2986ccd525af0915cd4ecaf8123f48be.ppt
- Количество слайдов: 103

BIOINFORMATICS AND SYSTEMS BIOLOGY, MSC PROGR Sequence analysis, UMF 018, 2010 Databases in bioinformatics II Marcela Davila-Lopez Department of Medical Biochemistry and Cell Biology Institute of Biomedicine

Overview – Genome sequencing – Sequencing methods • • Sanger, Maxam Next generation methods (2 nd, 3 rd) Uses Implications – Ref. Seq vs Gen. Bank – Trace. Archive – Refining searches at Entrez – e. Utilis (programer utilities) Databases in bioinformatics II 2

Genome sequencing Databases in bioinformatics II 3

Why Sequencing Genomes Databases in bioinformatics II 4

Why Sequencing Genomes Remarkable similar molecular level despite their obvious outward differences genes similar DNA sequence tend to perform ≈ functions Understanding the function of a gene in one organism we may get an idea of what function that gene may perform in a more complex organism (humans) Databases in bioinformatics II 5

Archon X Prize "the first Team that can build a device and use it to sequence 100 human genomes within 10 days or less, with an accuracy of no more than one error in every 100, 000 bases sequenced, with sequences accurately covering at least 98% of the genome, and at a recurring cost of no more than $10, 000 (US) per genome. " $10 million HGP 1993 1 st draft 2000 final 2003 ($3 billion) Craig Venter 2007 70 million ~10 years James Watson 2008 1 million ~ 2 months Databases in bioinformatics II 6

Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 7
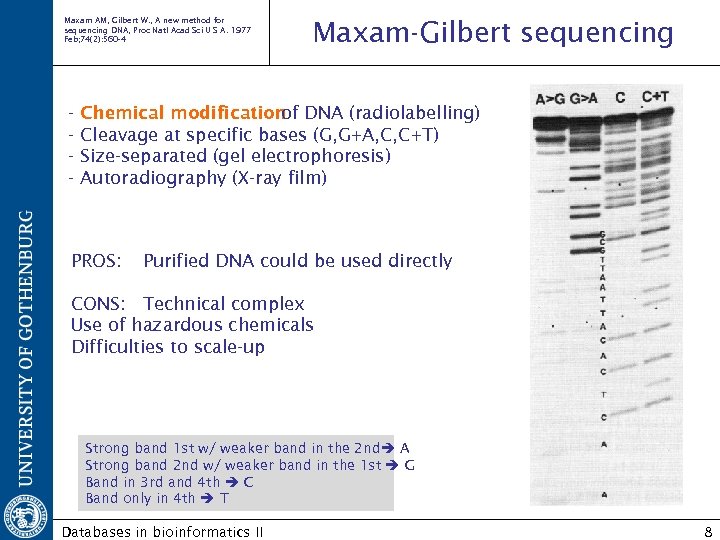
Maxam AM, Gilbert W. , A new method for sequencing DNA, Proc Natl Acad Sci U S A. 1977 Feb; 74(2): 560 -4 - Maxam-Gilbert sequencing Chemical modification DNA (radiolabelling) of Cleavage at specific bases (G, G+A, C, C+T) Size-separated (gel electrophoresis) Autoradiography (X-ray film) PROS: Purified DNA could be used directly CONS: Technical complex Use of hazardous chemicals Difficulties to scale-up Strong band 1 st w/ weaker band in the 2 nd A Strong band 2 nd w/ weaker band in the 1 st G Band in 3 rd and 4 th C Band only in 4 th T Databases in bioinformatics II 8
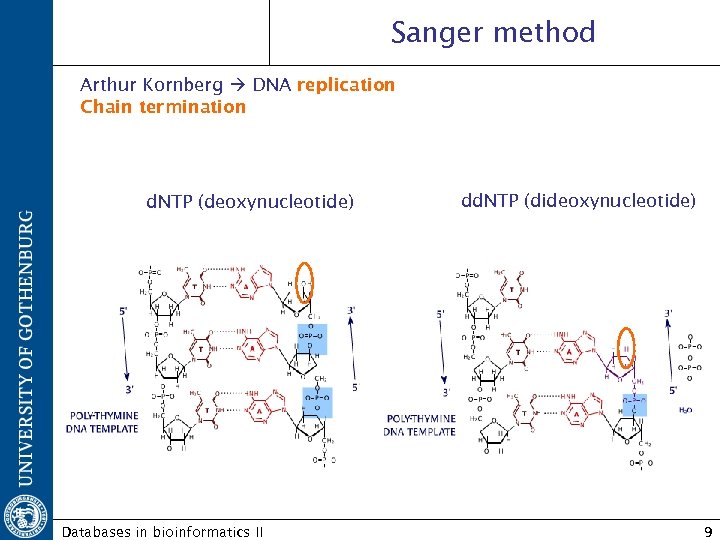
Sanger method Arthur Kornberg DNA replication Chain termination d. NTP (deoxynucleotide) Databases in bioinformatics II dd. NTP (dideoxynucleotide) 9
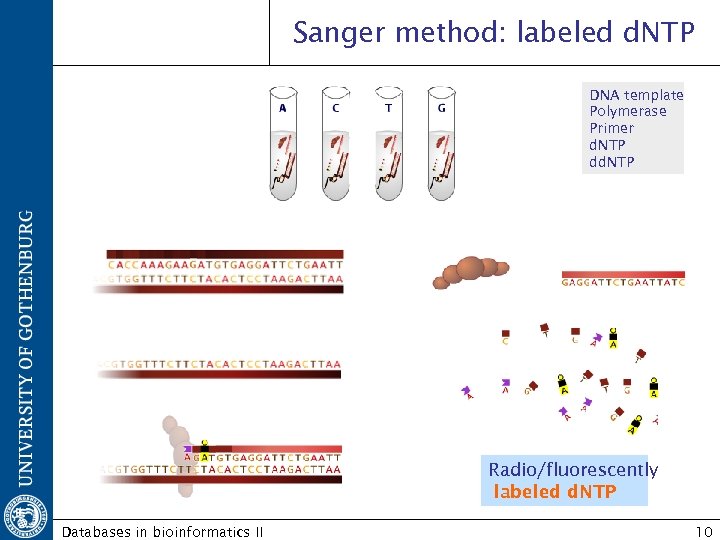
Sanger method: labeled d. NTP DNA template Polymerase Primer d. NTP dd. NTP Radio/fluorescently labeled d. NTP Databases in bioinformatics II 10
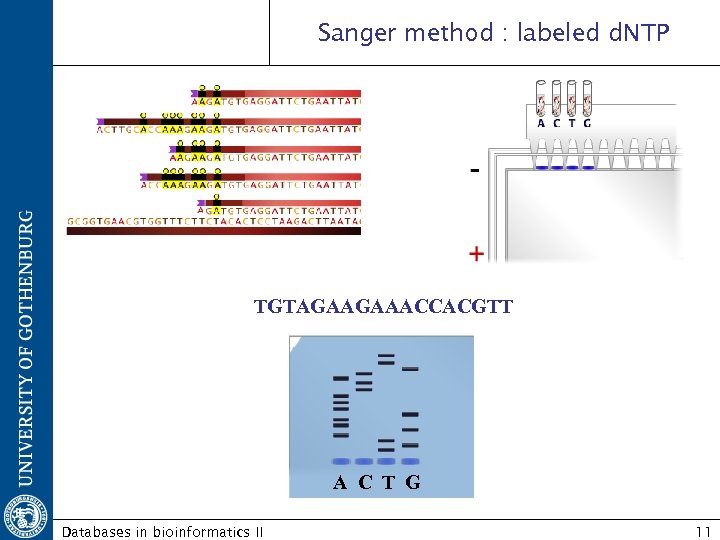
Sanger method : labeled d. NTP TGTAGAAGAAACCACGTT A C T G Databases in bioinformatics II 11
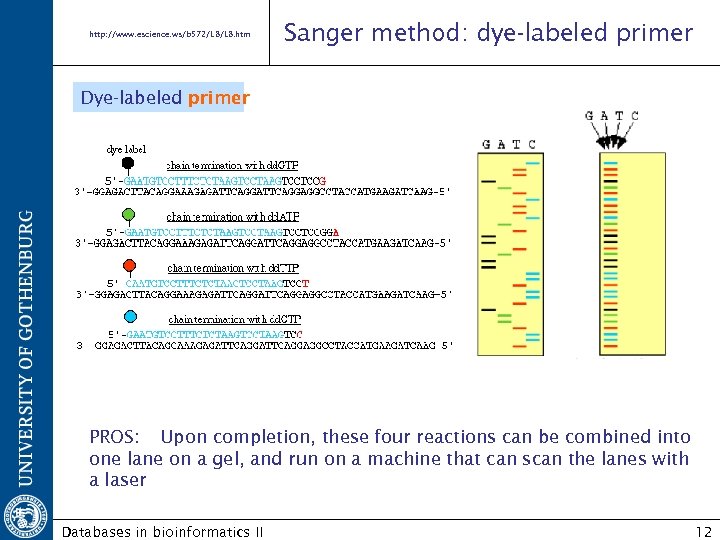

