0dd6036afd1d8bfe6892be420651c564.ppt
- Количество слайдов: 41

A Systems Biology Approach to Cancer Therapy and Early Detection Avijit Ghosh Dhruv Pant, Rui Zou, Dave Miller Yihua Wang, Hanbing Lin Travis Hoppe, Aparna Kumar Department of Physics Drexel University

Outline l Signal Transduction l l l Mutations, Oncogenes, Proto-Oncogenes Mathematical Models l l l l D. Pant A. Ghosh (2005) Biophys. Chem 113(3) 275 -288 The Results l l Enzymes Mutations Predicting Putative Oncogenes using Systems Biology l l Overview Experimentally Validated Results New Predictions Drugs! Spatio-Temporal Modeling Cell. Sim Overview Special topics on Electrical and Systems Engineering 2007 Systems Biology
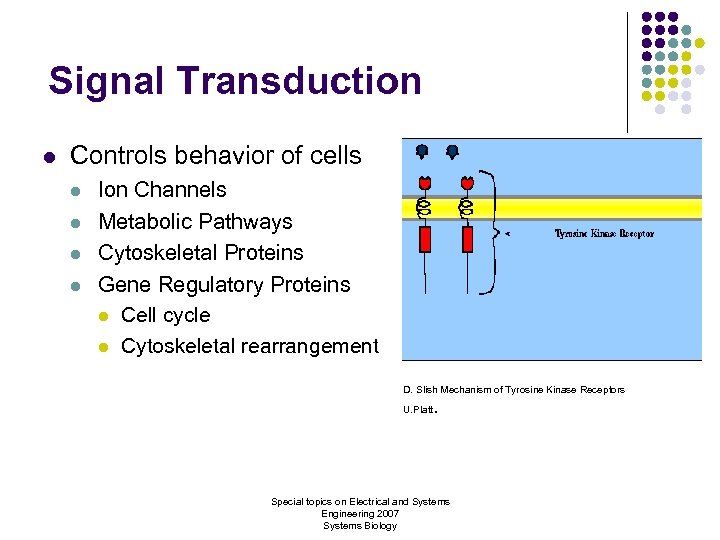
Signal Transduction l Controls behavior of cells l l Ion Channels Metabolic Pathways Cytoskeletal Proteins Gene Regulatory Proteins l Cell cycle l Cytoskeletal rearrangement D. Slish Mechanism of Tyrosine Kinase Receptors . U. Platt Special topics on Electrical and Systems Engineering 2007 Systems Biology
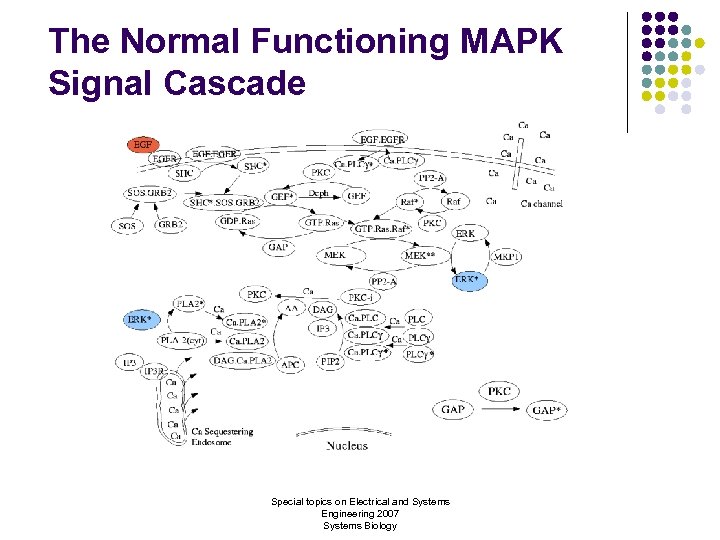
The Normal Functioning MAPK Signal Cascade Special topics on Electrical and Systems Engineering 2007 Systems Biology
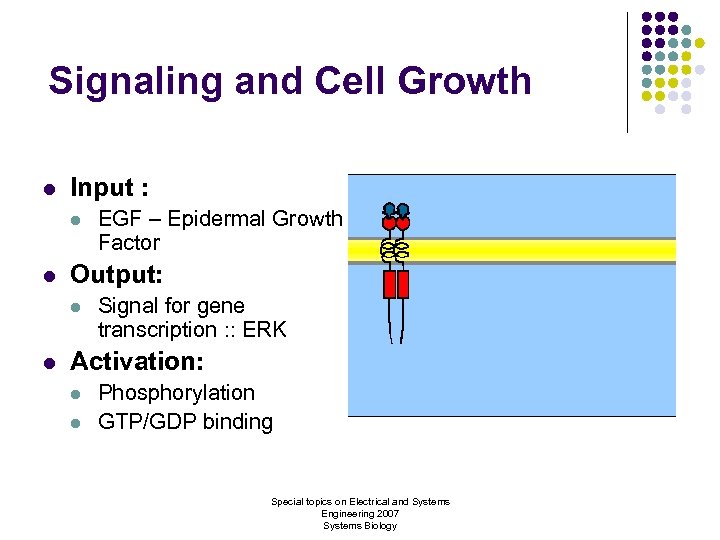
Signaling and Cell Growth l Input : l l Output: l l EGF – Epidermal Growth Factor Signal for gene transcription : : ERK Activation: l l Phosphorylation GTP/GDP binding Special topics on Electrical and Systems Engineering 2007 Systems Biology
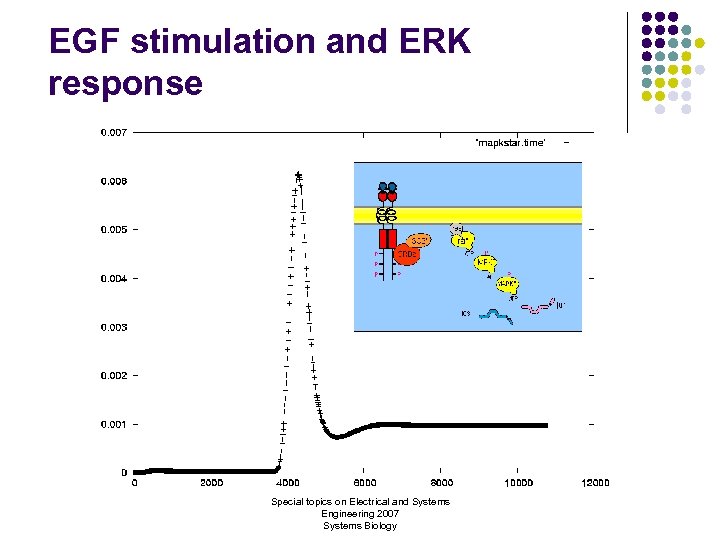
EGF stimulation and ERK response Special topics on Electrical and Systems Engineering 2007 Systems Biology

