1e778fd6d627a6c07733dbc89ae44300.ppt
- Количество слайдов: 1
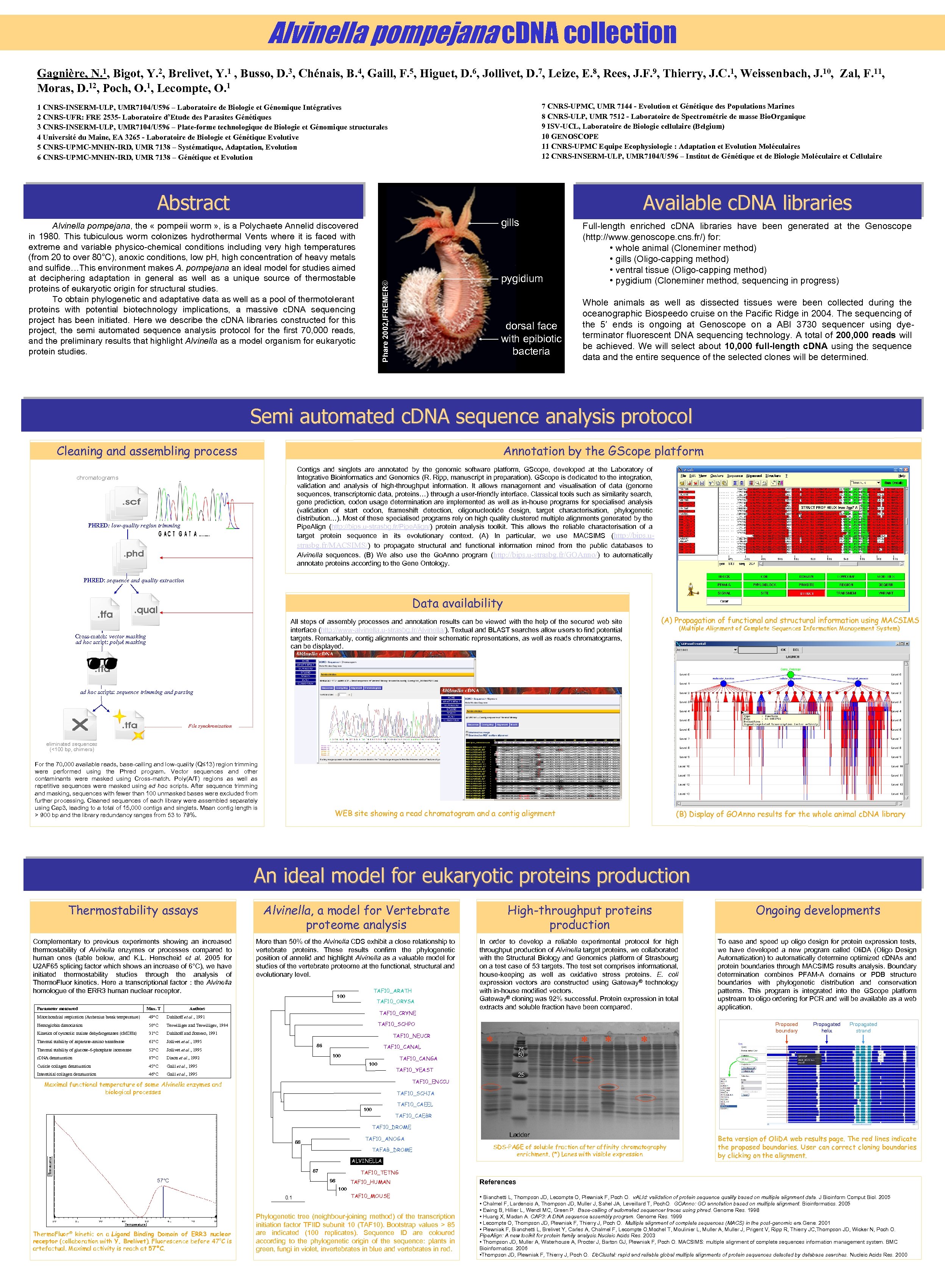 Alvinella pompejana c. DNA collection Gagnière, N. 1, Bigot, Y. 2, Brelivet, Y. 1 , Busso, D. 3, Chénais, B. 4, Gaill, F. 5, Higuet, D. 6, Jollivet, D. 7, Leize, E. 8, Rees, J. F. 9, Thierry, J. C. 1, Weissenbach, J. 10, Zal, F. 11, Moras, D. 12, Poch, O. 1, Lecompte, O. 1 7 CNRS-UPMC, UMR 7144 - Evolution et Génétique des Populations Marines 8 CNRS-ULP, UMR 7512 - Laboratoire de Spectrométrie de masse Bio. Organique 9 ISV-UCL, Laboratoire de Biologie cellulaire (Belgium) 10 GENOSCOPE 11 CNRS-UPMC Equipe Ecophysiologie : Adaptation et Evolution Moléculaires 12 CNRS-INSERM-ULP, UMR 7104/U 596 – Institut de Génétique et de Biologie Moléculaire et Cellulaire 1 CNRS-INSERM-ULP, UMR 7104/U 596 – Laboratoire de Biologie et Génomique Intégratives 2 CNRS-UFR: FRE 2535 - Laboratoire d’Etude des Parasites Génétiques 3 CNRS-INSERM-ULP, UMR 7104/U 596 – Plate-forme technologique de Biologie et Génomique structurales 4 Université du Maine, EA 3265 - Laboratoire de Biologie et Génétique Evolutive 5 CNRS-UPMC-MNHN-IRD, UMR 7138 – Systématique, Adaptation, Evolution 6 CNRS-UPMC-MNHN-IRD, UMR 7138 – Génétique et Evolution Abstract Available c. DNA libraries gills pygidium Phare 2002, IFREMER© Alvinella pompejana, the « pompeii worm » , is a Polychaete Annelid discovered in 1980. This tubiculous worm colonizes hydrothermal Vents where it is faced with extreme and variable physico-chemical conditions including very high temperatures (from 20 to over 80°C), anoxic conditions, low p. H, high concentration of heavy metals and sulfide…This environment makes A. pompejana an ideal model for studies aimed at deciphering adaptation in general as well as a unique source of thermostable proteins of eukaryotic origin for structural studies. To obtain phylogenetic and adaptative data as well as a pool of thermotolerant proteins with potential biotechnology implications, a massive c. DNA sequencing project has been initiated. Here we describe the c. DNA libraries constructed for this project, the semi automated sequence analysis protocol for the first 70, 000 reads, and the preliminary results that highlight Alvinella as a model organism for eukaryotic protein studies. dorsal face with epibiotic bacteria Full-length enriched c. DNA libraries have been generated at the Genoscope (http: //www. genoscope. cns. fr/) for: • whole animal (Cloneminer method) • gills (Oligo-capping method) • ventral tissue (Oligo-capping method) • pygidium (Cloneminer method, sequencing in progress) Whole animals as well as dissected tissues were been collected during the oceanographic Biospeedo cruise on the Pacific Ridge in 2004. The sequencing of the 5’ ends is ongoing at Genoscope on a ABI 3730 sequencer using dyeterminator fluorescent DNA sequencing technology. A total of 200, 000 reads will be achieved. We will select about 10, 000 full-length c. DNA using the sequence data and the entire sequence of the selected clones will be determined. Semi automated c. DNA sequence analysis protocol Annotation by the GScope platform Cleaning and assembling process Contigs and singlets are annotated by the genomic software platform, GScope, developed at the Laboratory of Integrative Bioinformatics and Genomics (R. Ripp, manuscript in preparation). GScope is dedicated to the integration, validation and analysis of high-throughput information. It allows management and visualisation of data (genome sequences, transcriptomic data, proteins…) through a user-friendly interface. Classical tools such as similarity search, gene prediction, codon usage