d9f177bef2805684bb5a264d04fefad0.ppt
- Количество слайдов: 35

BL 5203: Molecular Recognition & Interaction Lecture 5: Drug Design Methods Ligand-Protein Docking (part II) Prof. Chen Yu Zong Tel: 6874 -6877 Email: csccyz@nus. edu. sg http: //xin. cz 3. nus. edu. sg Room 07 -24, level 7, SOC 1, National University of Singapore

Conformational Ensembles Docking

Conformational Ensembles Docking Observations: 1. Generating an orientation of a ligand in a binding site may be separated from calculating a conformation of the ligand in that particular orientation. 2. Multiple conformations of a given ligand usually have some portion in common (internally rigid atoms such as ring systems), and therefore, contain redundancies. 3
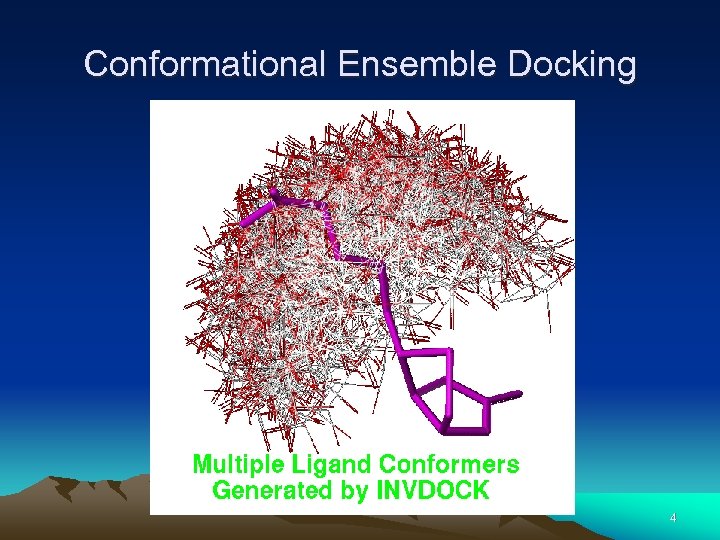
Conformational Ensemble Docking 4

Conformational Ensemble Docking • • Conformational ensembles are generated by overlaying all conformations of a given molecule onto its largest rigid fragment. Only atoms within this largest rigid fragment are used during the distance matching step. The RT matrix is defined. Each of the conformers is oriented into the site and scored. The score measures steric and electrostatic complementarity. One matching steps - all the conformers are docked and scored in the selected orientation. 5
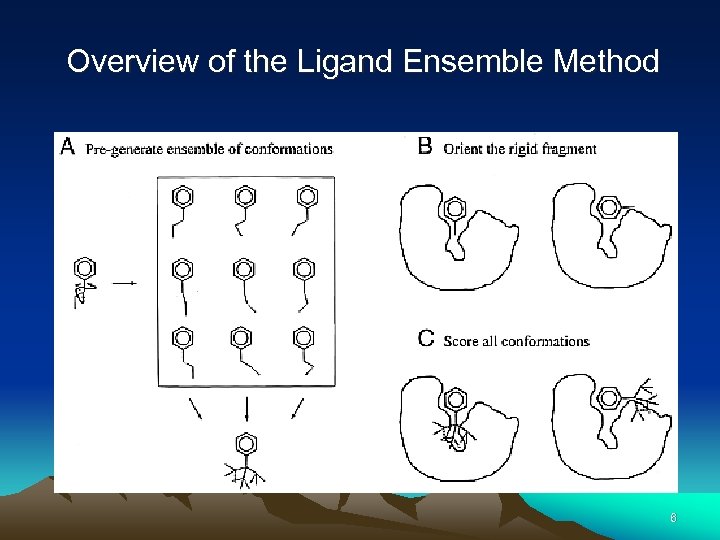
Overview of the Ligand Ensemble Method 6

Advantages of Conformational Ensemble Docking Speed increase due to: • One matching step for all of the conformers. • The largest rigid fragment usually has fewer atoms (less potential matches are examined). 7

Disadvantages of Conformational Ensemble Docking • Loss of information when the orientations are guided only by a subset of the atoms in molecule. Orientations may be missed because potential distance matches from nonrigid portions of the molecule are not considered. • The ensemble method will fail for ligands that lack internally rigid atoms. • The use of chemical matching and critical clusters is limited. 8
Pharmacophore-Based Docking 9

