c0cfed8d18c37be42ba79a865bd1b748.ppt
- Количество слайдов: 21

Alexander N. Glazer and Hiroshi Nikaido Microbial Biotechnology Fundamentals of Applied Microbiology, Second Edition Chapter 1: Picturing Distributions with Graphs Copyright © 2007 by Alexander N. Glazer and Hiroshi Nikaido
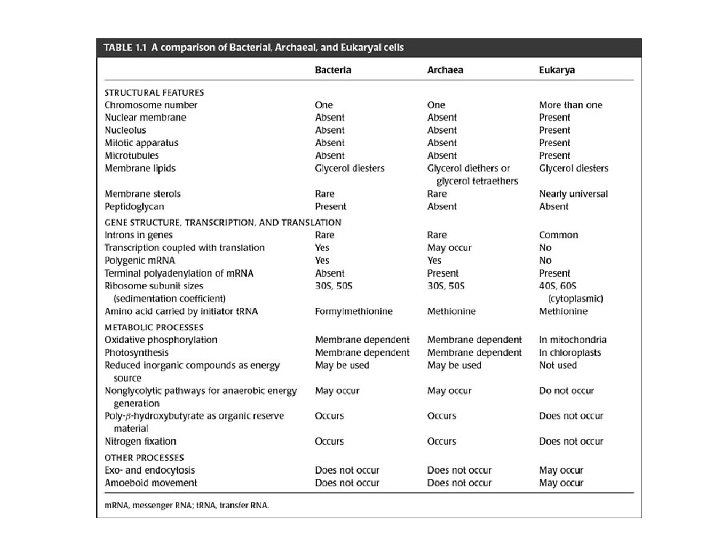
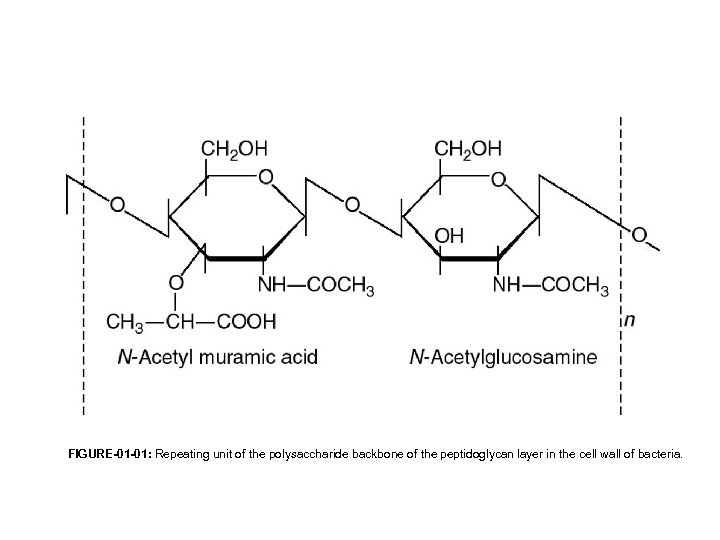
FIGURE-01 -01: Repeating unit of the polysaccharide backbone of the peptidoglycan layer in the cell wall of bacteria.
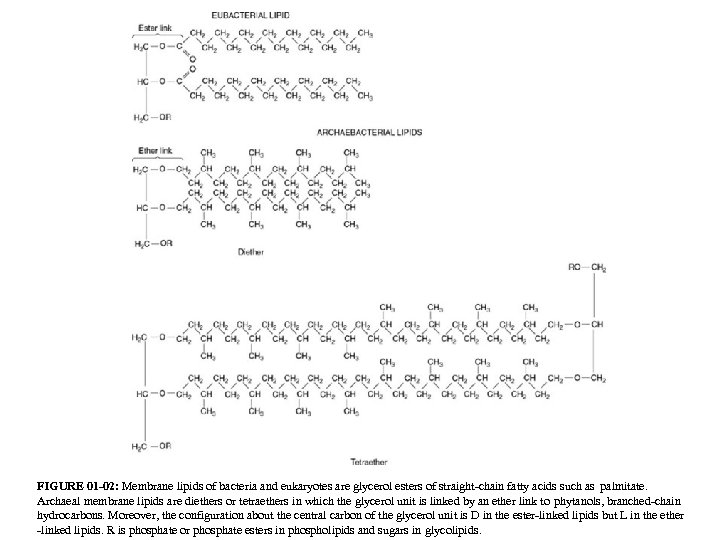
FIGURE 01 -02: Membrane lipids of bacteria and eukaryotes are glycerol esters of straight-chain fatty acids such as palmitate. Archaeal membrane lipids are diethers or tetraethers in which the glycerol unit is linked by an ether link to phytanols, branched-chain hydrocarbons. Moreover, the configuration about the central carbon of the glycerol unit is D in the ester-linked lipids but L in the ether -linked lipids. R is phosphate or phosphate esters in phospholipids and sugars in glycolipids.
