964c75d150140f3c98387cd3548088bb.ppt
- Количество слайдов: 37
 Automated collection and processing of macromolecular diffraction data with DNA • Project started in 2001 following an ESRF user meeting • Currently involves scientists at Diamond, ESRF, EMBL Grenoble, EMBL Hamburg, SRS and SOLEIL. • Funded (indirectly) by AUTOSTRUCT, BIOXHIT, e. HTPX, CCP 4 • Aim is to automate data collection and processing at synchrotron beamlines. Data Not Analysed Dog. Patch Neighbourhood Association Deer Natural Aroma Disco Night Action Drugz N Alchohol (Thanks to acronym. com) Definately Not Attractive Distributed Network Attack Dark Native Apostle Dawn of the New Age Douglas Noel Adams Does Not at All Deadly Nocturnal Assassins Drink No Alcohol Desire and Needs Assessment
Automated collection and processing of macromolecular diffraction data with DNA • Project started in 2001 following an ESRF user meeting • Currently involves scientists at Diamond, ESRF, EMBL Grenoble, EMBL Hamburg, SRS and SOLEIL. • Funded (indirectly) by AUTOSTRUCT, BIOXHIT, e. HTPX, CCP 4 • Aim is to automate data collection and processing at synchrotron beamlines. Data Not Analysed Dog. Patch Neighbourhood Association Deer Natural Aroma Disco Night Action Drugz N Alchohol (Thanks to acronym. com) Definately Not Attractive Distributed Network Attack Dark Native Apostle Dawn of the New Age Douglas Noel Adams Does Not at All Deadly Nocturnal Assassins Drink No Alcohol Desire and Needs Assessment
 Participants • DNA developers • Collaborating sites – Karen Ackroyd (a) – Alun Ashton (b) – (a) CLRC Daresbury, UK – Gleb Bourenkov (c) – (b) Diamond light source, UK – Sandor Brockhauser (d) – (c) EMBL Hamburg, Germany – Marie-Francoise Incardona (f) – (d) EMBL Grenoble, France – Steve Kinder (a) – (e) Synchrotron Soleil, France – Pierre Legrand (e) – Karl Levik (b) – (f) ESRF, France – Romeu Pieritz (f) – (g) MRC LMB, Cambridge, UK – Sasha Popov – (h) Global Phasing, Cambridge, UK – Harry Powell (g) – NSLS Brookhaven – Darren Spruce (f) – Olof Svensson (f) – Graeme Winter (a) • DNA Exec. Committee: – Gérard Bricogne (h) Funding from: EU (AUTOSTRUCT, – Andrew Leslie (g) BIOXHIT), BBSRC (EHTPX), CCP 4 – Sean Mc. Sweeney (f) – Colin Nave (a) – Alexander Popov (c) – Raimond Ravelli (d) – Andrew Thompson (e)
Participants • DNA developers • Collaborating sites – Karen Ackroyd (a) – Alun Ashton (b) – (a) CLRC Daresbury, UK – Gleb Bourenkov (c) – (b) Diamond light source, UK – Sandor Brockhauser (d) – (c) EMBL Hamburg, Germany – Marie-Francoise Incardona (f) – (d) EMBL Grenoble, France – Steve Kinder (a) – (e) Synchrotron Soleil, France – Pierre Legrand (e) – Karl Levik (b) – (f) ESRF, France – Romeu Pieritz (f) – (g) MRC LMB, Cambridge, UK – Sasha Popov – (h) Global Phasing, Cambridge, UK – Harry Powell (g) – NSLS Brookhaven – Darren Spruce (f) – Olof Svensson (f) – Graeme Winter (a) • DNA Exec. Committee: – Gérard Bricogne (h) Funding from: EU (AUTOSTRUCT, – Andrew Leslie (g) BIOXHIT), BBSRC (EHTPX), CCP 4 – Sean Mc. Sweeney (f) – Colin Nave (a) – Alexander Popov (c) – Raimond Ravelli (d) – Andrew Thompson (e)
 Current Structure Exec - policy decisions, conflict resolution. One representative from each institution that hosts a developer. Project Coordinator - (Alun Ashton). Arranges VCs, ensures actions are carried out. Developers - Do the work Full DNA meetings twice a year. Additional developers meetings twothree times a year. Video conferencing ~ every two weeks. Bugzilla for bug tracking. CVS at D/L (moving to Diamond)
Current Structure Exec - policy decisions, conflict resolution. One representative from each institution that hosts a developer. Project Coordinator - (Alun Ashton). Arranges VCs, ensures actions are carried out. Developers - Do the work Full DNA meetings twice a year. Additional developers meetings twothree times a year. Video conferencing ~ every two weeks. Bugzilla for bug tracking. CVS at D/L (moving to Diamond)
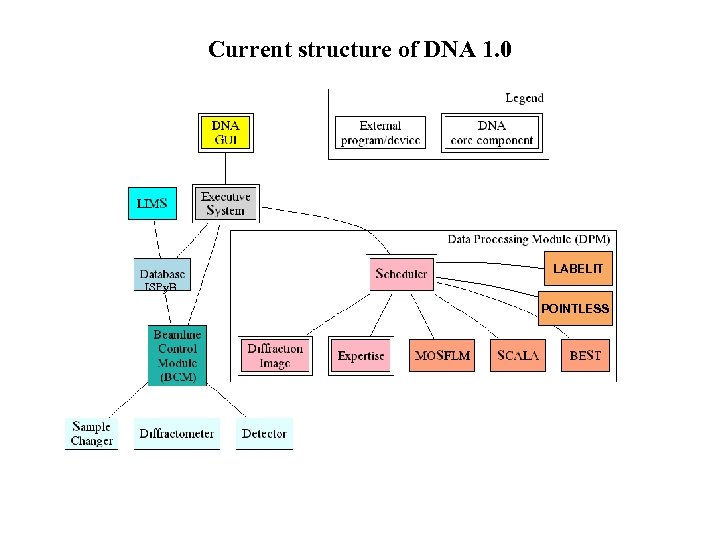 Current structure of DNA 1. 0 LABELIT ISPy. B POINTLESS
Current structure of DNA 1. 0 LABELIT ISPy. B POINTLESS
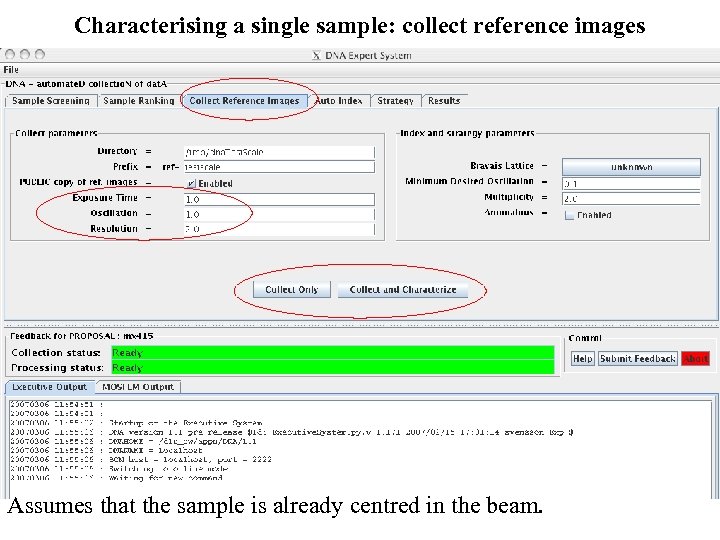 Characterising a single sample: collect reference images Assumes that the sample is already centred in the beam.