http: //www. escience. ws/b 572/L 8. htm Sanger method: dye-labeled primer Dye-labeled primer PROS: Upon completion, these four reactions can be combined into one lane on a gel, and run on a machine that can scan the lanes with a laser Databases in bioinformatics II 12
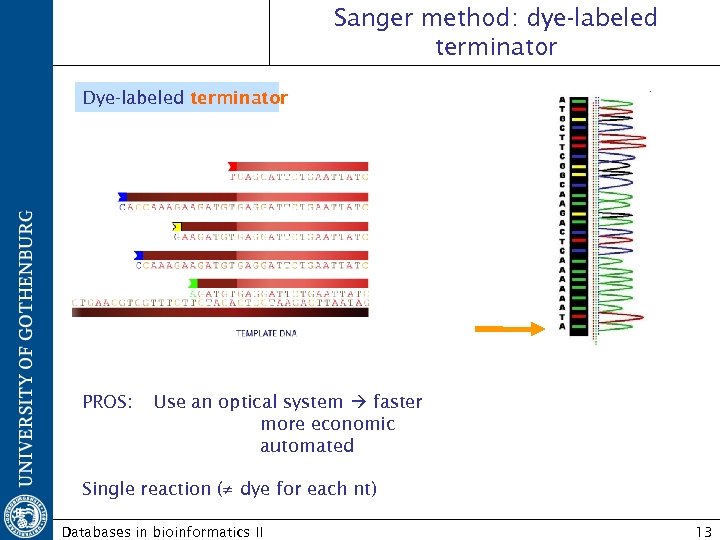
Sanger method: dye-labeled terminator Dye-labeled terminator PROS: Use an optical system faster more economic automated Single reaction (≠ dye for each nt) Databases in bioinformatics II 13
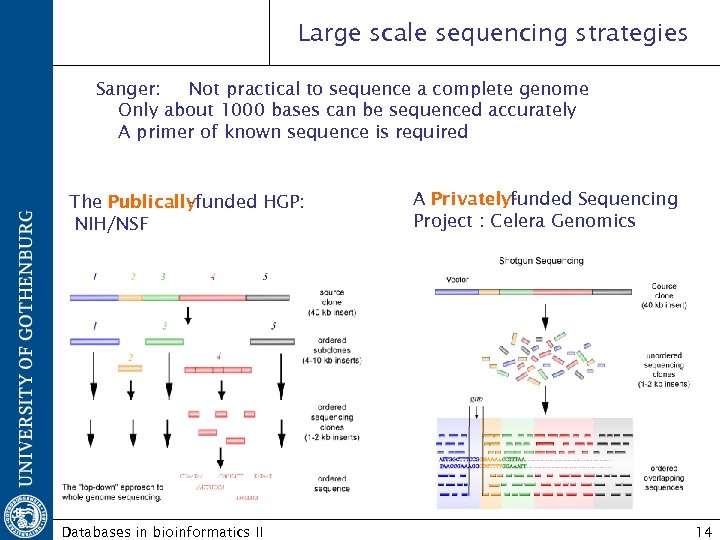
Large scale sequencing strategies Sanger: Not practical to sequence a complete genome Only about 1000 bases can be sequenced accurately A primer of known sequence is required The Publically -funded HGP: NIH/NSF Databases in bioinformatics II A Privately-unded Sequencing f Project : Celera Genomics 14
Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 15
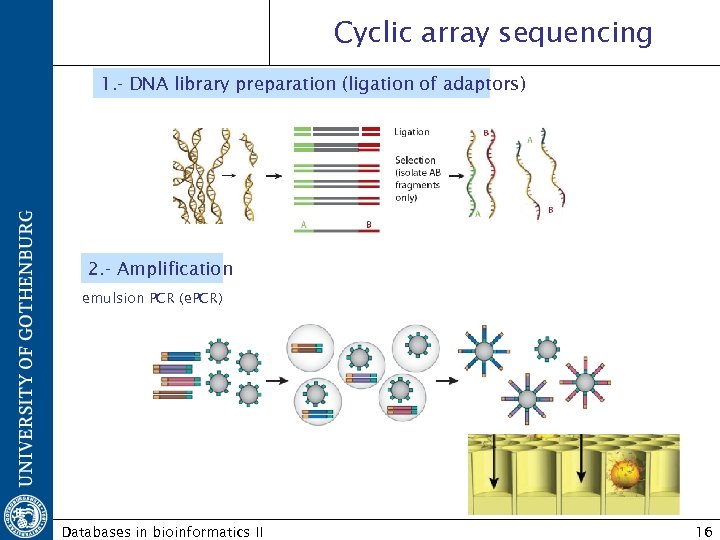
Cyclic array sequencing 1. - DNA library preparation (ligation of adaptors) 2. - Amplification emulsion PCR (e. PCR) Databases in bioinformatics II 16
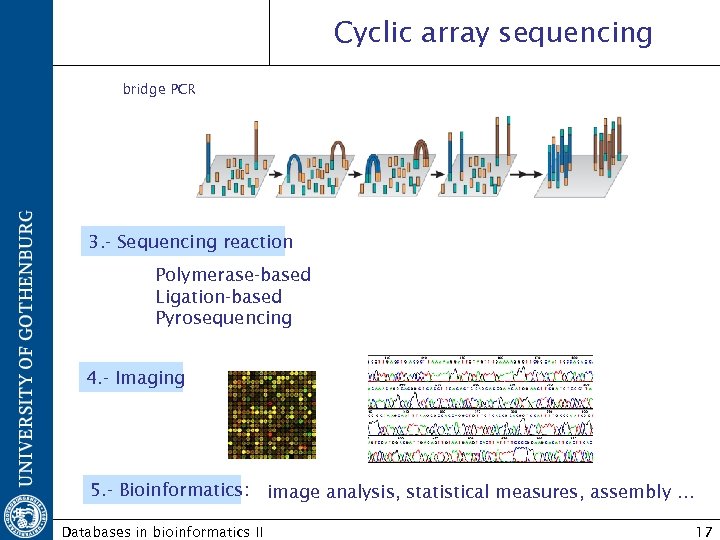
Cyclic array sequencing bridge PCR 3. - Sequencing reaction Polymerase-based Ligation-based Pyrosequencing 4. - Imaging 5. - Bioinformatics: image analysis, statistical measures, assembly … Databases in bioinformatics II 17
Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 18

Illumina/Solexa genome analyzer http: //www. illumina. com/technology/sequencing_technology. ilmn Sequencing by synthesis Detects the fluorescence of the added nucleotide at each position while synthesizing the complementary strand. Reverse terminator Databases in bioinformatics II 19
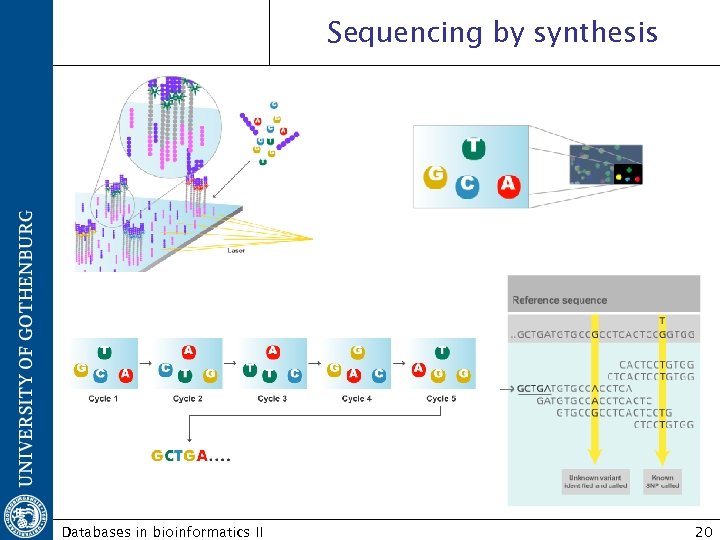
Sequencing by synthesis Databases in bioinformatics II 20
Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 21

Roche/454 FLX http: //www. roche-applied-science. com/index. jsp http: //www. roche-applied-science. com/publications/multimedia/genome_sequencer/flx_multimedia/wbt. h Pyrosequencing Detects the activity of DNA polymerase with a chemiluminescent enzyme by synthesizing the complementary strand. Databases in bioinformatics II 22
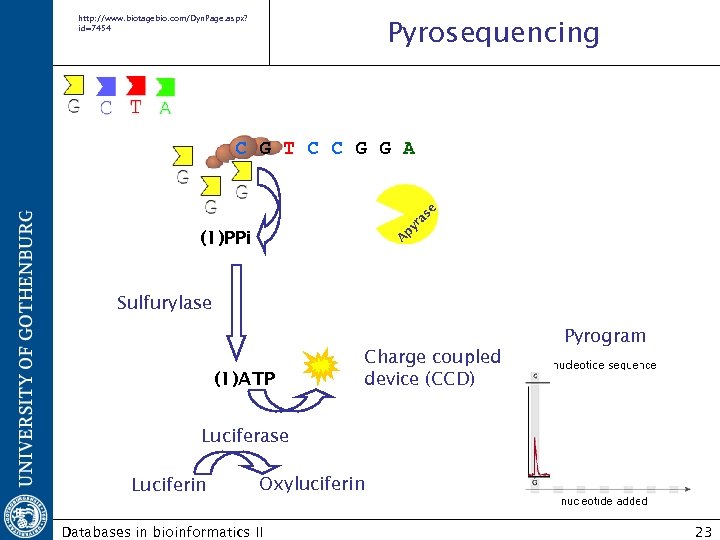
http: //www. biotagebio. com/Dyn. Page. aspx? id=7454 Pyrosequencing A py ra se C G T C C G G A (1)PPi Sulfurylase Pyrogram (1)ATP Charge coupled device (CCD) Luciferase Luciferin Oxyluciferin Databases in bioinformatics II 23
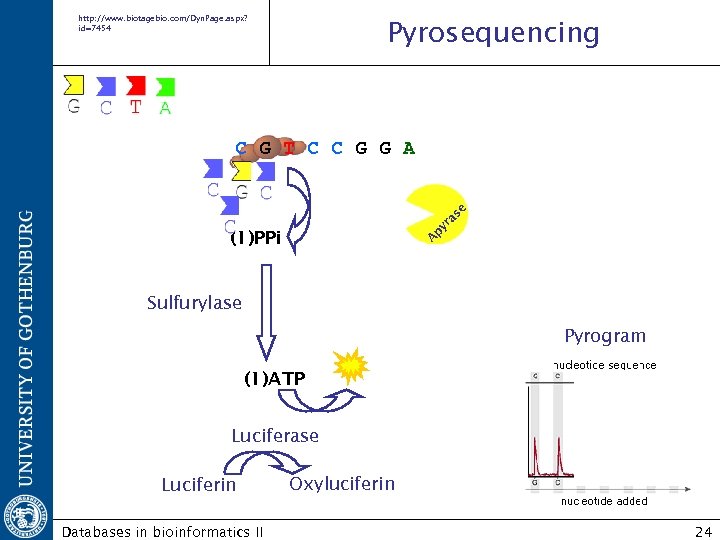
http: //www. biotagebio. com/Dyn. Page. aspx? id=7454 Pyrosequencing A py ra se C G T C C G G A (1)PPi Sulfurylase Pyrogram (1)ATP Luciferase Luciferin Databases in bioinformatics II Oxyluciferin 24
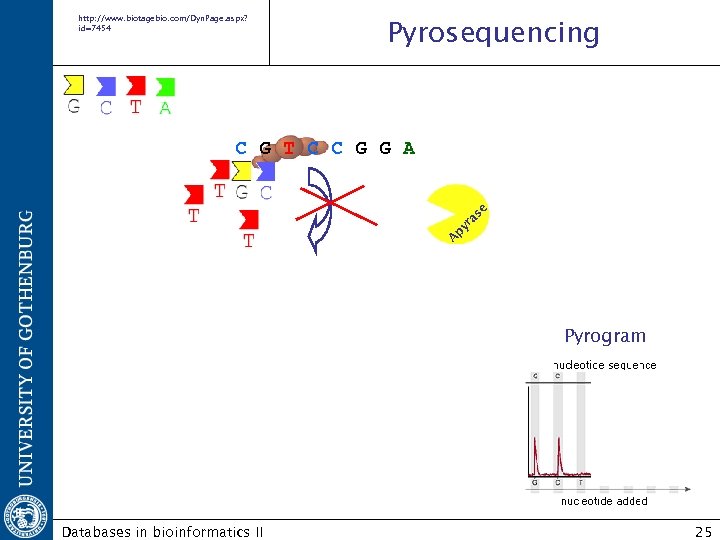
http: //www. biotagebio. com/Dyn. Page. aspx? id=7454 Pyrosequencing A py ra se C G T C C G G A Pyrogram Databases in bioinformatics II 25
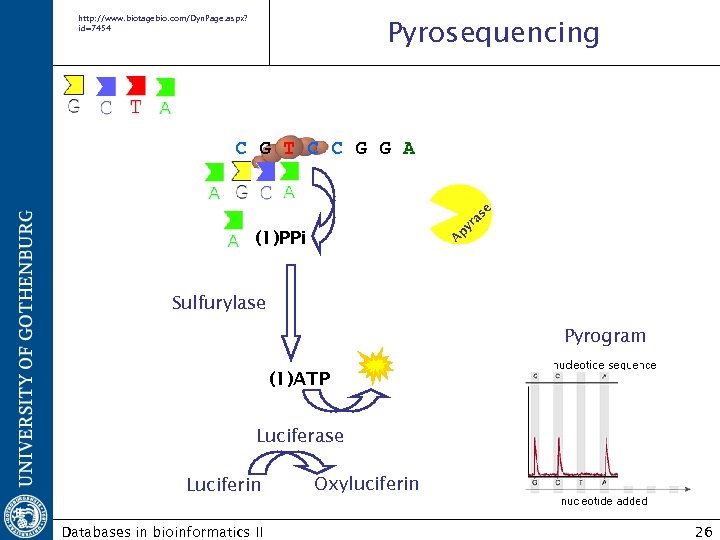
http: //www. biotagebio. com/Dyn. Page. aspx? id=7454 Pyrosequencing A py ra se C G T C C G G A (1)PPi Sulfurylase Pyrogram (1)ATP Luciferase Luciferin Databases in bioinformatics II Oxyluciferin 26
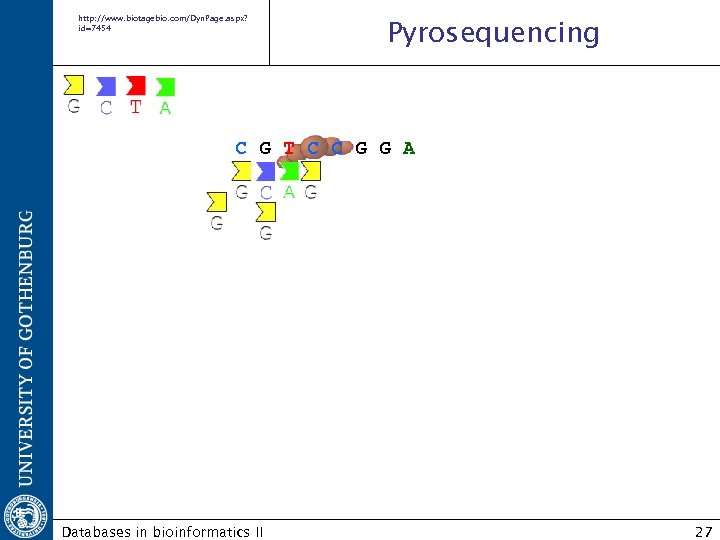
http: //www. biotagebio. com/Dyn. Page. aspx? id=7454 Pyrosequencing C G T C C G G A Databases in bioinformatics II 27
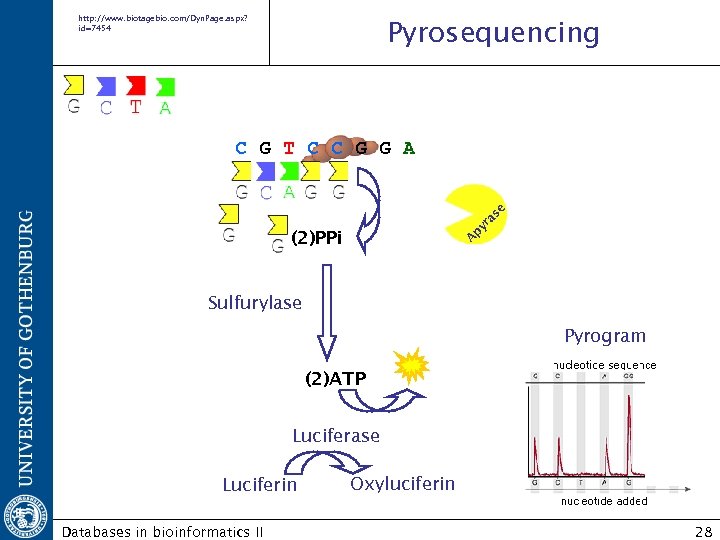
http: //www. biotagebio. com/Dyn. Page. aspx? id=7454 Pyrosequencing A py ra se C G T C C G G A (2)PPi Sulfurylase Pyrogram (2)ATP Luciferase Luciferin Databases in bioinformatics II Oxyluciferin 28
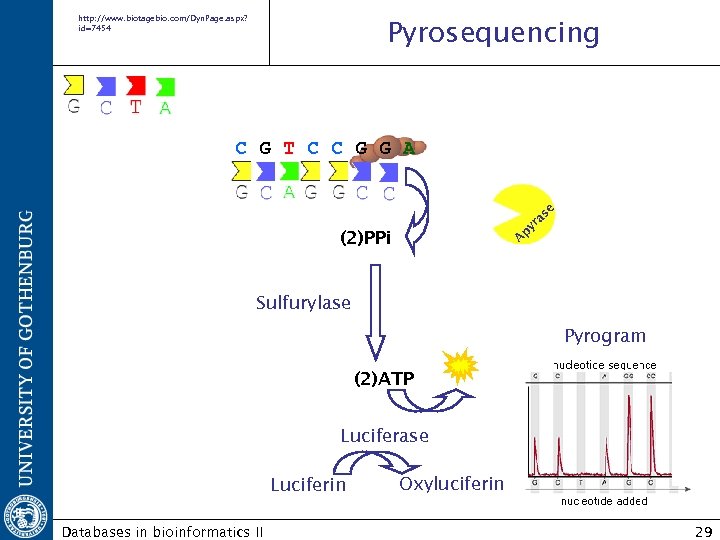
http: //www. biotagebio. com/Dyn. Page. aspx? id=7454 Pyrosequencing A py ra se C G T C C G G A (2)PPi Sulfurylase Pyrogram (2)ATP Luciferase Luciferin Databases in bioinformatics II Oxyluciferin 29
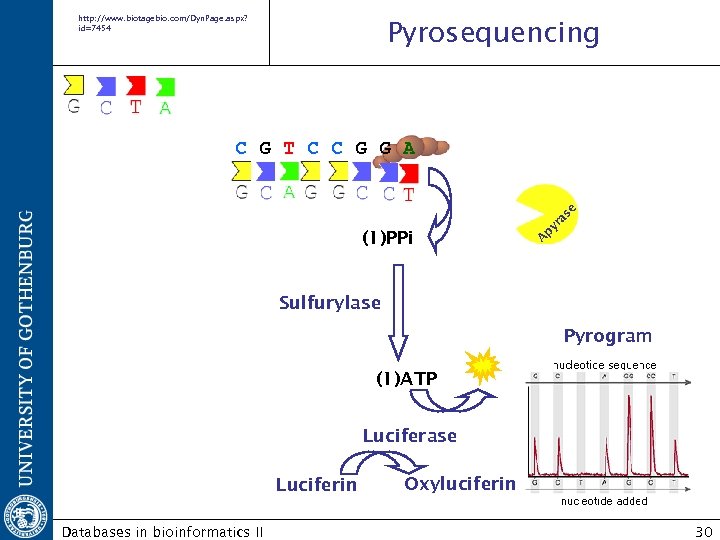
http: //www. biotagebio. com/Dyn. Page. aspx? id=7454 Pyrosequencing (1)PPi A py ra se C G T C C G G A Sulfurylase Pyrogram (1)ATP Luciferase Luciferin Databases in bioinformatics II Oxyluciferin 30
Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 31

