00a4f7f49fda10d031861a95961d3548.ppt
- Количество слайдов: 33

Biochem 503 December 3, 2008 Protein Ser/Thr Phosphatases (PPP family) Controlling 99% of cellular phosphorylation I. III. IV. V. History Type-1 Protein Phosphatase (PP 1) Type-2 A Protein phosphatase (PP 2 A) Calcineurin (PP 2 B) Ca 2+/Ca. M dependent PP 2 C or Mg 2+-dependent PPase (MPP)

History of Ser/Thr Protein Phosphatases 1950’s 1. PR Enzyme Gerty Cori glycogen phosphorylase a to b conversion (phosphorylase phosphatase) 1960’s a lost decade? Ed Fischer students struggle S. Tamura Tohoku Univ. Sendai, Japan 1970’s 2. Glycogen synthase PPase - Tsuiki, Tamura, Kikuchi C’ville to Sendai connection… 3 different enzymes separated later known as PP 1, PP 2 A and PP 2 C (MPP) 3. The Et. OH ppt method a grad student mistake becomes “standard” method - 35 k. Da PP-ase (C subunit) E. Y. C. Lee U. Miami, now NYMC 4. Isolation of inhibitor proteins (1976). . the C + I = CI idea of regulators 5. Same phosphatase for every substrate? ……. histone, casein, myosin, tropomyosin, spectrin, ribosomes, enzymes…… “broad specificity” (i. e. non-specific) 35 k. Da phosphatase

History of Ser/Thr Protein Phosphatases 1980’s 6. Separation of two phosphatases in 35 k. Da fraction (Ingebritsen) Classification (1983) Type 1 vs. type 2 A, B, C based on sensitivity to inhibitor proteins 7. Targeting hypothesis (Cohen, 1985) - glycogen-bound PP 1 a dimer of [ PP 1 C plus glycogen-subunit] R subunits modulate activity and anchor C 8. PP 1 binds to multiple different R subunits: GM and MYPT 1 myosin phosphatase PP 1 C plus MYPT 1 9. PP 2 A binds 60 and 55 k. Da subunits to form ABC trimer common AC, different Bs (B 55, B 56, B 72) NIH + Sendai + Hiroshima Sir Phillip Cohen Dundee, Scotland

