8d0e2dd000aaa4c972c07168839a1a58.ppt
- Количество слайдов: 23

Benefits from Cooperation in Genomics Paul Van. Raden, George Wiggans, Animal Improvement Programs Laboratory Curt Van Tassell, Tad Sonstegard, Bovine Functional Genomics Laboratory USDA Agricultural Research Service, Beltsville, MD, USA Flavio Shenkel CGIL, University of Guelph, ON, Canada Paul. Van. Raden@ars. usda. gov 2009 2007

Topics Ø Genomic cooperation • • Ø Simulation of very large population Proposals for genotype sharing Country border issues and North American experience Genomic MACE equations USA update • • Actual HOL, JER, and BSW results Database and implementation Interbull Genomics Workshop, Jan. 2009 (2) Paul Van. Raden 2009

Cooperative International Projects Ø Traditional genetic evaluations • • • Ø Ø MACE instead of merging phenotypes Small benefits expected from data merger Proven bulls only, not cows or young bulls Parentage testing, genetic recessives, pedigrees done by breed associations Genomics: what role for Interbull? Interbull Genomics Workshop, Jan. 2009 (3) Paul Van. Raden 2009
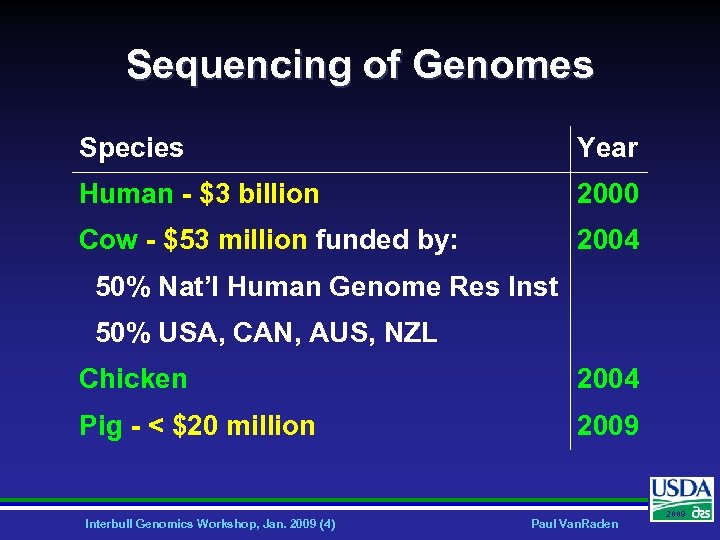

Sequencing of Genomes Species Year Human - $3 billion 2000 Cow - $53 million funded by: 2004 50% Nat’l Human Genome Res Inst 50% USA, CAN, AUS, NZL Chicken 2004 Pig - < $20 million 2009 Interbull Genomics Workshop, Jan. 2009 (4) Paul Van. Raden 2009

Human DNA Data Sharing "The highest priority of the International Human Genome Sequencing Consortium is ensuring that sequencing data from the human genome is available to the world's scientists rapidly, freely and without restriction. " National Human Genome Research Institute, 2008 "The principle of rapid pre-publication release should apply to other types of data from other large-scale production centers. " Wellcome Trust, 2003 Interbull Genomics Workshop, Jan. 2009 (5) Paul Van. Raden 2009
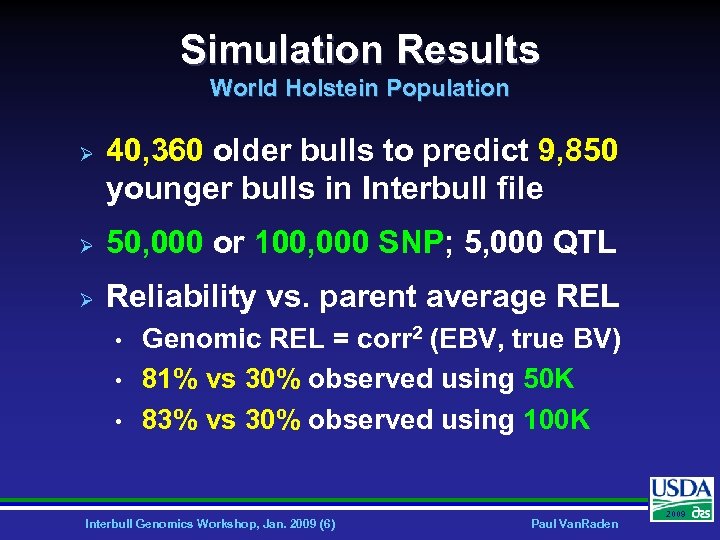