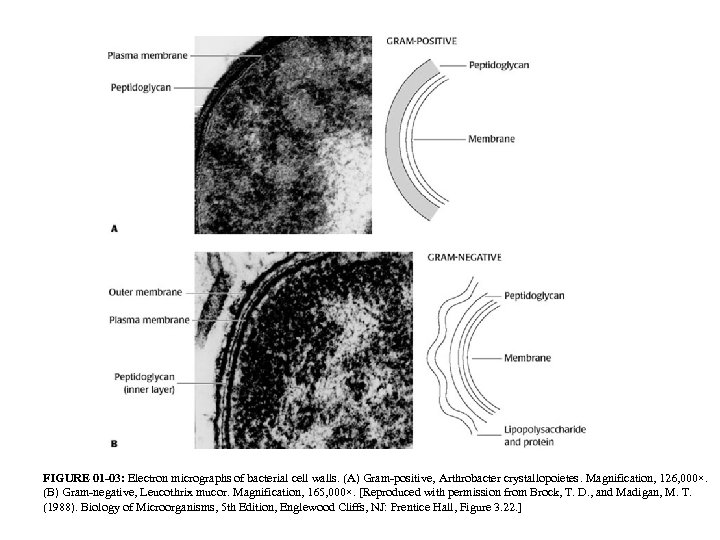
FIGURE 01 -03: Electron micrographs of bacterial cell walls. (A) Gram-positive, Arthrobacter crystallopoietes. Magnification, 126, 000×. (B) Gram-negative, Leucothrix mucor. Magnification, 165, 000×. [Reproduced with permission from Brock, T. D. , and Madigan, M. T. (1988). Biology of Microorganisms, 5 th Edition, Englewood Cliffs, NJ: Prentice Hall, Figure 3. 22. ]
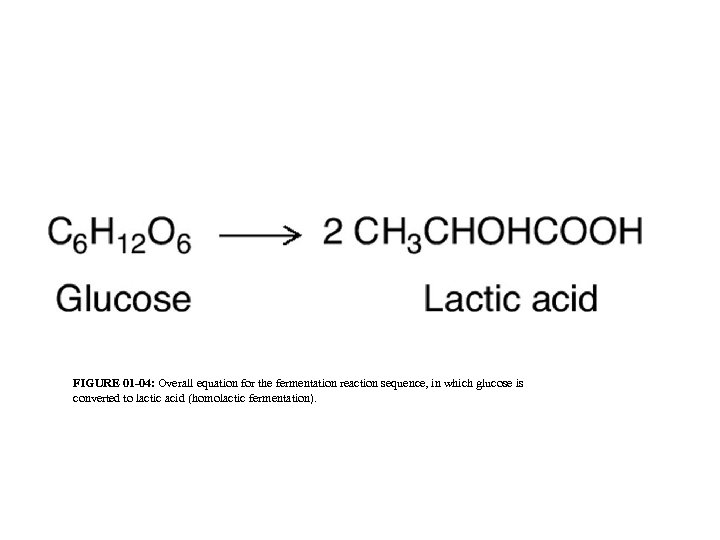
FIGURE 01 -04: Overall equation for the fermentation reaction sequence, in which glucose is converted to lactic acid (homolactic fermentation).