Characterising a single sample: collect reference images Assumes that the sample is already centred in the beam.
 Project Status DNA 1. 0 released December 2004. Basically it didn’t work. DNA 1. 1 planned release April 2007, although something very close to 1. 1 is already installed on 7 MX beamlines at ESRF. Improvements: • Robustness improved significantly • Datasets integrated and scaled “on the fly” • Allows a “sample ranking” mode for multiple crystals of the same type • Significant improvement in autoindexing success • Includes an interface to the EMBL/ESRF mini kappa goniostat. Now at a critical phase. . People are starting to use it !
Project Status DNA 1. 0 released December 2004. Basically it didn’t work. DNA 1. 1 planned release April 2007, although something very close to 1. 1 is already installed on 7 MX beamlines at ESRF. Improvements: • Robustness improved significantly • Datasets integrated and scaled “on the fly” • Allows a “sample ranking” mode for multiple crystals of the same type • Significant improvement in autoindexing success • Includes an interface to the EMBL/ESRF mini kappa goniostat. Now at a critical phase. . People are starting to use it !
 Future plans DNA 2. 0 involves a complete rewrite of the code. Project manager Olof Svensson. Currently a “spike” development is underway to establish which tools to use. A “scientific case” is being prepared which will define the scientific (user) objectives. Objectives: • More sophisticated data collection strategies (multiple wavelength, multiple crystals, multiple sweeps) • Feedback from “downstream” processing to improve data quality • More sophisticated treatment of radiation damage • Facilitate incorporation of other new/replacement modules, eg alternative data processing programs, absorption corrections etc • Improved summary of experimental results in database (ISPy. B) • Automated selection of wavelength for anomalous data (peak).
Future plans DNA 2. 0 involves a complete rewrite of the code. Project manager Olof Svensson. Currently a “spike” development is underway to establish which tools to use. A “scientific case” is being prepared which will define the scientific (user) objectives. Objectives: • More sophisticated data collection strategies (multiple wavelength, multiple crystals, multiple sweeps) • Feedback from “downstream” processing to improve data quality • More sophisticated treatment of radiation damage • Facilitate incorporation of other new/replacement modules, eg alternative data processing programs, absorption corrections etc • Improved summary of experimental results in database (ISPy. B) • Automated selection of wavelength for anomalous data (peak).
 The End
The End
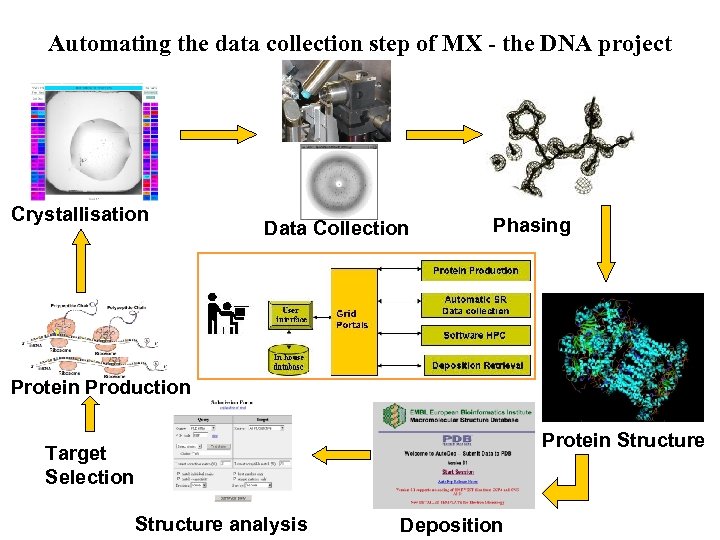 Automating the data collection step of MX - the DNA project Crystallisation Data Collection Phasing Protein Production Protein Structure Target Selection Structure analysis Deposition
Automating the data collection step of MX - the DNA project Crystallisation Data Collection Phasing Protein Production Protein Structure Target Selection Structure analysis Deposition
 Objective of DNA Given some basic information about the project and the available crystals, to determine and carry out the best possible diffraction experiment from the available crystals in the available time, by a procedure that requires little or no intervention from the user during the experiment. Make it easier for inexperienced users to collect good data. Should facilitate both “Fedex” crystallography and remote operation.
Objective of DNA Given some basic information about the project and the available crystals, to determine and carry out the best possible diffraction experiment from the available crystals in the available time, by a procedure that requires little or no intervention from the user during the experiment. Make it easier for inexperienced users to collect good data. Should facilitate both “Fedex” crystallography and remote operation.
