024e9a8a4e7ff78e44a3d2dbcbbb9deb.ppt
- Количество слайдов: 22

Af. CS Microscopy Laboratory Department of Molecular Pharmacology Stanford University Tobias Meyer Lead Scientists: Nancy O’Rourke Grischa Chandy Mary Verghese Liz Gehrig Wei. Sun Park Sarah Lim Takako Mukai Won Do Heo Jen Liou

The two Goals of the Af. CS Microscopy Core 1. To identify the location of signal transduction processes by investigating subcellular localization and by monitoring receptor triggered translocation processes 2. To dissect dynamic properties of signaling networks by using perturbation strategies combined with fluorescence microscopy based screening tools (antibodies, fluorescent probes, FRET and protein translocation assays)

Fluorescence microscopy strategies Why fluorescence microscopy? 1. To measure subcellular localization of signaling proteins 2. To monitor receptor triggered translocation of signaling proteins (GFP conjugation) 3. To measure dynamic signaling parameters such as delays, cooperativity, coincidence 4. Microscopy-based readouts of signaling processes enable one to perform large scale perturbations screens using RNAi, dominant negative and constitutively active signaling constructs (for example translocation or FRET as readout)

Requirements for effective microscopy studies in the Af. CS cell model A. Cells require distinguishable nuclei, cytosol, organelles, membrane structures (flat cells and morphologically differentiated cells are best suited for microscopy) B. Cells have to be adherent C. Cells have to be readily transfectable with expression constructs and RNAi molecules (or constructs) D. Single cell assays have to be available to measure important receptor-triggered cell functions in these cells Raw cells seem to be OK

