38923564bf719b9b073d05d6c5cf9aed.ppt
- Количество слайдов: 17

Biological Language Modeling Toolkit “Graphing Utilities” by: Danny Lam

Overview • BLMT Ex: Computes association measures in protein sequences • Graphing Utilities – Display how well the association measures or other data (known or surmised) feature boundaries • Step 1: Automatic extraction of feature boundaries from given source files • Step 2: Plot data along with feature positions along a sequence

BLMT : Mutual Information • Mutual Information -> Computes "mutual information”, which is a measure of association between adjacent amino acids. – Input: amino acid sequence file(s) • (ex) Swiss prot SW datasets – Output: file. mi. out. av -> • first column is position in sequence • second column is mutual information value associated with that position

Feature Positions • Extract feature position information (via Swiss -prot) – Extracellular (EC), – Cytoplasmic (CP), – Helices (H) --> label where the EC, CP, and H regions are in the sequence.

DR PROSITE; PS 00238; OPSIN; 1. KW Photoreceptor; Retinal protein; Transmembrane; Glycoprotein; Vision; KW Phosphorylation; Lipoprotein; Palmitate; G-protein coupled receptor; KW Acetylation; Retinitis pigmentosa; Disease mutation. FT DOMAIN 1 36 EXTRACELLULAR. FT TRANSMEM 37 61 1 (POTENTIAL). FT DOMAIN 62 73 CYTOPLASMIC. FT TRANSMEM 74 98 2 (POTENTIAL). FT DOMAIN 99 113 EXTRACELLULAR. FT TRANSMEM 114 133 3 (POTENTIAL). FT DOMAIN 134 152 CYTOPLASMIC. FT TRANSMEM 153 176 4 (POTENTIAL). FT DOMAIN 177 202 EXTRACELLULAR. FT TRANSMEM 203 230 5 (POTENTIAL). FT DOMAIN 231 252 CYTOPLASMIC. FT TRANSMEM 253 276 6 (POTENTIAL). FT DOMAIN 277 284 EXTRACELLULAR. FT TRANSMEM 285 309 7 (POTENTIAL). FT DOMAIN 310 348 CYTOPLASMIC. FT MOD_RES 1 1 ACETYLATION (BY SIMILARITY). FT CARBOHYD 2 2 N-LINKED (GLCNAC. . . ) (BY SIMILARITY). FT CARBOHYD 15 15 N-LINKED (GLCNAC. . . ) (BY SIMILARITY). FT DISULFID 110 187 BY SIMILARITY. FT BINDING 296 RETINAL CHROMOPHORE.
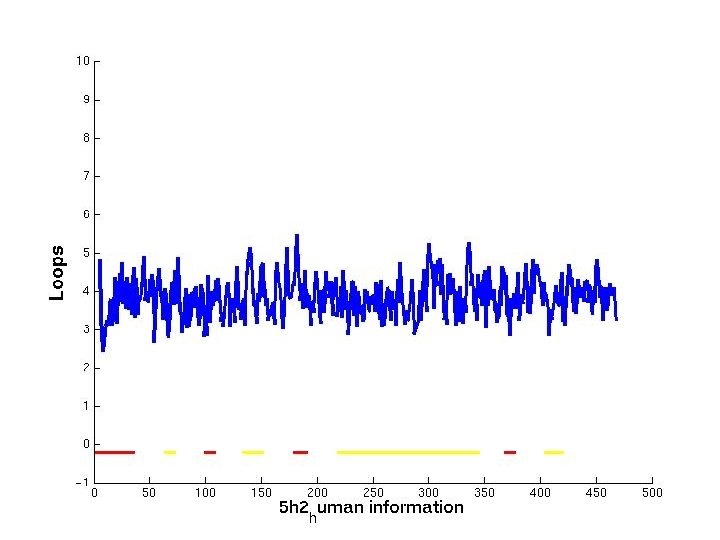
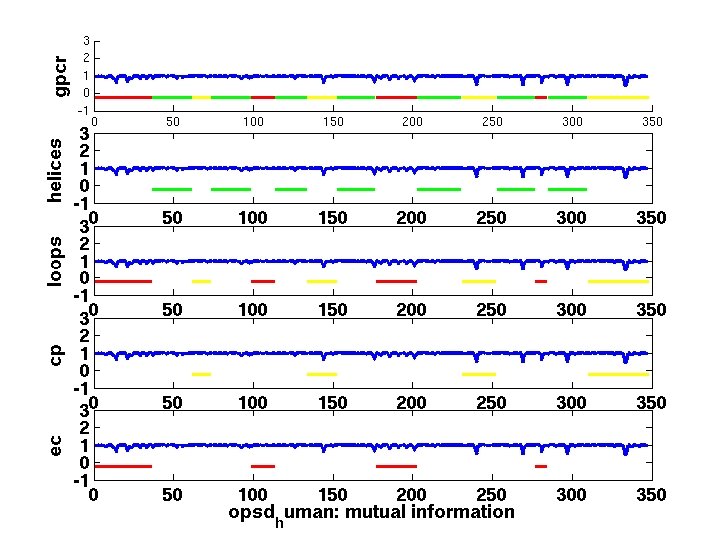