determination are implemented as well as in-house programs for specialised analysis (validation of start codon, frameshift detection, oligonucleotide design, target characterisation, phylogenetic distribution…). Most of these specialised programs rely on high quality clustered multiple alignments generated by the Pipe. Align (http: //bips. u-strasbg. fr/Pipe. Align/) protein analysis toolkit. This allows the reliable characterisation of a target protein sequence in its evolutionary context. (A) In particular, we use MACSIMS (http: //bips. ustrasbg. fr/MACSIMS/) to propagate structural and functional information mined from the public databases to Alvinella sequences. (B) We also use the Go. Anno program (http: //bips. u-strasbg. fr/GOAnno/) to automatically annotate proteins according to the Gene Ontology. chromatograms PHRED: low-quality region trimming PHRED: sequence and quality extraction Data availability (A) Propagation of functional and structural information using MACSIMS All steps of assembly processes and annotation results can be viewed with the help of the secured web site interface (http: //www-alvinella. u-strasbg. fr/Alvinella/). Textual and BLAST searches allow users to find potential targets. Remarkably, contig alignments and their schematic representations, as well as reads chromatograms, can be displayed. Cross-match: vector masking ad hoc script: poly. A masking (Multiple Alignment of Complete Sequences Information Management System) ad hoc scripts: sequence trimming and parsing File synchronization eliminated sequences (<100 bp, chimera) For the 70, 000 available reads, base-calling and low-quality (Q≤ 13) region trimming were performed using the Phred program. Vector sequences and other contaminants were masked using Cross-match. Poly(A/T) regions as well as repetitive sequences were masked using ad hoc scripts. After sequence trimming and masking, sequences with fewer than 100 unmasked bases were excluded from further processing. Cleaned sequences of each library were assembled separately using Cap 3, leading to a total of 15, 000 contigs and singlets. Mean contig length is > 900 bp and the library redundancy ranges from 53 to 79%. WEB site showing a read chromatogram and a contig alignment (B) Display of GOAnno results for the whole animal c. DNA library An ideal model for eukaryotic proteins production Thermostability assays Alvinella, a model for Vertebrate proteome analysis High-throughput proteins production Ongoing developments Complementary to previous experiments showing an increased thermostability of Alvinella enzymes or processes compared to human ones (table below, and K. L. Henscheid et al. 2005 for U 2 AF 65 splicing factor which shows an increase of 6°C), we have initiated thermostability studies through the analysis of Thermo. Fluor kinetics. Here a transcriptional factor : the Alvinella homologue of the ERR 3 human nuclear receptor. More than 50% of the Alvinella CDS exhibit a close relationship to vertebrate proteins. These results confirm the phylogenetic position of annelid and highlight Alvinella as a valuable model for studies of the vertebrate proteome at the functional, structural and evolutionary level. In order to develop a reliable experimental protocol for high throughput production of Alvinella target proteins, we collaborated with the Structural Biology and Genomics platform of Strasbourg on a test case of 53 targets. The test set comprises informational, house-keeping as well as oxidative stress proteins. E. coli expression vectors are constructed using Gateway® technology with in-house modified vectors. Gateway® cloning