The strategy for measuring protein localization is based on dual fluorescence imaging using proteins of interest versus marker proteins Subcellular markers are based on expression constructs of CFP and YFP conjugated marker and signaling proteins as well as on dual immunostaining using antibodies A marker set of expressable CFP-YFPconstructs has been developed (and keeps being refined) Quantitative co-localization tools still need to be further developed
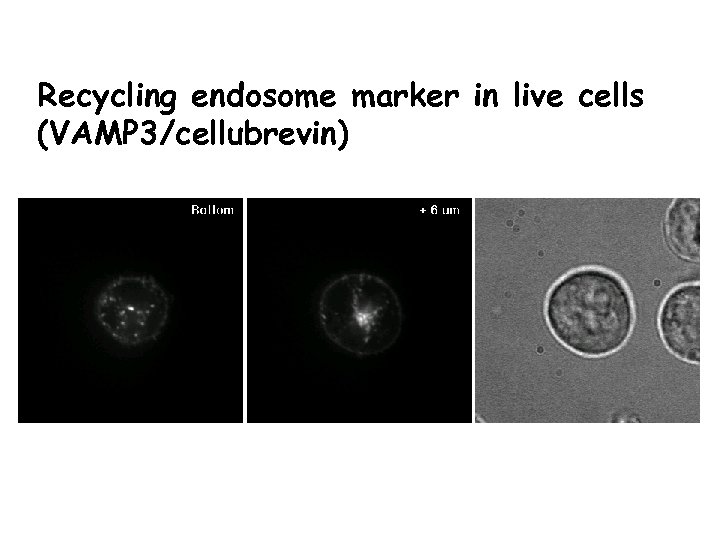
Recycling endosome marker in live cells (VAMP 3/cellubrevin)
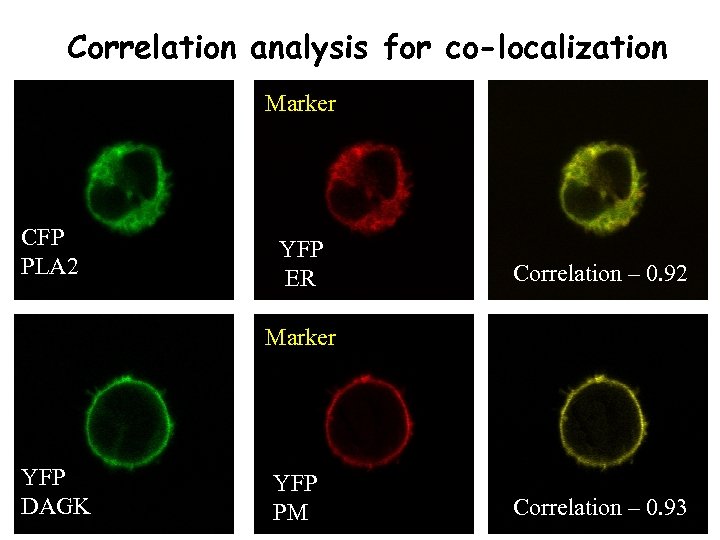
Correlation analysis for co-localization Marker CFP PLA 2 YFP ER Correlation – 0. 92 Marker YFP DAGK YFP PM Correlation – 0. 93
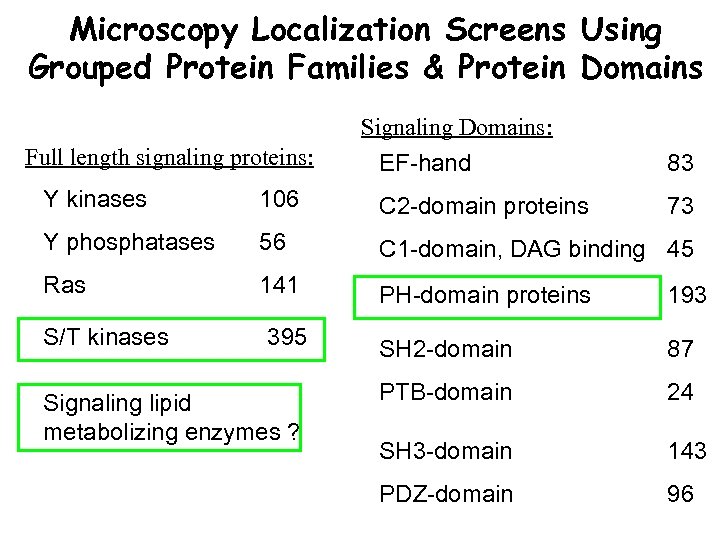
Microscopy Localization Screens Using Grouped Protein Families & Protein Domains Signaling Domains: Full length signaling proteins: EF-hand 83 73 Y kinases 106 C 2 -domain proteins Y phosphatases 56 C 1 -domain, DAG binding 45 Ras 141 PH-domain proteins 193 395 SH 2 -domain 87 Signaling lipid metabolizing enzymes ? PTB-domain 24 SH 3 -domain 143 PDZ-domain 96 S/T kinases
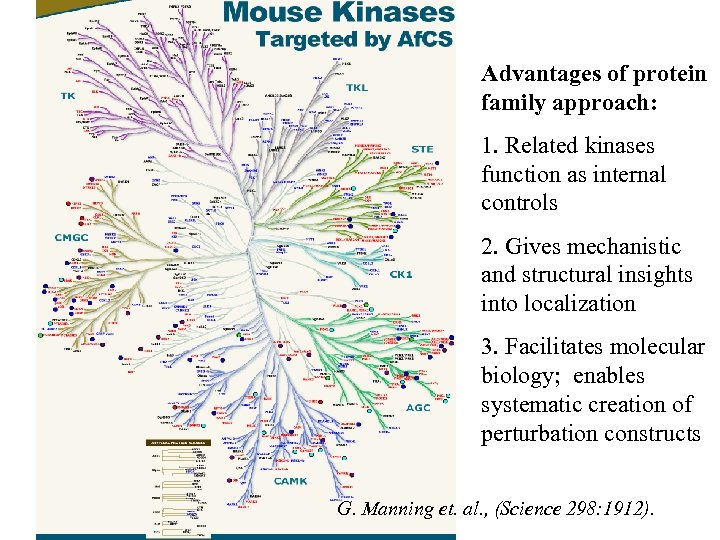
Advantages of protein family approach: 1. Related kinases function as internal controls 2. Gives mechanistic and structural insights into localization 3. Facilitates molecular biology; enables systematic creation of perturbation constructs G. Manning et. al. , (Science 298: 1912).
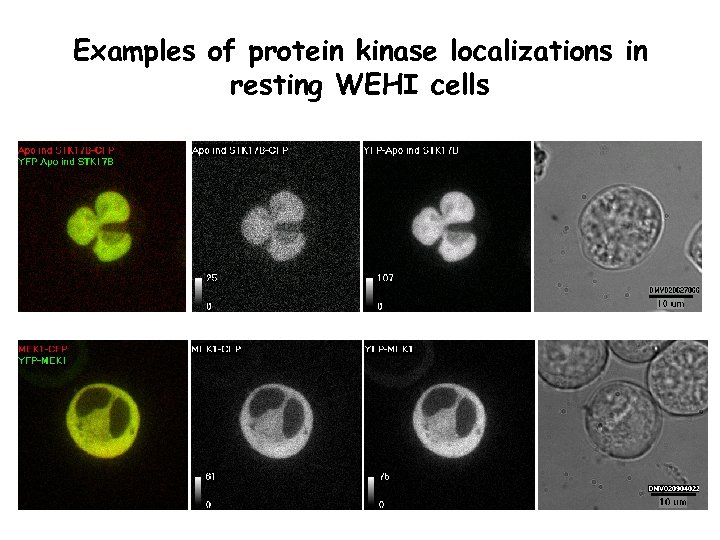
Examples of protein kinase localizations in resting WEHI cells
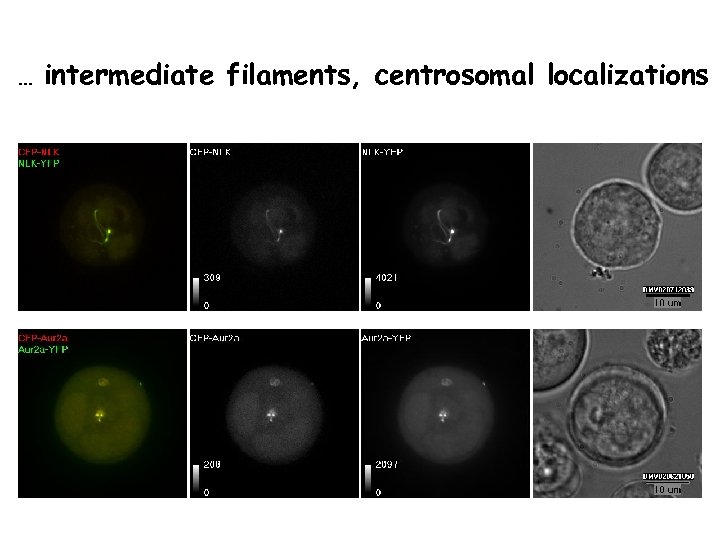
… intermediate filaments, centrosomal localizations