 Input to DNA • The diffraction plan (supplied by user) • The type of experiment (MAD/SAD phasing, sulphur phasing, high resolution data for refinement). • Resolution: ideal, minimum acceptable, maximum required. • Type of anomalous scatterers for phasing experiments • Unit cell and space group (if known) • Crystal lifetime (if known) • X-ray source properties (supplied by beamline scientists) • Typical crystal lifetime (in seconds or photons) • Maximum rotation rate for spindle • Minimum safe exposure time (depends on shutter) • Accessible wavelength range
Input to DNA • The diffraction plan (supplied by user) • The type of experiment (MAD/SAD phasing, sulphur phasing, high resolution data for refinement). • Resolution: ideal, minimum acceptable, maximum required. • Type of anomalous scatterers for phasing experiments • Unit cell and space group (if known) • Crystal lifetime (if known) • X-ray source properties (supplied by beamline scientists) • Typical crystal lifetime (in seconds or photons) • Maximum rotation rate for spindle • Minimum safe exposure time (depends on shutter) • Accessible wavelength range
 Output from DNA • A summary of the diffraction characteristics for every crystal tested • diffraction limit (based on analysis of spot finding/ BEST)) • unit cell, probable Laue group/lattice type (autoindexing) • mosaicity (MOSFLM) • assessment of crystal perfection (based on presence of multiple lattices) • assessment of spot shape (single, streaky, split, multiple) • When defined criteria have been met (resolution, best sample of those tested), one or more (MAD) data sets (h, k, l, F, sig(F)) of scaled and (optionally) merged structure factor amplitudes. The data processing may be preliminary (to save time) but will give a realistic estimate of data quality.
Output from DNA • A summary of the diffraction characteristics for every crystal tested • diffraction limit (based on analysis of spot finding/ BEST)) • unit cell, probable Laue group/lattice type (autoindexing) • mosaicity (MOSFLM) • assessment of crystal perfection (based on presence of multiple lattices) • assessment of spot shape (single, streaky, split, multiple) • When defined criteria have been met (resolution, best sample of those tested), one or more (MAD) data sets (h, k, l, F, sig(F)) of scaled and (optionally) merged structure factor amplitudes. The data processing may be preliminary (to save time) but will give a realistic estimate of data quality.
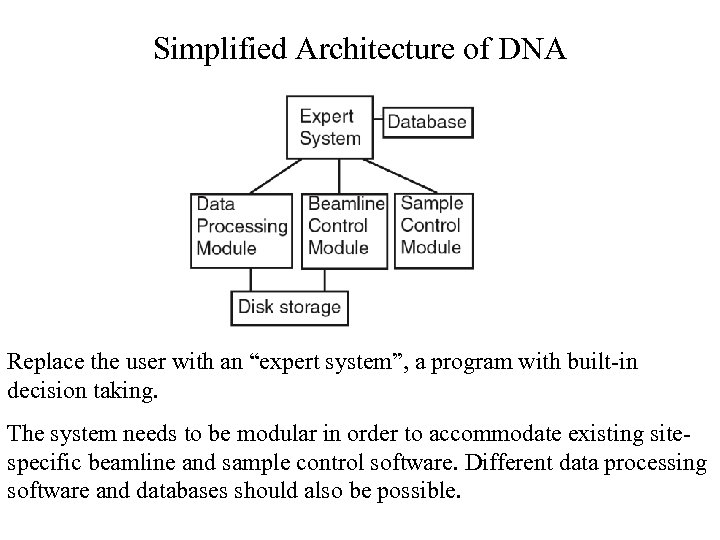 Simplified Architecture of DNA Replace the user with an “expert system”, a program with built-in decision taking. The system needs to be modular in order to accommodate existing sitespecific beamline and sample control software. Different data processing software and databases should also be possible.
Simplified Architecture of DNA Replace the user with an “expert system”, a program with built-in decision taking. The system needs to be modular in order to accommodate existing sitespecific beamline and sample control software. Different data processing software and databases should also be possible.
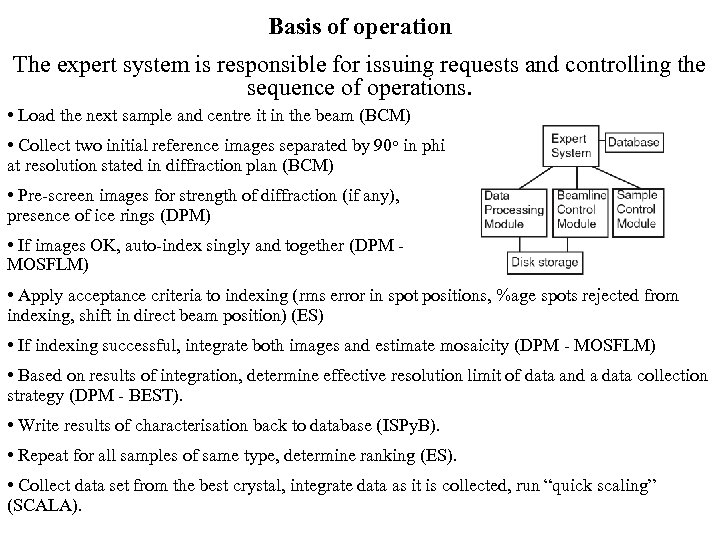 Basis of operation The expert system is responsible for issuing requests and controlling the sequence of operations. • Load the next sample and centre it in the beam (BCM) • Collect two initial reference images separated by 90 o in phi at resolution stated in diffraction plan (BCM) • Pre-screen images for strength of diffraction (if any), presence of ice rings (DPM) • If images OK, auto-index singly and together (DPM MOSFLM) • Apply acceptance criteria to indexing (rms error in spot positions, %age spots rejected from indexing, shift in direct beam position) (ES) • If indexing successful, integrate both images and estimate mosaicity (DPM - MOSFLM) • Based on results of integration, determine effective resolution limit of data and a data collection strategy (DPM - BEST). • Write results of characterisation back to database (ISPy. B). • Repeat for all samples of same type, determine ranking (ES). • Collect data set from the best crystal, integrate data as it is collected, run “quick scaling” (SCALA).