http: //www 3. appliedbiosystems. com/AB_Home/ index. htm Applied biosystems / SOLi. D System TM http: //appliedbiosystems. cnpg. com/Video/flat. Files/699/index. aspx Sequencing by ligation Uses the enzyme DNA ligase to identify the nucleotide present at a given position in a DNA sequence. Databases in bioinformatics II 32
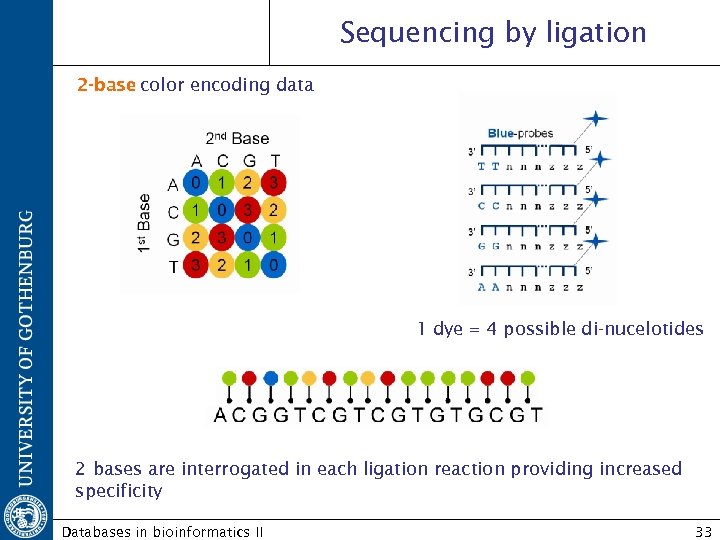
Sequencing by ligation 2 -base color encoding data 1 dye = 4 possible di-nucelotides 2 bases are interrogated in each ligation reaction providing increased specificity Databases in bioinformatics II 33
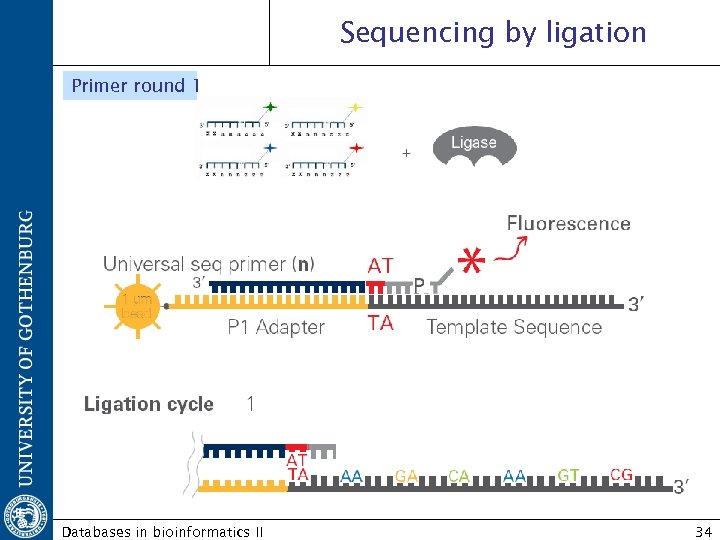
Sequencing by ligation Primer round 1 Databases in bioinformatics II 34
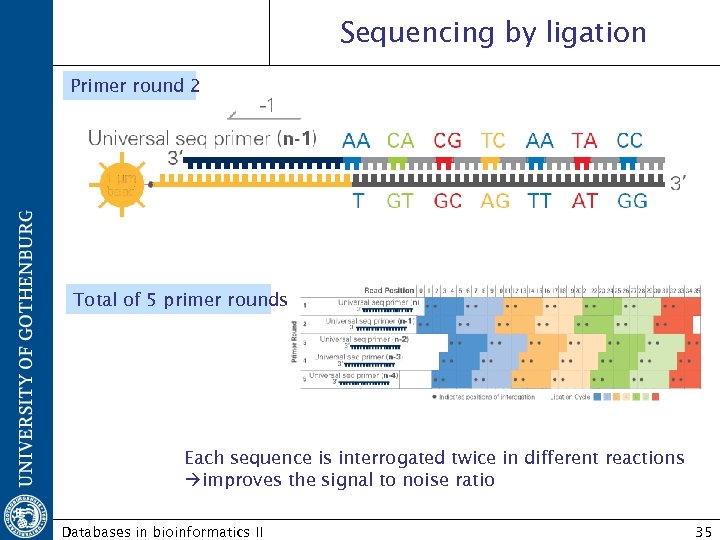
Sequencing by ligation Primer round 2 Total of 5 primer rounds Each sequence is interrogated twice in different reactions improves the signal to noise ratio Databases in bioinformatics II 35
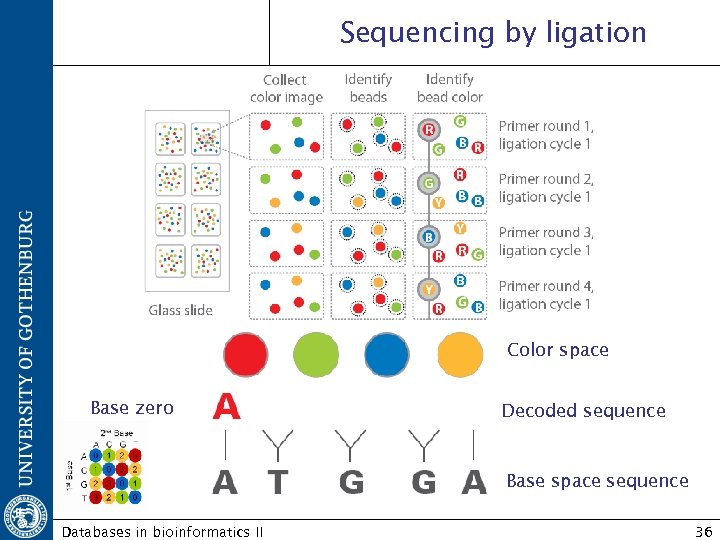
Sequencing by ligation Color space Base zero Decoded sequence Base space sequence Databases in bioinformatics II 36
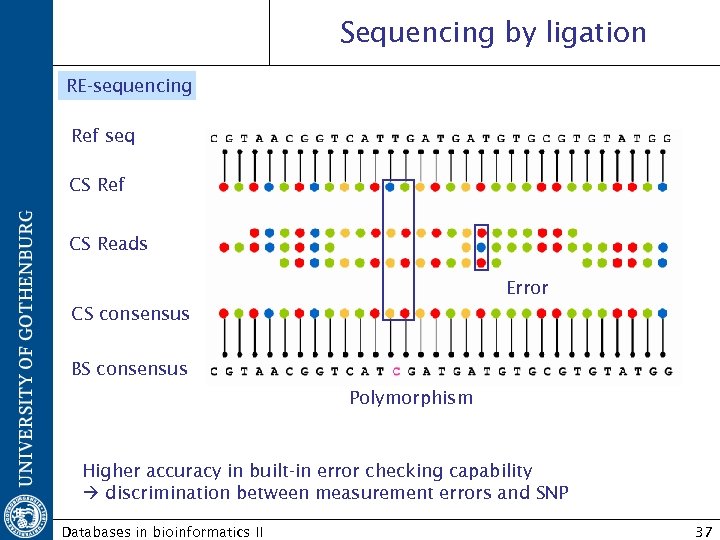
Sequencing by ligation RE-sequencing Ref seq CS Ref CS Reads Error CS consensus BS consensus Polymorphism Higher accuracy in built-in error checking capability discrimination between measurement errors and SNP Databases in bioinformatics II 37
Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 38

