590f8144bfaebfdad065632124927179.ppt
- Количество слайдов: 31

Approaches to Computation A Biomolecular Transducer Employing Ternary Language and Rendering a Biological Output Mark Chaskes and Paul Lazarescu Mentor: Tamar Ratner The Schulich Faculty of Chemistry Technion, Haifa, Israel, 32000

Objective Design a theoretical biomolecular transducer to solve consecutive mathematical equations in ternary. -First divide an input by three and then divide the yeild of that by two
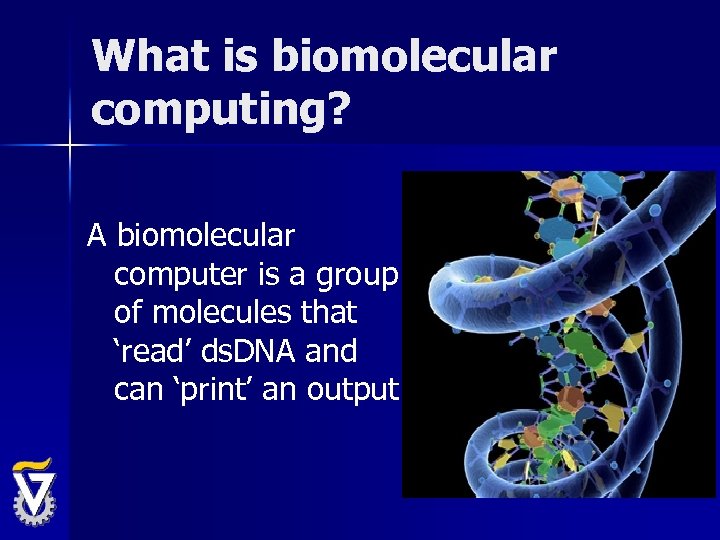

What is biomolecular computing? A biomolecular computer is a group of molecules that ‘read’ ds. DNA and can ‘print’ an output.

What is a DNA based ? transducer A transducer is not a PC; it has unique capabilities that an ordinary computer does not. Advantages include: • Direct interface with a biological system • Can release a biological output • Able to compute in parallel • Store large amounts of data in a small volume
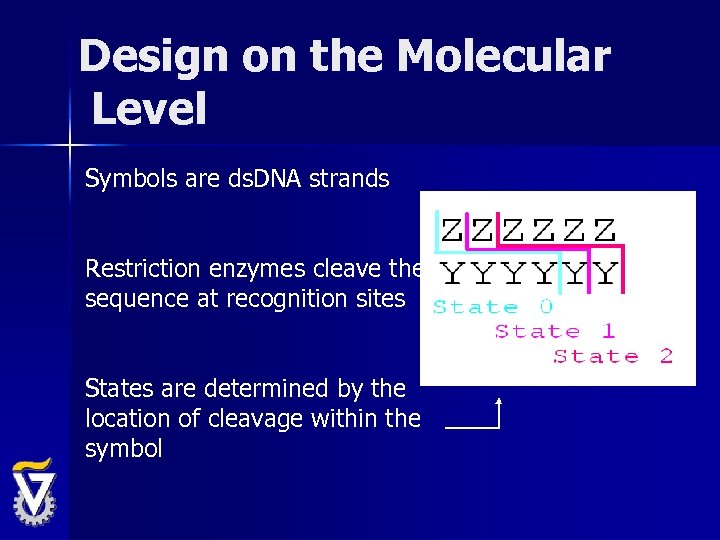
Design on the Molecular Level Symbols are ds. DNA strands Restriction enzymes cleave the sequence at recognition sites States are determined by the location of cleavage within the symbol
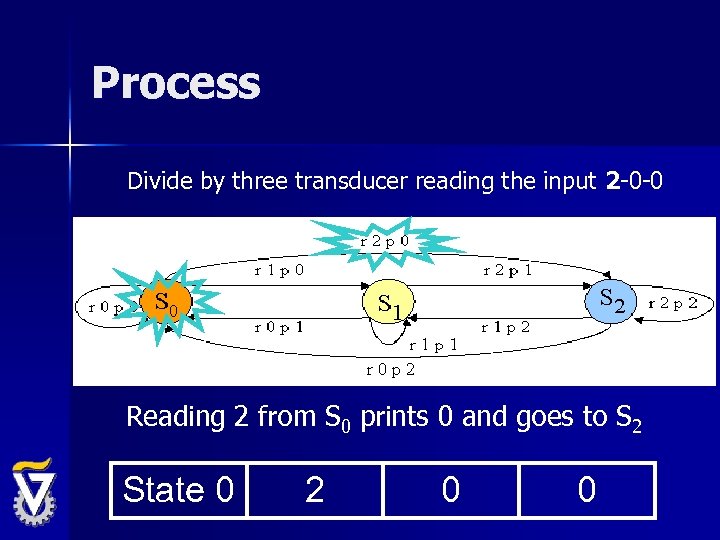
Process Divide by three transducer reading the input 2 -0 -0 Reading 2 from S 0 prints 0 and goes to S 2 State 0 2 0 0
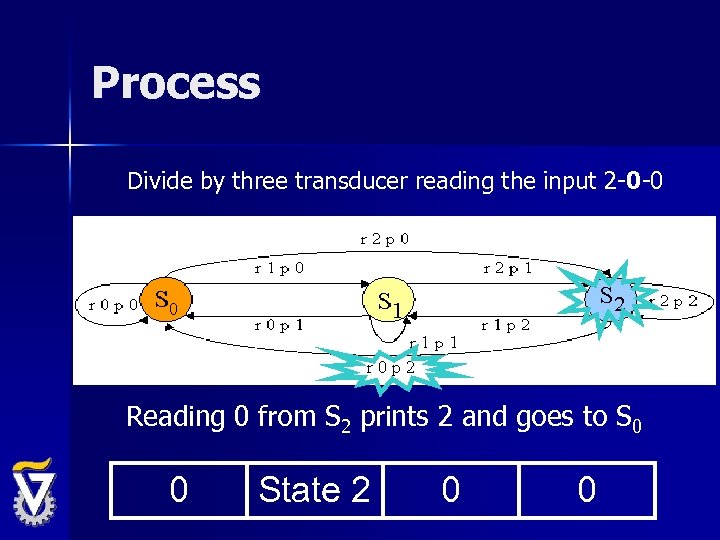
Process Divide by three transducer reading the input 2 -0 -0 Reading 0 from S 2 prints 2 and goes to S 0 0 State 2 0 0
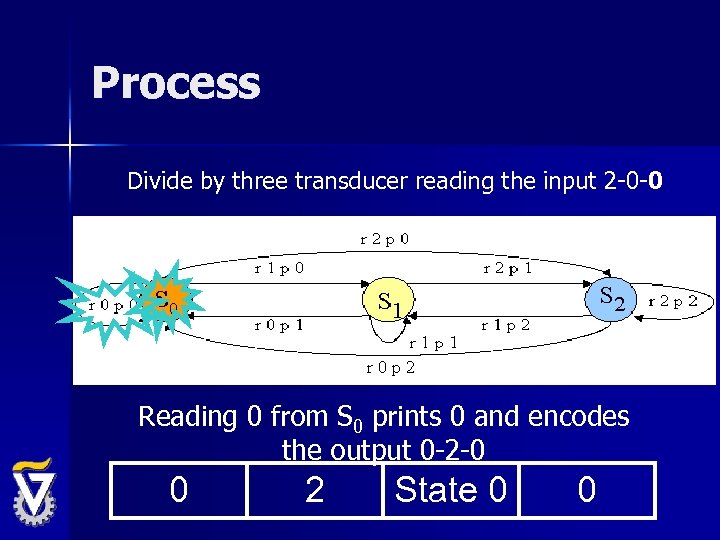
Process Divide by three transducer reading the input 2 -0 -0 Reading 0 from S 0 prints 0 and encodes the output 0 -2 -0 0 2 State 0 0
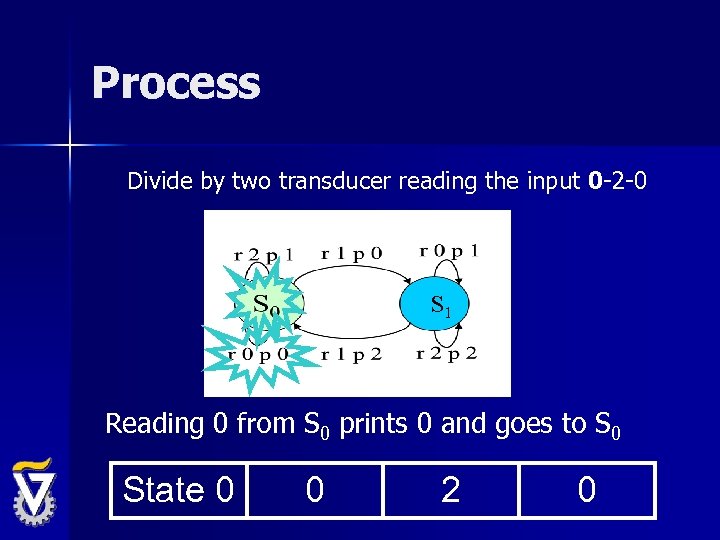
Process Divide by two transducer reading the input 0 -2 -0 S 1 Reading 0 from S 0 prints 0 and goes to S 0 State 0 0 2 0