History Marc Mumby UTSW Edgar de Cruz e Silva Dundee, NYC, Portugal 10. Cloning of C subunits of PP 2 A and PP 1 unrelated to PTPs, but similar to purple acid PP-ase 11. Yeast-fungal genetics show functional conservation GLC 7=PP 1 12. Discovery of Okadaic Acid as selective PP 1 -PP 2 A inhibitor 1990’s 13. Cloning of multiple PPP catalytic subunits PP 4, PP 5, PP 6 Family Conservation & Diversity in Catalytic subunits (see PTW Cohen TIBS 22: 245 -251) 14. Many PP 2 A B subunits in Families- B, B’, B” 15. Genomics - over 40 human genes, 24 in yeast David Virshup Univ. Utah, now Singapore NUS-Duke Med Sch. B Tricia (PTW) Cohen (Lady Cohen) Dundee, Scotland Anna De. Paoli-Roach Indiana U Med. Sch.
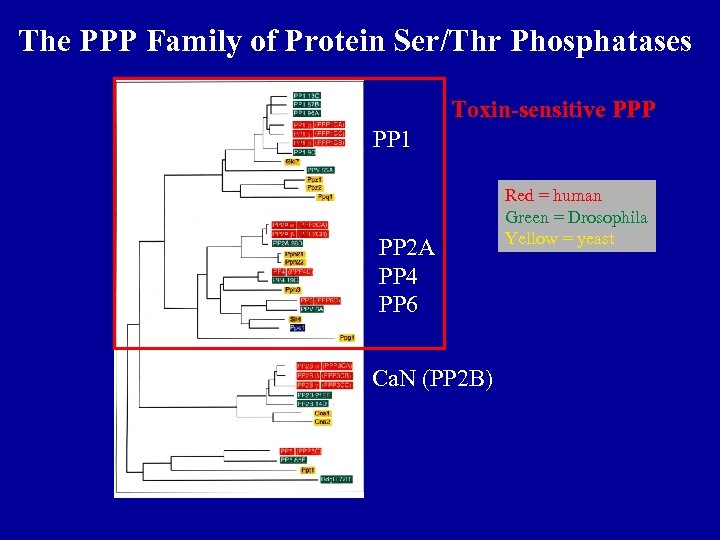
The PPP Family of Protein Ser/Thr Phosphatases Toxin-sensitive PPP PP 1 PP 2 A PP 4 PP 6 Ca. N (PP 2 B) Red = human Green = Drosophila Yellow = yeast
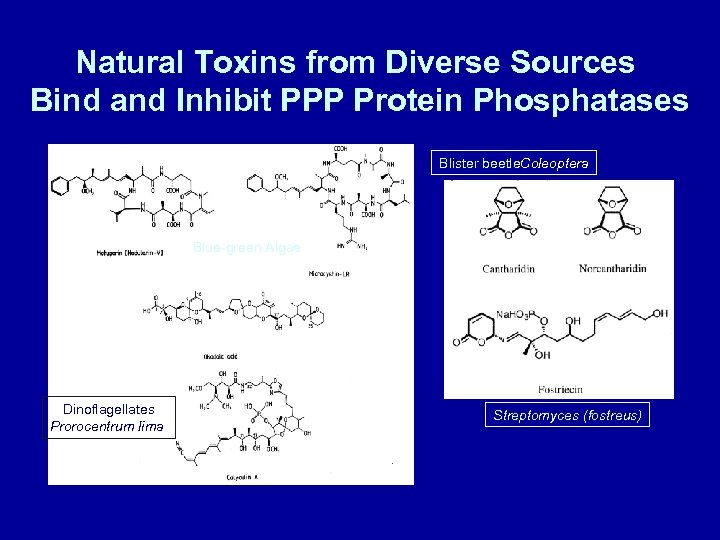
Natural Toxins from Diverse Sources Bind and Inhibit PPP Protein Phosphatases Blister beetle. Coleoptera Blue-green Algae Dinoflagellates Prorocentrum lima Streptomyces (fostreus)
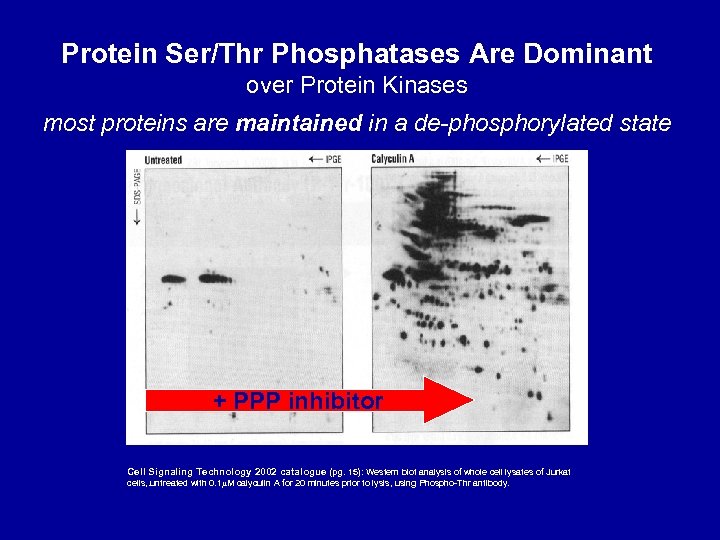
Protein Ser/Thr Phosphatases Are Dominant over Protein Kinases most proteins are maintained in a de-phosphorylated state + PPP inhibitor Cell Signaling Technology 2002 catalogue (pg. 15): Western blot analysis of whole cell lysates of Jurkat cells, . untreated with 0. 1 M calyculin A for 20 minutes prior to lysis, using Phospho-Thr antibody.
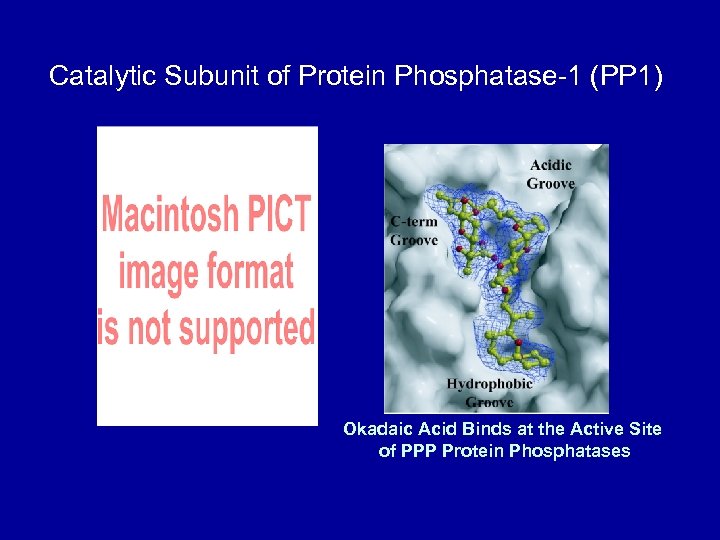