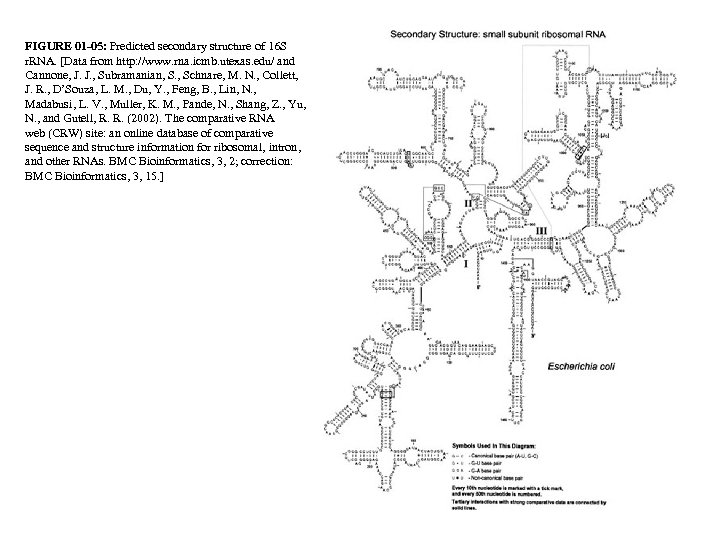
FIGURE 01 -05: Predicted secondary structure of 16 S r. RNA. [Data from http: //www. rna. icmb. utexas. edu/ and Cannone, J. J. , Subramanian, S. , Schnare, M. N. , Collett, J. R. , D’Souza, L. M. , Du, Y. , Feng, B. , Lin, N. , Madabusi, L. V. , Muller, K. M. , Pande, N. , Shang, Z. , Yu, N. , and Gutell, R. R. (2002). The comparative RNA web (CRW) site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. BMC Bioinformatics, 3, 2; correction: BMC Bioinformatics, 3, 15. ]
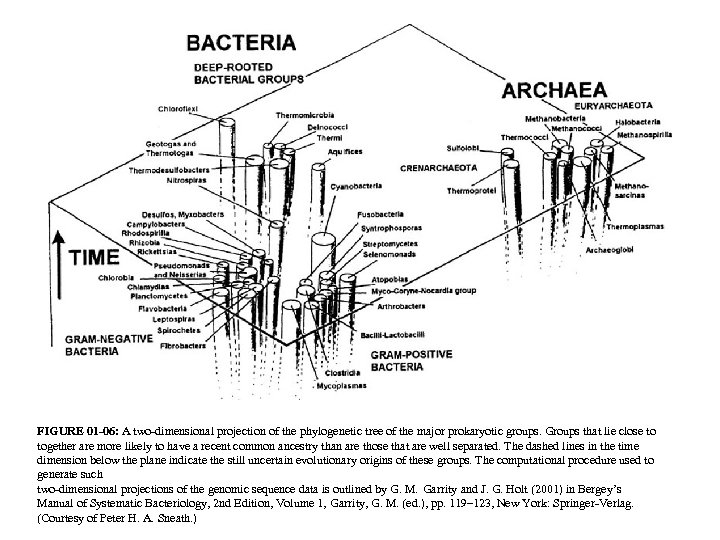
FIGURE 01 -06: A two-dimensional projection of the phylogenetic tree of the major prokaryotic groups. Groups that lie close to together are more likely to have a recent common ancestry than are those that are well separated. The dashed lines in the time dimension below the plane indicate the still uncertain evolutionary origins of these groups. The computational procedure used to generate such two-dimensional projections of the genomic sequence data is outlined by G. M. Garrity and J. G. Holt (2001) in Bergey’s Manual of Systematic Bacteriology, 2 nd Edition, Volume 1, Garrity, G. M. (ed. ), pp. 119– 123, New York: Springer-Verlag. (Courtesy of Peter H. A. Sneath. )
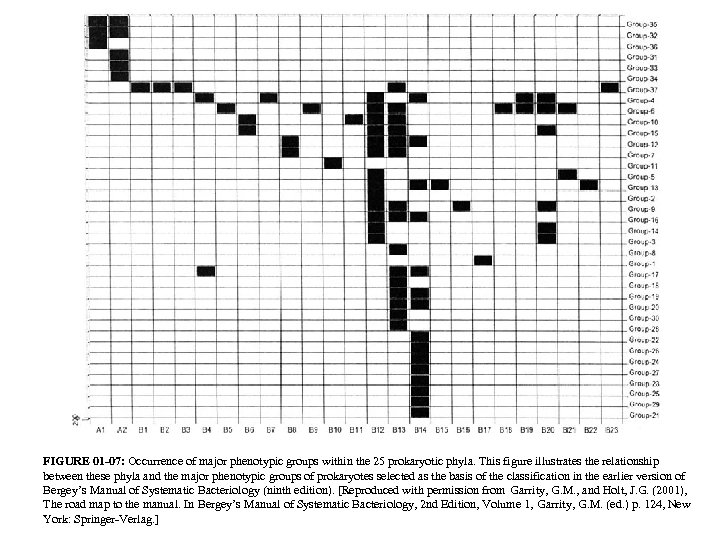
FIGURE 01 -07: Occurrence of major phenotypic groups within the 25 prokaryotic phyla. This figure illustrates the relationship between these phyla and the major phenotypic groups of prokaryotes selected as the basis of the classification in the earlier version of Bergey’s