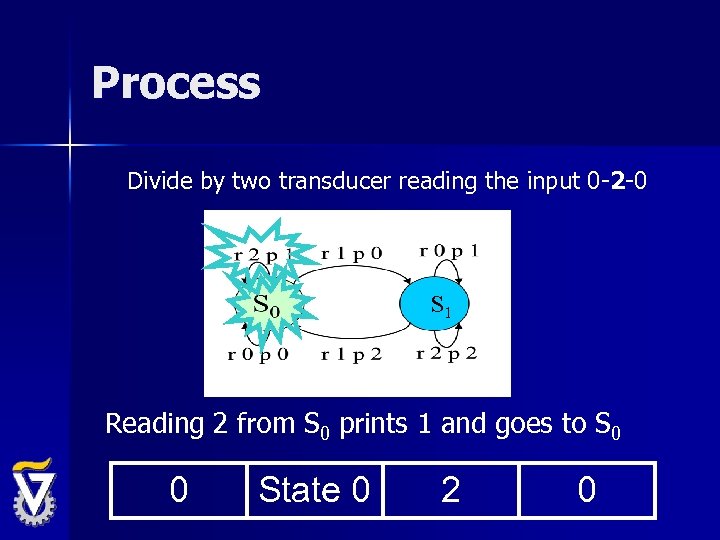
Process Divide by two transducer reading the input 0 -2 -0 S 1 Reading 2 from S 0 prints 1 and goes to S 0 0 State 0 2 0
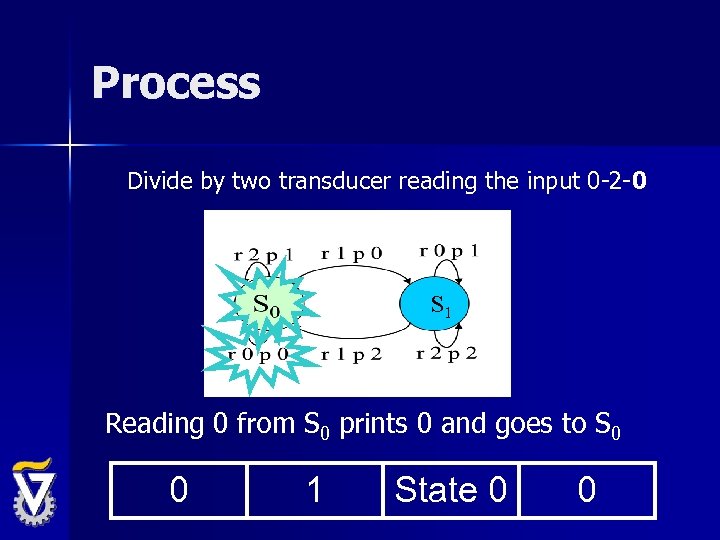
Process Divide by two transducer reading the input 0 -2 -0 S 1 Reading 0 from S 0 prints 0 and goes to S 0 0 1 State 0 0
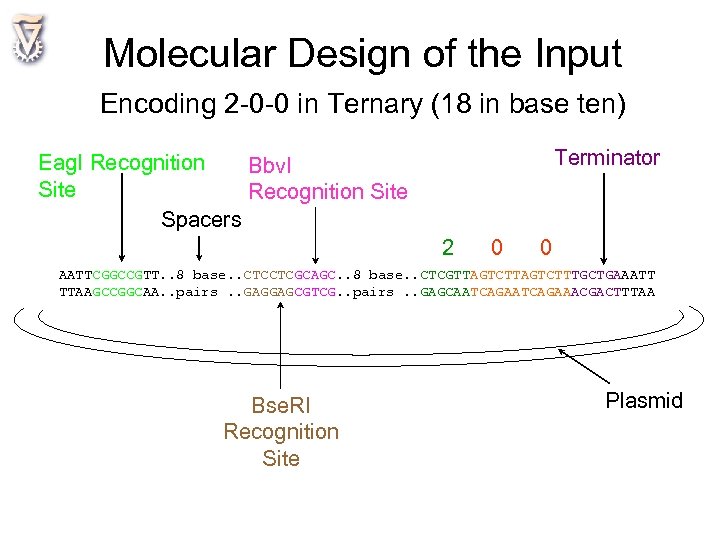
Molecular Design of the Input Encoding 2 -0 -0 in Ternary (18 in base ten) Terminator Eag. I Recognition Bbv. I Site Recognition Site Spacers 2 0 0 AATTCGGCCGTT. . 8 base. . CTCCTCGCAGC. . 8 base. . CTCGTTAGTCTTTGCTGAAATT TTAAGCCGGCAA. . pairs. . GAGGAGCGTCG. . pairs. . GAGCAATCAGAAACGACTTTAA Bse. RI Recognition Site Plasmid
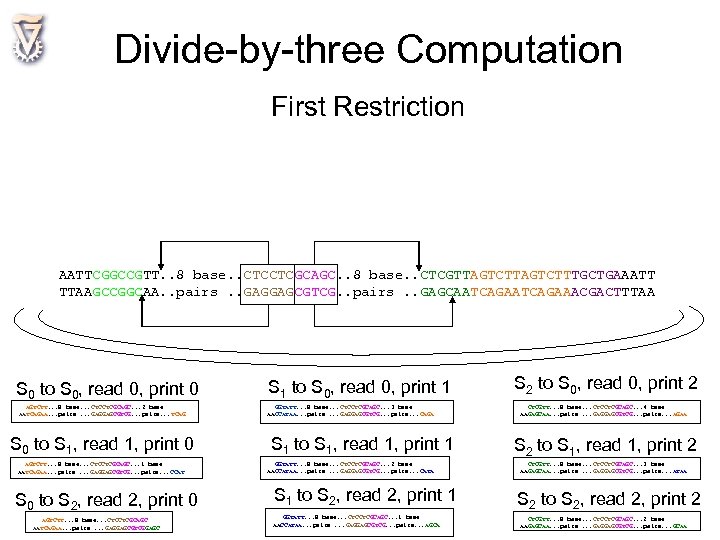
Divide-by-three Computation First Restriction AATTCGGCCGTT. . 8 base. . CTCCTCGCAGC. . 8 base. . CTCGTTAGTCTTTGCTGAAATT TTAAGCCGGCAA. . pairs. . GAGGAGCGTCG. . pairs. . GAGCAATCAGAAACGACTTTAA S 0 to S 0, read 0, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AATCAGAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . TCAG S 0 to S 1, read 1, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC. . . 1 base AATCAGAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CCAT S 0 to S 2, read 2, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC AATCAGAA. . . pairs. . . GAGGAGCGTCGGAGC S 1 to S 0, read 0, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 3 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CAGA S 1 to S 1, read 1, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CATA S 1 to S 2, read 2, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 1 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . AGCA S 2 to S 0, read 0, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 4 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . AGAA S 2 to S 1, read 1, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 3 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . ATAA S 2 to S 2, read 2, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . GCAA
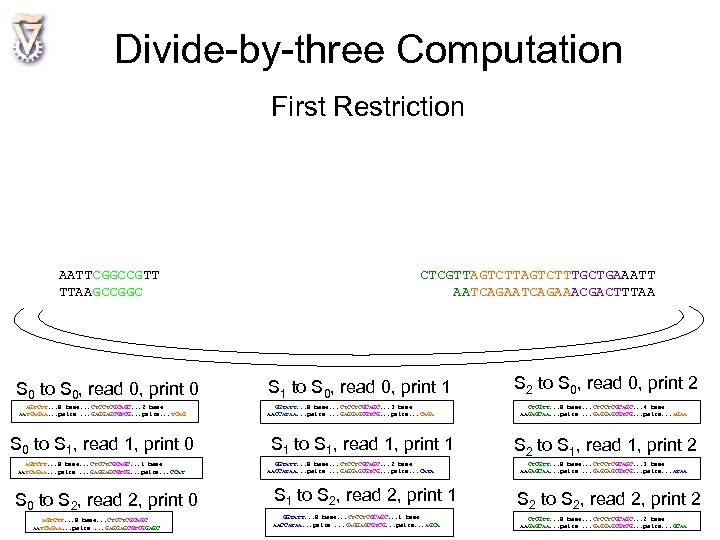
Divide-by-three Computation First Restriction AATTCGGCCGTT TTAAGCCGGC S 0 to S 0, read 0, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AATCAGAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . TCAG S 0 to S 1, read 1, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC. . . 1 base AATCAGAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CCAT S 0 to S 2, read 2, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC AATCAGAA. . . pairs. . . GAGGAGCGTCGGAGC CTCGTTAGTCTTTGCTGAAATT AATCAGAAACGACTTTAA S 1 to S 0, read 0, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 3 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CAGA S 1 to S 1, read 1, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CATA S 1 to S 2, read 2, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 1 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . AGCA S 2 to S 0, read 0, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 4 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . AGAA S 2 to S 1, read 1, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 3 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . ATAA S 2 to S 2, read 2, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . GCAA
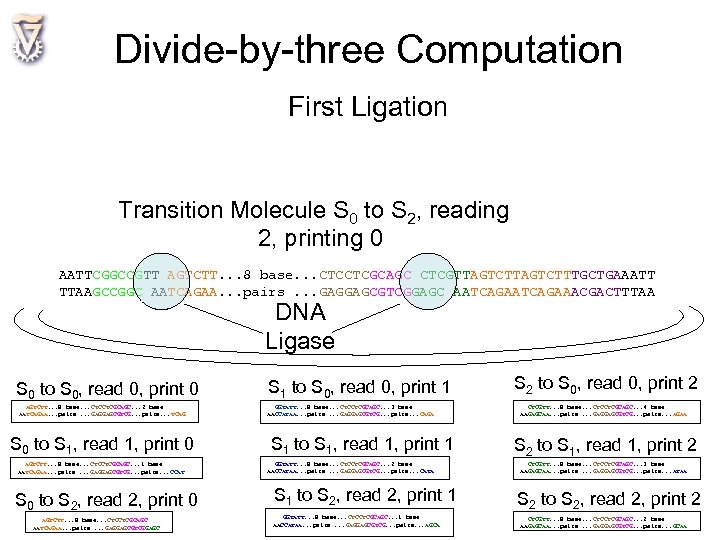