Catalytic Subunit of Protein Phosphatase-1 (PP 1) Okadaic Acid Binds at the Active Site of PPP Protein Phosphatases
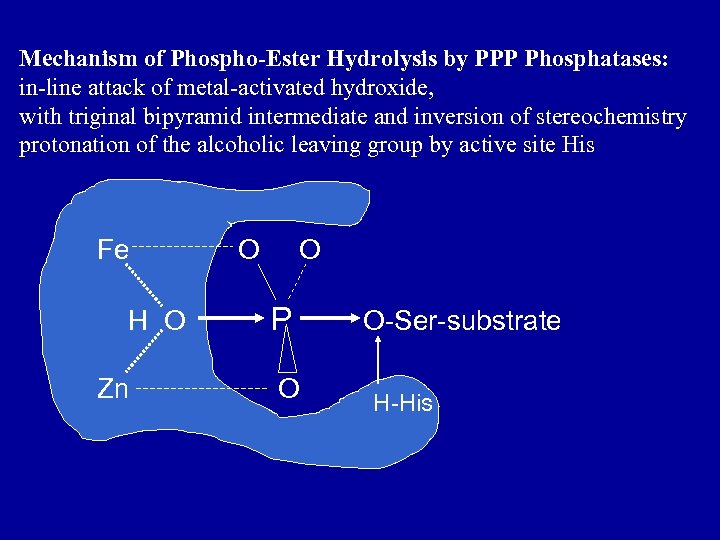
Mechanism of Phospho-Ester Hydrolysis by PPP Phosphatases: in-line attack of metal-activated hydroxide, with triginal bipyramid intermediate and inversion of stereochemistry protonation of the alcoholic leaving group by active site His Fe H O Zn O O P O O-Ser-substrate H-His
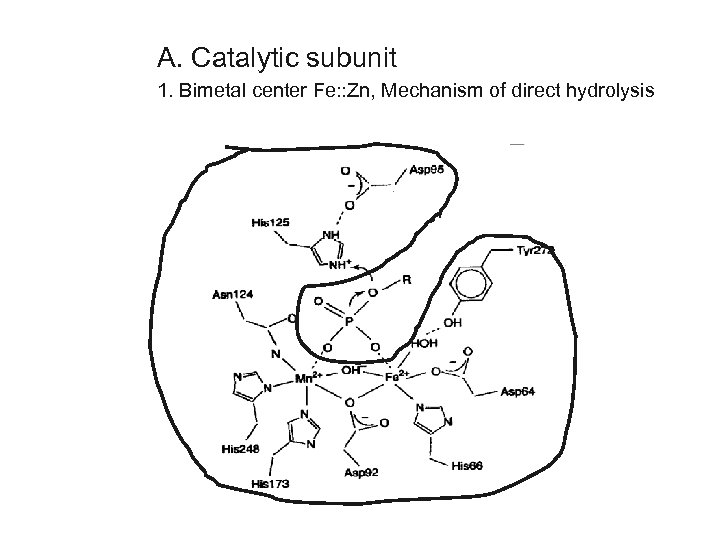
A. Catalytic subunit 1. Bimetal center Fe: : Zn, Mechanism of direct hydrolysis

Type-1 Protein Phosphatase (PP 1) 1. Bi-metallic active site with Fe and Zn 2. 3 D structure - beta sheet and alpha helix clusters 3. Isoforms ifferences mostly in C terminal, allow specific antibodies d alpha gamma delta NPGGRPITPPRN--SAKAKK --ATRPVTPPRGMITKQAKK NSG-RPVTPPRTANPPK-KR 4. Phosphorylation in RPITPPR motif first found in yeast sds 22 cyclin-dep kinases (CDK) phosphorylate to inactivate; reversed by auto-dephosphorylation 5. Toxins - microcystin, okadaic acid, calyculin A bind at active site (3 D structures)