Oncogenes and Tumor Suppressor Genes l Oncogene - A gene (protein) that when mutated or inappropriately expressed contributes to cancer formation l Mutations in signal transduction genes Mutations in Tumor Suppressor genes l INHERITED l l Activation l l Point Mutation Chromosomal rearrangement Gene amplification Viral Insertion Special topics on Electrical and Systems Engineering 2007 Systems Biology
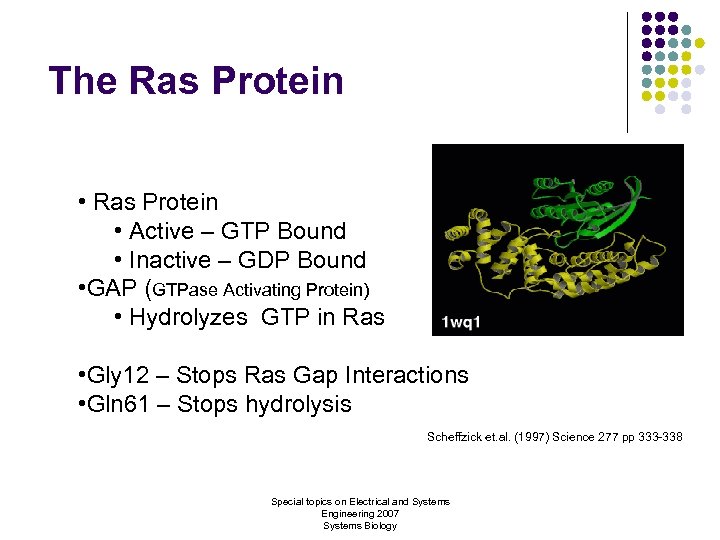

The Ras Protein • Active – GTP Bound • Inactive – GDP Bound • GAP (GTPase Activating Protein) • Hydrolyzes GTP in Ras • Gly 12 – Stops Ras Gap Interactions • Gln 61 – Stops hydrolysis Scheffzick et. al. (1997) Science 277 pp 333 -338 Special topics on Electrical and Systems Engineering 2007 Systems Biology

Interactions Special topics on Electrical and Systems Engineering 2007 Systems Biology

EGF / MAPK / PLC-γ/ PKC / Calcium / PLA 2 Pathways # # locations # the volume is a optional parameter # our volume units are L location extracellular 1 E-10 Ca-ext location cytosol 1 E-12 location endosome 1. 6 E-13 Ca-sequester # Channels channel capacitive_Ca_entry* Ca-ext -> Ca : : 0. 005 channel Ca-leak-from-extracell Ca-ext -> Ca : : 0. 004 channel Ca-leak-to-cytoplasm Ca-sequester->Ca : : 8 channel IP 3 R* Ca-sequester -> Ca : : 19. 2 # conservation. Reactions #conserve Ca. EPump : : Ca_Ca. EPump #conserve Ca. Transp-2 Ca : : Ca. Transp #conserve m. Glu. R : : Rec-Glu-Gq Rec-Gq Blocked-rec-Gq #conserve PLC : : match /PLC-/ #conserve PLA 2 -cytosolic : : match /PLA 2/ # sum up #sum PKC-active : : PKC-DAG-AA* PKC-Ca-memb* PKC-Ca-AA* PKC-DAG-memb* # sum. Reactions # (native units are #/compartment) Ca + Ca. EPump <> Ca_Ca. EPump : : 0. 003 288 Ca_Ca. EPump -> Ca. EPump +Ca-ext : : 72 Ca. PLC_g + L. EGFR <> Ca. PLC_g_L. EGFR : : 5 e-06 0. 8 Ca. PLC_g_L. EGFR -> L. EGFR +Ca. PLC_g* : : 0. 