Divide-by-three Computation First Ligation Transition Molecule S 0 to S 2, reading 2, printing 0 AATTCGGCCGTT AGTCTT. . . 8 base. . . CTCCTCGCAGC CTCGTTAGTCTTTGCTGAAATT TTAAGCCGGC AATCAGAA. . . pairs. . . GAGGAGCGTCGGAGC AATCAGAAACGACTTTAA DNA Ligase S 0 to S 0, read 0, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AATCAGAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . TCAG S 0 to S 1, read 1, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC. . . 1 base AATCAGAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CCAT S 0 to S 2, read 2, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC AATCAGAA. . . pairs. . . GAGGAGCGTCGGAGC S 1 to S 0, read 0, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 3 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CAGA S 1 to S 1, read 1, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CATA S 1 to S 2, read 2, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 1 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . AGCA S 2 to S 0, read 0, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 4 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . AGAA S 2 to S 1, read 1, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 3 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . ATAA S 2 to S 2, read 2, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . GCAA
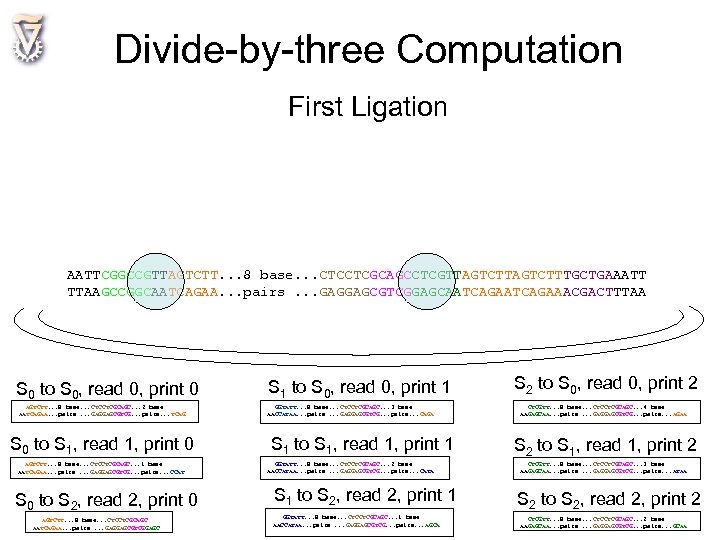
Divide-by-three Computation First Ligation AATTCGGCCGTTAGTCTT. . . 8 base. . . CTCCTCGCAGCCTCGTTAGTCTTAGTCTTTGCTGAAATT TTAAGCCGGCAATCAGAA. . . pairs. . . GAGGAGCGTCGGAGCAATCAGAATCAGAAACGACTTTAA S 0 to S 0, read 0, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AATCAGAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . TCAG S 0 to S 1, read 1, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC. . . 1 base AATCAGAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CCAT S 0 to S 2, read 2, print 0 AGTCTT. . . 