B. Regulatory/targeting subunits for PP 1 C 1. The pioneer…. . glycogen-binding GM 2. The RVx. F Motif as primary recognition site 3. Myosin phosphatase MYPT 1 as regulator & scaffold 4. Dozens more, >200 total…. . phospho regulation C. Inhibitor phospho-proteins 1. Inh 1 and DARPP-32 2. Inh 2, Inh 4 3. CPI-17 4. Linking kinases to PP 1 inhibitioninhibitors target selective R-C complexes D. Substrates 1. abundant phosphatase, handles large capacity, abundant substrates… myosin, lamins, glycogen enzymes, histones
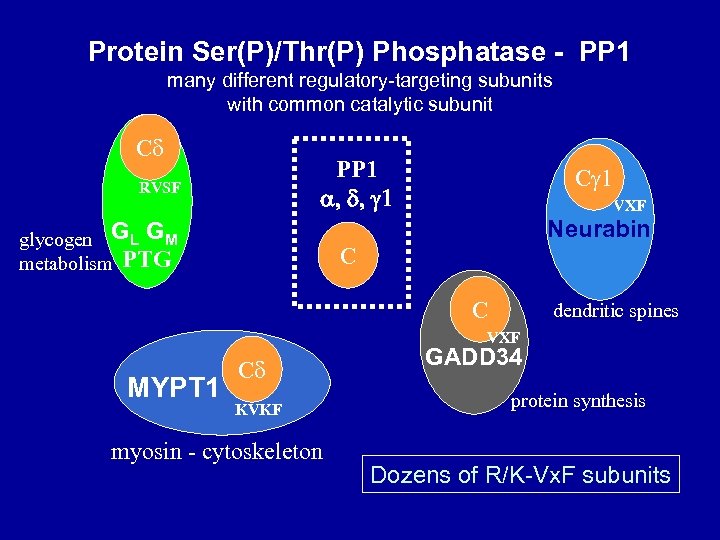
Protein Ser(P)/Thr(P) Phosphatase - PP 1 many different regulatory-targeting subunits with common catalytic subunit C PP 1 a, d, g 1 RVSF C VXF Neurabin glycogen GL GM metabolism PTG C C dendritic spines VXF MYPT 1 C KVKF myosin - cytoskeleton GADD 34 protein synthesis Dozens of R/K-Vx. F subunits
Myosin Phosphatase inhibition by phosphorylation-dependent inhibitor protein CPI-17. PKC P MYPT 1 CPI-17 Myosin Phosphatase: PP 1 -MYPT 1 First example of an inhibitor specific for a PP 1 Holoenzyme
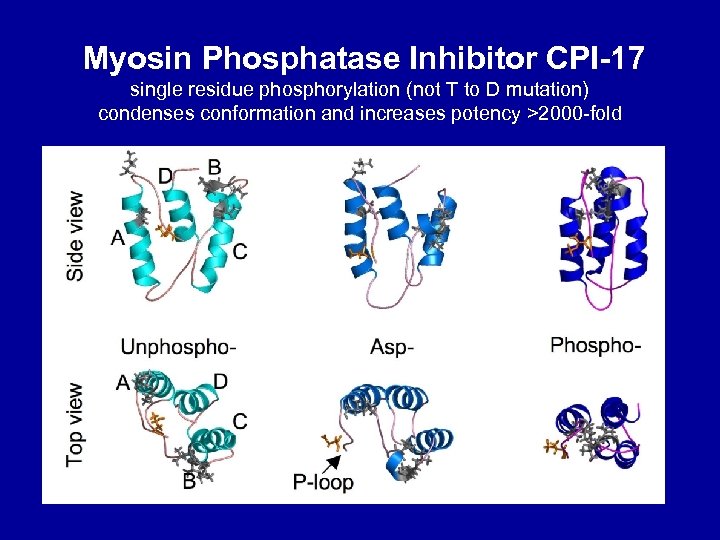
Myosin Phosphatase Inhibitor CPI-17 single residue phosphorylation (not T to D mutation) condenses conformation and increases potency >2000 -fold
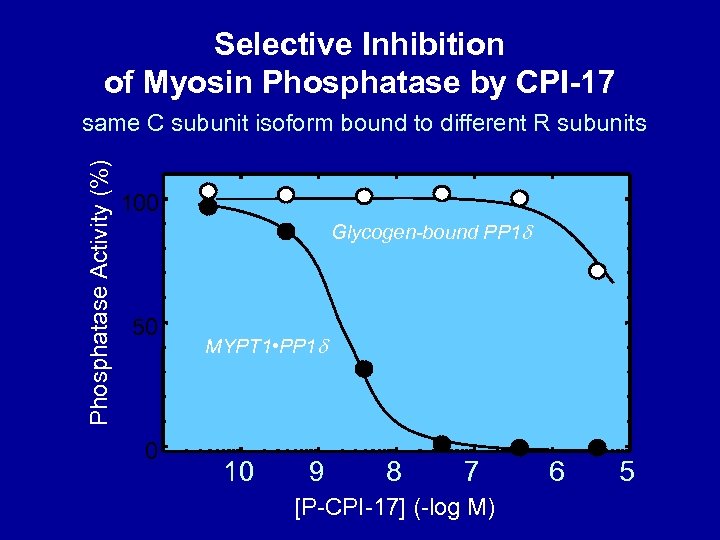
Selective Inhibition of Myosin Phosphatase by CPI-17 Phosphatase Activity (%) same C subunit isoform bound to different R subunits 100 Glycogen-bound PP 1 d 50 0 MYPT 1 • PP 1 d 10 9 8 7 [P-CPI-17] (-log M) 6 5