Why GFP conjugated protein domains? 1. To understand mechanisms of localization 2. To identify minimal fluorescent translocation biosensors 3. Also useful as perturbation constructs C 1 -domains monitor localized diacylglycerol signals C 2 -domains monitor local Ca 2+/PS-signals PH-domains monitor local changes in phosphoinositide concentration SH 2 -domains monitor local tyrosine phosphorylation
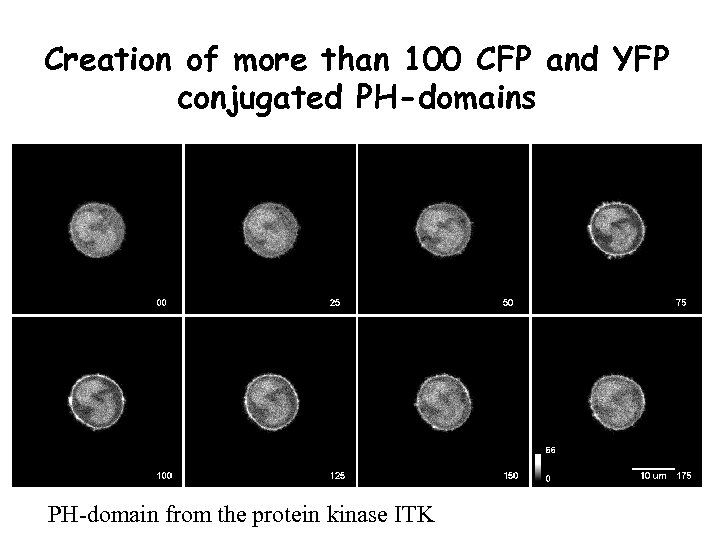
Creation of more than 100 CFP and YFP conjugated PH-domains PH-domain from the protein kinase ITK