8 base. . . CTCCTCGCAGC AATCAGAA. . . pairs. . . GAGGAGCGTCGGAGC S 1 to S 0, read 0, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 3 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CAGA S 1 to S 1, read 1, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . CATA S 1 to S 2, read 2, print 1 GGTATT. . . 8 base. . . CTCCTCGCAGC. . . 1 base AACCATAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . AGCA S 2 to S 0, read 0, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 4 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . AGAA S 2 to S 1, read 1, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 3 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . ATAA S 2 to S 2, read 2, print 2 CTCGTT. . . 8 base. . . CTCCTCGCAGC. . . 2 base AAGAGCAA. . . pairs. . . GAGGAGCGTCG. . . pairs. . . GCAA
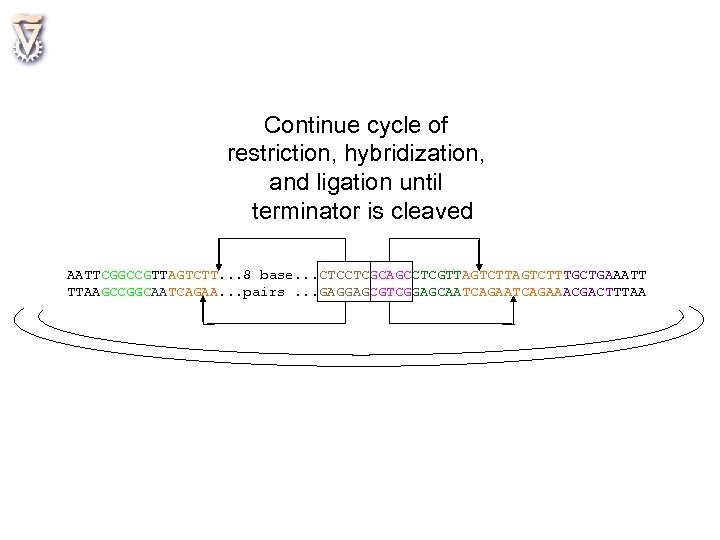
Continue cycle of restriction, hybridization, and ligation until terminator is cleaved AATTCGGCCGTTAGTCTT. . . 8 base. . . CTCCTCGCAGCCTCGTTAGTCTTAGTCTTTGCTGAAATT TTAAGCCGGCAATCAGAA. . . pairs. . . GAGGAGCGTCGGAGCAATCAGAATCAGAAACGACTTTAA
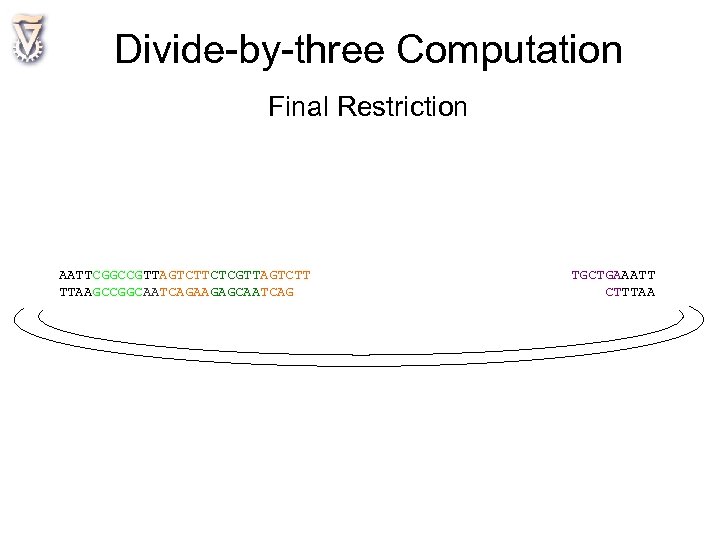

Divide-by-three Computation Final Restriction AATTCGGCCGTTAGTCTTCTCGTTAGTCTT TTAAGCCGGCAATCAGAAGAGCAATCAG TGCTGAAATT CTTTAA
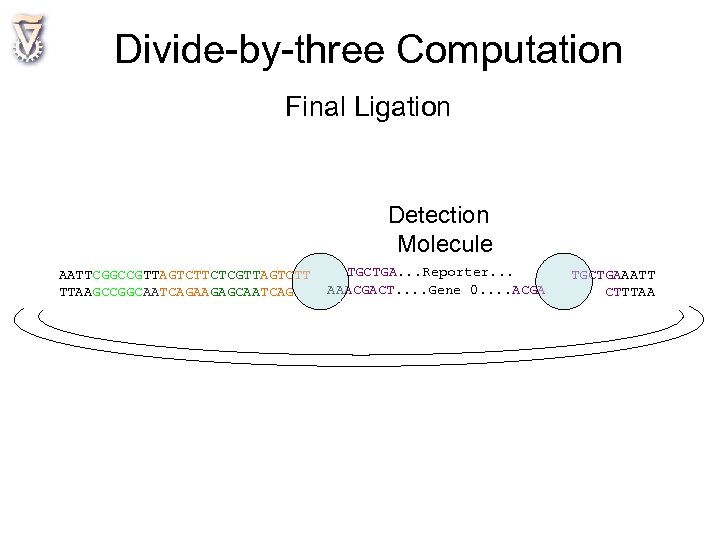
Divide-by-three Computation Final Ligation Detection Molecule AATTCGGCCGTTAGTCTTCTCGTTAGTCTT TTAAGCCGGCAATCAGAAGAGCAATCAG TGCTGA. . . Reporter. . . AAACGACT. . Gene 0. . ACGA TGCTGAAATT CTTTAA
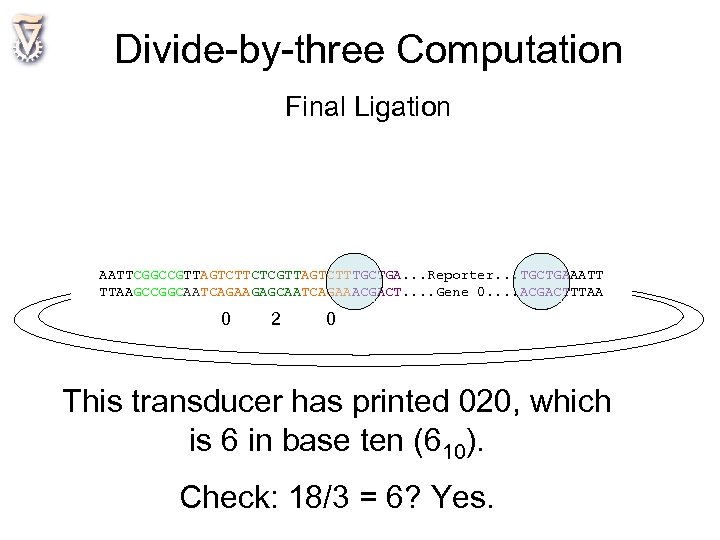
Divide-by-three Computation Final Ligation AATTCGGCCGTTAGTCTTCTCGTTAGTCTTTGCTGA. . . Reporter. . . TGCTGAAATT TTAAGCCGGCAATCAGAAGAGCAATCAGAAACGACT. . Gene 0. . ACGACTTTAA 0 2 0 This transducer has printed 020, which is 6 in base ten (610). Check: 18/3 = 6? Yes.
