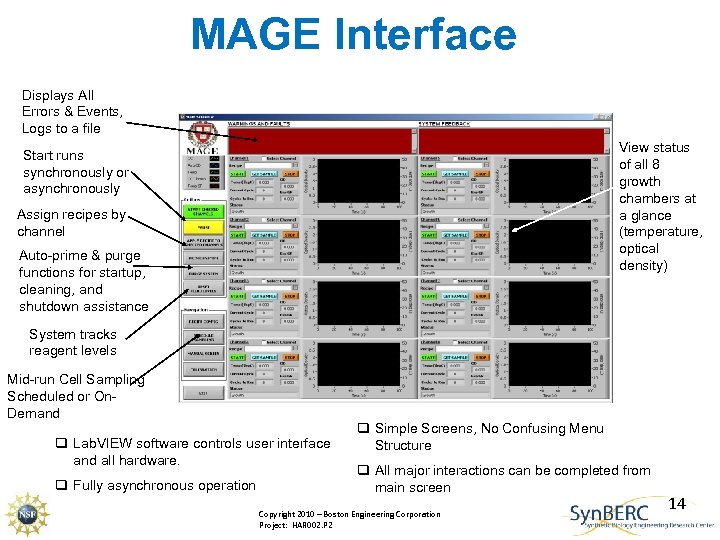
MAGE Interface Displays All Errors & Events, Logs to a file View status of all 8 growth chambers at a glance (temperature, optical density) Start runs synchronously or asynchronously Assign recipes by channel Auto-prime & purge functions for startup, cleaning, and shutdown assistance System tracks reagent levels Mid-run Cell Sampling Scheduled or On. Demand q Lab. VIEW software controls user interface and all hardware. q Fully asynchronous operation q Simple Screens, No Confusing Menu Structure q All major interactions can be completed from main screen Copyright 2010 – Boston Engineering Corporation Project: HAR 002. P 2 14
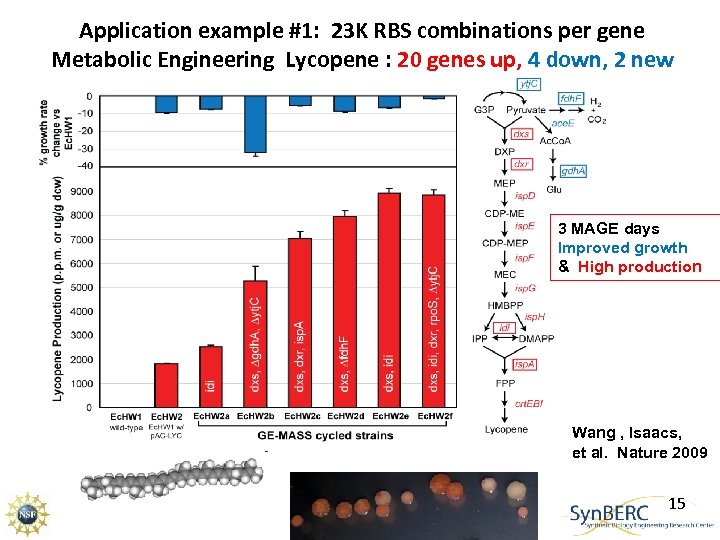
Application example #1: 23 K RBS combinations per gene Metabolic Engineering Lycopene : 20 genes up, 4 down, 2 new 3 MAGE days Improved growth & High production Wang , Isaacs, et al. Nature 2009 15
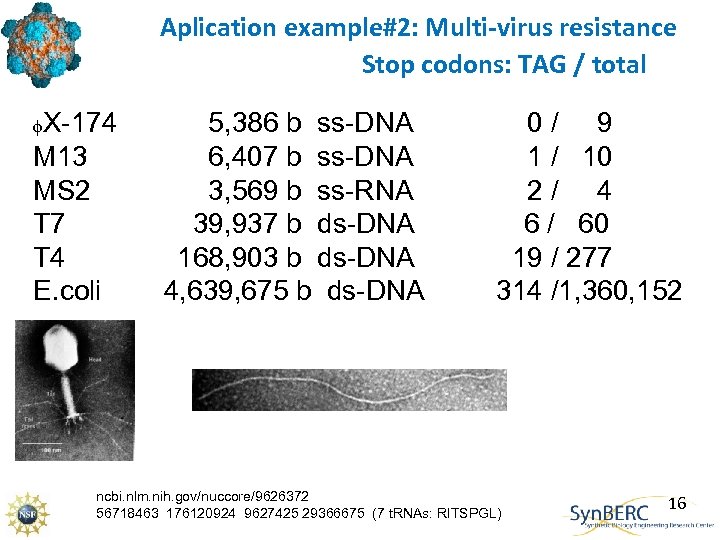