Pharmacophore-based Docking Basic idea: • Appropriate spatial disposition of a small number of functional groups in a molecule is sufficient for achieving a desired biological effect. • The ensemble formation will be guided by these functional groups. 10
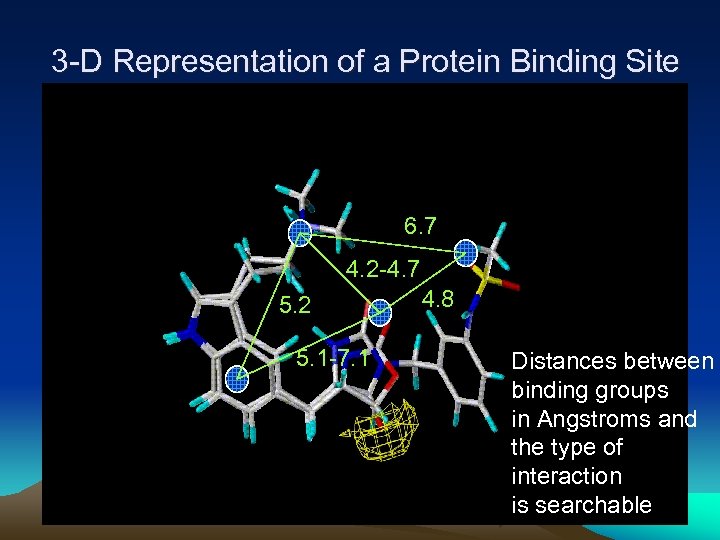
3 -D Representation of a Protein Binding Site 6. 7 4. 2 -4. 7 5. 2 5. 1 -7. 1 4. 8 Distances between binding groups in Angstroms and the type of interaction is searchable 11

Pharmacophore Fingerprint • Pharmacophore fingerprint - a set of pharmacophore features and their relative position. • Typical pharmacophore features: – Hydrogen-bond donors and acceptors – Positive and negative ionizable atoms/groups – Hydrophobes and ring centroids • Implemented in DOCK 4. 0. 1 – Hydrogen-bond donors – Hydrogen-bond acceptors – Dual hydrogen-bond donor and acceptor – 5 or 6 membered ring centroids 12

Notes on Pharmacophore Fingerprint • Each conformer has pharmacophore fingerprint. • Different conformers of the same molecule can have identical pharmacophore fingerprints. 13
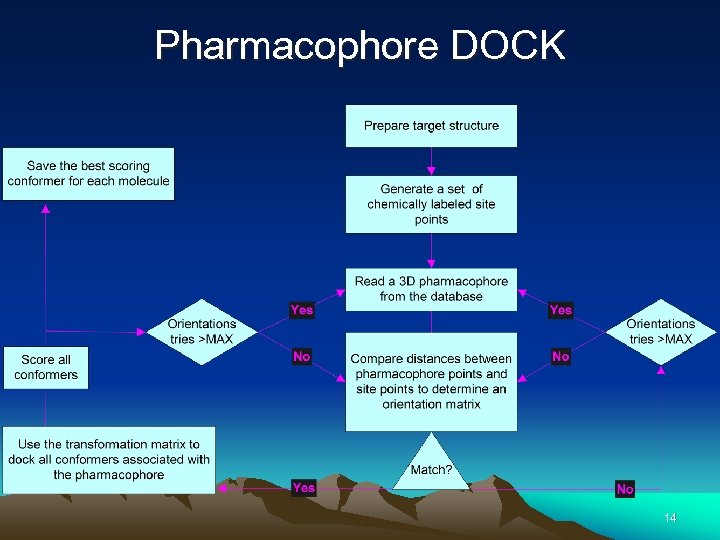
Pharmacophore DOCK 14

Advantages of Pharmacophore-based Docking • Rapid elimination of ligands containing functional groups which would interfere with binding. • Speed increase over docking of individual molecules. • More information pertaining to the entire molecule is retained (no rigid portions). • Chemical matching and critical clusters are encouraged. 15

Speed Comparison Between Ensemble and Pharmacophore-based Docking. Pharmacophore-based advantage: • Reduced number of ligand points considered during distance matching. Ensemble docking advantage: • The average number of conformers per molecule is higher than the average number of conformers per fingerprint. The one step matching speed reduction is slightly higher. 16

Speed Reduction Cont. • Ensemble docking: the average number of conformers per molecule is 297. • Pharmacophore-based: 50 -100 conformers per pharmacophore 17

Database Preparation • Generating molecular conformations – • Systematic search method with SYBYL. Overlaying molecular conformers onto pharmacophores 1. Extract 3 D pharmacophore from the first molecule of a cluster 2. Use it to perform a rigid 3 D UNITY search of the rest of that cluster to find matches 3. Save the pharmacophore query with the associated molecules 4. Process until all molecules are associated with a pharmacophore 18

Site Points Generation • Chemically labeled site point are generated in an automated fashion using the script MCSS 2 SPTS. • The script runs a series of MCSS (Multiple Copy Simultaneous Searches) calculations. • MCSS – methodology for finding energetically favorable positions and orientations of small functional group in a binding site. • Uses CHARMM potential energy function to determine the preferred locations or potential energy minima simultaneously for thousands of copies of a given chemical group. 19