Myosin LC Phosphatase: A complex of PP 1 C with MYPT 1 Subunit C subunit bound to specific R subunit

Type-2 A Protein phosphatase (PP 2 A) A. Catalytic subunit 1. Bimetal center Fe: : Zn and catalytic Mechanism same as PP 1 2. 3 D structure…known in complex with A and in ABC 3. Isoforms 10: 1 ratio, essential for development DYFLCOOH motif at C terminus conserved phosphorylation - PTKs, eg. Src, JAK methylation - PMT and PME, alters subunit association 5. Toxins - MCLR, OA bind at active site. Differences between PP 1 and PP 2 A in 12 - 13 loop
![Protein Ser(P)/Thr(P) Phosphatase - PP 2 A ABC = { [A] Scaffold + [C] Protein Ser(P)/Thr(P) Phosphatase - PP 2 A ABC = { [A] Scaffold + [C]](https://present5.com/presentation/00a4f7f49fda10d031861a95961d3548/image-19.jpg)
Protein Ser(P)/Thr(P) Phosphatase - PP 2 A ABC = { [A] Scaffold + [C] Catalytic} + [B] Regulatory ABC B A Tpd 3 HEAT Helical repeats A subunit C Pph 21 Pph 22 Binding of C to A decreases Vmax and alters KM
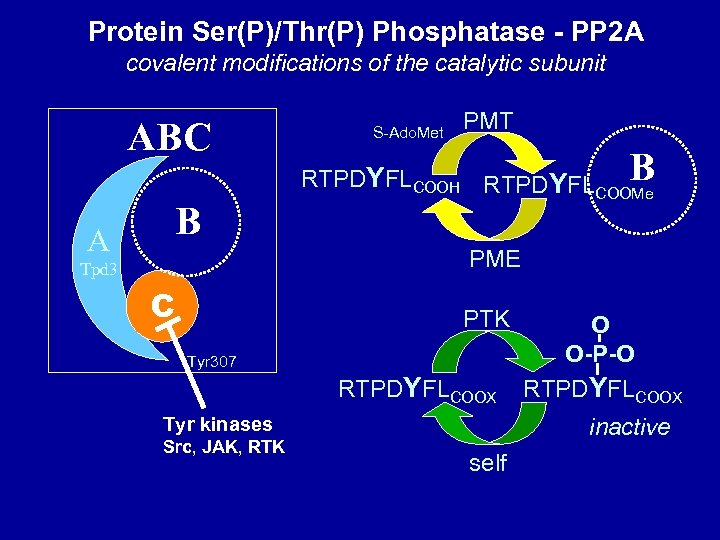
Protein Ser(P)/Thr(P) Phosphatase - PP 2 A covalent modifications of the catalytic subunit ABC B A S-Ado. Met PMT B RTPDYFLCOOH RTPDYFL COOMe PME Tpd 3 C PTK Tyr 307 RTPDYFLCOOX Tyr kinases Src, JAK, RTK O O-P-O RTPDYFLCOOX inactive self