Basis of operation The expert system is responsible for issuing requests and controlling the sequence of operations. • Load the next sample and centre it in the beam (BCM) • Collect two initial reference images separated by 90 o in phi at resolution stated in diffraction plan (BCM) • Pre-screen images for strength of diffraction (if any), presence of ice rings (DPM) • If images OK, auto-index singly and together (DPM MOSFLM) • Apply acceptance criteria to indexing (rms error in spot positions, %age spots rejected from indexing, shift in direct beam position) (ES) • If indexing successful, integrate both images and estimate mosaicity (DPM - MOSFLM) • Based on results of integration, determine effective resolution limit of data and a data collection strategy (DPM - BEST). • Write results of characterisation back to database (ISPy. B). • Repeat for all samples of same type, determine ranking (ES). • Collect data set from the best crystal, integrate data as it is collected, run “quick scaling” (SCALA).
 Implementation of DNA at ESRF DNA is installed on all MX beamlines except ID 13. Started with a desktop icon (after starting Mx. Cube) Currently two modes of operation: • Characterising a single sample • Sample screening (pipeline mode)
Implementation of DNA at ESRF DNA is installed on all MX beamlines except ID 13. Started with a desktop icon (after starting Mx. Cube) Currently two modes of operation: • Characterising a single sample • Sample screening (pipeline mode)
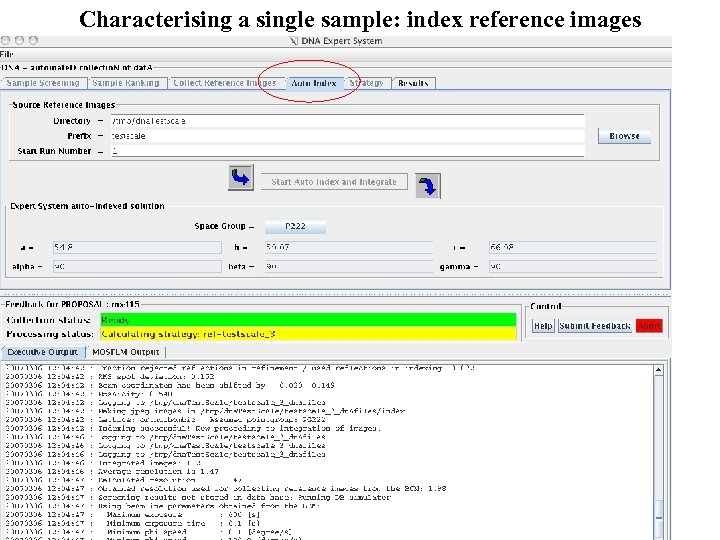 Characterising a single sample: index reference images
Characterising a single sample: index reference images
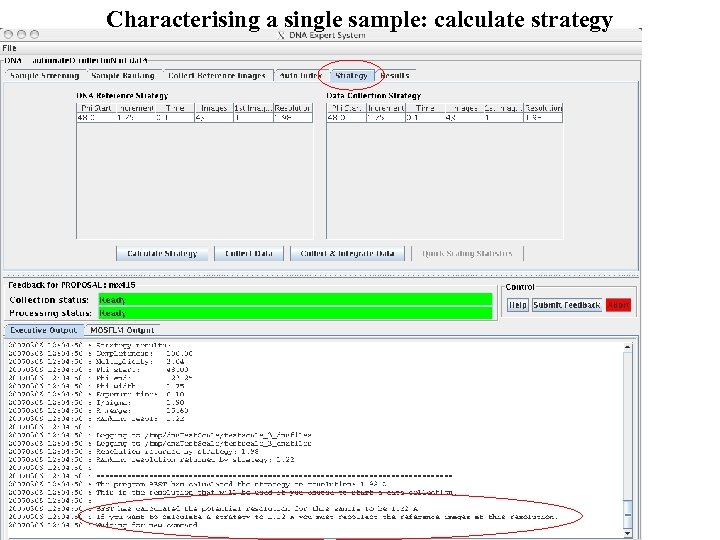 Characterising a single sample: calculate strategy
Characterising a single sample: calculate strategy
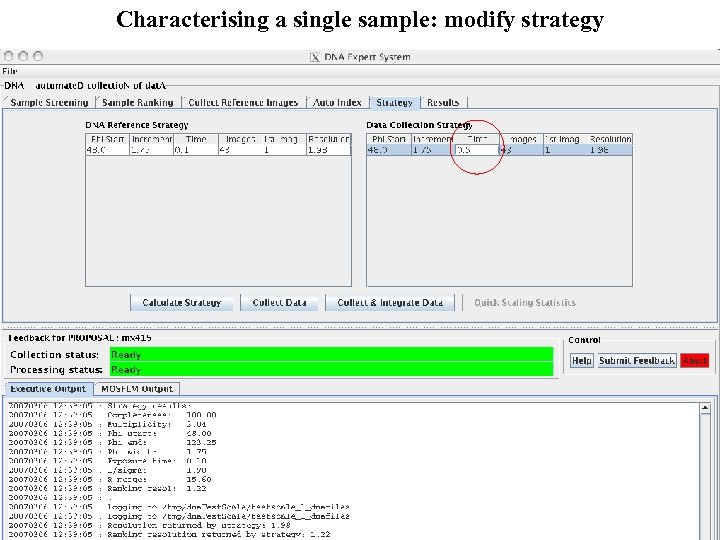 Characterising a single sample: modify strategy
Characterising a single sample: modify strategy
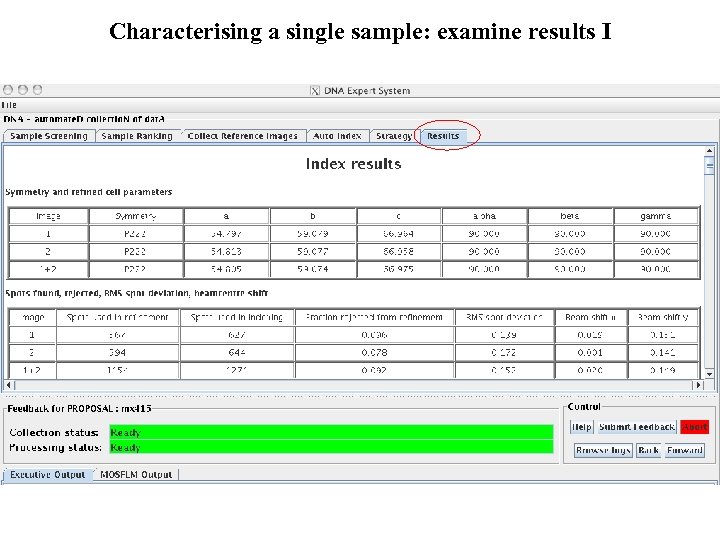 Characterising a single sample: examine results I