Helicos Heliscope TM http: //www. helicosbio. com/Default. aspx? base http: //www. pacificbiosciences. com/aboutus/videogallery? video. Image=pac_bio_lg. jpg Sequencing by synthesis Databases in bioinformatics II 39
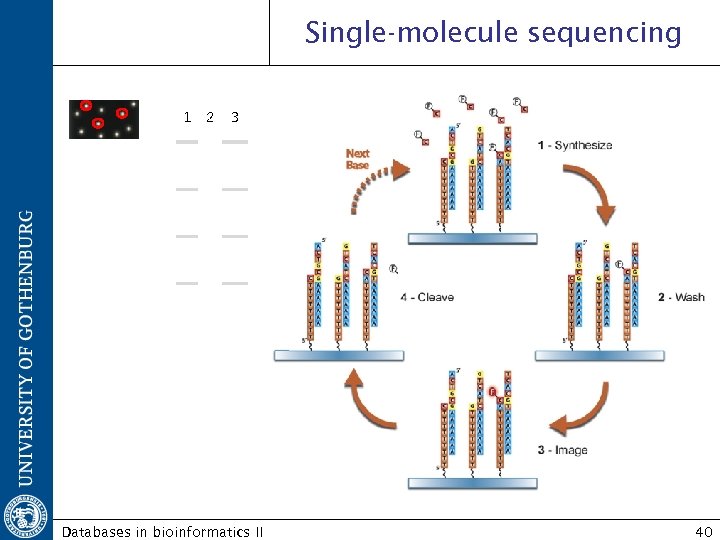
Single-molecule sequencing 1 2 3 A A C C G G T T A C G T Databases in bioinformatics II 40
Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 41
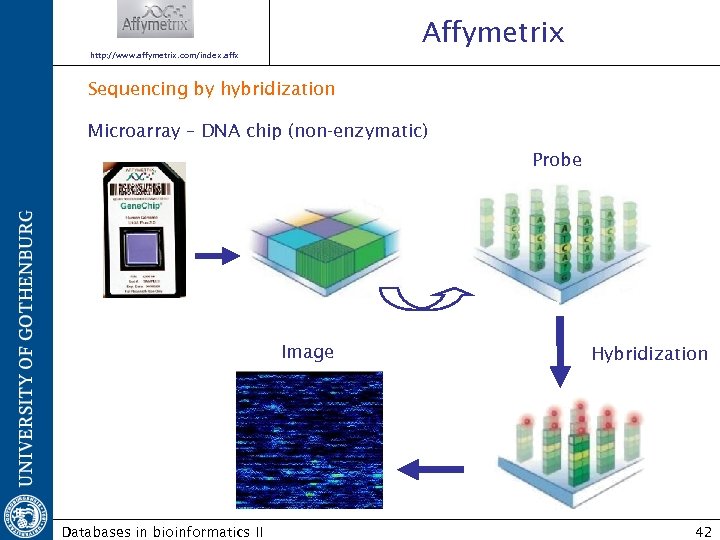
Affymetrix http: //www. affymetrix. com/index. affx Sequencing by hybridization Microarray – DNA chip (non-enzymatic) Probe Image Databases in bioinformatics II Hybridization 42
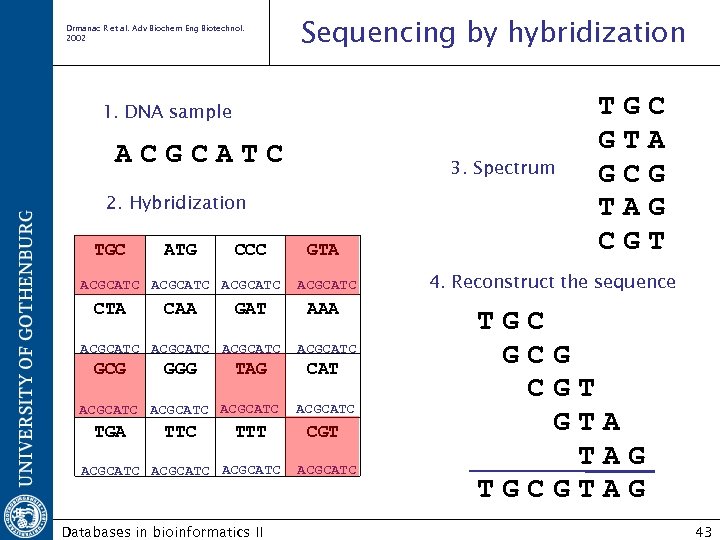
Drmanac R et al. Adv Biochem Eng Biotechnol. 2002 Sequencing by hybridization 1. DNA sample ACGCATC 3. Spectrum 2. Hybridization TGC ATG CCC ACGCATC CTA CAA GAT ACGCATC GCG GGG TAG ACGCATC TGA TTC TTT ACGCATC Databases in bioinformatics II GTA ACGCATC AAA ACGCATC CAT ACGCATC CGT ACGCATC T G G T C A G G T 4. Reconstruct the sequence TGC GCG CGT GTA TAG TGCGTAG 43
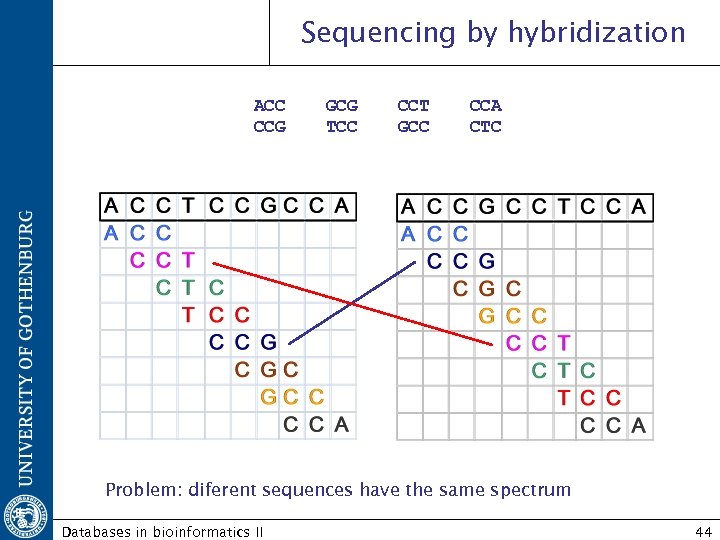
Sequencing by hybridization ACC CCG GCG TCC CCT GCC CCA CTC Problem: diferent sequences have the same spectrum Databases in bioinformatics II 44
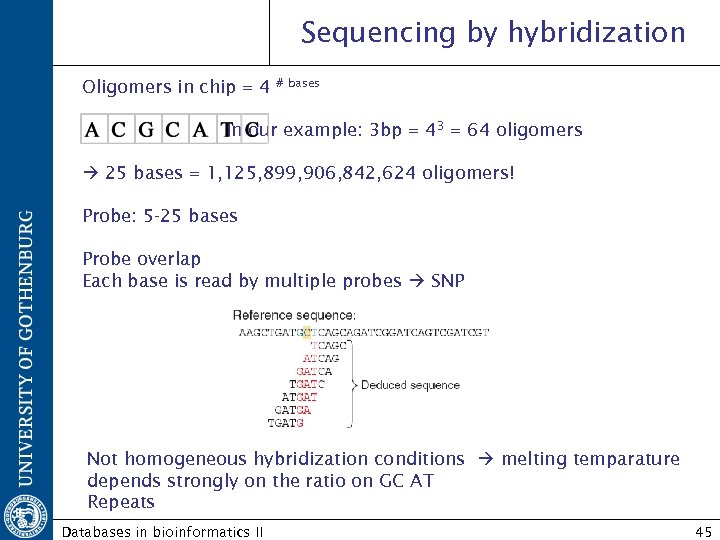

Sequencing by hybridization Oligomers in chip = 4 # bases In our example: 3 bp = 43 = 64 oligomers 25 bases = 1, 125, 899, 906, 842, 624 oligomers! Probe: 5 -25 bases Probe overlap Each base is read by multiple probes SNP Not homogeneous hybridization conditions melting temparature depends strongly on the ratio on GC AT Repeats Databases in bioinformatics II 45

Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 46

http: //www. pacificbiosciences. com/ Pacific Biosciences / SMRTTM technology http: //www. pacificbiosciences. com/video_lg. html Single Molecule Real Time Not commercially available Platform for single molecule real time detection based on DNA Polymerase activity. Databases in bioinformatics II 47
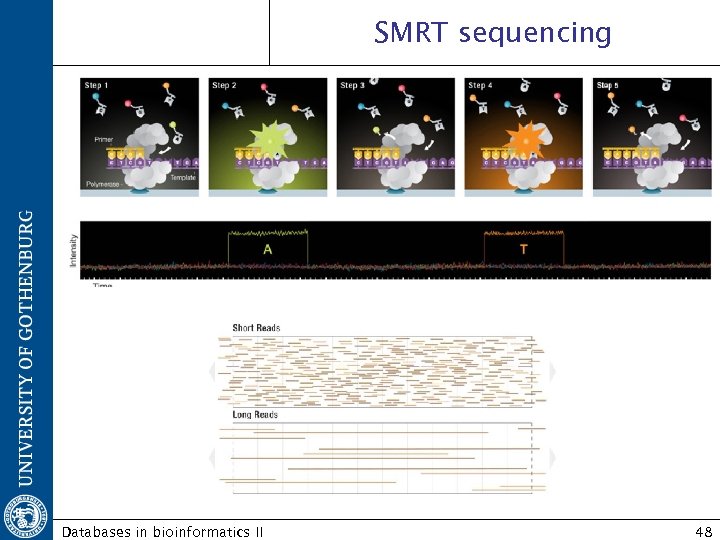
SMRT sequencing Databases in bioinformatics II 48
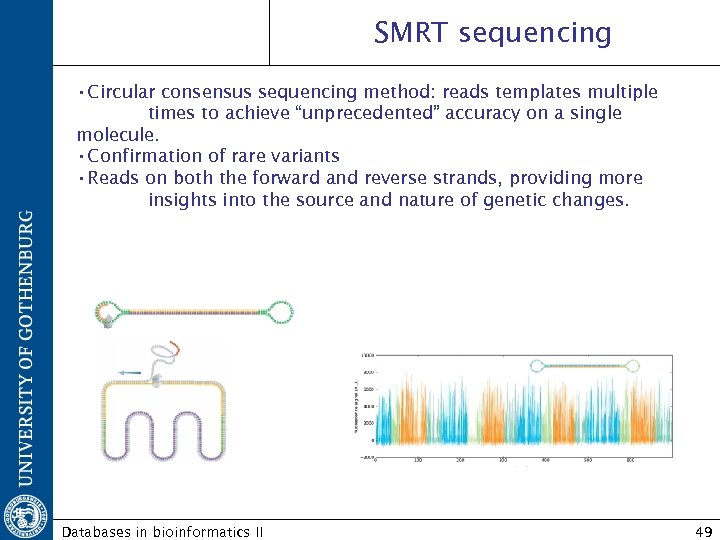
SMRT sequencing • Circular consensus sequencing method: reads templates multiple times to achieve “unprecedented” accuracy on a single molecule. • Confirmation of rare variants • Reads on both the forward and reverse strands, providing more insights into the source and nature of genetic changes. Databases in bioinformatics II 49