Limitations of Pharmacophore-based Searching • A limited subset of key interactions (typically 4 -6) which must be extracted from the target site with dozens of potential interactions. • Complex queries are extremely slow. • The majority of the information contained in the target structure is not considered during the search. There is no scoring function beyond the binary (match/no match). Any steric or electronic constraints imposed by the target, but not defined by the target are ignored. 20
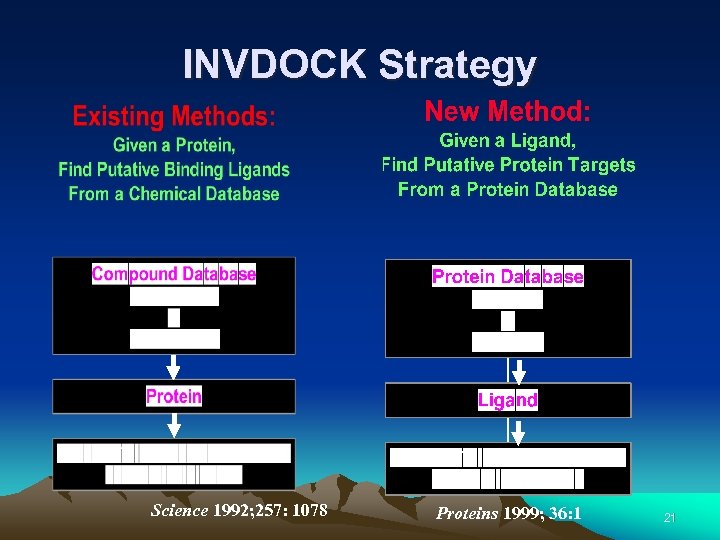

INVDOCK Strategy Science 1992; 257: 1078 Proteins 1999; 36: 1 21
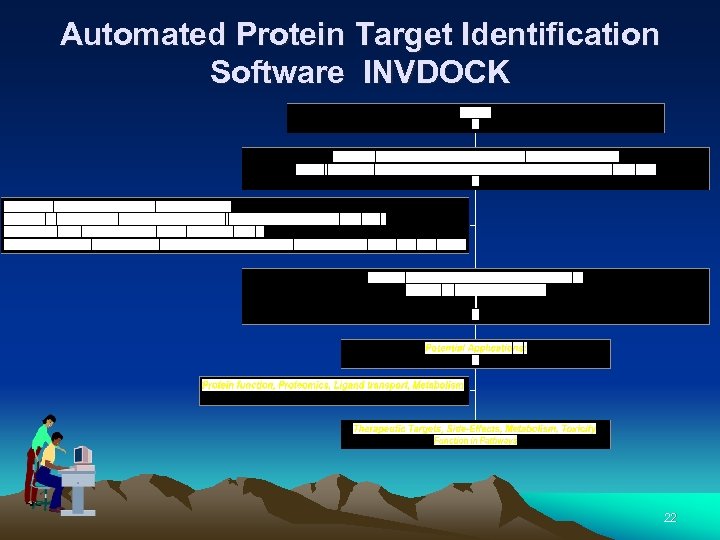
Automated Protein Target Identification Software INVDOCK 22
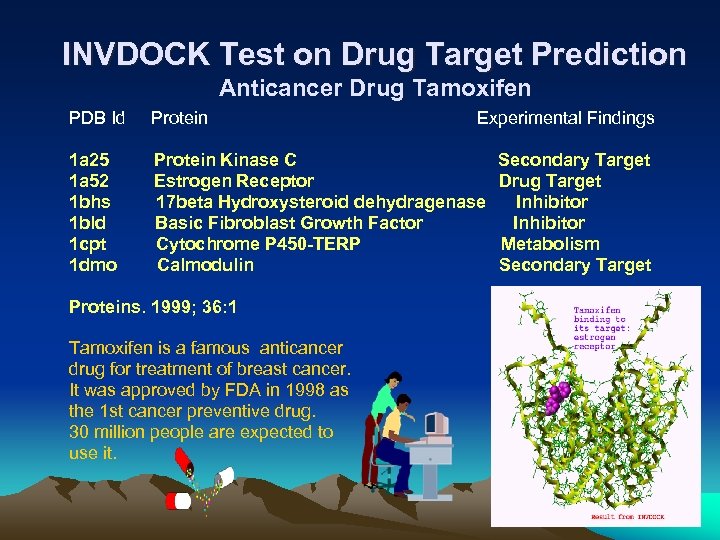