Manual of Systematic Bacteriology (ninth edition). [Reproduced with permission from Garrity, G. M. , and Holt, J. G. (2001), The road map to the manual. In Bergey’s Manual of Systematic Bacteriology, 2 nd Edition, Volume 1, Garrity, G. M. (ed. ) p. 124, New York: Springer-Verlag. ]
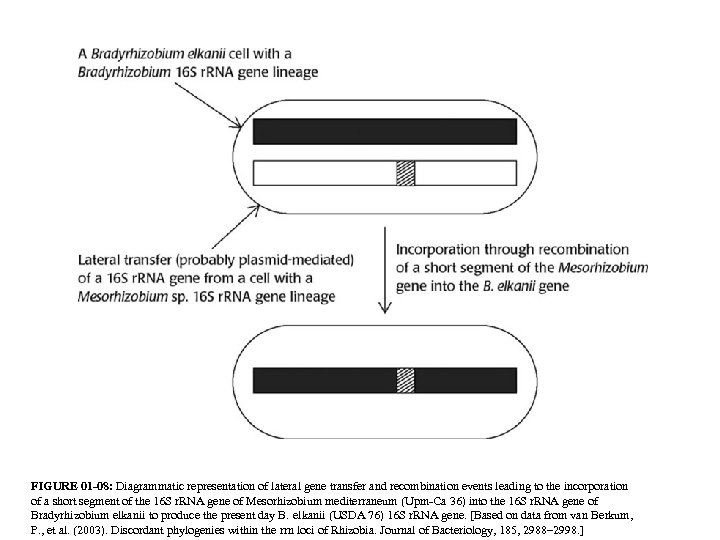
FIGURE 01 -08: Diagrammatic representation of lateral gene transfer and recombination events leading to the incorporation of a short segment of the 16 S r. RNA gene of Mesorhizobium mediterraneum (Upm-Ca 36) into the 16 S r. RNA gene of Bradyrhizobium elkanii to produce the present day B. elkanii (USDA 76) 16 S r. RNA gene. [Based on data from van Berkum, P. , et al. (2003). Discordant phylogenies within the rrn loci of Rhizobia. Journal of Bacteriology, 185, 2988– 2998. ]
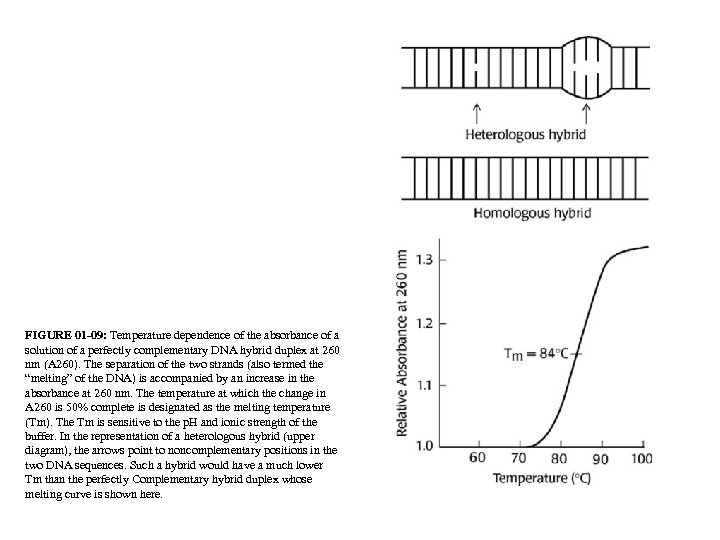
FIGURE 01 -09: Temperature dependence of the absorbance of a solution of a perfectly complementary DNA hybrid duplex at 260 nm (A 260). The separation of the two strands (also termed the “melting” of the DNA) is accompanied by an increase in the absorbance at 260 nm. The temperature at which the change in A 260 is 50% complete is designated as the melting temperature (Tm). The Tm is sensitive to the p. H and ionic strength of the buffer. In the representation of a heterologous hybrid (upper diagram), the arrows point to noncomplementary positions in the two DNA sequences. Such a hybrid would have a much lower Tm than the perfectly Complementary hybrid duplex whose melting curve is shown here.