Problems/Solution • Problems: -Making one subplot graph (MATLAB) requires program customization - Generation of multiple subplots together requires more tedious work. Waste of time and effort. • Solution: -Need clear interface to generate subplot graphs for you w/o writing tedious matlab code.
![[a 1, b 1]=textread(’test. out', '%d %f'); hold on subplot(1, 1, 1); hold on [a 1, b 1]=textread(’test. out', '%d %f'); hold on subplot(1, 1, 1); hold on](https://present5.com/presentation/38923564bf719b9b073d05d6c5cf9aed/image-9.jpg)
[a 1, b 1]=textread(’test. out', '%d %f'); hold on subplot(1, 1, 1); hold on hh 1 = plot(a 1, b 1, 'linewidth', 2. 5); hold on ylabel('yaxis', 'fontsize', 16, 'Color', 'k', 'fontweight', 'bold'); set(hh 1, 'Marker. Size', 5); set(gca, 'YLim', [-1, 3]); %set(gca, 'ytick', [-. 6, -. 2, . 2] xdash = [Na. N, 62, 73, Na. N, 134, 152, Na. N, 231, 252, Na. N, 310, 348]; %cp ydash = (-. 2)*(ones(size(xdash))); line(xdash, ydash, 'color', 'y', 'linewidth', 3); xdash = [1, 36, Na. N, 99, 113, Na. N, 177, 202, Na. N, 277, 284, Na. N]; %ec ydash = (-. 2)*(ones(size(xdash))); line(xdash, ydash, 'color', 'linewidth', 3); hold on xlabel('x_axis', 'fontsize', 16, 'Color', 'k'); print -dpsc -r 0 sample;

Design Capabilities • Access multiple mutual information output datasets • Display combination of EC/CP/H position information on MI datasets (color coded) • Specify range (Y limits) and naming conventions (X axis) • Output into convenient picture files (ex: . tiff file).

Subplotter • Version 1: (In house only) -Initially the program takes as input: -->. SW file: (EC/CP/H) -->. m file: (MATLAB file that code will be generated in)

Subplotter ( Version 1) ****************** How many output files to textread: 1 What is the file to be textread into matlab program [output file 1]: opsdh_1 gpcr. out How many TOTAL subplots do you request? : 1 ******************

Subplotter ( Version 1) *********** Subplot(1, 1, 1) *********** Which file do you want results to be graphed on this subplot? : 0: opsdh_1 gpcr. out Make selection (0): 0 +++++++++++++++++++++++ How many items (EC, CP, H) do you want plotted (1, 2, 3: GPCR, 4: Loops)? : ++++++++++++++++++++++++ --> 3

Subplotter ( Version 1) Specify Y-Axis Label? (y/n): n Y-Axis Label: GPCR Specify YLim? (y, n): n Give name to X-Axis: sample Give name to. tiff file for output (no extension!): sample Matlab Program completed! wait. . .
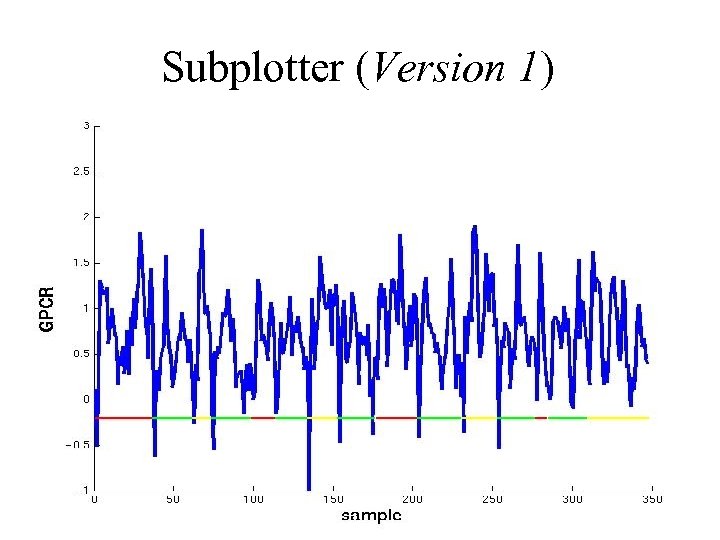
Subplotter (Version 1)

Current/Future Work • Generate graphing utility for every tool on the BLMT website.

Questions?
38923564bf719b9b073d05d6c5cf9aed.ppt