INVDOCK Test on Drug Target Prediction Anticancer Drug Tamoxifen PDB Id Protein Experimental Findings 1 a 25 1 a 52 1 bhs 1 bld 1 cpt 1 dmo Protein Kinase C Estrogen Receptor 17 beta Hydroxysteroid dehydragenase Basic Fibroblast Growth Factor Cytochrome P 450 -TERP Calmodulin Secondary Target Drug Target Inhibitor Metabolism Secondary Target Proteins. 1999; 36: 1 Tamoxifen is a famous anticancer drug for treatment of breast cancer. It was approved by FDA in 1998 as the 1 st cancer preventive drug. 30 million people are expected to use it. 23
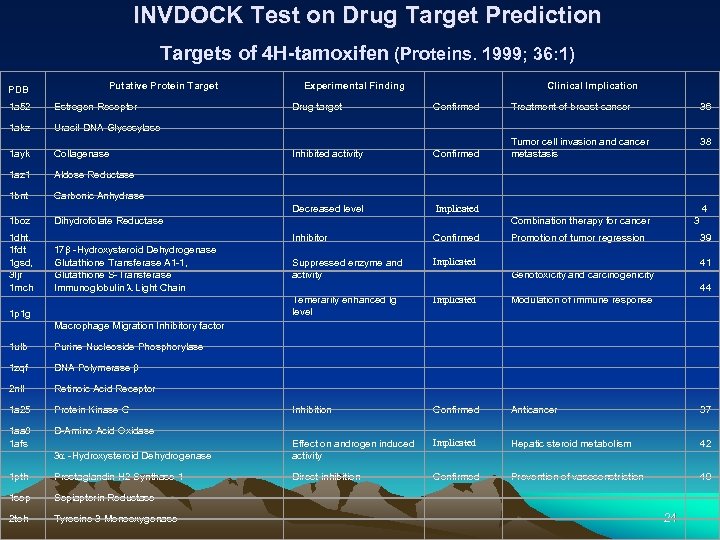
INVDOCK Test on Drug Target Prediction Targets of 4 H-tamoxifen (Proteins. 1999; 36: 1) PDB Putative Protein Target Clinical Implication Experimental Finding 1 a 52 Estrogen Receptor Drug target Confirmed Treatment of breast cancer 1 akz Uracil-DNA Glycosylase 1 ayk Collagenase Inhibited activity Confirmed Tumor cell invasion and cancer metastasis 1 az 1 Aldose Reductase 1 bnt Carbonic Anhydrase Decreased level Implicated 1 boz Dihydrofolate Reductase Combination therapy for cancer 3 1 dht, 1 fdt 1 gsd, 3 ljr 1 mch 1 p 1 g 17 b -Hydroxysteroid Dehydrogenase Glutathione Transferase A 1 -1, Glutathione S-Transferase Immunoglobulin l Light Chain Inhibitor Suppressed enzyme and activity 36 38 4 Confirmed Implicated Promotion of tumor regression 39 41 44 Genotoxicity and carcinogenicity Macrophage Migration Inhibitory factor Temerarily enhanced Ig level Modulation of immune response 1 ulb Purine Nucleoside Phosphorylase 1 zqf DNA Polymerase b 2 nll Retinoic Acid Receptor 1 a 25 Protein Kinase C Inhibition Confirmed Anticancer 1 aa 8 1 afs D-Amino Acid Oxidase 3 a -Hydroxysteroid Dehydrogenase Effect on androgen induced activity Implicated Hepatic steroid metabolism 1 pth Prostaglandin H 2 Synthase-1 Direct inhibition Confirmed Prevention of vasoconstriction 1 sep Sepiapterin Reductase 2 toh Tyrosine 3 -Monooxygenase 37 42 40 24
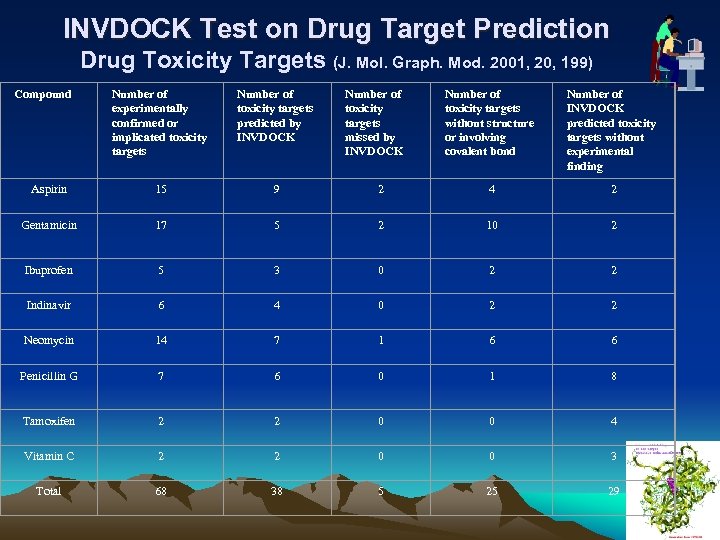
INVDOCK Test on Drug Target Prediction Drug Toxicity Targets (J. Mol. Graph. Mod. 2001, 20, 199) Compound Number of experimentally confirmed or implicated toxicity targets Number of toxicity targets predicted by INVDOCK Aspirin 15 9 Gentamicin 17 Ibuprofen Number of toxicity targets missed by INVDOCK Number of toxicity targets without structure or involving covalent bond Number of INVDOCK predicted toxicity targets without experimental finding 2 4 2 5 2 10 2 5 3 0 2 2 Indinavir 6 4 0 2 2 Neomycin 14 7 1 6 6 Penicillin G 7 6 0 1 8 Tamoxifen 2 2 0 0 4 Vitamin C 2 2 0 0 3 Total 68 38 5 25 29 25
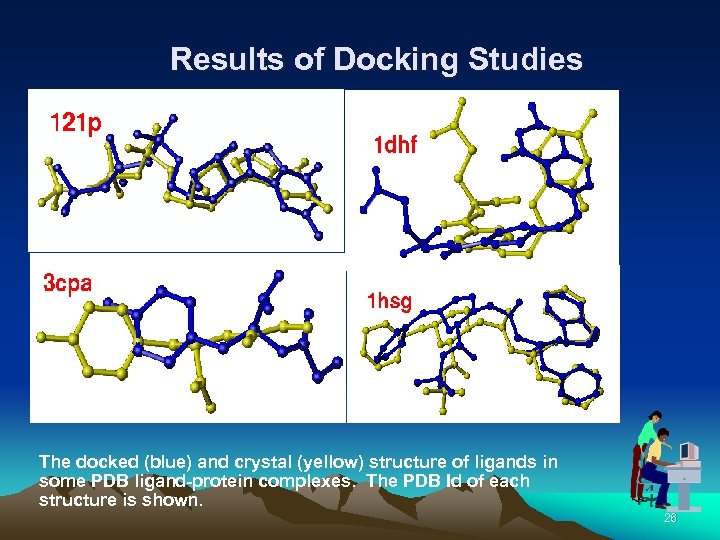
Results of Docking Studies The docked (blue) and crystal (yellow) structure of ligands in some PDB ligand-protein complexes. The PDB Id of each structure is shown. 26