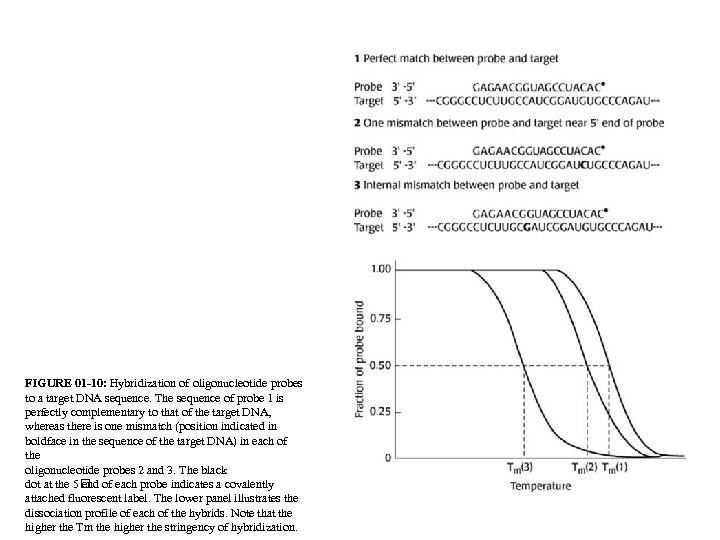
FIGURE 01 -10: Hybridization of oligonucleotide probes to a target DNA sequence. The sequence of probe 1 is perfectly complementary to that of the target DNA, whereas there is one mismatch (position indicated in boldface in the sequence of the target DNA) in each of the oligonucleotide probes 2 and 3. The black dot at the 5 of each probe indicates a covalently end attached fluorescent label. The lower panel illustrates the dissociation profile of each of the hybrids. Note that the higher the Tm the higher the stringency of hybridization.
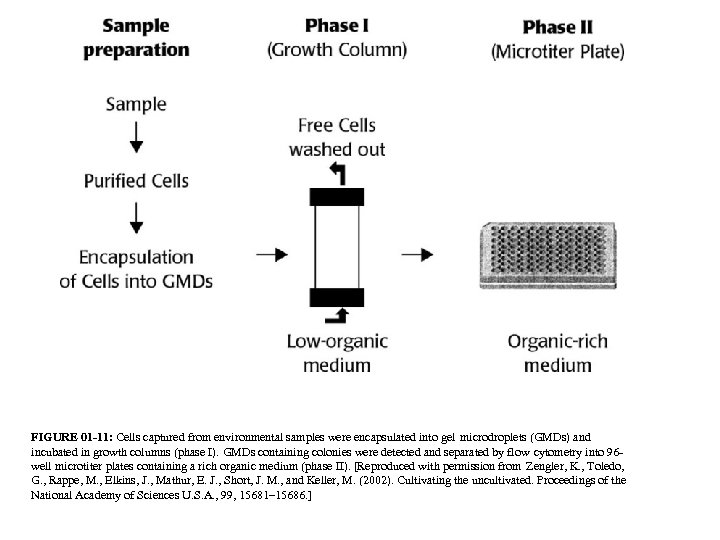
FIGURE 01 -11: Cells captured from environmental samples were encapsulated into gel microdroplets (GMDs) and incubated in growth columns (phase I). GMDs containing colonies were detected and separated by flow cytometry into 96 well microtiter plates containing a rich organic medium (phase