Sequencing methods 1 Whitfeld PR. - Sequencing by degradation 1975 -1977 W. Gilbert – A. Maxam (chemical modification) F. Sanger (chain termination) Next generation Cyclic array sequencing Illumina/Solexa Roche/454 AB SOLi. D Helicos/Heli. Scope Sequencing by Hybridization Affymetrix Sequencing in real time (3 rd generation) Oxford Nanopore Technologies Pacific Bioscience SMRT Databases in bioinformatics II 50
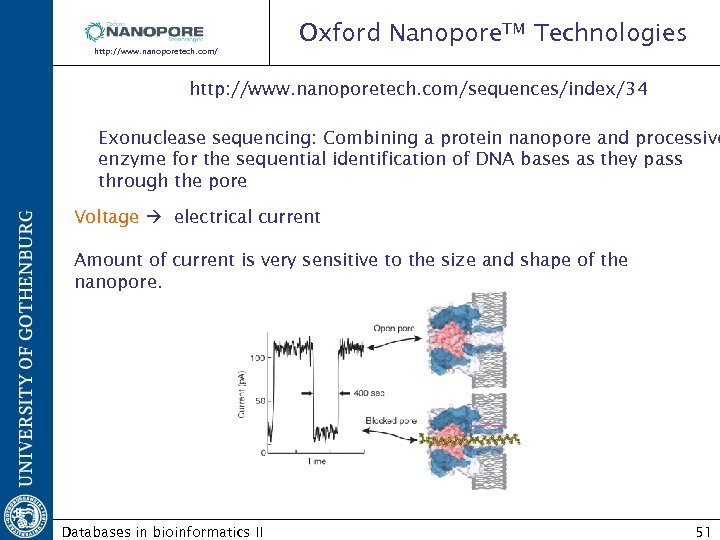

Oxford Nanopore. TM Technologies http: //www. nanoporetech. com/sequences/index/34 Exonuclease sequencing: Combining a protein nanopore and processive enzyme for the sequential identification of DNA bases as they pass through the pore Voltage electrical current Amount of current is very sensitive to the size and shape of the nanopore. Databases in bioinformatics II 51
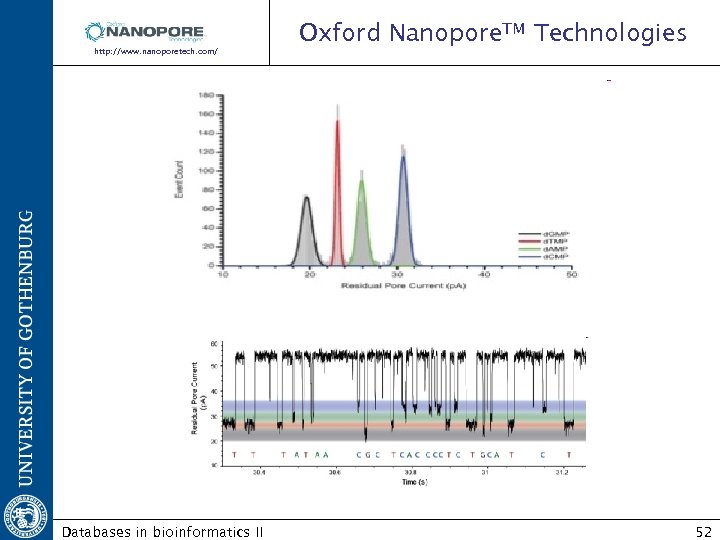
Oxford Nanopore. TM Technologies http: //www. nanoporetech. com/ Databases in bioinformatics II 52
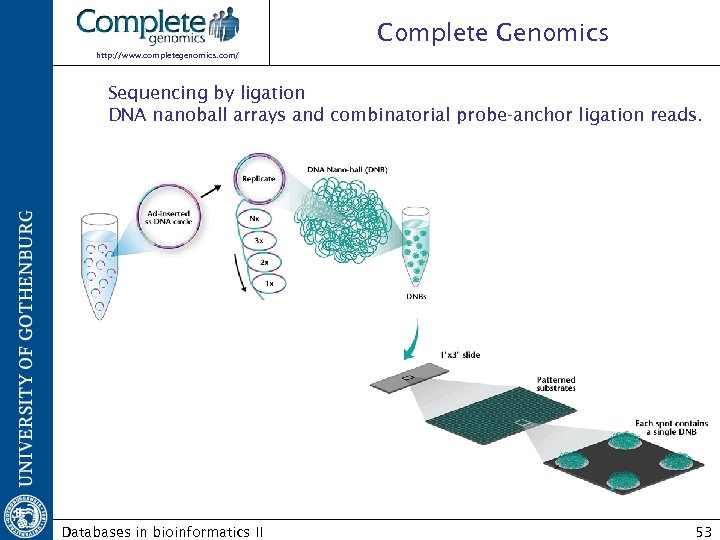
Complete Genomics http: //www. completegenomics. com/ Sequencing by ligation DNA nanoball arrays and combinatorial probe-anchor ligation reads. Databases in bioinformatics II 53
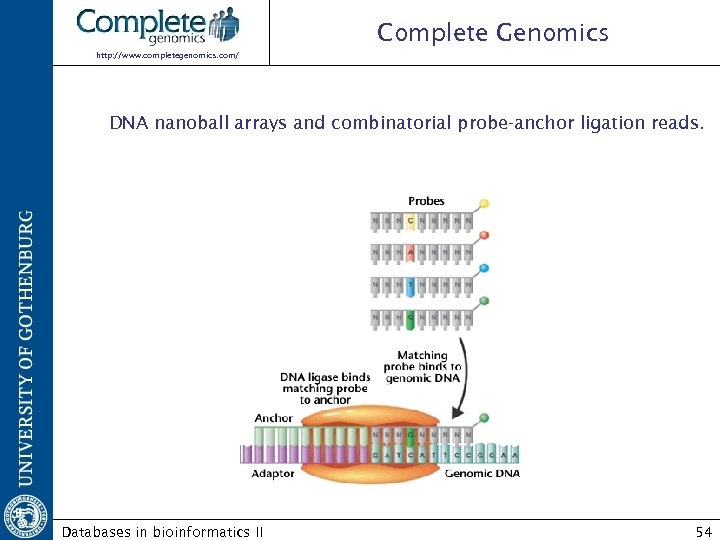
Complete Genomics http: //www. completegenomics. com/ DNA nanoball arrays and combinatorial probe-anchor ligation reads. Databases in bioinformatics II 54
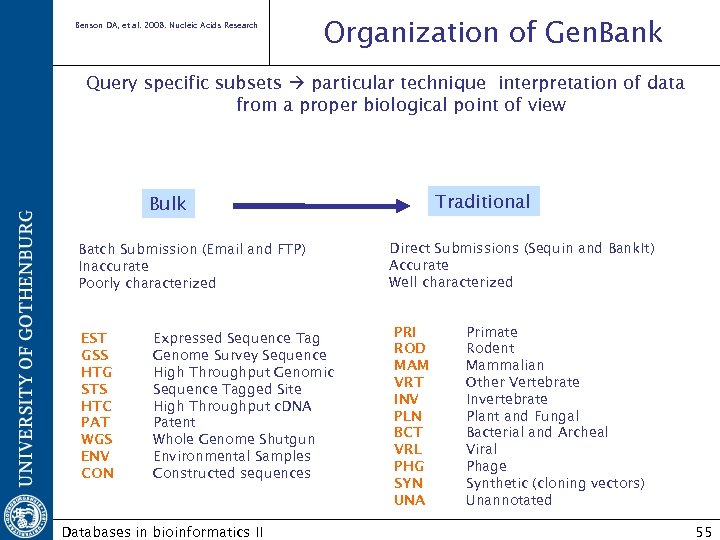

Benson DA, et al. 2008. Nucleic Acids Research Organization of Gen. Bank Query specific subsets particular technique interpretation of data from a proper biological point of view Traditional Bulk Batch Submission (Email and FTP) Inaccurate Poorly characterized EST GSS HTG STS HTC PAT WGS ENV CON Expressed Sequence Tag Genome Survey Sequence High Throughput Genomic Sequence Tagged Site High Throughput c. DNA Patent Whole Genome Shutgun Environmental Samples Constructed sequences Databases in bioinformatics II Direct Submissions (Sequin and Bank. It) Accurate Well characterized PRI ROD MAM VRT INV PLN BCT VRL PHG SYN UNA Primate Rodent Mammalian Other Vertebrate Invertebrate Plant and Fungal Bacterial and Archeal Viral Phage Synthetic (cloning vectors) Unannotated 55

Redundancy at Gen. Bank Many sequences are represented more than once in Gen. Bank huge degrees of Redundancy 2003 Ref. Seq collection : curated secondary database non-redundant selected organisms • Genome DNA (assemblies) • Transcripts (RNA) • Protein Databases in bioinformatics II 56
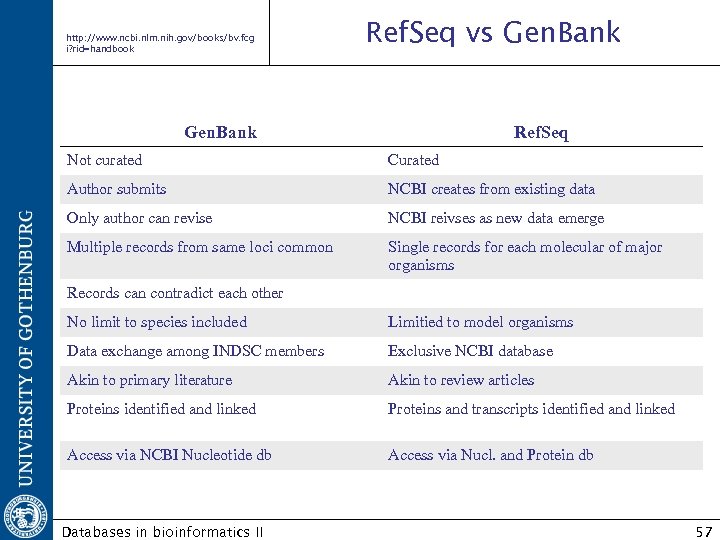

http: //www. ncbi. nlm. nih. gov/books/bv. fcg i? rid=handbook Ref. Seq vs Gen. Bank Ref. Seq Not curated Curated Author submits NCBI creates from existing data Only author can revise NCBI reivses as new data emerge Multiple records from same loci common Single records for each molecular of major organisms Records can contradict each other No limit to species included Limitied to model organisms Data exchange among INDSC members Exclusive NCBI database Akin to primary literature Akin to review articles Proteins identified and linked Proteins and transcripts identified and linked Access via NCBI Nucleotide db Access via Nucl. and Protein db Databases in bioinformatics II 57
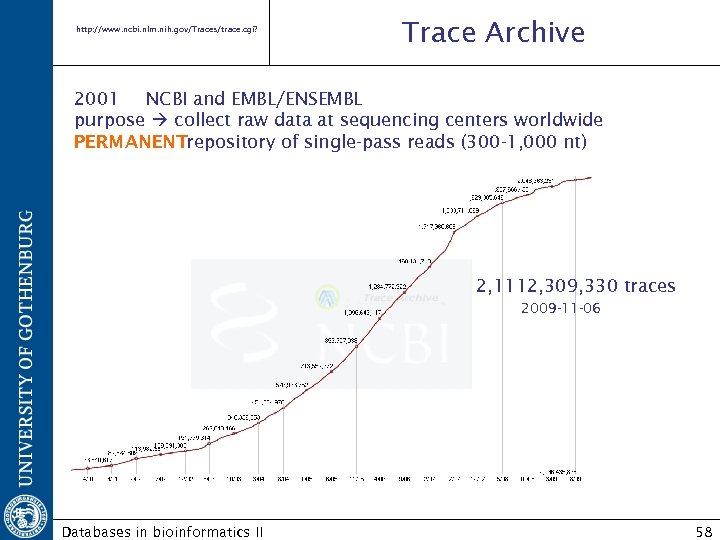
http: //www. ncbi. nlm. nih. gov/Traces/trace. cgi? Trace Archive 2001 NCBI and EMBL/ENSEMBL purpose collect raw data at sequencing centers worldwide PERMANENTrepository of single-pass reads (300 -1, 000 nt) 2, 1112, 309, 330 traces 2009 -11 -06 Databases in bioinformatics II 58