Challenges for the Af. CS Now that the parts list of cells and connection maps start to exist, two things are needed: 1. Measurements of dynamic systems parameters 2. Measurement protocols of sufficiently high throughput to dissect the modular structure of signaling network 3. Single-cell fluorescence measurements are ideally suited for: 1. Measuring rates, bistability, cooperativity, delays, coincidence and other dynamic signaling parameters 2. High Throughput Perturbation Screens to define pathways, cross-talk and modular structures in signaling networks
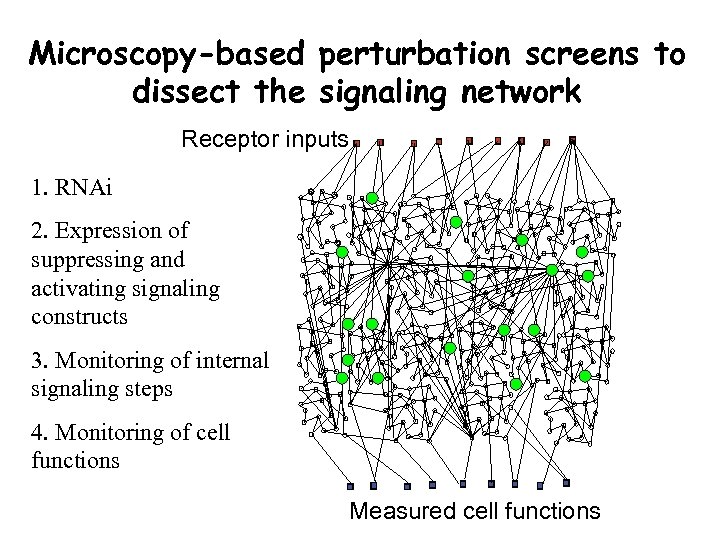
Microscopy-based perturbation screens to dissect the signaling network Receptor inputs 1. RNAi 2. Expression of suppressing and activating signaling constructs 3. Monitoring of internal signaling steps 4. Monitoring of cell functions Measured cell functions