Simulation Results World Holstein Population Ø 40, 360 older bulls to predict 9, 850 younger bulls in Interbull file Ø 50, 000 or 100, 000 SNP; 5, 000 QTL Ø Reliability vs. parent average REL • • • Genomic REL = corr 2 (EBV, true BV) 81% vs 30% observed using 50 K 83% vs 30% observed using 100 K Interbull Genomics Workshop, Jan. 2009 (6) Paul Van. Raden 2009

Genotype Exchange Options Ø Ø Give away for free (not likely) Genotype own bulls, then trade? • • • Ø Ø Trade an equal number or all bulls? Country A has 5000 and B has 1000 Proportional to population size? Trade among organization pairs or create central genomic database? Service fee for young animals to pay for ancestor genotyping? Interbull Genomics Workshop, Jan. 2009 (7) Paul Van. Raden 2009

Problems of Not Sharing Ø Ø Ø Genetic progress not as fast as with full access to genotypes Severe limits on researcher access to genotypes (secrecy) Genomics may lead to natural monopoly, similar to railroads • Small companies / countries can’t afford to buy sufficient genotypes Interbull Genomics Workshop, Jan. 2009 (8) Paul Van. Raden 2009

Share Young Bull, Cow Genotypes? Ø Ø May be marketed in >1 country Exchange of young animals and females more important as their REL increases with genomics Ø Helps to synchronize databases Ø Could lead to joint evaluation Interbull Genomics Workshop, Jan. 2009 (9) Paul Van. Raden 2009

North American Cooperation Ø 174 markers, 1068 USA and CAN bulls • • Ø 367 markers, 1415 USA and CAN bulls • • Ø Illinois, Israel, and USDA researchers 1991 -1999 USDA, Illinois, and Israel 1995 -2004 38, 416 markers, 19, 464 animals • • USDA, Missouri, Canada, and Illumina Oct 2007 - Dec 2008 Interbull Genomics Workshop, Jan. 2009 (10) Paul Van. Raden 2009

Country Borders Ø Ø Most phenotypic data collected and stored within country Genomic data allows simple, accurate prediction across borders • • • Need traditional EBV or PA foreign animals, but not available for young bulls, cows, or heifers May need full foreign pedigrees Genomic evaluations official on USA scale for many foreign animals (not just CAN) Interbull Genomics Workshop, Jan. 2009 (11) Paul Van. Raden 2009
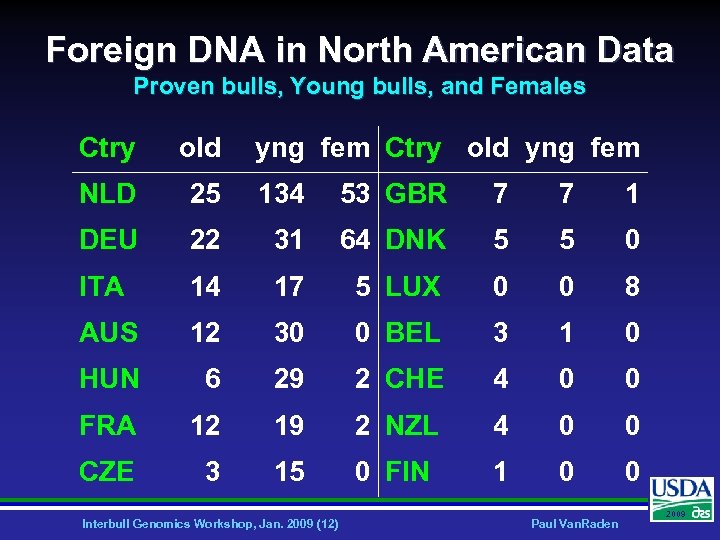
Foreign DNA in North American Data Proven bulls, Young bulls, and Females Ctry old yng fem NLD 25 134 53 GBR 7 7 1 DEU 22 31 64 DNK 5 5 0 ITA 14 17 5 LUX 0 0 8 AUS 12 30 0 BEL 3 1 0 HUN 6 29 2 CHE 4 0 0 FRA 12 19 2 NZL 4 0 0 CZE 3 15 0 FIN 1 0 0 Interbull Genomics Workshop, Jan. 2009 (12) Paul Van. Raden 2009

USA Update Ø Genomic PTAs official in January • • Traditional PTAs sent to Interbull MACE used if foreign dtrs included Genomic info used for most bulls Genomic PTA transferred to descendants (to ancestors in future) Ø Jersey results also are official Ø More Brown Swiss needed (CHE) Interbull Genomics Workshop, Jan. 2009 (13) Paul Van. Raden 2009