More advanced queries Databases in bioinformatics II 59

Entrez Databases in bioinformatics II 60
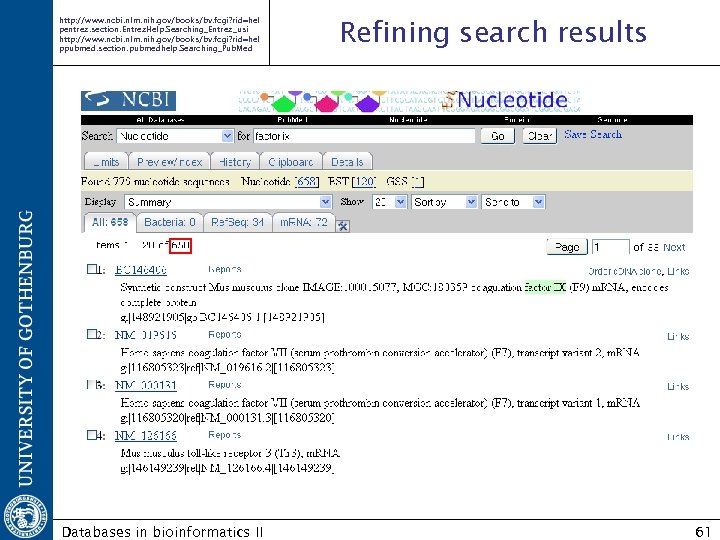
http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid=hel pentrez. section. Entrez. Help. Searching_Entrez_usi http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid=hel ppubmed. section. pubmedhelp. Searching_Pub. Med Databases in bioinformatics II Refining search results 61
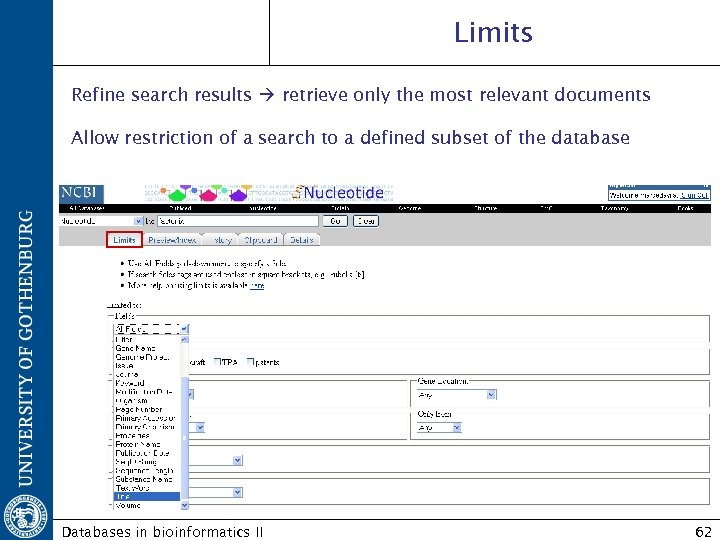
Limits Refine search results retrieve only the most relevant documents Allow restriction of a search to a defined subset of the database Databases in bioinformatics II 62
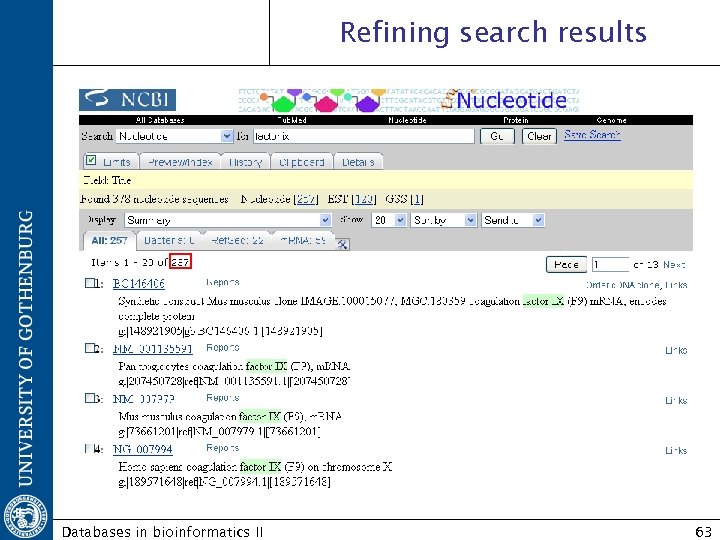
Refining search results Databases in bioinformatics II 63
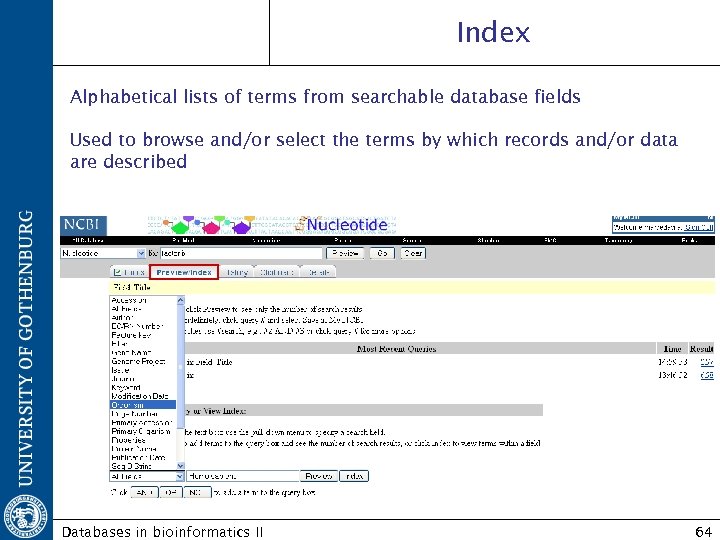
Index Alphabetical lists of terms from searchable database fields Used to browse and/or select the terms by which records and/or data are described Databases in bioinformatics II 64
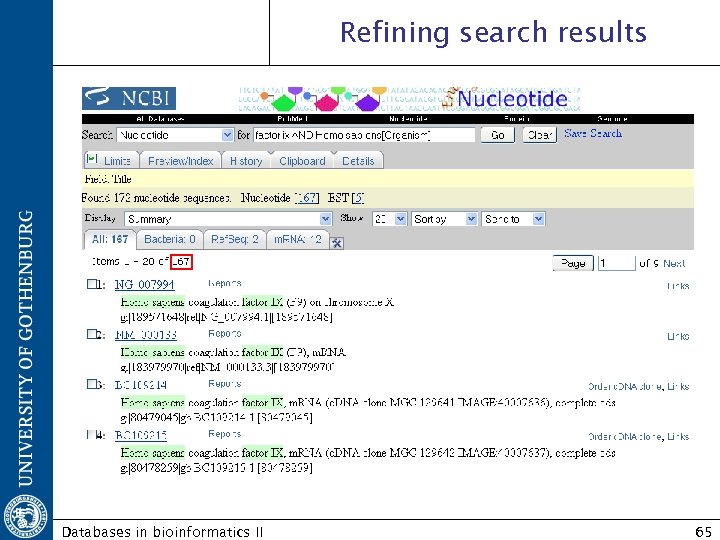
Refining search results Databases in bioinformatics II 65
![Advanced search statements term [field] OPERATOR term [field] Find all human nucleotide sequences with Advanced search statements term [field] OPERATOR term [field] Find all human nucleotide sequences with](https://present5.com/presentation/2986ccd525af0915cd4ecaf8123f48be/image-66.jpg)
Advanced search statements term [field] OPERATOR term [field] Find all human nucleotide sequences with D-loop annotations D-loop[FKEY] AND human[ORGN] in Nucleotide database Find Drosophila population studies published in the Journal of Molecular Evolution j mol evol[JOUR] AND drosophila[ORGN] Databases in bioinformatics II in Pop. Set database 66
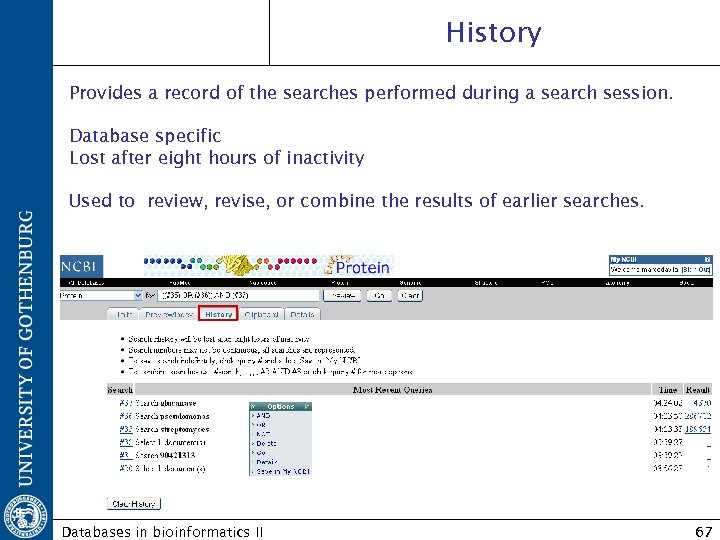
History Provides a record of the searches performed during a search session. Database specific Lost after eight hours of inactivity Used to review, revise, or combine the results of earlier searches. Databases in bioinformatics II 67
Combining results Databases in bioinformatics II 68
Query translation Databases in bioinformatics II 69
Details Display your search strategy as translated using Entrez's search and syntax rules Error messages, when applicable Databases in bioinformatics II 70
Author search Databases in bioinformatics II 71
Example - author Databases in bioinformatics II 72
Example - journal Databases in bioinformatics II 73

e. Utils Databases in bioinformatics II 74
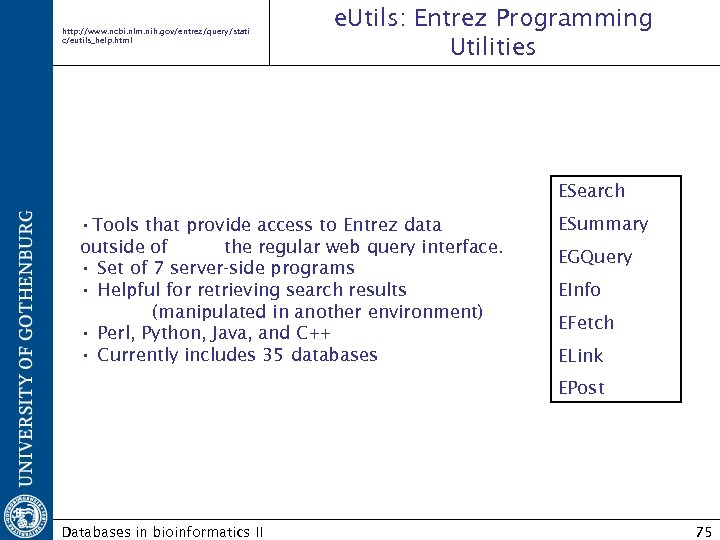