Characterising a single sample: examine results I
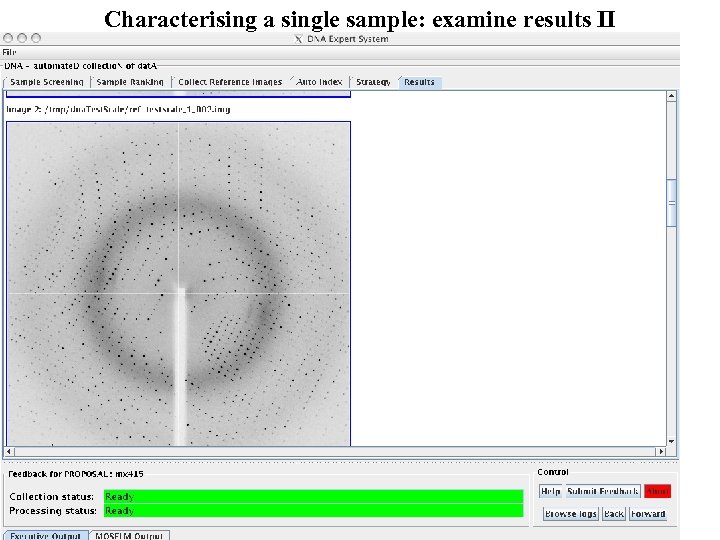 Characterising a single sample: examine results II
Characterising a single sample: examine results II
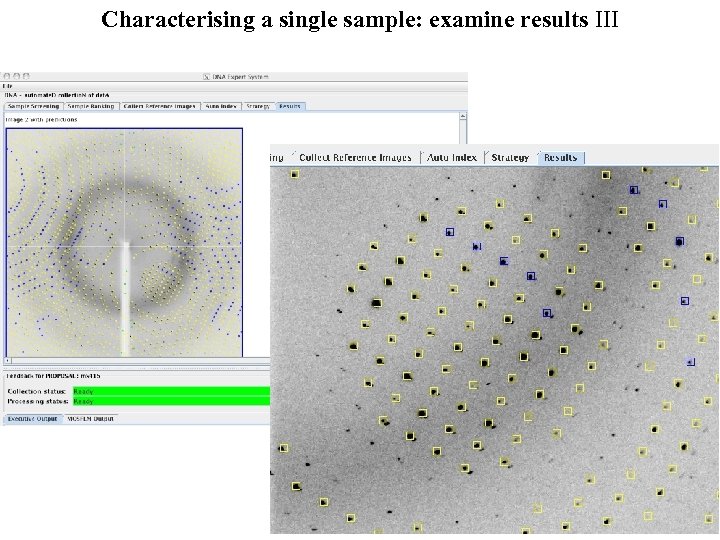 Characterising a single sample: examine results III
Characterising a single sample: examine results III
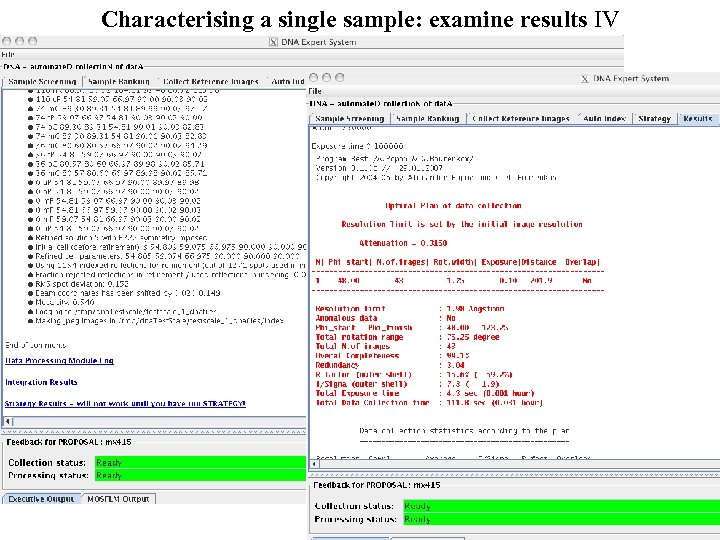 Characterising a single sample: examine results IV
Characterising a single sample: examine results IV
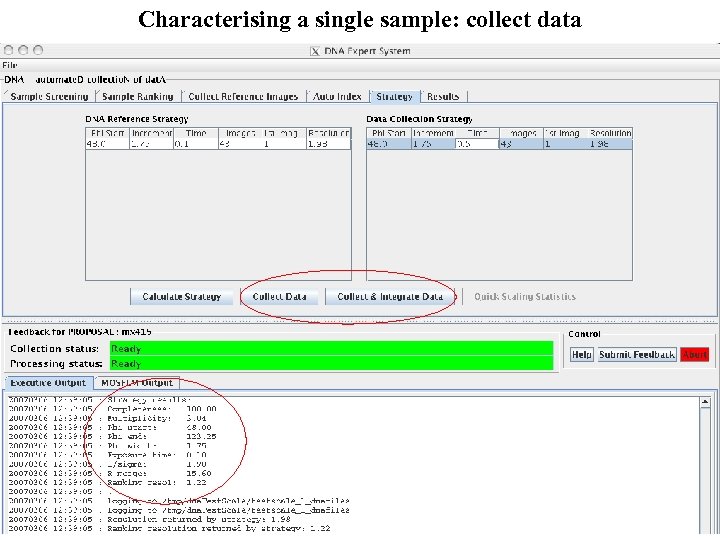 Characterising a single sample: collect data
Characterising a single sample: collect data
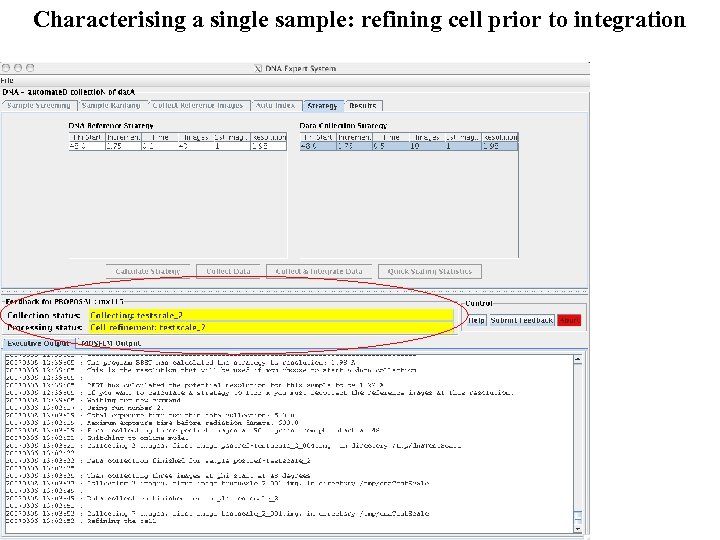 Characterising a single sample: refining cell prior to integration
Characterising a single sample: refining cell prior to integration
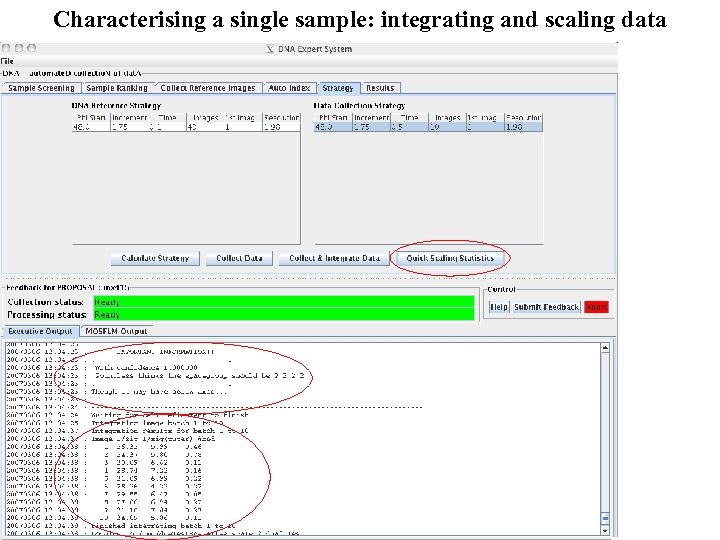 Characterising a single sample: integrating and scaling data
Characterising a single sample: integrating and scaling data
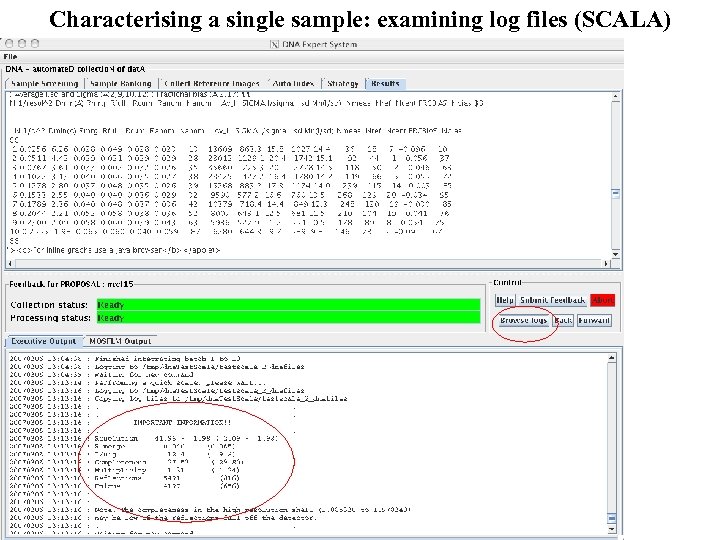 Characterising a single sample: examining log files (SCALA)