![Dataset and Testing Results Protein-Protein cases from protein-protein docking benchmark [6]: Enzyme-inhibitor – 22 Dataset and Testing Results Protein-Protein cases from protein-protein docking benchmark [6]: Enzyme-inhibitor – 22](https://present5.com/presentation/d9f177bef2805684bb5a264d04fefad0/image-27.jpg)
Dataset and Testing Results Protein-Protein cases from protein-protein docking benchmark [6]: Enzyme-inhibitor – 22 cases Antibody-antigen – 16 cases Protein-DNA docking: 2 unbound-bound cases Protein-drug docking: tens of bound cases (Estrogen receptor, HIV protease, COX) Performance: Several minutes for large protein molecules and seconds for small drug molecules on standard PC computer. Estrogen receptor Estradiol molecule from complex docking solution DNA endonuclease docking solution Endonuclease I-Ppo. I (1 EVX) with DNA (1 A 73). RMSD 0. 87Å, rank 2 Estrogen receptor with estradiol (1 A 52). RMSD 0. 9Å, rank 1, running time: 11 27 seconds
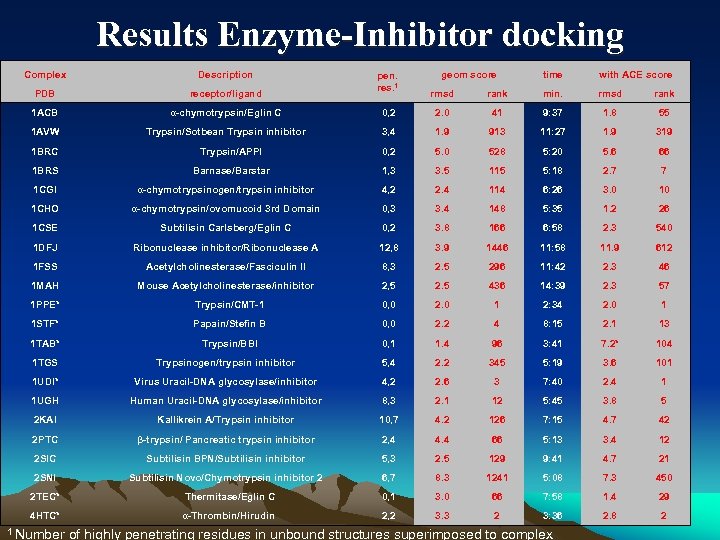
Results Enzyme-Inhibitor docking Complex Description PDB receptor/ligand 1 ACB α-chymotrypsin/Eglin C 1 AVW Trypsin/Sotbean Trypsin inhibitor pen. res. 1 geom score time with ACE score rmsd rank min. rmsd rank 0, 2 2. 0 41 9: 37 1. 8 55 3, 4 1. 9 913 11: 27 1. 9 319 1 BRC Trypsin/APPI 0, 2 5. 0 528 5: 20 5. 6 66 1 BRS Barnase/Barstar 1, 3 3. 5 115 5: 18 2. 7 7 1 CGI α-chymotrypsinogen/trypsin inhibitor 4, 2 2. 4 114 6: 26 3. 0 10 1 CHO α-chymotrypsin/ovomucoid 3 rd Domain 0, 3 3. 4 148 5: 35 1. 2 26 1 CSE Subtilisin Carlsberg/Eglin C 0, 2 3. 8 166 6: 58 2. 3 540 1 DFJ Ribonuclease inhibitor/Ribonuclease A 12, 8 3. 9 1446 11: 58 11. 9 612 1 FSS Acetylcholinesterase/Fasciculin II 8, 3 2. 5 296 11: 42 2. 3 46 1 MAH Mouse Acetylcholinesterase/inhibitor 2, 5 2. 5 436 14: 39 2. 3 57 1 PPE* Trypsin/CMT-1 0, 0 2. 0 1 2: 34 2. 0 1 1 STF* Papain/Stefin B 0, 0 2. 2 4 8: 15 2. 1 13 1 TAB* Trypsin/BBI 0, 1 1. 4 96 3: 41 7. 2* 104 1 TGS Trypsinogen/trypsin inhibitor 5, 4 2. 2 345 5: 19 3. 6 101 1 UDI* Virus Uracil-DNA glycosylase/inhibitor 4, 2 2. 6 3 7: 40 2. 4 1 1 UGH Human Uracil-DNA glycosylase/inhibitor 8, 3 2. 1 12 5: 45 3. 8 5 2 KAI Kallikrein A/Trypsin inhibitor 10, 7 4. 2 126 7: 15 4. 7 42 2 PTC β-trypsin/ Pancreatic trypsin inhibitor 2, 4 4. 4 66 5: 13 3. 4 12 2 SIC Subtilisin BPN/Subtilisin inhibitor 5, 3 2. 5 129 9: 41 4. 7 21 2 SNI Subtilisin Novo/Chymotrypsin inhibitor 2 6, 7 8. 3 1241 5: 08 7. 3 450 2 TEC* Thermitase/Eglin C 0, 1 3. 0 66 7: 58 1. 4 29 4 HTC* α-Thrombin/Hirudin 2, 2 3. 3 2 3: 36 2. 8 2 28 1 Number of highly penetrating residues in unbound structures superimposed to complex