http: //www. ncbi. nlm. nih. gov/entrez/query/stati c/eutils_help. html e. Utils: Entrez Programming Utilities ESearch • Tools that provide access to Entrez data outside of the regular web query interface. • Set of 7 server-side programs • Helpful for retrieving search results (manipulated in another environment) • Perl, Python, Java, and C++ • Currently includes 35 databases ESummary EGQuery EInfo EFetch ELink EPost Databases in bioinformatics II 75
http: //www. ncbi. nlm. nih. gov/entrez/query/stati c/eutils_help. html URL e. Utils: Entrez Programming Utilities Result (XML) • Perform searches on large datasets • Implement data pipelines for genomic, proteomic, or microarray analysis • Create automated searches to keep local databases current • Create and download customized datasets • Seamlessly combine local data with NCBI data • Develop a focused interface to NCBI data Databases in bioinformatics II 76
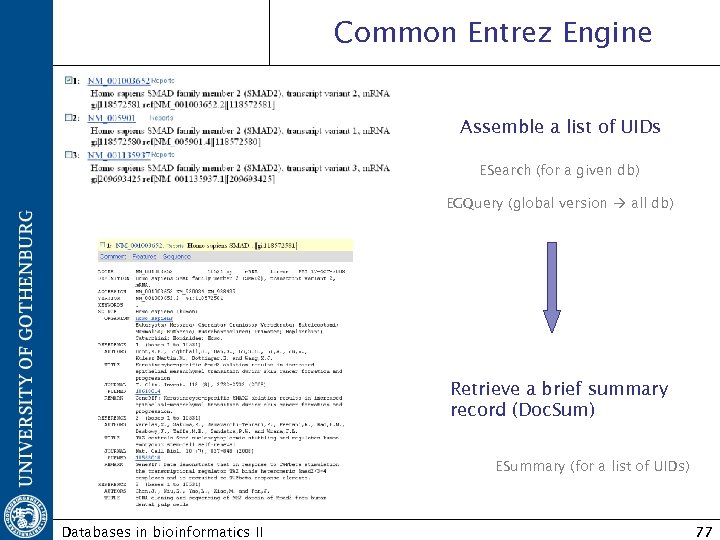

Common Entrez Engine Assemble a list of UIDs ESearch (for a given db) EGQuery (global version all db) Retrieve a brief summary record (Doc. Sum) ESummary (for a list of UIDs) Databases in bioinformatics II 77
![URL http: //www. ncbi. nlm. nih. gov/sites/gquery? term=cancer+stem+cells [Base_URL] [Eutils_URL] [Query] http: //eutils. ncbi. URL http: //www. ncbi. nlm. nih. gov/sites/gquery? term=cancer+stem+cells [Base_URL] [Eutils_URL] [Query] http: //eutils. ncbi.](https://present5.com/presentation/2986ccd525af0915cd4ecaf8123f48be/image-78.jpg)
URL http: //www. ncbi. nlm. nih. gov/sites/gquery? term=cancer+stem+cells [Base_URL] [Eutils_URL] [Query] http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/esummary. fcgi? db=taxonomy&id=9913&retmode=xml [Base_URL] Databases in bioinformatics II [Eutils_URL] [DB] [Query] 78
![URL: DB e. Search = [Base_URL] [Eutils_URL] [DB] [Query] Entrez Database E-Utility Database Name URL: DB e. Search = [Base_URL] [Eutils_URL] [DB] [Query] Entrez Database E-Utility Database Name](https://present5.com/presentation/2986ccd525af0915cd4ecaf8123f48be/image-79.jpg)
URL: DB e. Search = [Base_URL] [Eutils_URL] [DB] [Query] Entrez Database E-Utility Database Name 3 D Domains domains Domains cdd Genome genome Nucleotide nucleotide OMIM omim Pop. Set popset Protein protein Probe. Set geo Pub. Med pubmed Structure structure SNP snp Taxonomy taxonomy Uni. Gene unigene Uni. STS unists Databases in bioinformatics II Each Entrez DB has an E-Utility name (used instead of its original name) 79
![URL: Query e. Search = [Base_URL] [Eutils_URL] [DB] [Query] EFetch EGQuery term X EInfo URL: Query e. Search = [Base_URL] [Eutils_URL] [DB] [Query] EFetch EGQuery term X EInfo](https://present5.com/presentation/2986ccd525af0915cd4ecaf8123f48be/image-80.jpg)
URL: Query e. Search = [Base_URL] [Eutils_URL] [DB] [Query] EFetch EGQuery term X EInfo ESearch ESummary X db Espell X X Tax Seq Lit X X ELink EPost X X X field X reldate X X mindate X X maxdate X X datatype X X retstart X X retmax X X retmode X X X rettype X X history X X X X Web. Env X X X X query_key X X X X id report X strand X seq_start X seq_stop X dbfrom X cmd X Databases in bioinformatics II 80

EInfo Provides detailed information about a given database: term counts, last update and available links http: //eutils. ncbi. nlm. nih. gov/ entrez/eutils/einfo. fcgi? db=pubmed Databases in bioinformatics II 81
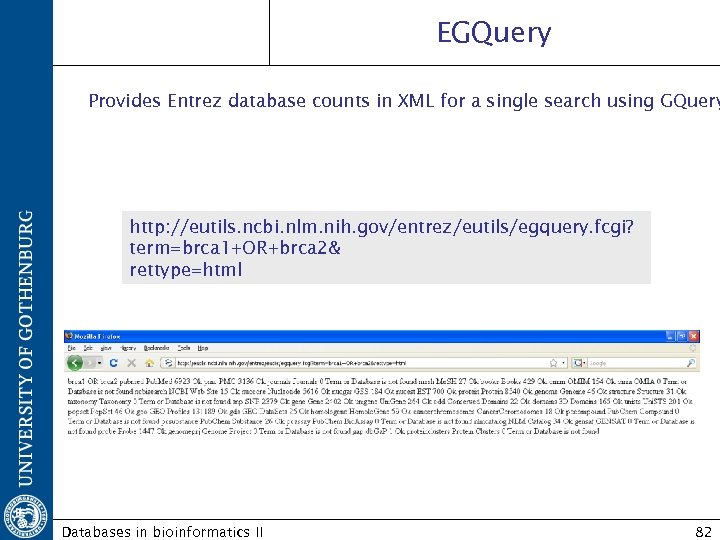

EGQuery Provides Entrez database counts in XML for a single search using GQuery http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/egquery. fcgi? term=brca 1+OR+brca 2& rettype=html Databases in bioinformatics II 82
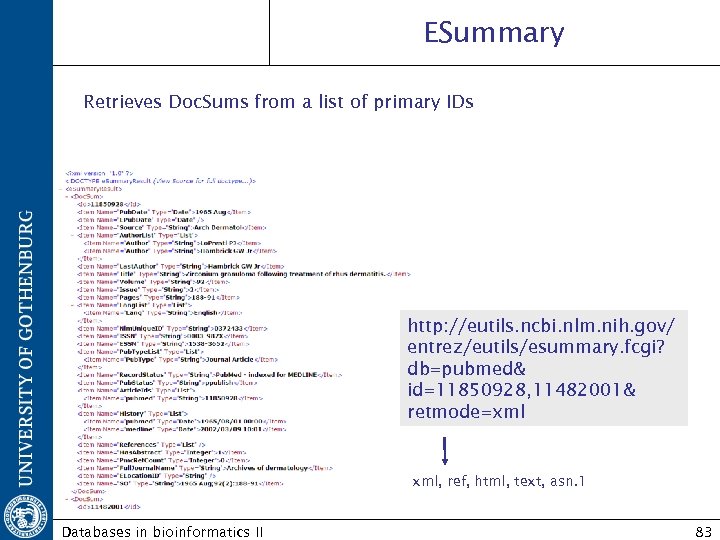

ESummary Retrieves Doc. Sums from a list of primary IDs http: //eutils. ncbi. nlm. nih. gov/ entrez/eutils/esummary. fcgi? db=pubmed& id=11850928, 11482001& retmode=xml xml, ref, html, text, asn. 1 Databases in bioinformatics II 83

ELink Existence of an external/Related Articles link from a list of UIDs Retrieves related IDs to a list of UIDs (same db, external db) http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/e link. fcgi? dbfrom=nuccore& db=protein& id=7140346 Databases in bioinformatics II 84
ELink Creates a hyperlink to the primary Link. Out provider for a specific ID Lists Link. Out URLs and attributes for multiple IDs. http: //eutils. ncbi. nlm. nih. gov/ entrez/eutils/elink. fcgi? dbfrom=pubmed& id=10611131& retmode=ref&cmd=prlinks Databases in bioinformatics II 85

ESearch Returns a list of matching UIDs (text search) in a given Entrez database http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/e search. fcgi? db=pubmed &term=cancer &reldate=60 &datetype=edat, mdat, pdat &retmax=10 Databases in bioinformatics II 86

EFetch Generates formatted output for a list of input IDs: abstracts from Pub. Med FASTA format from Protein http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/efetch. fcgi? DBs: Literature Database Pub. Med, Journals, Pub. Med Central, OMIM Sequence and other Molecular Biology Databases Nucleotide, Protein, Gene, etc. Taxonomy Databases in bioinformatics II 87

EFetch - Literature http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/efetch. fcgi? db=pubmed&i d=12091962, 9997&retmode=html&rettype=abstract Databases in bioinformatics II 88
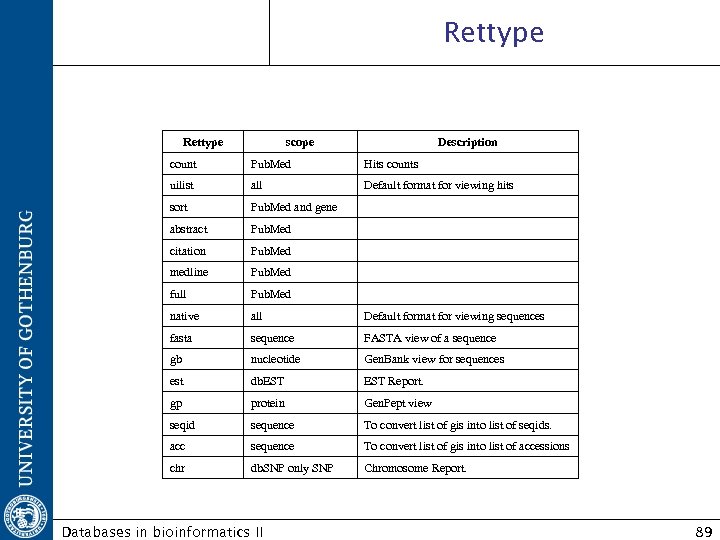
Rettype scope Description count Pub. Med Hits counts uilist all Default format for viewing hits sort Pub. Med and gene abstract Pub. Med citation Pub. Med medline Pub. Med full Pub. Med native all Default format for viewing sequences fasta sequence FASTA view of a sequence gb nucleotide Gen. Bank view for sequences est db. EST Report. gp protein Gen. Pept view seqid sequence To convert list of gis into list of seqids. acc sequence To convert list of gis into list of accessions chr db. SNP only SNP Chromosome Report. Databases in bioinformatics II 89

EFetch - Sequences http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/efetch. fcgi? db=nucleotide& id=5&rettype=fasta strand 1(+), 2(-) Databases in bioinformatics II 90