Characterising a single sample: examining log files (SCALA)
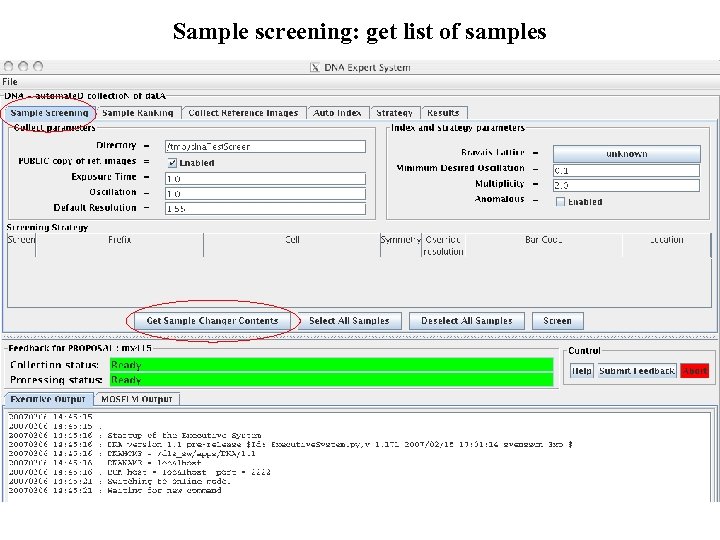 Sample screening: get list of samples
Sample screening: get list of samples
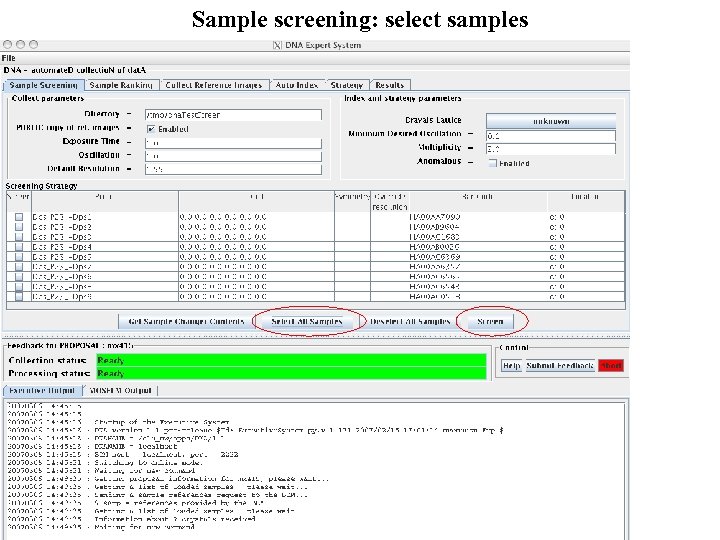 Sample screening: select samples
Sample screening: select samples
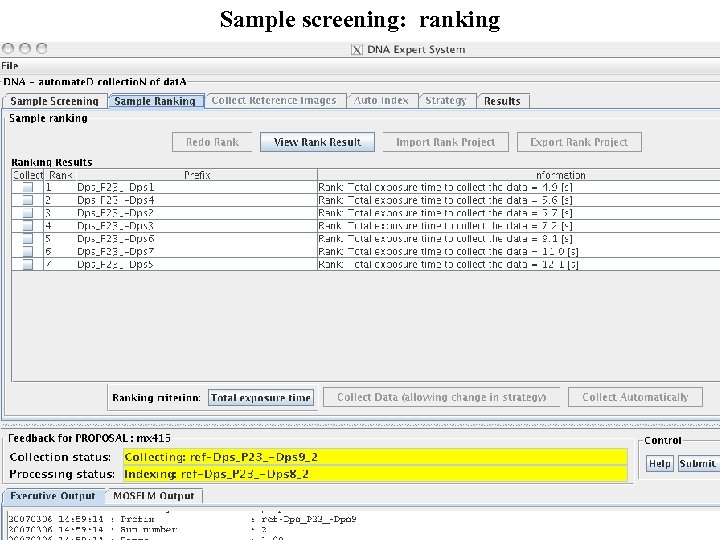 Sample screening: ranking
Sample screening: ranking
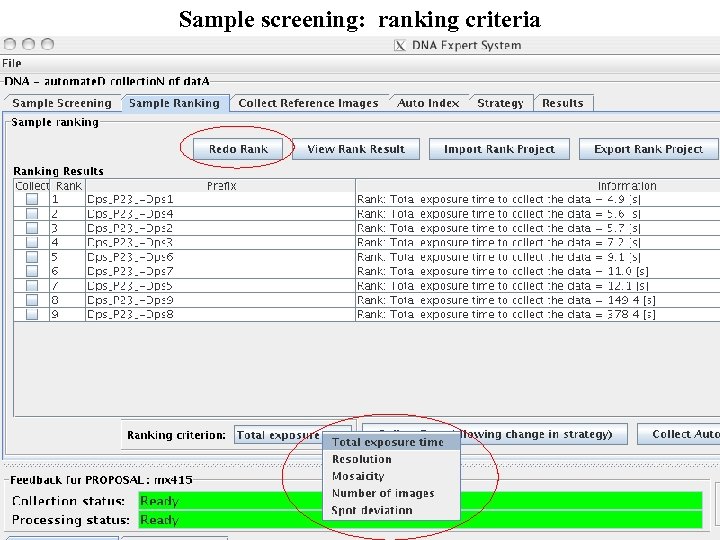 Sample screening: ranking criteria
Sample screening: ranking criteria
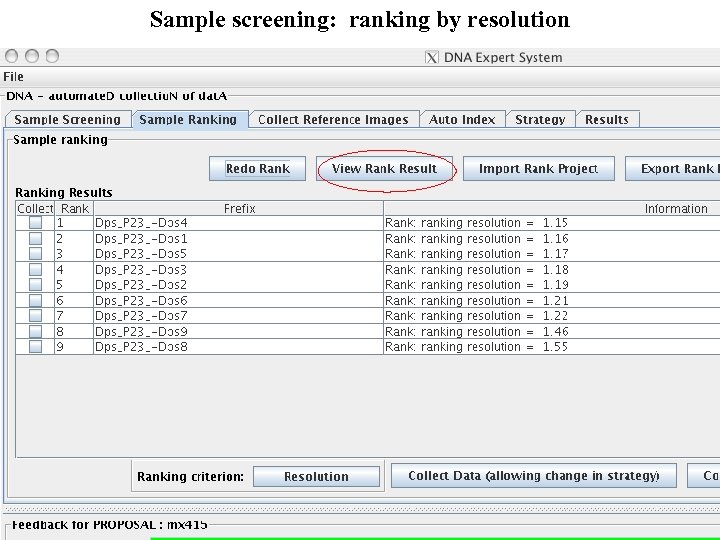 Sample screening: ranking by resolution
Sample screening: ranking by resolution
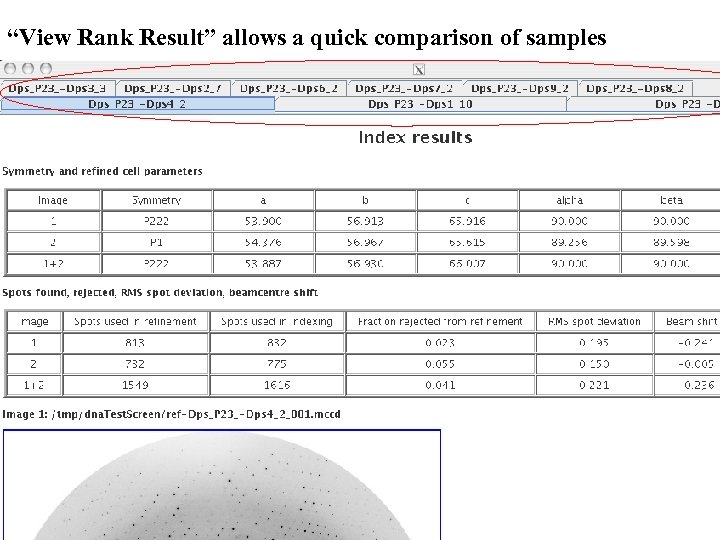 “View Rank Result” allows a quick comparison of samples
“View Rank Result” allows a quick comparison of samples
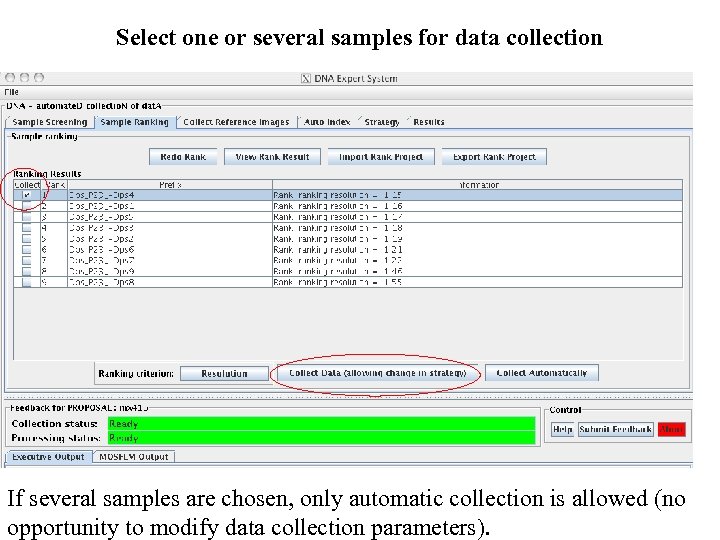 Select one or several samples for data collection If several samples are chosen, only automatic collection is allowed (no opportunity to modify data collection parameters).
