609886fd9fb907b90d8bad8435b4a325.ppt
- Количество слайдов: 23

Biology 3. 3 Describe the role of DNA in relation to gene expression Dr Hayley Ridgway Ms Dalin Dore

Things to know 1. 2. 3. 4. 5. The role of DNA includes DNA structure and replication, the control of gene expression, protein synthesis, and the determination of phenotype. The structure of DNA includes the molecular components and their role in carrying the genetic code. The replication of DNA includes the processes involved in replication and the role that enzymes have in producing accurate copies. Control of gene expression is limited to factors that operate at transcription level: 1. feedback in prokaryotes (repressors, inducers) 2. enhancers and transcription factors in eukaryotes. Protein synthesis includes the role of DNA in determining the structure of a protein and how that protein is produced (transcription and translation). The determination of phenotype includes: – allele interactions: dominance, incomplete dominance, co-dominance, multiple alleles, lethal alleles – linkage and sex linkage – gene-gene interactions: epistasis, collaboration, polygenes – pleiotropy – mutations: gene mutations, chromosomal mutations – control of metabolic pathways by gene expression.

Previous questions 2005 - Munchkin cat Key concepts to know: • Dominance • Expected progeny from crosses • Selective breeding (genome) • Transgenics • Cloning (genome)

2006 - A 1 and A 2 Milk Key concepts to know: • Codominance • Alleles • Mutation (single nucleotide polymorphism) • Frequency and what affects it eg. – founder, linkage, selective breeding, selective advantage

2007 - Huntington’s disease Key concepts to know: • Mutation • Dominance • Autosomal - therefore how is it passed on? • Factors affecting disease expression – what is disease expression? – number of repeats – heterozygous vs homozygous, any difference? – female vs male • Needed to carefully read the material provided The neuron in the center (yellow) contains an abnormal accumulation of huntingtin (orange). Studies demonstrated that neurons with huntingtin survive longer than those that do not.

Key Concepts Dominance & Co-dominance • How can you tell the difference? Frequency • What is it • What could affect frequency? What is an allele? Mutation • How can it arise? • What are the consequences? Linkage Breeding systems • Selective • Cloning • Transgenics

“Superbugs” MRSA or Methicillin Resistant Staphyloccocus aureus What is MRSA? A common bacterium that can causes infections in different parts of the body. Usually no problem but it has become resistant to some commonly used antibiotics.

Staphylococcus aureus - a short history • Bacteria have very good adaptive capabilities • 1940 s: Penicillin was introduced - a strong selective pressure, induces mutation • Bacteria can transfer traits by mobile DNA such as heavy metal tolerance etc • Penicillin was virtually useless as an antibiotic within a decade because a plasmid spread the penicillinase (ßlactamase ) gene through the entire species of S. aureus • New antibiotics such as methicillin which were not degraded by the product of the ß-lactamase gene were used

• By 1960 methicillin resistant S. aureus (MRSA) strains were identified • By the 1980 s, epidemic clones of MRSA acquired multidrug resistant traits and spread worldwide to become one of the most important causes of hospital acquired infections • In the early 2000 s, MRSA strains carrying the additional Tn 1546 transposon-based vancomycin resistant mechanism were identified, bringing the possibility of a totally resistant bacterial pathogen closer to reality

How did this happen? • Selection pressure applied • S. aureus acquired and mutated a gene from another species of Staphylococcus (S. sciuri) – the penicillinase gene (via a plasmid) – then the methicillin resistance gene mec Mec: • Originated in S. scuiri another species of Staphylococcus • Located in a mobile piece of DNA that contains its own enzymes for moving it around the genome • This piece of DNA is called the Staphylococcal cassette chromosome mec (SCCmec) • Has about 100 ORF on this element – so also contains other genes