Efetch - Taxonomy http: //www. ncbi. nlm. nih. gov/entrez/eutils/efetch. fcgi? d b=taxonomy&id=44689&report=docsum uilist, brief, docsum, xml 1818 Databases in bioinformatics II 91

Excercise Search in Journals for the term obstetrics: http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/esearch. fcgi? db=journals&term=obstetrics In Pub. Med display PMIDs 12091962 and 9997 in html retrieval mode and abstract retrieval type: http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/efetch. fcgi? db=pubmed&id=12091962, 9997&retmode=html&rettype=abstract From Entrez Gene display as xml the Genome. ID 2: http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/efetch. fcgi? db=gene&id=2&retmode=xml To retrieve Pub. Med related articles for proteins 61742829 with a publication date from 1995 to the present: http: //eutils. ncbi. nlm. nih. gov/entrez/eutils/elink. fcgi? dbfrom=protein&id=61742829&db=pubmed& mindate=1995&datetype=pdat Databases in bioinformatics II 92

Combining e. Utils calls The e. Utils are useful when used by themselves in single URLs; however their full potential is reached when successive e. Utils URLs are combined to create a data pipeline • Retrieving data records matching an Entrez query ESearch → ESummary ESearch → EFetch • Finding IDs linked to records matching an Entrez query ESearch → ELink • Retrieving data records in database B linked to records in database A matching an Entrez query ESearch → ELink → ESummary ESearch → ELink → EFetch Databases in bioinformatics II 93
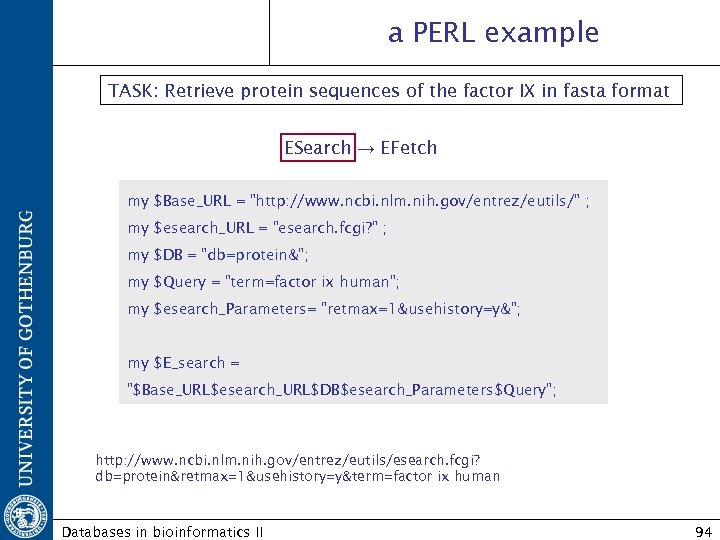

a PERL example TASK: Retrieve protein sequences of the factor IX in fasta format ESearch → EFetch my $Base_URL = "http: //www. ncbi. nlm. nih. gov/entrez/eutils/" ; my $esearch_URL = "esearch. fcgi? " ; my $DB = "db=protein&"; my $Query = "term=factor ix human"; my $esearch_Parameters= "retmax=1&usehistory=y&"; my $E_search = "$Base_URL$esearch_URL$DB$esearch_Parameters$Query"; http: //www. ncbi. nlm. nih. gov/entrez/eutils/esearch. fcgi? db=protein&retmax=1&usehistory=y&term=factor ix human Databases in bioinformatics II 94

Output from ESearch Databases in bioinformatics II 95

Query. Key - Web. Env $Query. Key: value used for a history search number (label) $Web. Env: cookie value used with EFetch in place of primary ID result list (encoded server address) corresponds to a UID list for subsequent search strategies Databases in bioinformatics II 96
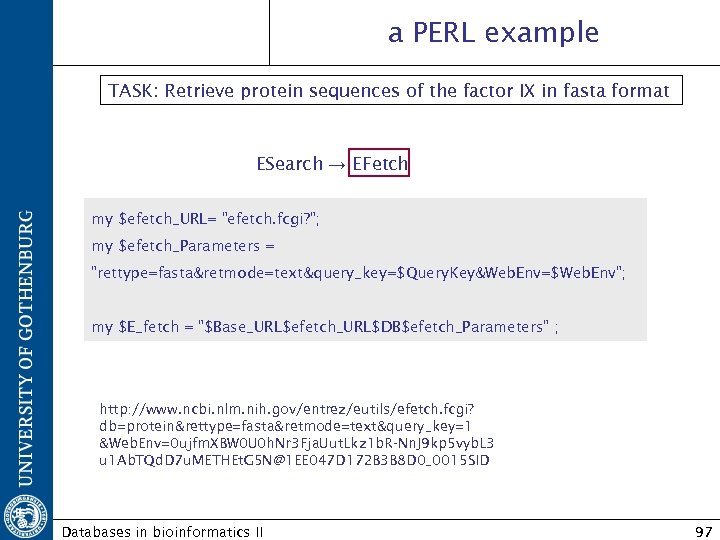

a PERL example TASK: Retrieve protein sequences of the factor IX in fasta format ESearch → EFetch my $efetch_URL= "efetch. fcgi? "; my $efetch_Parameters = "rettype=fasta&retmode=text&query_key=$Query. Key&Web. Env=$Web. Env"; my $E_fetch = "$Base_URL$efetch_URL$DB$efetch_Parameters" ; http: //www. ncbi. nlm. nih. gov/entrez/eutils/efetch. fcgi? db=protein&rettype=fasta&retmode=text&query_key=1 &Web. Env=0 ujfm. XBW 0 U 0 h. Nr 3 Fja. Uut. Lkz 1 b. R-Nn. J 9 kp 5 vyb. L 3 u 1 Ab. TQd. D 7 u. METHEt. G 5 N@1 EE 047 D 172 B 3 B 8 D 0_0015 SID Databases in bioinformatics II 97

Output from EFetch Databases in bioinformatics II 98


So. . . – Genome sequencing – Sequencing methods • • Sanger, Maxam Next generation methods (2 nd, 3 rd) Uses Implications – Ref. Seq vs Gen. Bank – Trace. Archive – Refining searches at Entrez – e. Utilis (programer utilities) Databases in bioinformatics II 100
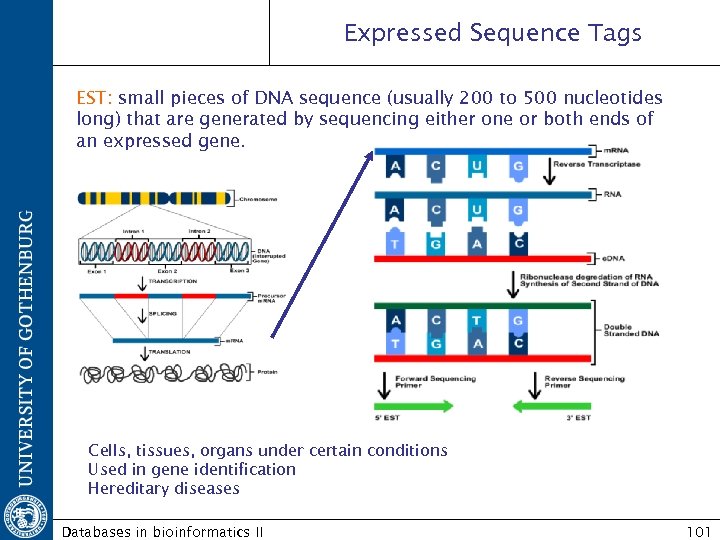
Expressed Sequence Tags EST: small pieces of DNA sequence (usually 200 to 500 nucleotides long) that are generated by sequencing either one or both ends of an expressed gene. Cells, tissues, organs under certain conditions Used in gene identification Hereditary diseases Databases in bioinformatics II 101
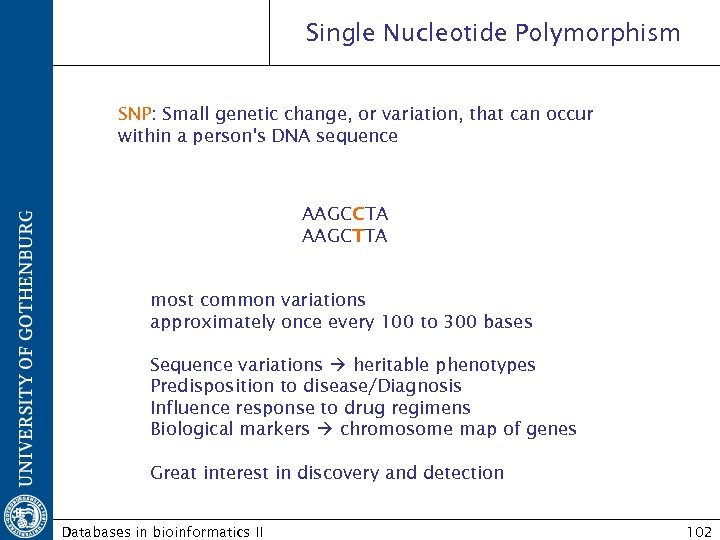

Single Nucleotide Polymorphism SNP: Small genetic change, or variation, that can occur within a person's DNA sequence AAGCCTA AAGCTTA most common variations approximately once every 100 to 300 bases Sequence variations heritable phenotypes Predisposition to disease/Diagnosis Influence response to drug regimens Biological markers chromosome map of genes Great interest in discovery and detection Databases in bioinformatics II 102
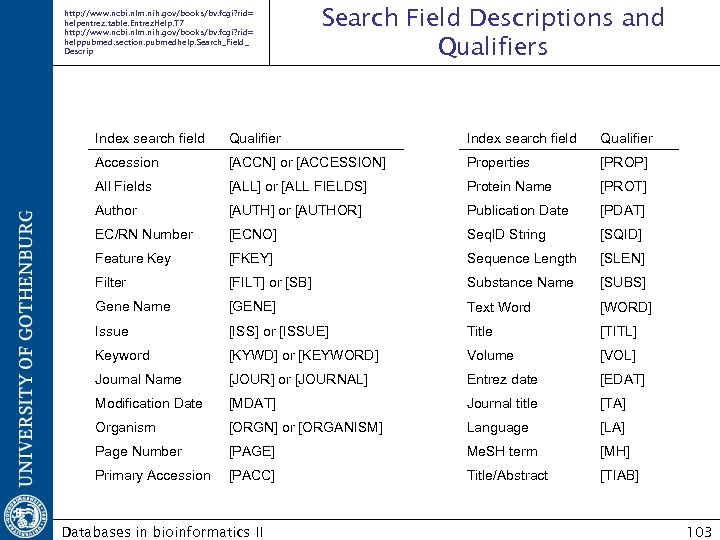
http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid= helpentrez. table. Entrez. Help. T 7 http: //www. ncbi. nlm. nih. gov/books/bv. fcgi? rid= helppubmed. section. pubmedhelp. Search_Field_ Descrip Search Field Descriptions and Qualifiers Index search field Qualifier Accession [ACCN] or [ACCESSION] Properties [PROP] All Fields [ALL] or [ALL FIELDS] Protein Name [PROT] Author [AUTH] or [AUTHOR] Publication Date [PDAT] EC/RN Number [ECNO] Seq. ID String [SQID] Feature Key [FKEY] Sequence Length [SLEN] Filter [FILT] or [SB] Substance Name [SUBS] Gene Name [GENE] Text Word [WORD] Issue [ISS] or [ISSUE] Title [TITL] Keyword [KYWD] or [KEYWORD] Volume [VOL] Journal Name [JOUR] or [JOURNAL] Entrez date [EDAT] Modification Date [MDAT] Journal title [TA] Organism [ORGN] or [ORGANISM] Language [LA] Page Number [PAGE] Me. SH term [MH] Primary Accession [PACC] Title/Abstract [TIAB] Databases in bioinformatics II 103
2986ccd525af0915cd4ecaf8123f48be.ppt