was 92% successful. Protein expression in total extracts and soluble fraction have been compared. To ease and speed up oligo design for protein expression tests, we have developed a new program called Oli. DA (Oligo Design Automatization) to automatically determine optimized c. DNAs and protein boundaries through MACSIMS results analysis. Boundary determination combines PFAM-A domains or PDB structure boundaries with phylogenetic distribution and conservation patterns. This program is integrated into the GScope platform upstream to oligo ordering for PCR and will be available as a web application. Parameter measured Max. T TAF 10_ARATH 100 TAF 10_ORYSA Authors Mitochondrial respiration (Arrhenius break temperature) 49°C Hemoglobin dissociation 50°C Terwilliger and Terwilliger, 1984 Kinetics of cytosolic malate dehydrogenases (c. MDHs) 31°C Dahlhoff and Somero, 1991 Thermal stability of aspartate-amino transferase 61°C Jollivet et al. , 1995 Thermal stability of glucose-6 -phosphate isomerase 52°C Jollivet et al. , 1995 r. DNA denaturation 87°C TAF 10_CRYNE Dahlhoff et al. , 1991 Dixon et al. , 1992 Cuticle collagen denaturation 45°C 46°C TAF 10_NEUCR 86 TAF 10_CANAL 100 Gaill et al. , 1995 Interstitial collagen denaturation TAF 10_SCHPO Gaill et al. , 1995 * 75 50 TAF 10_CANGA TAF 10_YEAST * * Propagated helix Propagated strand 25 TAF 10_ENCCU Maximal functional temperature of some Alvinella enzymes and biological processes * Proposed boundary TAF 10_SCHJA TAF 10_CAEEL 100 TAF 10_CAEBR TAF 10_DROME TAF 10_ANOGA 88 TAFAB_DROME ALVINELLA 87 57°C Ladder SDS-PAGE of soluble fraction after affinity chromatography enrichment. (*) Lanes with visible expression Beta version of Oli. DA web results page. The red lines indicate the proposed boundaries. User can correct cloning boundaries by clicking on the alignment. TAF 10_TETNG 98 TAF 10_HUMAN References TAF 10_MOUSE • Bianchetti L, Thompson JD, Lecompte O, Plewniak F, Poch O. v. ALId: validation of protein sequence quality based on multiple alignment data. J Bioinform Comput Biol. 2005 100 0. 1 Thermo. Fluor® kinetic on a Ligand Binding Domain of ERR 3 nuclear receptor (collaboration with Y. Brelivet). Fluorescence before 47°C is artefactual. Maximal activity is reach at 57°C. Phylogenetic tree (neighbour-joining method) of the transcription initiation factor TFIID subunit 10 (TAF 10). Bootstrap values > 85 are indicated (100 replicates). Sequence ID are coloured according to the phylogenetic origin of the sequence: plants in green, fungi in violet, invertebrates in blue and vertebrates in red. • Chalmel F, Lardenois A, Thompson JD, Muller J, Sahel JA, Leveillard T, Poch. O. GOAnno: GO annotation based on multiple alignment. Bioinformatics. 2005 • Ewing B, Hillier L, Wendl MC, Green P. Base-calling of automated sequencer traces using phred. Genome Res. 1998 • Huang X, Madan A. CAP 3: A DNA sequence assembly program. Genome Res. 1999 • Lecompte O, Thompson JD, Plewniak F, Thierry J, Poch O. Multiple alignment of complete sequences (MACS) in the post-genomic era. Gene. 2001 • Plewniak F, Bianchetti L, Brelivet Y, Carles A, Chalmel F, Lecompte O, Mochel T, Moulinier L, Muller A, Muller J, Prigent V, Ripp R, Thierry JC, Thompson JD, Wicker N, Poch O. Pipe. Align: A new toolkit for protein family analysis. Nucleic Acids Res. 2003 • Thompson JD, Muller A, Waterhouse A, Procter J, Barton GJ, Plewniak F, Poch O. MACSIMS: multiple alignment of complete sequences information management system. BMC Bioinformatics. 2006 • Thompson JD, Plewniak F, Thierry J, Poch O. Db. Clustal: rapid and reliable global multiple alignments of protein sequences detected by database searches. Nucleic Acids Res. 2000