Mobile DNA • Common in bacteria • Two general types: 1. Plasmids – extrachromosomal circular or linear DNA molecules which are not part of the bacterial genome – carry functions advantageous to the host such as eg antibiotics or heavy metal resistance 2. Transposons – jumping genes, mobile genetic elements – composed of a gene coding for a special enzyme and two short flanking segments of DNA called inverted repeats
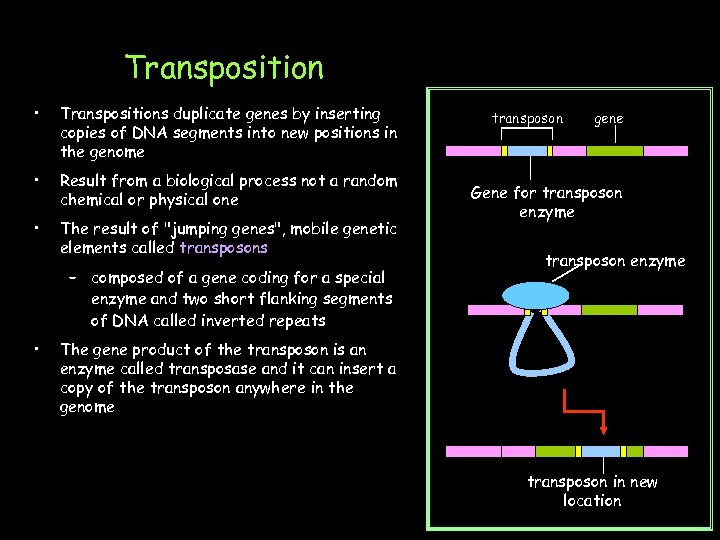
Transposition • Transpositions duplicate genes by inserting copies of DNA segments into new positions in the genome • Result from a biological process not a random chemical or physical one • The result of "jumping genes", mobile genetic elements called transposons – composed of a gene coding for a special enzyme and two short flanking segments of DNA called inverted repeats • transposon gene Gene for transposon enzyme The gene product of the transposon is an enzyme called transposase and it can insert a copy of the transposon anywhere in the genome transposon in new location

How do antibiotics work? Penicillin (β-lactam) • Inhibits formation of peptidoglycan cross-links (major part of bacterial cell wall) in the cell wall • Normally crosslinking is done by PBPs (penicillin binding proteins or transpeptidases ) • The β-lactam moiety (functional group) of penicillin binds to the enzyme (PBPs) • Weakens the cell wall of the bacterium and causes lysis Resistance to Penicillin • Bacteria evolved Penicillinases which hydrolysing penicillin

Methicillin (also β-lactam) • Acts the same way as Penicillin • BUT insensitive to penicillinases Resistance to Methicillin • Expresses a different PBP, called PBP 2 a, that is resistant to methicillin • Often resistant S. aureus are resistant to other classes of antibiotics (through different mechanisms) • Accessory factors (genes) influence the level and nature of methicillin resistance

Selection pressures • Selection pressure – killing bacterium • One mechanism of action – only need to find a away around one thing • Bacteria multiply rapidly • Over prescription of antibiotics, • People not finishing the course of antibiotics • And others……

Added info…. • Mutation • Genes

Evolution depends on accidents and mistakes! • In general cells do not have mechanisms for creating changes in their genomes – Normally replication, recombination and repair are high fidelity processes – 1/1000 bp randomly changed every 200, 000 years – So if n = 10, 000, every SNP tried out 50 x over 1 million years – relatively short time evolutionarily speaking • Most changes result from mistakes in normal copy and repair mechanisms • Transposable elements play a role • Can vary from SNPs to large scale rearrangements such as deletions, duplications, inversions and translocations

• Genome differences have accumulated over 3 billion years • Comparisons of genomes allow reconstruction of evolutionary process • Balanced process – genome stabilty and evolutionary change • Mutation/variation are rare because DNA is a very stable molecule for several reasons: 1. DNA is made of complementary strands and repairs can be made when one side is damaged 2. The nucleotides are protected inside the sugar-phosphate backbone and secured by hydrogen bonds. 3. DNA structure (nucleosomes etc) provide a tight structure which restricts access.

Point mutations • Once a nucleotide or nucleotides have been altered the genetic code will be subject to one of the following mutations. – Base substitution • Single base pair changes or deletions in DNA • Diallelic • Approx 2, 000 -3, 000 in every chromosome (1 evry 1000 bp) • silent mutation - the new base pair codes for the same amino acid • neutral mutation - the new base pair codes for a different amino acid but the shape of the resulting protein is unchanged • MUTATION - most are disastrous, but some can benefit the organism
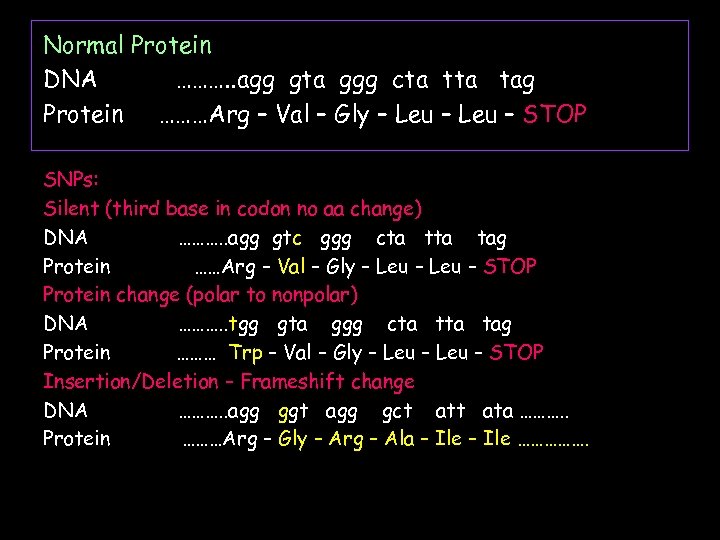
Normal Protein DNA ………. . agg gta ggg cta tag Protein ………Arg – Val – Gly – Leu – STOP SNPs: Silent (third base in codon no aa change) DNA ………. . agg gtc ggg cta tag Protein ……Arg – Val – Gly – Leu – STOP Protein change (polar to nonpolar) DNA ………. . tgg gta ggg cta tag Protein ……… Trp – Val – Gly – Leu – STOP Insertion/Deletion – Frameshift change DNA ………. . agg ggt agg gct ata ………. . Protein ………Arg – Gly – Arg – Ala – Ile …………….