2 SHC + L. EGFR <> SHC_L. EGFR : : 2 e-06 0. 8 SHC_L. EGFR -> L. EGFR +SHC* : : 0. 2 MAPK + MAPKK* <> MAPK_MAPKK* : : 2. 7 e-05 0. 6 MAPK_MAPKK* -> MAPKK* +MAPK-tyr : : 0. 15 MAPK-tyr + MAPKK* <> MAPK-tyr_MAPKK* : : 2. 7 e-05 0. 6 MAPK-tyr_MAPKK* -> MAPKK* +MAPK* : : 0. 15 MAPKK + Raf-GTP-Ras* <> MAPKK_Raf-GTP-Ras* : : 5. 5 e-06 0. 42 MAPKK_Raf-GTP-Ras* -> Raf-GTP-Ras* +MAPKK-ser : : 0. 105 MAPKK-ser + Raf-GTP-Ras* <> MAPKK-ser_Raf-GTP-Ras* : : 5. 5 e-06 0. 42 MAPKK-ser_Raf-GTP-Ras* -> Raf-GTP-Ras* +MAPKK* : : 0. 105 #craf-1 + PKC-active <> craf-1_PKC-active : : 5 e-07 16 #craf-1_PKC-active -> PKC-active +craf-1* : : 4 #GAP + PKC-active <> GAP_PKC-active : : 1 e-05 16 #GAP_PKC-active -> PKC-active +GAP* : : 4 #inact-GEF + PKC-active <> inact-GEF_PKC-active : : 1 e-05 16 #inact-GEF_PKC-active -> PKC-active +GEF* : : 4 craf-1 + PKC-DAG-AA* <> craf-1_PKC-DAG-AA* : : 5 e-07 16 craf-1_PKC-DAG-AA* -> PKC-DAG-AA* +craf-1* : : 4 GAP + PKC-DAG-AA* <> GAP_PKC-DAG-AA* : : 1 e-05 16 GAP_PKC-DAG-AA* -> PKC-DAG-AA* +GAP* : : 4 inact-GEF + PKC-DAG-AA* <> inact-GEF_PKC-DAG-AA* : : 1 e-05 16 inact-GEF_PKC-DAG-AA* -> PKC-DAG-AA* +GEF* : : 4 craf-1 + PKC-Ca-memb* <> craf-1_PKC-Ca-memb* : : 5 e-07 16 craf-1_PKC-Ca-memb* -> PKC-Ca-memb* +craf-1* : : 4 GAP + PKC-Ca-memb* <> GAP_PKC-Ca-memb* : : 1 e-05 16 GAP_PKC-Ca-memb* -> PKC-Ca-memb* +GAP* : : 4 inact-GEF + PKC-Ca-memb* <> inact-GEF_PKC-Ca-memb* : : 1 e-05 16 inact-GEF_PKC-Ca-memb* -> PKC-Ca-memb* +GEF* : : 4 craf-1 + PKC-Ca-AA* <> craf-1_PKC-Ca-AA* : : 5 e-07 16 craf-1_PKC-Ca-AA* -> PKC-Ca-AA* +craf-1* : : 4 GAP + PKC-Ca-AA* <> GAP_PKC-Ca-AA* : : 1 e-05 16 GAP_PKC-Ca-AA* -> PKC-Ca-AA* +GAP* : : 4 inact-GEF + PKC-Ca-AA* <> inact-GEF_PKC-Ca-AA* : : 