Genomic Methods Ø Direct genomic evaluation • • Ø Combined genomic evaluation • Ø Sum of effects for 38, 416 genetic markers Not published Include phenotypes of non-genotyped ancestors by selection index Transferred genomic evaluation • Propagate info from genotyped animals to non-genotyped relatives by selection index Interbull Genomics Workshop, Jan. 2009 (14) Paul Van. Raden 2009
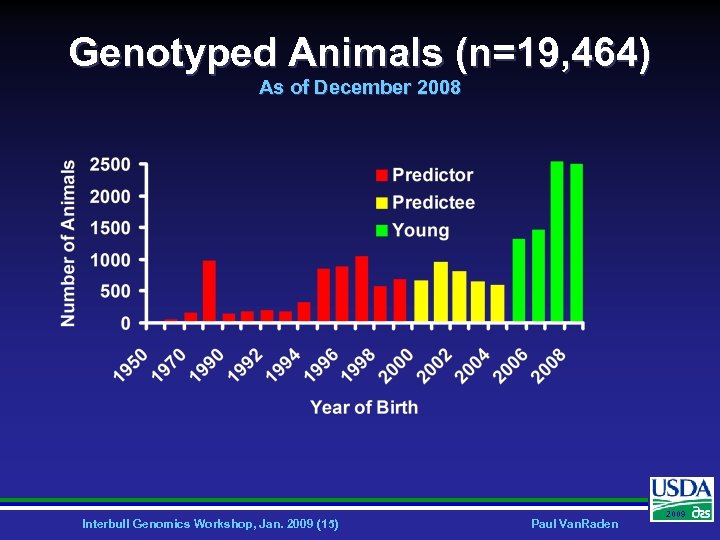
Genotyped Animals (n=19, 464) As of December 2008 Interbull Genomics Workshop, Jan. 2009 (15) Paul Van. Raden 2009
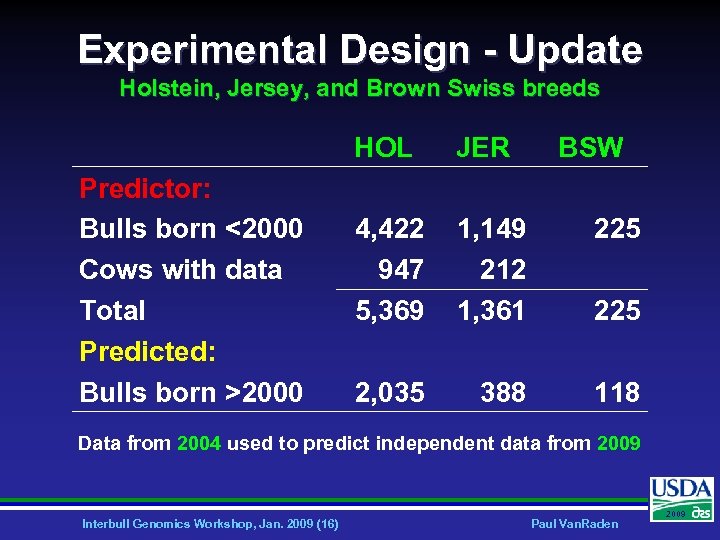
Experimental Design - Update Holstein, Jersey, and Brown Swiss breeds HOL Predictor: Bulls born <2000 Cows with data Total Predicted: Bulls born >2000 JER BSW 4, 422 947 5, 369 1, 149 212 1, 361 225 2, 035 388 118 225 Data from 2004 used to predict independent data from 2009 Interbull Genomics Workshop, Jan. 2009 (16) Paul Van. Raden 2009
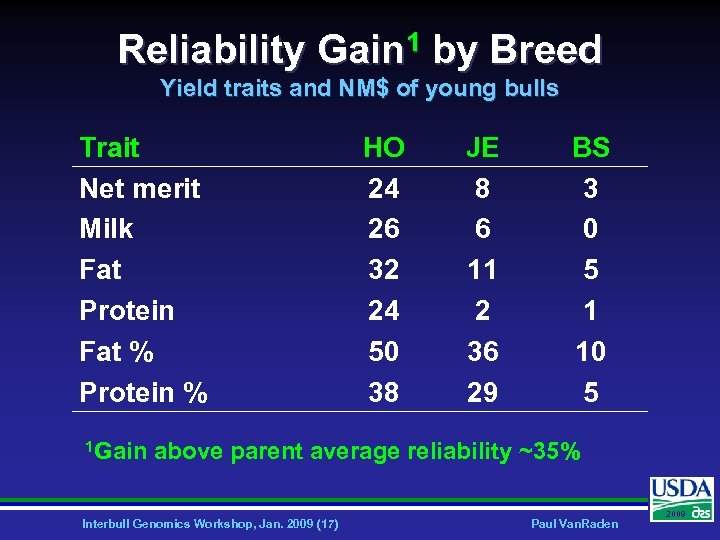
Reliability Gain 1 by Breed Yield traits and NM$ of young bulls Trait Net merit Milk Fat Protein Fat % Protein % 1 Gain HO 24 26 32 24 50 38 JE 8 6 11 2 36 29 BS 3 0 5 1 10 5 above parent average reliability ~35% Interbull Genomics Workshop, Jan. 2009 (17) Paul Van. Raden 2009
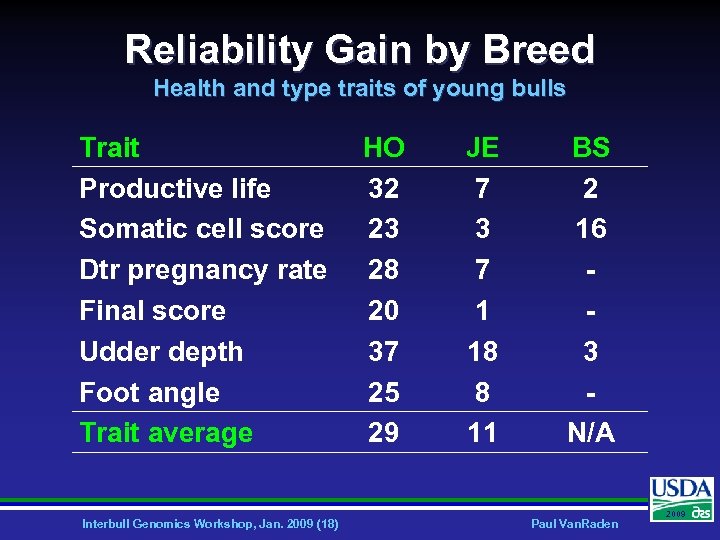