Alvinella pompejana c. DNA collection Gagnière, N. 1, Bigot, Y. 2, Brelivet, Y. 1 , Busso, D. 3, Chénais, B. 4, Gaill, F. 5, Higuet, D. 6, Jollivet, D. 7, Leize, E. 8, Rees, J. F. 9, Thierry, J. C. 1, Weissenbach, J. 10, Zal, F. 11, Moras, D. 12, Poch, O. 1, Lecompte, O. 1 7 CNRS-UPMC, UMR 7144 - Evolution et Génétique des Populations Marines 8 CNRS-ULP, UMR 7512 - Laboratoire de Spectrométrie de masse Bio. Organique 9 ISV-UCL, Laboratoire de Biologie cellulaire (Belgium) 10 GENOSCOPE 11 CNRS-UPMC Equipe Ecophysiologie : Adaptation et Evolution Moléculaires 12 CNRS-INSERM-ULP, UMR 7104/U 596 – Institut de Génétique et de Biologie Moléculaire et Cellulaire 1 CNRS-INSERM-ULP, UMR 7104/U 596 – Laboratoire de Biologie et Génomique Intégratives 2 CNRS-UFR: FRE 2535 - Laboratoire d’Etude des Parasites Génétiques 3 CNRS-INSERM-ULP, UMR 7104/U 596 – Plate-forme technologique de Biologie et Génomique structurales 4 Université du Maine, EA 3265 - Laboratoire de Biologie et Génétique Evolutive 5 CNRS-UPMC-MNHN-IRD, UMR 7138 – Systématique, Adaptation, Evolution 6 CNRS-UPMC-MNHN-IRD, UMR 7138 – Génétique et Evolution Abstract Available c. DNA libraries gills pygidium Phare 2002, IFREMER© Alvinella pompejana, the « pompeii worm » , is a Polychaete Annelid discovered in 1980. This tubiculous worm colonizes hydrothermal Vents where it is faced with extreme and variable physico-chemical conditions including very high temperatures (from 20 to over 80°C), anoxic conditions, low p. H, high concentration of heavy metals and sulfide…This environment makes A. pompejana an ideal model for studies aimed at deciphering adaptation in general as well as a unique source of thermostable proteins of eukaryotic origin for structural studies. To obtain phylogenetic and adaptative data as well as a pool of thermotolerant proteins with potential biotechnology implications, a massive c. DNA sequencing project has been initiated. Here we describe the c. DNA libraries constructed for this project, the semi automated sequence analysis protocol for the first 70, 000 reads, and the preliminary results that highlight Alvinella as a model organism for eukaryotic protein studies. dorsal face with epibiotic bacteria Full-length enriched c. DNA libraries have been generated at the Genoscope (http: //www. genoscope. cns. fr/) for: • whole animal (Cloneminer method) • gills (Oligo-capping method) • ventral tissue (Oligo-capping method) • pygidium (Cloneminer method, sequencing in progress) Whole animals as well as dissected tissues were been collected during the oceanographic Biospeedo cruise on the Pacific Ridge in 2004. The sequencing of the 5’ ends is ongoing at Genoscope on a ABI 3730 sequencer using dyeterminator fluorescent DNA sequencing technology. A total of 200, 000 reads will be achieved. We will select about 10, 000 full-length c. DNA using the sequence data and the entire sequence of the selected clones will be determined. Semi automated c. DNA sequence analysis protocol Annotation by the GScope platform Cleaning and assembling process Contigs and singlets are annotated by the genomic software platform, GScope, developed at the Laboratory of Integrative Bioinformatics and Genomics (R. Ripp, manuscript in preparation). GScope is dedicated to the integration, validation and analysis of high-throughput information. It allows management and visualisation of data (genome sequences, transcriptomic data, proteins…) through a user-friendly interface. Classical tools such as similarity search, gene prediction, codon usage determination are implemented as well as in-house