Results Antibody-Antigen docking Complex Description PDB receptor/ligand 1 AHW Antibody Fab 5 G 9/Tissue factor 1 BQL* pen. res. 1 geom score time ACE score rmsd rank min. rmsd rank 3, 3 2. 5 29 10: 12 2. 5 10 Hyhel - 5 Fab/Lysozyme 0, 0 2. 5 13 6: 21 1. 4 7 1 BVK Antibody Hulys 11 Fv/Lysozyme 0, 0 3. 8 1301 6: 25 3. 5 809 1 DQJ Hyhel - 63 Fab/Lysozyme 18, 7 4. 3 773 5: 30 5. 1 953 1 EO 8* Bh 151 Fab/Hemagglutinin 3, 1 1. 8 567 9: 45 1. 6 292 1 FBI* Ig. G 1 Fab fragment/Lysozyme 2, 5 5. 0 536 10: 13 5. 0 2416 1 IAI* Ig. G 1 Idiotypic Fab/Igg 2 A Anti-Idiotypic Fab 5, 6 4. 8 1302 9: 13 3. 4 1304 1 JHL* Ig. G 1 Fv Fragment/Lysozyme 0, 0 1. 6 282 13: 15 1. 3 143 1 MEL* Vh Single-Domain Antibody/Lysozyme 0, 1 1. 8 3 2: 40 2. 0 2 1 MLC Ig. G 1 D 44. 1 Fab fragment/Lysozyme 8, 3 4. 0 136 5: 29 2. 6 123 1 NCA* Fab NC 41/Neuraminidase 0, 0 2. 6 114 17: 50 2. 8 66 1 NMB* Fab NC 10/Neuraminidase 0, 0 2. 7 2593 28: 10 2. 4 1734 1 QFU* Igg 1 -k Fab/Hemagglutinin 0, 0 2. 7 44 5: 42 2. 7 23 1 WEJ Ig. G 1 E 8 Fab fragment/Cytochrome C 0, 0 4. 3 232 7: 44 2. 6 87 2 JEL* Jel 42 Fab Fragment/A 06 Phosphotransferase 0, 2 4. 7 114 5: 02 4. 7 50 2 VIR* Igg 1 -lamda Fab/Hemagglutinin 0, 0 3. 1 258 7: 34 3. 5 306 1 Number of highly penetrating residues in unbound structures superimposed to complex 29
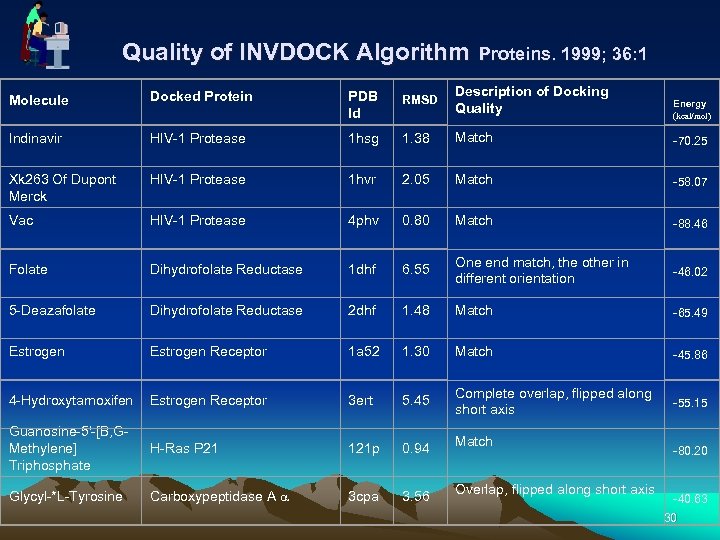
Quality of INVDOCK Algorithm Proteins. 1999; 36: 1 Molecule Docked Protein PDB Id RMSD Description of Docking Quality Energy (kcal/mol) Indinavir HIV-1 Protease 1 hsg 1. 38 Match -70. 25 Xk 263 Of Dupont Merck HIV-1 Protease 1 hvr 2. 05 Match -58. 07 Vac HIV-1 Protease 4 phv 0. 80 Match -88. 46 Folate Dihydrofolate Reductase 1 dhf 6. 55 One end match, the other in different orientation -46. 02 5 -Deazafolate Dihydrofolate Reductase 2 dhf 1. 48 Match -65. 49 Estrogen Receptor 1 a 52 1. 30 Match -45. 86 4 -Hydroxytamoxifen Estrogen Receptor 3 ert 5. 45 Complete overlap, flipped along short axis -55. 15 H-Ras P 21 121 p 0. 94 Match Carboxypeptidase A a 3 cpa 3. 56 Overlap, flipped along short axis Guanosine-5'-[B, GMethylene] Triphosphate Glycyl-*L-Tyrosine -80. 20 -40. 63 30
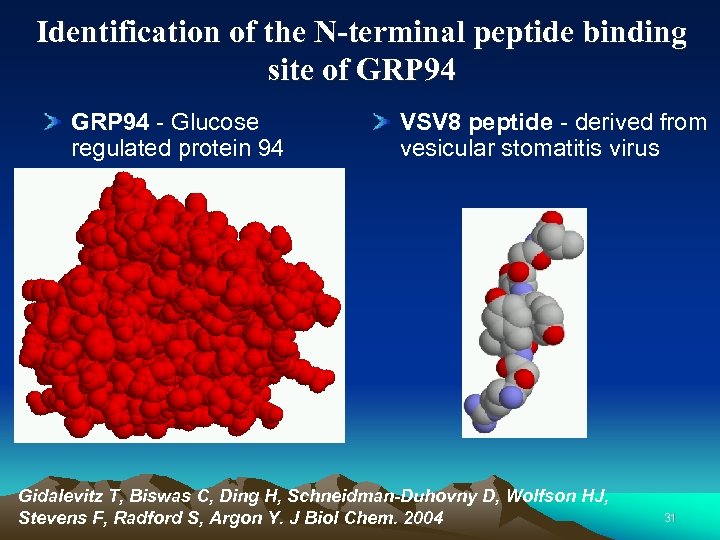