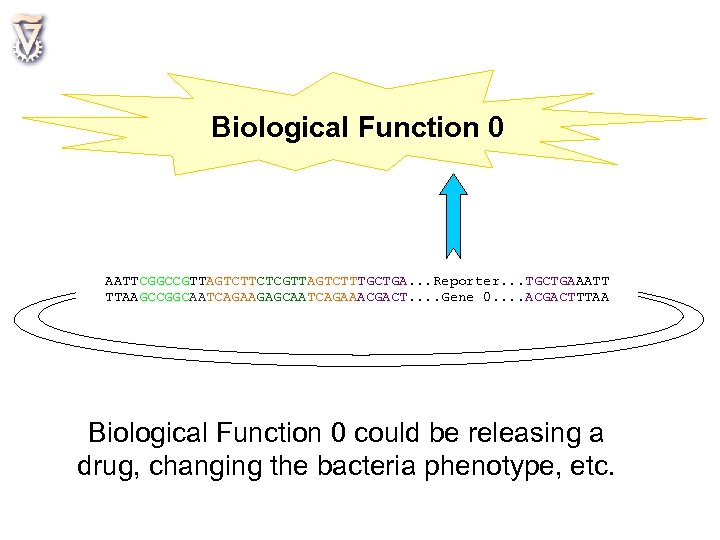
Biological Function 0 AATTCGGCCGTTAGTCTTCTCGTTAGTCTTTGCTGA. . . Reporter. . . TGCTGAAATT TTAAGCCGGCAATCAGAAGAGCAATCAGAAACGACT. . Gene 0. . ACGACTTTAA Biological Function 0 could be releasing a drug, changing the bacteria phenotype, etc.
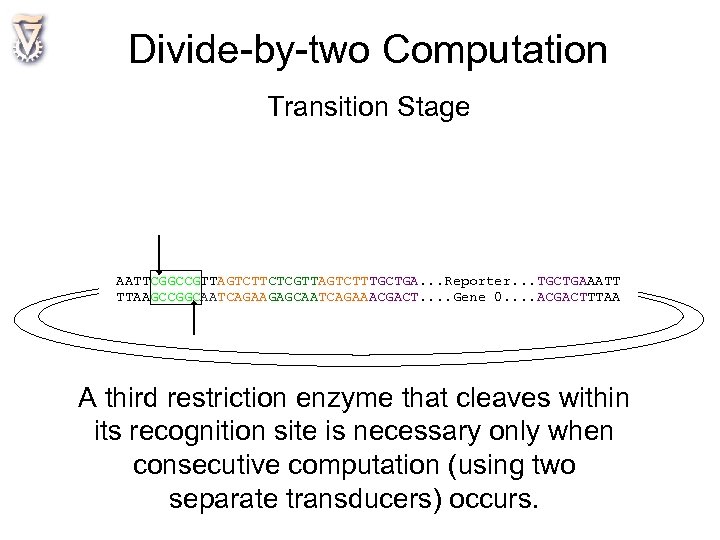
Divide-by-two Computation Transition Stage AATTCGGCCGTTAGTCTTCTCGTTAGTCTTTGCTGA. . . Reporter. . . TGCTGAAATT TTAAGCCGGCAATCAGAAGAGCAATCAGAAACGACT. . Gene 0. . ACGACTTTAA A third restriction enzyme that cleaves within its recognition site is necessary only when consecutive computation (using two separate transducers) occurs.