B. Scaffold and Regulatory subunitsthe ABC’s of PP 2 A. 1. The Scaffold "A" or PR 65 (HEAT repeats) the AC dimer A and A 85: 15, but A specific tumor suppressor 2. Many Bs…. are they for dedicated functions? Yeast Cdc 55 and Rts 1 - distinct phenotypes, not complemented (conclusion? ) now 3 B families cloned - more than 15 genes B B 55 B’ B 56 B’’ B 72(130) 3. Tumor antigens, s. T, m. T, E 4 orf replace B subunits to re-direct activity. 4. More and more Bs? , p 107, p 48, etc. proposed as alternates
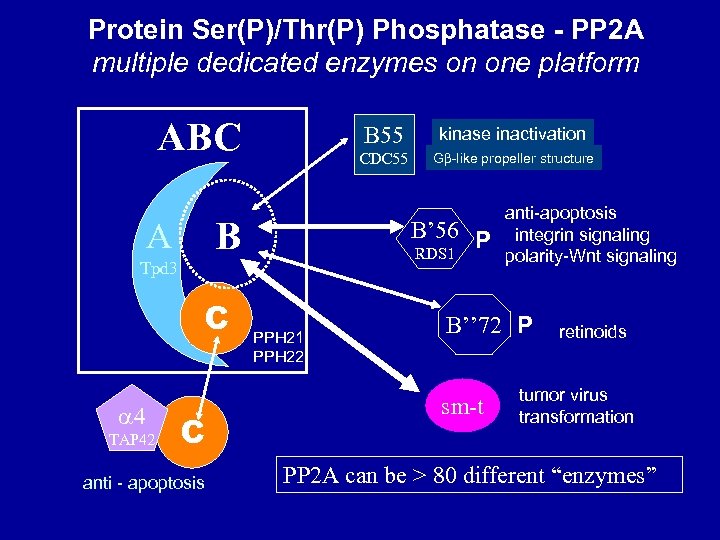
Protein Ser(P)/Thr(P) Phosphatase - PP 2 A multiple dedicated enzymes on one platform ABC B 55 CDC 55 B A 4 TAP 42 C anti - apoptosis G -like propeller structure B’ 56 P RDS 1 Tpd 3 C kinase inactivation PPH 21 PPH 22 anti-apoptosis integrin signaling polarity-Wnt signaling B’’ 72 P sm-t retinoids tumor virus transformation PP 2 A can be > 80 different “enzymes”
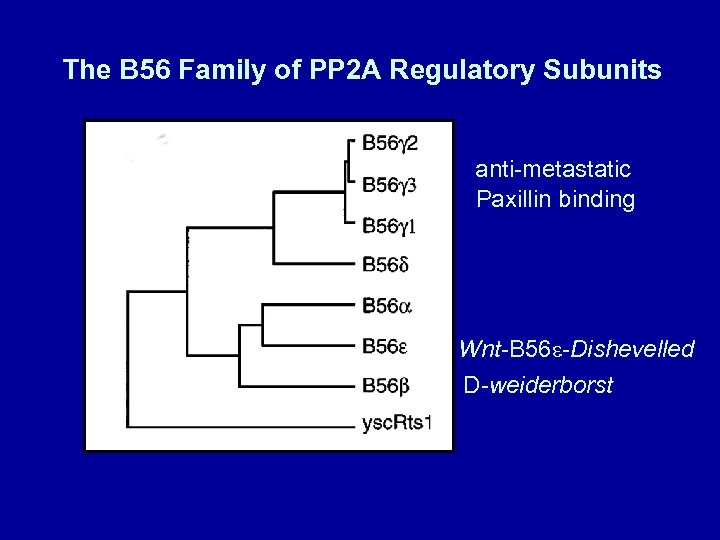
The B 56 Family of PP 2 A Regulatory Subunits anti-metastatic Paxillin binding Wnt-B 56 -Dishevelled D-weiderborst
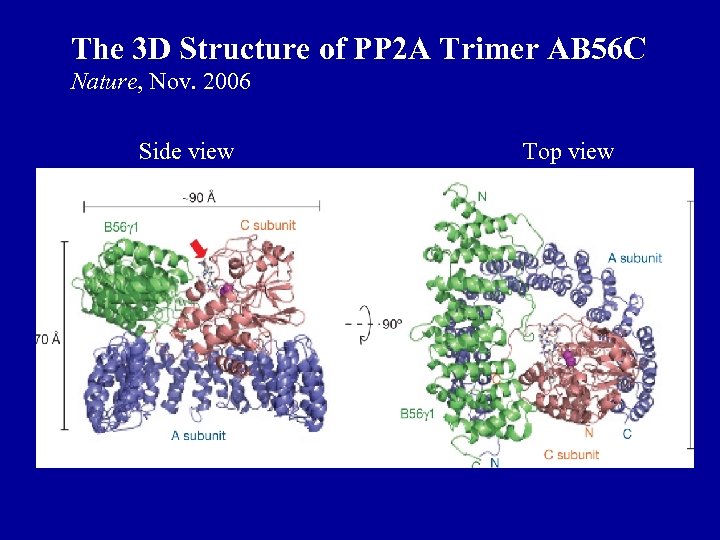
The 3 D Structure of PP 2 A Trimer AB 56 C Nature, Nov. 2006 Side view Top view