II). [Reproduced with permission from Zengler, K. , Toledo, G. , Rappe, M. , Elkins, J. , Mathur, E. J. , Short, J. M. , and Keller, M. (2002). Cultivating the uncultivated. Proceedings of the National Academy of Sciences U. S. A. , 99, 15681– 15686. ]
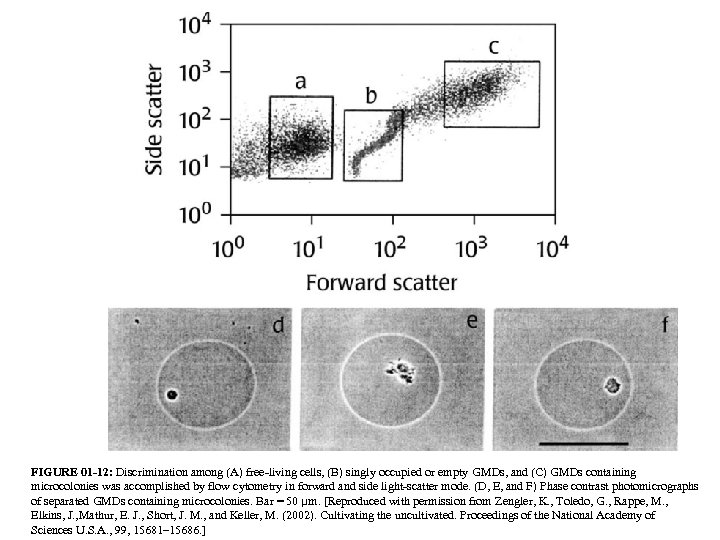
FIGURE 01 -12: Discrimination among (A) free-living cells, (B) singly occupied or empty GMDs, and (C) GMDs containing microcolonies was accomplished by flow cytometry in forward and side light-scatter mode. (D, E, and F) Phase contrast photomicrographs of separated GMDs containing microcolonies. Bar = 50 μm. [Reproduced with permission from Zengler, K. , Toledo, G. , Rappe, M. , Elkins, J. , Mathur, E. J. , Short, J. M. , and Keller, M. (2002). Cultivating the uncultivated. Proceedings of the National Academy of Sciences U. S. A. , 99, 15681– 15686. ]
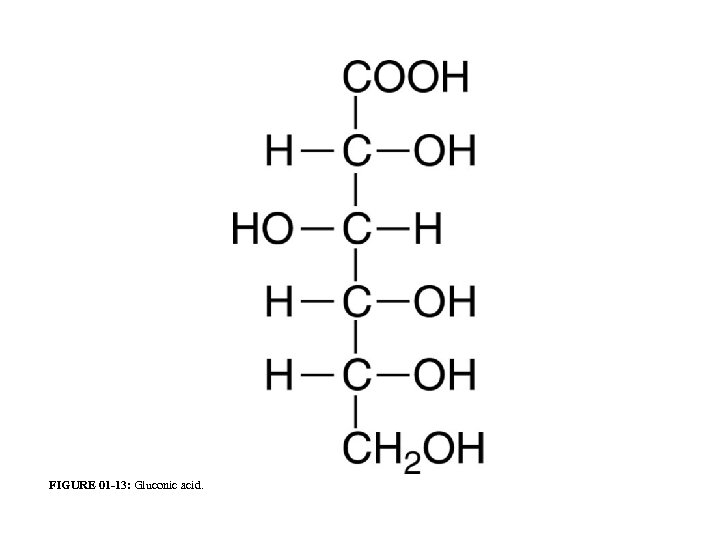
FIGURE 01 -13: Gluconic acid.