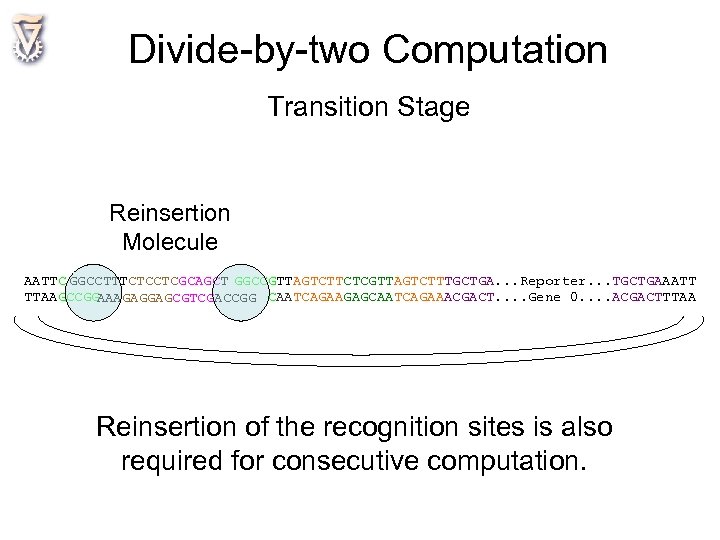
Divide-by-two Computation Transition Stage Reinsertion Molecule AATTC GGCCTTTCTCCTCGCAGCT GGCCGTTAGTCTTCTCGTTAGTCTTTGCTGA. . . Reporter. . . TGCTGAAATT TTAAGCCGG AAAGAGGAGCGTCGACCGG CAATCAGAAGAGCAATCAGAAACGACT. . Gene 0. . ACGACTTTAA Reinsertion of the recognition sites is also required for consecutive computation.
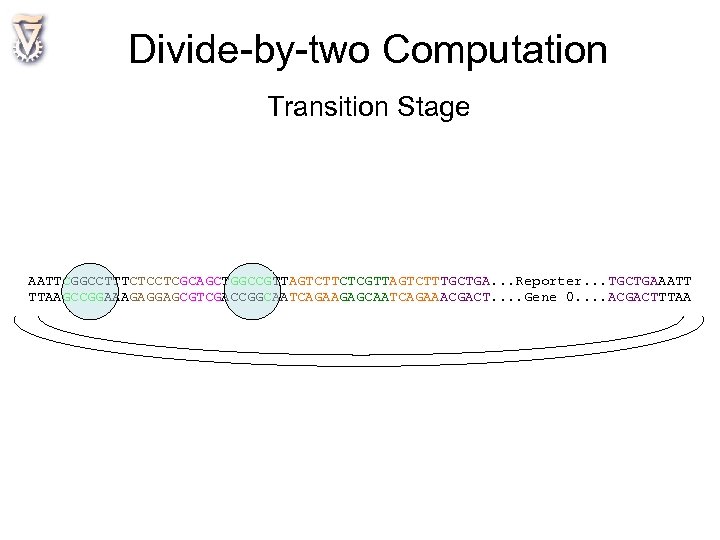
Divide-by-two Computation Transition Stage AATTCGGCCTTTCTCCTCGCAGCTGGCCGTTAGTCTTCTCGTTAGTCTTTGCTGA. . . Reporter. . . TGCTGAAATT TTAAGCCGGAAAGAGGAGCGTCGACCGGCAATCAGAAGAGCAATCAGAAACGACT. . Gene 0. . ACGACTTTAA
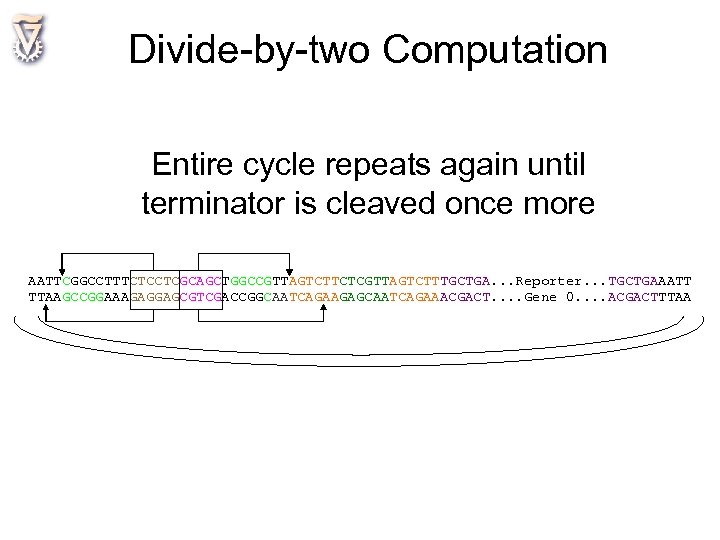
Divide-by-two Computation Entire cycle repeats again until terminator is cleaved once more AATTCGGCCTTTCTCCTCGCAGCTGGCCGTTAGTCTTCTCGTTAGTCTTTGCTGA. . . Reporter. . . TGCTGAAATT TTAAGCCGGAAAGAGGAGCGTCGACCGGCAATCAGAAGAGCAATCAGAAACGACT. . Gene 0. . ACGACTTTAA
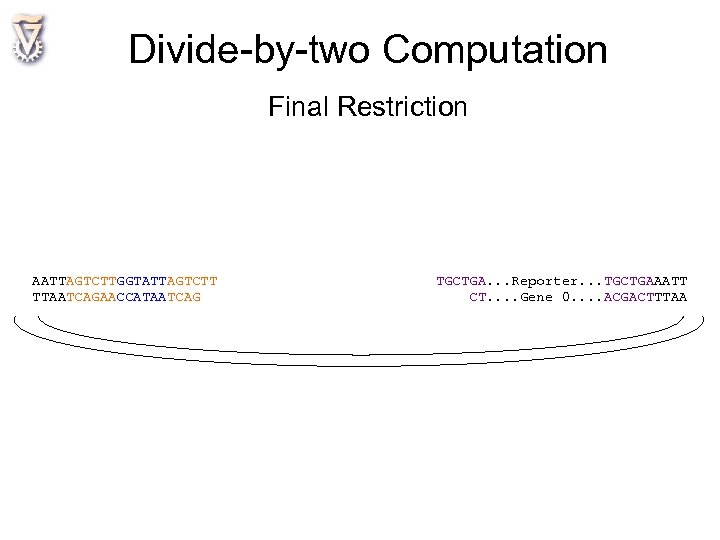

Divide-by-two Computation Final Restriction AATTAGTCTTGGTATTAGTCTT TTAATCAGAACCATAATCAG TGCTGA. . . Reporter. . . TGCTGAAATT CT. . Gene 0. . ACGACTTTAA
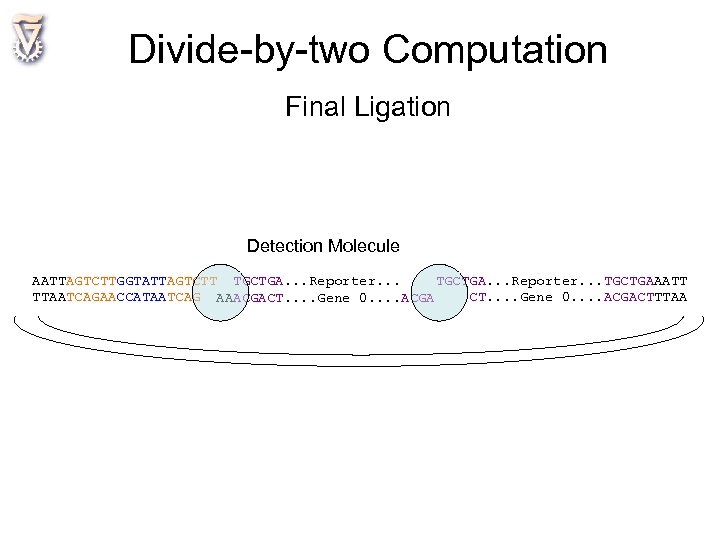