Fluorescent single cell perturbation screens Why is a perturbation strategy required to understand signaling networks? Perturbation screens are the only way to assess the relevance of different players in a signaling system What are the advantages of single cell assays? 1. Identification of signaling states, oscillations, delay times, coincidence measurements 2. Partial cell transfection does not reduce the quality of the data; improved signal-to-noise What are perturbation strategies that the Af. CS should employ? 1. RNAi 2. Expression of perturbation constructs 3. Small molecule inhibitors and activators
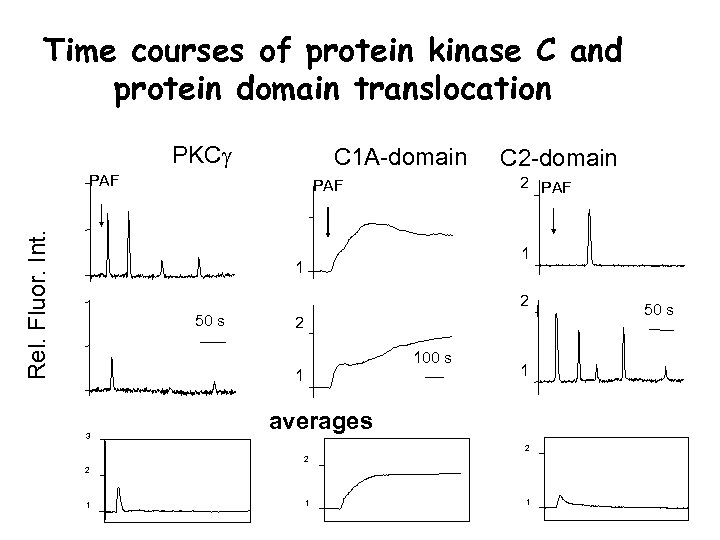
Time courses of protein kinase C and protein domain translocation PKCg C 1 A-domain Rel. Fluor. Int. PAF C 2 -domain 2 PAF 1 1 2 50 s 2 100 s 1 3 1 averages 2 2 2 1 1 1. 2 50 s
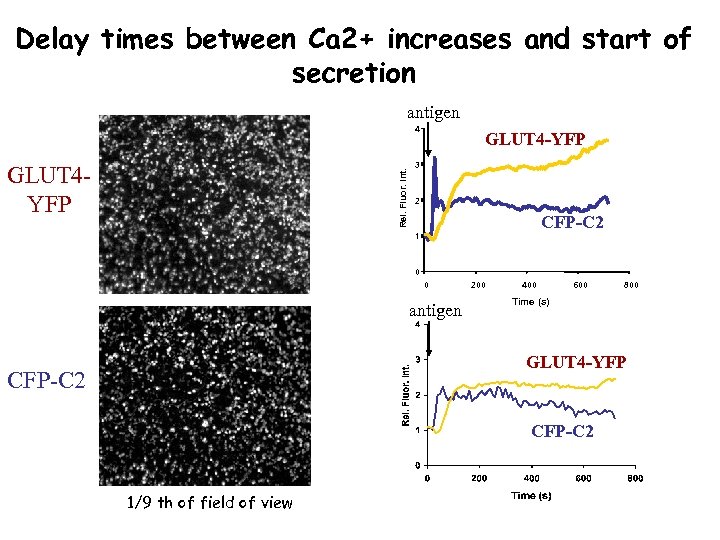
Delay times between Ca 2+ increases and start of secretion antigen 4 Rel. Fluor. Int. GLUT 4 YFP GLUT 4 -YFP 3 2 CFP-C 2 1 0 0 antigen 200 400 600 800 Time (s) GLUT 4 -YFP CFP-C 2 1/9 th of field of view

Readouts for a microscopy-based perturbation screen Fixed-cell assays (higher throughput) Using activation-dependent and other antibodies For example: phospecific antibodies, nuclear localization, increased protein expression, polymerized actin, cell size Live-cell assays (lower throughput) 1. Protein translocation (for example to nucleus or to plasma membrane) 2. FRET or small molecule biosensors 3. Functional assays such as cell migration, exocytosisendocytosis, phagacytosis

Overall strategy for a microscopy based perturbation screen Fixed and live cell fluorescence imaging assays: Existing assays as well as future assays developed by Cell lab, Antibody lab & Microscopy lab Perturbation expression constructs: These can be created by systematically mutating signaling proteins in a protein family (for example S/T protein kinase) as well as by creating sets of constructs that express signaling protein domains (for example PH-domain) RNAi screens: For microscopy based assays, this can be done using a new in vitro dicing technique developed by James Ferrell’s group that significantly cuts the cost of RNAi and enables high throughput microscopy-based perturbation screens

Dissecting the signaling network: Perturbation constructs Why using perturbation constructs based on proteins that are low abundant or absent in a cell type? Every perturbation construct that affects a pathway is (like a specific drug) a tool to dissect the structure of a signaling network Example: K to R mutation in the ATP binding pocket often yields a dominant negative enzyme; a large set of these protein kinase mutants has been created (320 currently planned) Constitutively active kinases can be created for many protein kinases and hundreds of other perturbation constructs can be made using available signaling literature

Specific steps to implement a microscopy-based perturbation screen for signaling networks Proposal: 1. Automated image acquisition and analysis. This microscope for fixed cell assays could be acquired immediately using existing funds in this years budget of the microscopy core ($170’ 000). 2. Creation of a large set of perturbation constructs (~ 20003000). These constructs can be made using existing literature. The time for completion is ~ 2 years using collaborative efforts. The cost is ~ $250 per construct. 3. Creation of a mouse library of in vitro diced RNAi for all known mouse signaling proteins (3000 -4000 constructs). This can be done by 3 people in 1 year at a cost of less than $300’ 000
024e9a8a4e7ff78e44a3d2dbcbbb9deb.ppt