Aplication example#2: Multi-virus resistance Stop codons: TAG / total f. X-174 M 13 MS 2 T 7 T 4 E. coli 5, 386 b ss-DNA 6, 407 b ss-DNA 3, 569 b ss-RNA 39, 937 b ds-DNA 168, 903 b ds-DNA 4, 639, 675 b ds-DNA 0/ 9 1 / 10 2/ 4 6 / 60 19 / 277 314 /1, 360, 152 ncbi. nlm. nih. gov/nuccore/9626372 56718463 176120924 9627425 29366675 (7 t. RNAs: RITSPGL) 16
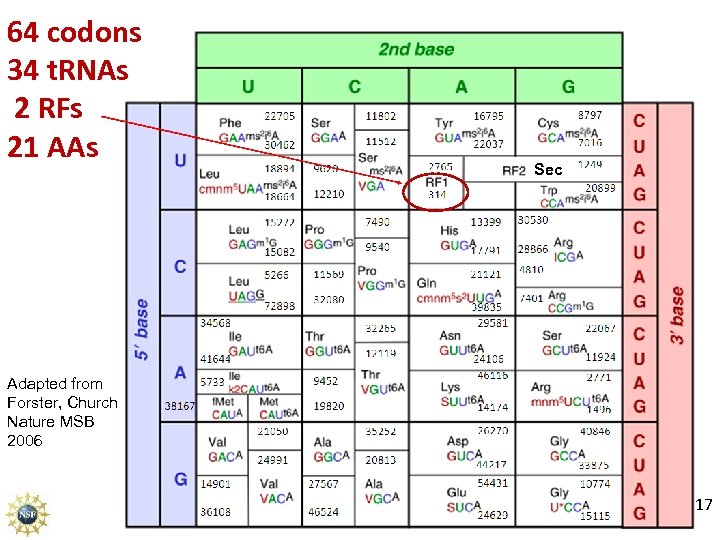
64 codons 34 t. RNAs 2 RFs 21 AAs Sec Adapted from Forster, Church Nature MSB 2006 17
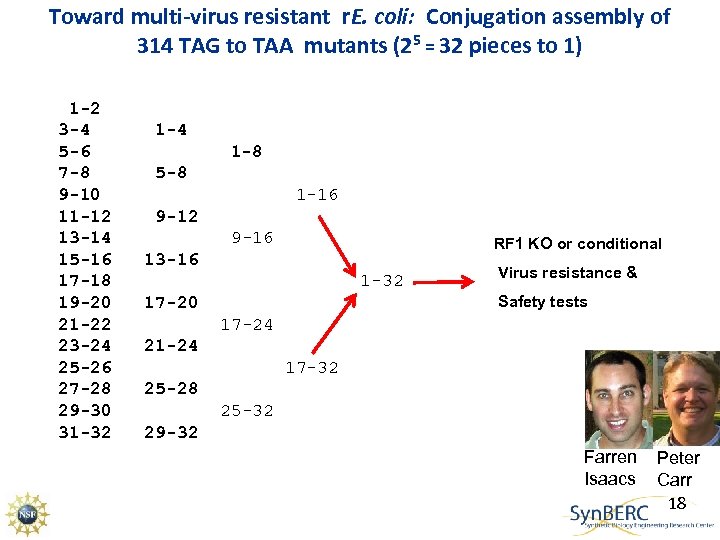
Toward multi-virus resistant r. E. coli: Conjugation assembly of 314 TAG to TAA mutants (25 = 32 pieces to 1) 1 -2 3 -4 5 -6 7 -8 9 -10 11 -12 13 -14 15 -16 17 -18 19 -20 21 -22 23 -24 25 -26 27 -28 29 -30 31 -32 1 -4 1 -8 5 -8 1 -16 9 -12 9 -16 RF 1 KO or conditional 13 -16 1 -32 Virus resistance & Safety tests 17 -20 17 -24 21 -24 17 -32 25 -28 25 -32 29 -32 Farren Isaacs Peter Carr 18

Chassis (&Bio. Fab) Goals Microbes, organelles, in vitro, plant, animal CAD-CAM: Agilent Chip Design / Assembly Opt. MAGE Genetic Design via Local Search (GDLS) Measurements: In situ sequencing Morphology & behavior Selection (evolution): Cell sorting Sensor set +/- Selector sets George Church, Carlos Bustamante, Tom Knight 19
7492bf1d2d8b1ce447300f4a92fe2b7a.ppt