Reliability Gain by Breed Health and type traits of young bulls Trait Productive life Somatic cell score Dtr pregnancy rate Final score Udder depth Foot angle Trait average Interbull Genomics Workshop, Jan. 2009 (18) HO 32 23 28 20 37 25 29 JE 7 3 7 1 18 8 11 BS 2 16 3 N/A Paul Van. Raden 2009
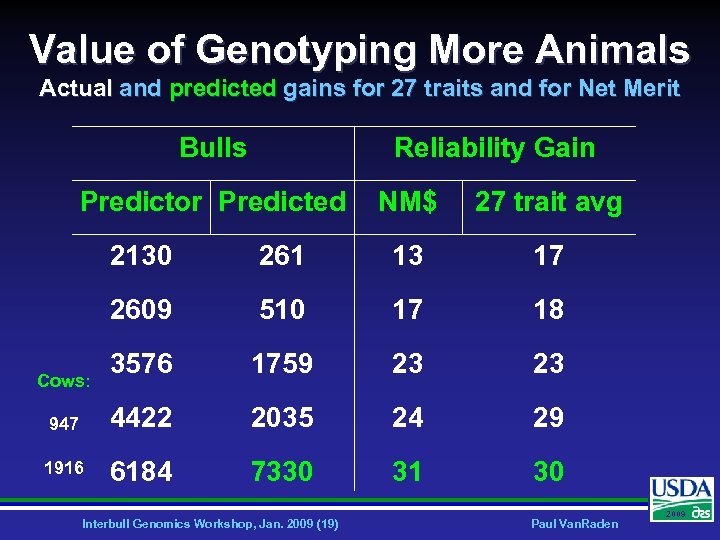
Value of Genotyping More Animals Actual and predicted gains for 27 traits and for Net Merit Bulls Reliability Gain Predictor Predicted NM$ 27 trait avg 2130 261 13 17 2609 510 17 18 3576 1759 23 23 947 4422 2035 24 29 1916 6184 7330 31 30 Cows: Interbull Genomics Workshop, Jan. 2009 (19) Paul Van. Raden 2009
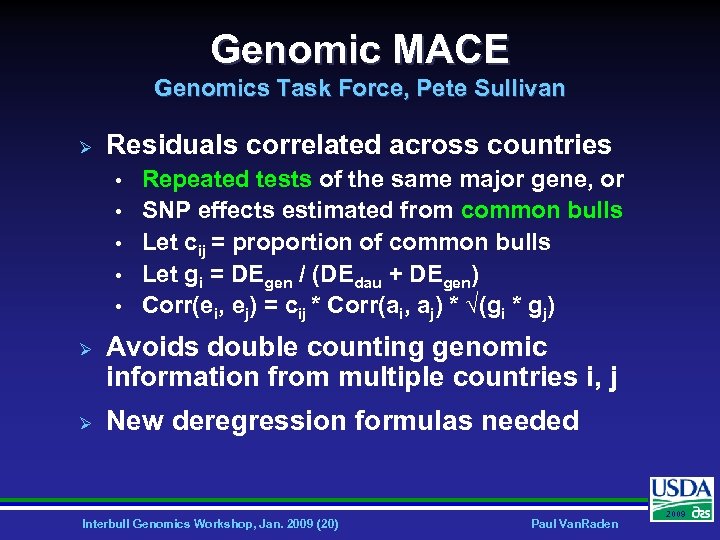