Select one or several samples for data collection If several samples are chosen, only automatic collection is allowed (no opportunity to modify data collection parameters).
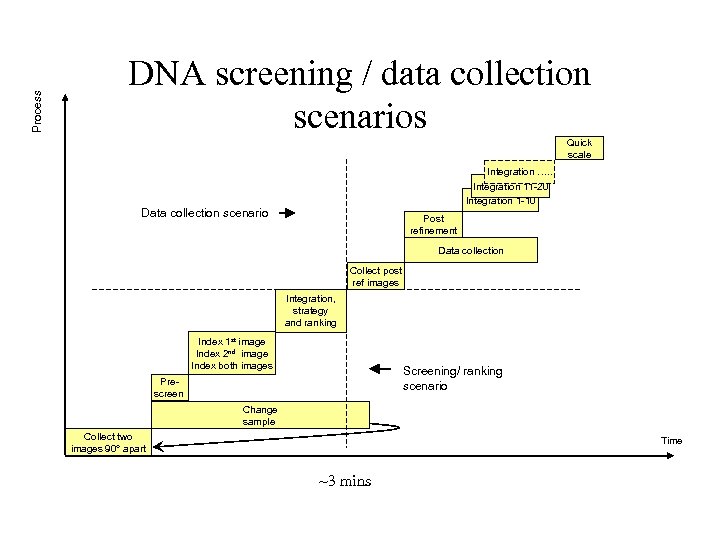 Process DNA screening / data collection scenarios Quick scale Integration …. . Integration 11 -20 Integration 1 -10 Data collection scenario Post refinement Data collection Collect post ref images Integration, strategy and ranking Index 1 st image Index 2 nd image Index both images Screening/ ranking scenario Prescreen Change sample Collect two images 90° apart Time ~3 mins
Process DNA screening / data collection scenarios Quick scale Integration …. . Integration 11 -20 Integration 1 -10 Data collection scenario Post refinement Data collection Collect post ref images Integration, strategy and ranking Index 1 st image Index 2 nd image Index both images Screening/ ranking scenario Prescreen Change sample Collect two images 90° apart Time ~3 mins
 Issues Conceptual • How ambitious to be in terms of handling “less than perfect” crystals. • How much user control should be accommodated ? Technical • Depends on success of crystal centring • Dependency of success of autoindexing • Choice of criteria for ranking • How to deal with user requested resolution • ** How to deal with radiation damage ** • How to make processing keep up with data collection
Issues Conceptual • How ambitious to be in terms of handling “less than perfect” crystals. • How much user control should be accommodated ? Technical • Depends on success of crystal centring • Dependency of success of autoindexing • Choice of criteria for ranking • How to deal with user requested resolution • ** How to deal with radiation damage ** • How to make processing keep up with data collection
 Future Plans • Scrap it all and start again • Extend to include collection and analysis of edge scans for design of anomalous diffraction data collection • Optimise beam size wrt crystal size • Allow more sophisticated data collection strategies (about more than one axis, variable exposure time/oscillation angle per image, using more than one position on a crystal or several different crystals) • Allow strategy to be updated if initial estimate of point group was incorrect • Automatic determination of crystal lifetime from a “sacrificial” crystal. • Improve level of information stored in the ISPy. B database • Improve robustness
Future Plans • Scrap it all and start again • Extend to include collection and analysis of edge scans for design of anomalous diffraction data collection • Optimise beam size wrt crystal size • Allow more sophisticated data collection strategies (about more than one axis, variable exposure time/oscillation angle per image, using more than one position on a crystal or several different crystals) • Allow strategy to be updated if initial estimate of point group was incorrect • Automatic determination of crystal lifetime from a “sacrificial” crystal. • Improve level of information stored in the ISPy. B database • Improve robustness
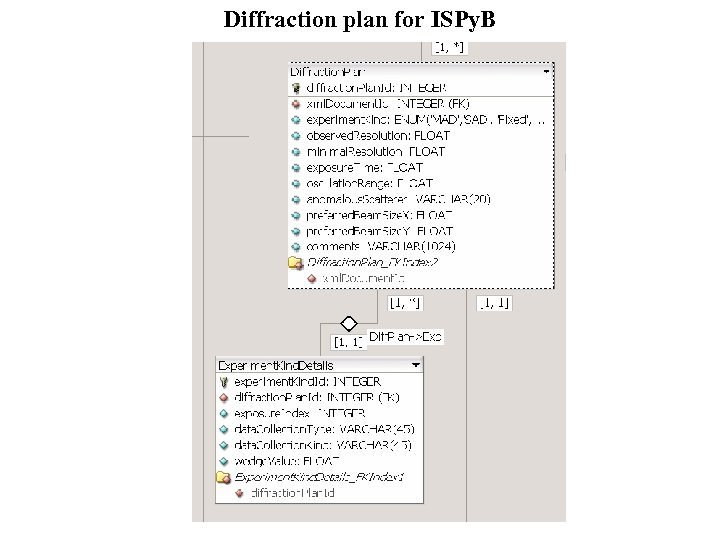 Diffraction plan for ISPy. B
Diffraction plan for ISPy. B