programs for specialised analysis (validation of start codon, frameshift detection, oligonucleotide design, target characterisation, phylogenetic distribution…). Most of these specialised programs rely on high quality clustered multiple alignments generated by the Pipe. Align (http: //bips. u-strasbg. fr/Pipe. Align/) protein analysis toolkit. This allows the reliable characterisation of a target protein sequence in its evolutionary context. (A) In particular, we use MACSIMS (http: //bips. ustrasbg. fr/MACSIMS/) to propagate structural and functional information mined from the public databases to Alvinella sequences. (B) We also use the Go. Anno program (http: //bips. u-strasbg. fr/GOAnno/) to automatically annotate proteins according to the Gene Ontology. chromatograms PHRED: low-quality region trimming PHRED: sequence and quality extraction Data availability (A) Propagation of functional and structural information using MACSIMS All steps of assembly processes and annotation results can be viewed with the help of the secured web site interface (http: //www-alvinella. u-strasbg. fr/Alvinella/). Textual and BLAST searches allow users to find potential targets. Remarkably, contig alignments and their schematic representations, as well as reads chromatograms, can be displayed. Cross-match: vector masking ad hoc script: poly. A masking (Multiple Alignment of Complete Sequences Information Management System) ad hoc scripts: sequence trimming and parsing File synchronization eliminated sequences (<100 bp, chimera) For the 70, 000 available reads, base-calling and low-quality (Q≤ 13) region trimming were performed using the Phred program. Vector sequences and other contaminants were masked using Cross-match. Poly(A/T) regions as well as repetitive sequences were masked using ad hoc scripts. After sequence trimming and masking, sequences with fewer than 100 unmasked bases were excluded from further processing. Cleaned sequences of each library were assembled separately using Cap 3, leading to a total of 15, 000 contigs and singlets. Mean contig length is > 900 bp and the library redundancy ranges from 53 to 79%. WEB site showing a read chromatogram and a contig alignment (B) Display of GOAnno results for the whole animal c. DNA library An ideal model for eukaryotic proteins production Thermostability assays Alvinella, a model for Vertebrate proteome analysis High-throughput proteins production Ongoing developments Complementary to previous experiments showing an increased thermostability of Alvinella enzymes or processes compared to human ones (table below, and K. L. Henscheid et al. 2005 for U 2 AF 65 splicing factor which shows an increase of 6°C), we have initiated thermostability studies through the analysis of Thermo. Fluor kinetics. Here a transcriptional factor : the Alvinella homologue of the ERR 3 human nuclear receptor. More than 50% of the Alvinella CDS exhibit a close relationship to vertebrate proteins. These results confirm the phylogenetic position of annelid and highlight Alvinella as a valuable model for studies of the vertebrate proteome at the functional, structural and evolutionary level. In order to develop a reliable experimental protocol for high throughput production of Alvinella target proteins, we collaborated with the Structural Biology and Genomics platform of Strasbourg on a test case of 53 targets. The test set comprises informational, house-keeping as well as oxidative stress proteins. E. coli expression vectors are constructed using Gateway® technology with in-house modified vectors. Gateway® cloning was 92% successful. Protein expression in total extracts