Divide-by-two Computation Final Ligation Detection Molecule AATTAGTCTTGGTATTAGTCTT TGCTGA. . . Reporter. . . TGCTGAAATT TTAATCAGAACCATAATCAG AAACGACT. . Gene 0. . ACGACTTTAA
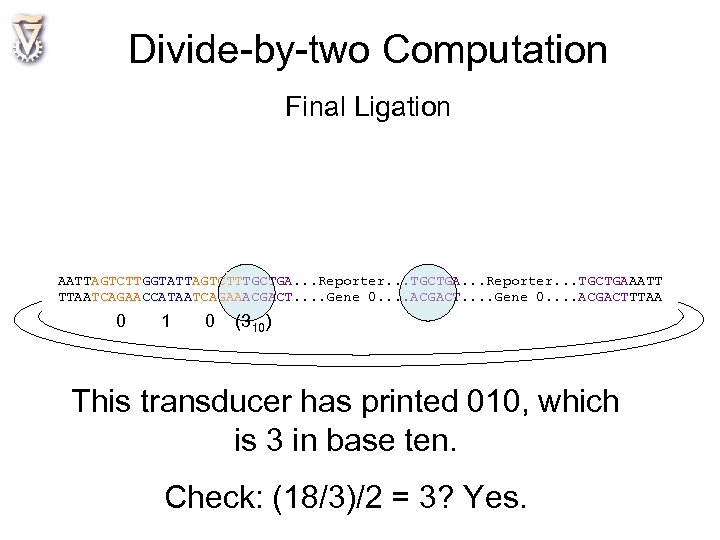
Divide-by-two Computation Final Ligation AATTAGTCTTGGTATTAGTCTTTGCTGA. . . Reporter. . . TGCTGAAATT TTAATCAGAACCATAATCAGAAACGACT. . . . Gene 0. . . . ACGACTTTAA 0 1 0 (310) This transducer has printed 010, which is 3 in base ten. Check: (18/3)/2 = 3? Yes.
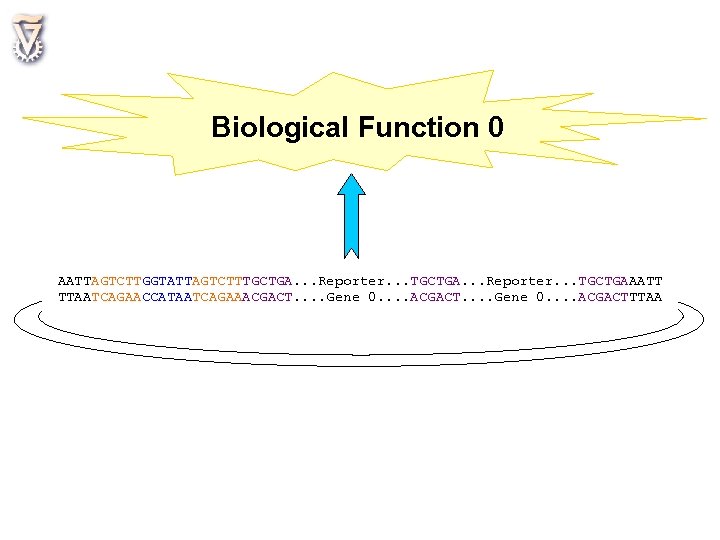
Biological Function 0 AATTAGTCTTGGTATTAGTCTTTGCTGA. . . Reporter. . . TGCTGAAATT TTAATCAGAACCATAATCAGAAACGACT. . . . Gene 0. . . . ACGACTTTAA

Discussion & Conclusions This project worked as expected. 18 ÷ 3= 6 ; 6 ÷ 2= 3 No molecule encoded the recognition site of an enzyme Proof of concept worked however not done in practicality. Transducers engineered functioned as coded

Acknowledgements n n We would like to sincerely thank Mr. Russell N. Stern for his generosity and donation. Thank you to the Louis Herman Israel Experience Fund for their contribution. We would also like to thank our mentor Tamar Ratner, for her continued dedication and help. Finally, we would like to thank Professor Ehud Keinan for allowing us to use his laboratory and his student.
590f8144bfaebfdad065632124927179.ppt