Genomic MACE Genomics Task Force, Pete Sullivan Ø Residuals correlated across countries • • • Ø Ø Repeated tests of the same major gene, or SNP effects estimated from common bulls Let cij = proportion of common bulls Let gi = DEgen / (DEdau + DEgen) Corr(ei, ej) = cij * Corr(ai, aj) * √(gi * gj) Avoids double counting genomic information from multiple countries i, j New deregression formulas needed Interbull Genomics Workshop, Jan. 2009 (20) Paul Van. Raden 2009
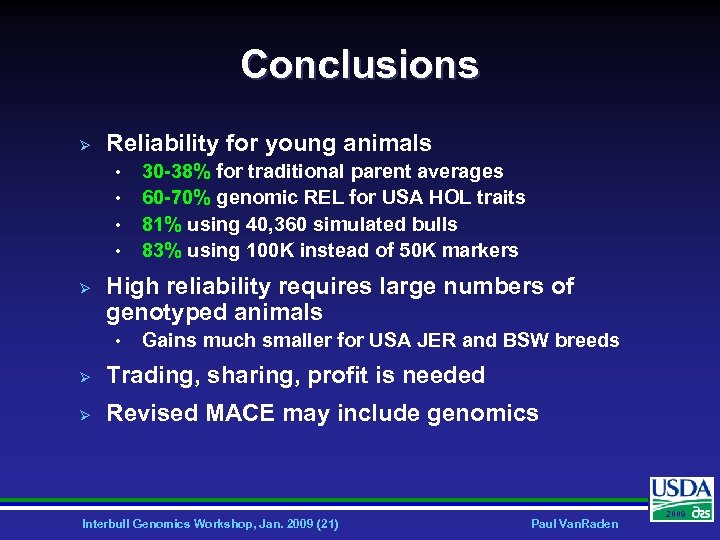

Conclusions Ø Reliability for young animals • • Ø 30 -38% for traditional parent averages 60 -70% genomic REL for USA HOL traits 81% using 40, 360 simulated bulls 83% using 100 K instead of 50 K markers High reliability requires large numbers of genotyped animals • Gains much smaller for USA JER and BSW breeds Ø Trading, sharing, profit is needed Ø Revised MACE may include genomics Interbull Genomics Workshop, Jan. 2009 (21) Paul Van. Raden 2009

Acknowledgments Ø Genotyping and DNA extraction: • Ø Computing: • Ø USDA Bovine Functional Genomics Lab, U. Missouri, U. Alberta, Gene. Seek, Genetics & IVF Institute, Genetic Visions, and Illumina AIPL staff (Mel Tooker, Leigh Walton, Jay Megonigal) Funding: • National Research Initiative grants – • • • 2006 -35205 -16888, 2006 -35205 -16701 Agriculture Research Service Holstein and Jersey breed associations Contributors to Cooperative Dairy DNA Repository (CDDR) Interbull Genomics Workshop, Jan. 2009 (22) Paul Van. Raden 2009

CDDR Contributors Ø National Association of Animal Breeders (NAAB, Columbia, MO) • ABS Global (De. Forest, WI) Accelerated Genetics (Baraboo, WI) Alta (Balzac, AB, Canada) Genex (Shawano, WI) New Generation Genetics (Fort Atkinson, WI) • Select Sires (Plain City, OH) • Semex Alliance (Guelph, ON, Canada) • Taurus-Service (Mehoopany, PA) • • Interbull Genomics Workshop, Jan. 2009 (23) Paul Van. Raden 2009
8d0e2dd000aaa4c972c07168839a1a58.ppt