Identification of the N-terminal peptide binding site of GRP 94 - Glucose regulated protein 94 VSV 8 peptide - derived from vesicular stomatitis virus Gidalevitz T, Biswas C, Ding H, Schneidman-Duhovny D, Wolfson HJ, Stevens F, Radford S, Argon Y. J Biol Chem. 2004 31

Biological motivation The complex between the two molecules highly stimulates the response of the T-cells of the immune system. The grp 94 protein alone does not have this property. The activity that stimulates the immune response is due to the ability of grp 94 to bind different peptides. Characterization of peptide binding site is highly important. 32
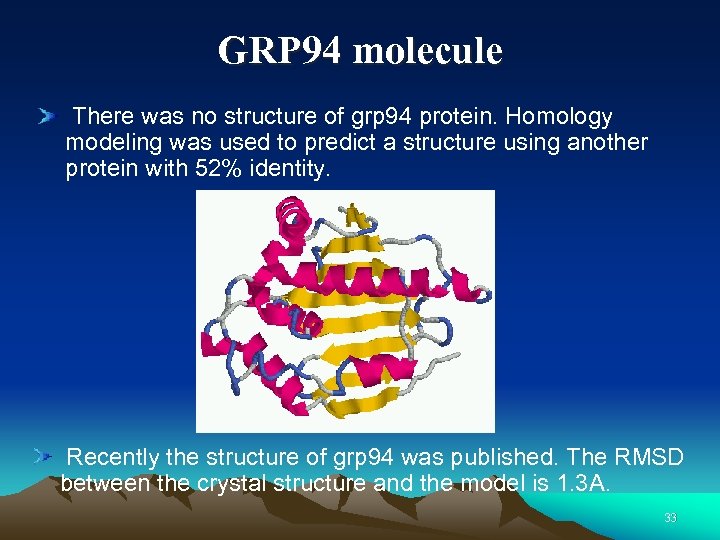