Deleterious effects: – Chain termination - Produce a stop codon, premature chain termination = non-functional proteins – Additions and Deletions - When the number of nucleotides added or removed is not equal to three, causes a frameshift – Point Mutations may also occur in non-coding regions - introns or regulatory regions, severity of the mutation depends upon the region it occurs in, mutations in introns have little or no effect.
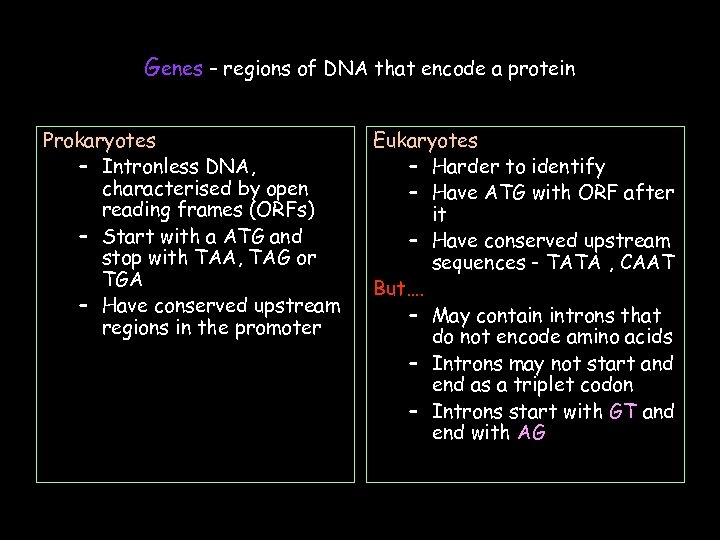

Genes – regions of DNA that encode a protein Prokaryotes – Intronless DNA, characterised by open reading frames (ORFs) – Start with a ATG and stop with TAA, TAG or TGA – Have conserved upstream regions in the promoter Eukaryotes – Harder to identify – Have ATG with ORF after it – Have conserved upstream sequences - TATA , CAAT But…. – May contain introns that do not encode amino acids – Introns may not start and end as a triplet codon – Introns start with GT and end with AG
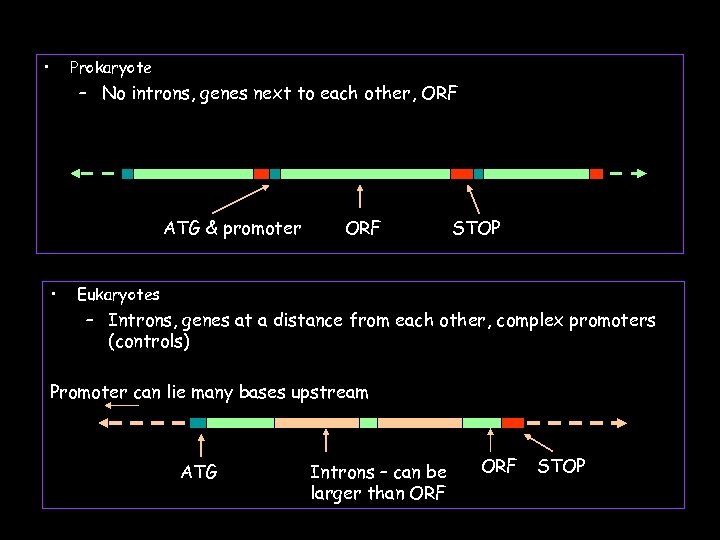
• Prokaryote – No introns, genes next to each other, ORF ATG & promoter • ORF STOP Eukaryotes – Introns, genes at a distance from each other, complex promoters (controls) Promoter can lie many bases upstream ATG Introns – can be larger than ORF STOP
609886fd9fb907b90d8bad8435b4a325.ppt