C. Inhibitor proteins for PP 2 A 1. Inh 1(PP 2 A) Damuni et al 2. Inh 2(PP 2 A) SET protein - important in solid tumors ceramide “receptor” for PP 2 A activation D. Alternative partners 1. A binds HSF 2 and PP 5 2. C binds alpha 4; in yeast part of TOR signaling E. Substrates 1. Signaling Kinases (MEK, PKB, PI 3 K, p 70 S 6 K, etc. ) 2. anti-apoptosis (PKB, Bad, Bcl) 3. integrin signaling - Paxillin B 56 deletions & increased invasion

Calcineurin (PP 2 B)(PPP 3) Ca 2+/Ca. M activated Phosphatase A. Catalytic subunit 1. C has Fe: : Zn active site like PP 1, PP 2 A 2. Suppressor domain, Ca. M dependent B. Regulatory subunits 1. B subunit (Ca 2+) 2. additional Ca. M 3. FKBP + FK-506 or cyclosporin ($B product) C. Inhibitor Proteins 1. induced in response to signaling have reported activity as activator-inhibitor. D. Substrates 1. NFAT (co-transport into nucleus) 2. Elk-1 (TCF transcription factor) 3. DARPP-32 (NMDA receptor)
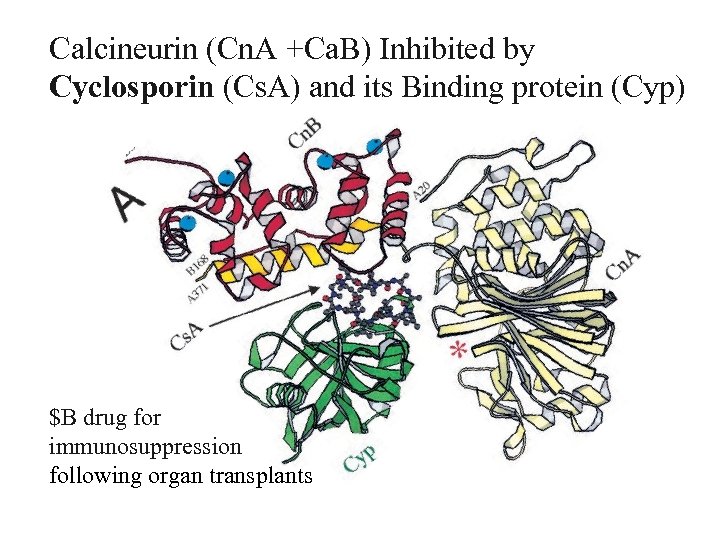
Calcineurin (Cn. A +Ca. B) Inhibited by Cyclosporin (Cs. A) and its Binding protein (Cyp) $B drug for immunosuppression following organ transplants

PP 2 C = Mg 2+-dependent PPase (MPP) A. Catalytic subunit 1. unrelated to PPP but bimetallic Mg: Mg active center 2. isoforms , etc. 3. many new family members in genome B. Regulatory subunits - none? C. Inhibitor Proteins - none? D. Substrates 1. CDKs 2. the kinase activation loop 3. PI 3 K 4. Glycogen synthase Maybe activated by small molecule second messenger? Lipids?
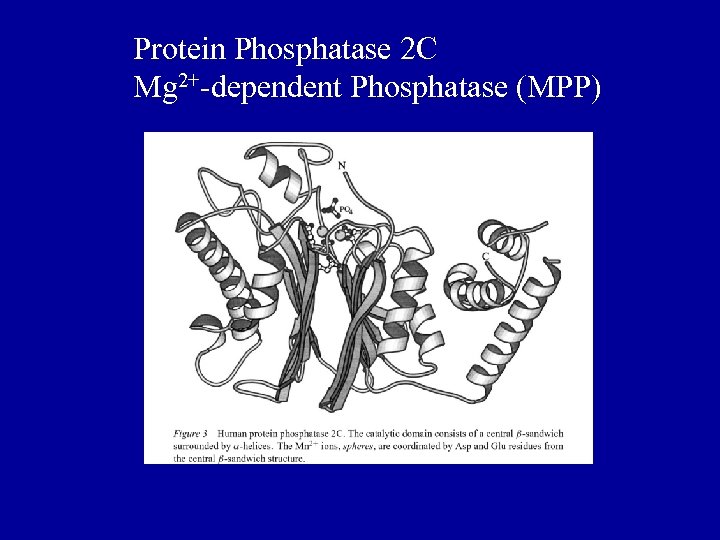