and soluble fraction have been compared. To ease and speed up oligo design for protein expression tests, we have developed a new program called Oli. DA (Oligo Design Automatization) to automatically determine optimized c. DNAs and protein boundaries through MACSIMS results analysis. Boundary determination combines PFAM-A domains or PDB structure boundaries with phylogenetic distribution and conservation patterns. This program is integrated into the GScope platform upstream to oligo ordering for PCR and will be available as a web application. Parameter measured Max. T TAF 10_ARATH 100 TAF 10_ORYSA Authors Mitochondrial respiration (Arrhenius break temperature) 49°C Hemoglobin dissociation 50°C Terwilliger and Terwilliger, 1984 Kinetics of cytosolic malate dehydrogenases (c. MDHs) 31°C Dahlhoff and Somero, 1991 Thermal stability of aspartate-amino transferase 61°C Jollivet et al. , 1995 Thermal stability of glucose-6 -phosphate isomerase 52°C Jollivet et al. , 1995 r. DNA denaturation 87°C TAF 10_CRYNE Dahlhoff et al. , 1991 Dixon et al. , 1992 Cuticle collagen denaturation 45°C 46°C TAF 10_NEUCR 86 TAF 10_CANAL 100 Gaill et al. , 1995 Interstitial collagen denaturation TAF 10_SCHPO Gaill et al. , 1995 * 75 50 TAF 10_CANGA TAF 10_YEAST * * Propagated helix Propagated strand 25 TAF 10_ENCCU Maximal functional temperature of some Alvinella enzymes and biological processes * Proposed boundary TAF 10_SCHJA TAF 10_CAEEL 100 TAF 10_CAEBR TAF 10_DROME TAF 10_ANOGA 88 TAFAB_DROME ALVINELLA 87 57°C Ladder SDS-PAGE of soluble fraction after affinity chromatography enrichment. (*) Lanes with visible expression Beta version of Oli. DA web results page. The red lines indicate the proposed boundaries. User can correct cloning boundaries by clicking on the alignment. TAF 10_TETNG 98 TAF 10_HUMAN References TAF 10_MOUSE • Bianchetti L, Thompson JD, Lecompte O, Plewniak F, Poch O. v. ALId: validation of protein sequence quality based on multiple alignment data. J Bioinform Comput Biol. 2005 100 0. 1 Thermo. Fluor® kinetic on a Ligand Binding Domain of ERR 3 nuclear receptor (collaboration with Y. Brelivet). Fluorescence before 47°C is artefactual. Maximal activity is reach at 57°C. Phylogenetic tree (neighbour-joining method) of the transcription initiation factor TFIID subunit 10 (TAF 10). Bootstrap values > 85 are indicated (100 replicates). Sequence ID are coloured according to the phylogenetic origin of the sequence: plants in green, fungi in violet, invertebrates in blue and vertebrates in red. • Chalmel F, Lardenois A, Thompson JD, Muller J, Sahel JA, Leveillard T, Poch. O. GOAnno: GO annotation based on multiple alignment. Bioinformatics. 2005 • Ewing B, Hillier L, Wendl MC, Green P. Base-calling of automated sequencer traces using phred. Genome Res. 1998 • Huang X, Madan A. CAP 3: A DNA sequence assembly program. Genome Res. 1999 • Lecompte O, Thompson JD, Plewniak F, Thierry J, Poch O. Multiple alignment of complete sequences (MACS) in the post-genomic era. Gene. 2001 • Plewniak F, Bianchetti L, Brelivet Y, Carles A, Chalmel F, Lecompte O, Mochel T, Moulinier L, Muller A, Muller J, Prigent V, Ripp R, Thierry JC, Thompson JD, Wicker N, Poch O. Pipe. Align: A new toolkit for protein family analysis. Nucleic Acids Res. 2003 • Thompson JD, Muller A, Waterhouse A, Procter J, Barton GJ, Plewniak F, Poch O. MACSIMS: multiple alignment of complete sequences information management system. BMC Bioinformatics. 2006 • Thompson JD, Plewniak F, Thierry J, Poch O. Db. Clustal: rapid and reliable global multiple alignments of protein sequences detected by database searches. Nucleic Acids Res. 2000