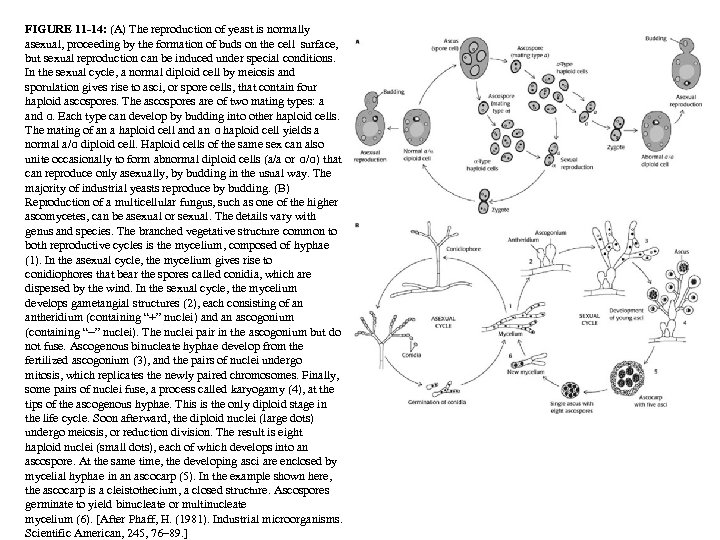
FIGURE 11 -14: (A) The reproduction of yeast is normally asexual, proceeding by the formation of buds on the cell surface, but sexual reproduction can be induced under special conditions. In the sexual cycle, a normal diploid cell by meiosis and sporulation gives rise to asci, or spore cells, that contain four haploid ascospores. The ascospores are of two mating types: a and α. Each type can develop by budding into other haploid cells. The mating of an a haploid cell and an α haploid cell yields a normal a/α diploid cell. Haploid cells of the same sex can also unite occasionally to form abnormal diploid cells (a/a or α/α) that can reproduce only asexually, by budding in the usual way. The majority of industrial yeasts reproduce by budding. (B) Reproduction of a multicellular fungus, such as one of the higher ascomycetes, can be asexual or sexual. The details vary with genus and species. The branched vegetative structure common to both reproductive cycles is the mycelium, composed of hyphae (1). In the asexual cycle, the mycelium gives rise to conidiophores that bear the spores called conidia, which are dispersed by the wind. In the sexual cycle, the mycelium develops gametangial structures (2), each consisting of an antheridium (containing “+” nuclei) and an ascogonium (containing “−” nuclei). The nuclei pair in the ascogonium but do not fuse. Ascogenous binucleate hyphae develop from the fertilized ascogonium (3), and the pairs of nuclei undergo mitosis, which replicates the newly paired chromosomes. Finally, some pairs of nuclei fuse, a process called karyogamy (4), at the tips of the ascogenous hyphae. This is the only diploid stage in the life cycle. Soon afterward, the diploid nuclei (large dots) undergo meiosis, or reduction division. The result is eight haploid nuclei (small dots), each of which develops into an ascospore. At the same time, the developing asci are enclosed by mycelial hyphae in an ascocarp (5). In the example shown here, the ascocarp is a cleistothecium, a closed structure. Ascospores germinate to yield binucleate or multinucleate mycelium (6). [After Phaff, H. (1981). Industrial microorganisms. Scientific American, 245, 76– 89. ]
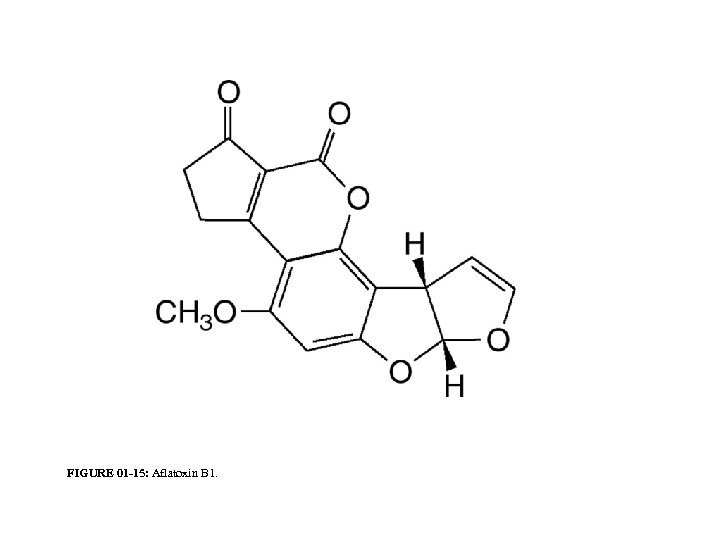
FIGURE 01 -15: Aflatoxin B 1.
c0cfed8d18c37be42ba79a865bd1b748.ppt