Protein Phosphatase 2 C Mg 2+-dependent Phosphatase (MPP)
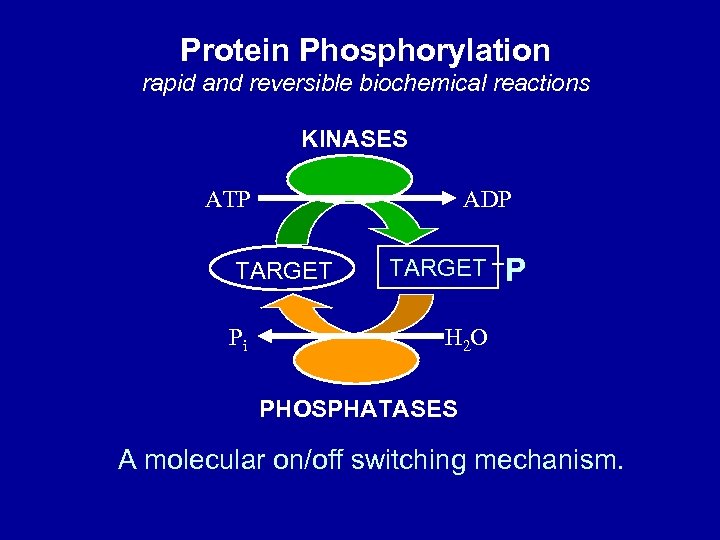
Protein Phosphorylation rapid and reversible biochemical reactions KINASES ATP ADP TARGET Pi TARGET P H 2 O PHOSPHATASES A molecular on/off switching mechanism.
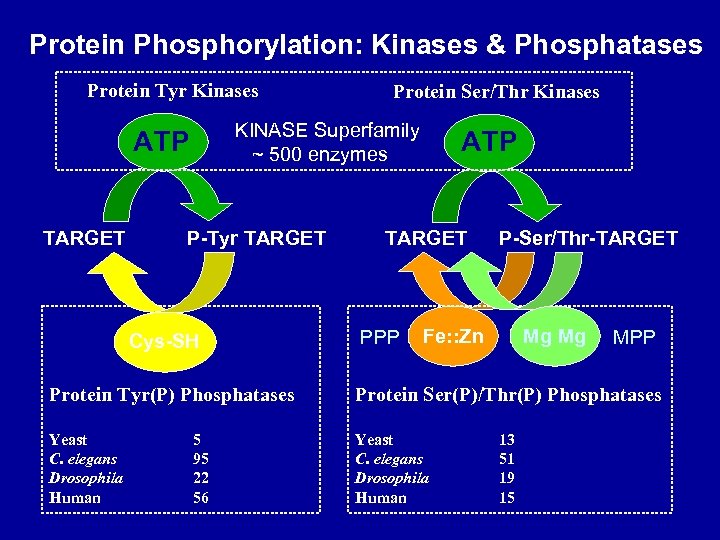
Protein Phosphorylation: Kinases & Phosphatases Protein Tyr Kinases KINASE Superfamily ~ 500 enzymes ATP TARGET Protein Ser/Thr Kinases P-Tyr TARGET Cys-SH ATP TARGET PPP P-Ser/Thr-TARGET Fe: : Zn Mg Mg MPP Protein Tyr(P) Phosphatases Protein Ser(P)/Thr(P) Phosphatases Yeast C. elegans Drosophila Human 5 95 22 56 13 51 19 15
Protein Phosphorylation: Kinases & Phosphatases ATP ATP ATP KINASE Superfamily ~ 500 enzymes ATP ATP ATP Cys-SH Fe: : Zn Cys-SH Phosphatase families and ensembles ~ 500 enzymes Fe: : Zn Mg Mg MPP

Protein Kinases and Protein Phosphatases ancient enzymes essential to cell signaling and cellular regulation New targets for Pharmaceuticals
00a4f7f49fda10d031861a95961d3548.ppt