GRP 94 molecule There was no structure of grp 94 protein. Homology modeling was used to predict a structure using another protein with 52% identity. Recently the structure of grp 94 was published. The RMSD between the crystal structure and the model is 1. 3 A. 33
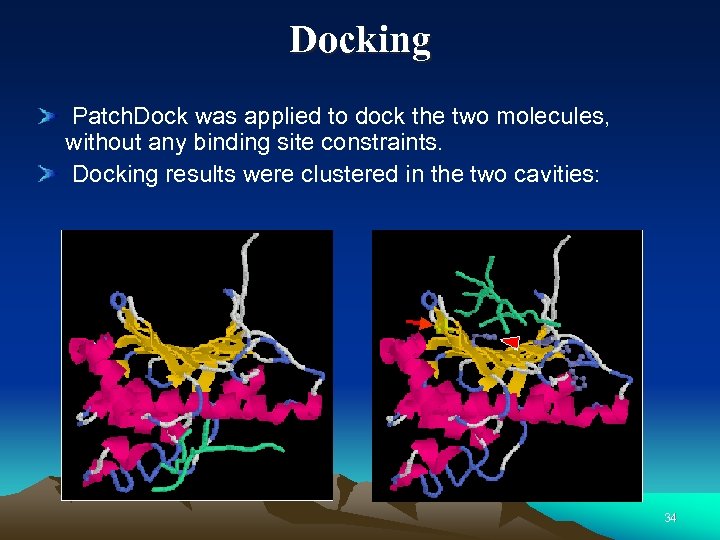
Docking Patch. Dock was applied to dock the two molecules, without any binding site constraints. Docking results were clustered in the two cavities: 34
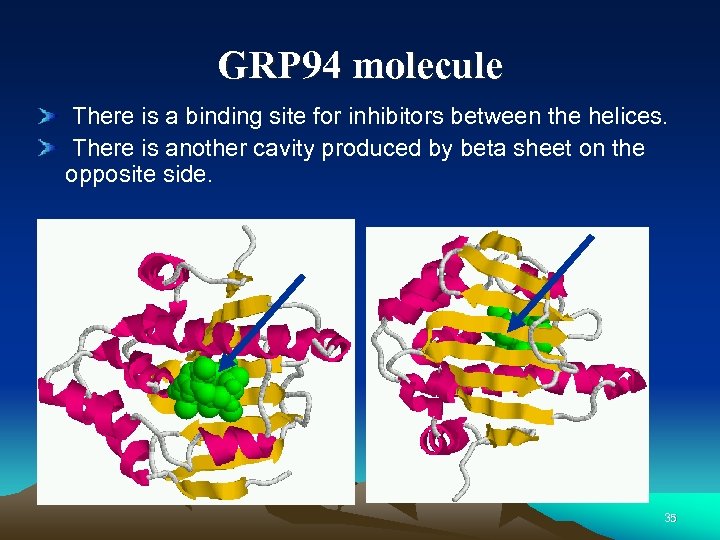
GRP 94 molecule There is a binding site for inhibitors between the helices. There is another cavity produced by beta sheet on the opposite side. 35
d9f177bef2805684bb5a264d04fefad0.ppt