1 e-05 16 inact-GEF_PKC-Ca-AA* -> PKC-Ca-AA* +GEF* : : 4 craf-1 + PKC-DAG-memb* <> craf-1_PKC-DAG-memb* : : 5 e-07 16 craf-1_PKC-DAG-memb* -> PKC-DAG-memb* +craf-1* : : 4 GAP + PKC-DAG-memb* <> GAP_PKC-DAG-memb* : : 1 e-05 16 GAP_PKC-DAG-memb* -> PKC-DAG-memb* +GAP* : : 4 inact-GEF + PKC-DAG-memb* <> inact-GEF_PKC-DAG-memb* : : 1 e-05 16 inact-GEF_PKC-DAG-memb* -> PKC-DAG-memb* +GEF* : : 4 craf-1 + PKC-basal* <> craf-1_PKC-basal* : : 5 e-07 16 craf-1_PKC-basal* -> PKC-basal* +craf-1* : : 4 GAP + PKC-basal* <> GAP_PKC-basal* : : 1 e-05 16 GAP_PKC-basal* -> PKC-basal* +GAP* : : 4 inact-GEF + PKC-basal* <> inact-GEF_PKC-basal* : : 1 e-05 16 inact-GEF_PKC-basal* -> PKC-basal* +GEF* : : 4 craf-1 + PKC-AA* <> craf-1_PKC-AA* : : 5 e-07 16 craf-1_PKC-AA* -> PKC-AA* +craf-1* : : 4 GAP + PKC-AA* <> GAP_PKC-AA* : : 1 e-05 16 GAP_PKC-AA* -> PKC-AA* +GAP* : : 4 inact-GEF + PKC-AA* <> inact-GEF_PKC-AA* : : 1 e-05 16 inact-GEF_PKC-AA* -> PKC-AA* +GEF* : : 4 # end expand APC + PLA 2 -Ca* <> APC_PLA 2 -Ca* : : 2. 25 e-06 21. 6 APC_PLA 2 -Ca* -> PLA 2 -Ca* +AA : : 5. 4 APC + PIP 2 -PLA 2* <> APC_PIP 2 -PLA 2* : : 4. 6 e-06 44. 16 APC_PIP 2 -PLA 2* -> PIP 2 -PLA 2* +AA : : 11. 04 APC + PIP 2 -Ca-PLA 2* <> APC_PIP 2 -Ca-PLA 2* : : 1. 5 e-05 144 APC_PIP 2 -Ca-PLA 2* -> PIP 2 -Ca-PLA 2* +AA : : 36 APC + DAG-Ca-PLA 2* <> APC_DAG-Ca-PLA 2* : : 2. 5 e-05 240 APC_DAG-Ca-PLA 2* -> DAG-Ca-PLA 2* +AA : : 60 APC + PLA 2*-Ca <> APC_PLA 2*-Ca : : 5 e-05 480 APC_PLA 2*-Ca -> PLA 2*-Ca +AA : : 120 PLA 2 -cytosolic + MAPK* <> PLA 2 -cytosolic_MAPK* : : 6. 5 e-06 80 PLA 2 -cytosolic_MAPK* -> MAPK* +PLA 2* : : 20 craf-1* + MAPK* <> craf-1*_MAPK* : : 3. 25 e-06 40 craf-1*_MAPK* -> MAPK* +craf-1** : : 10 Sos + MAPK* <> Sos_MAPK* : : 3. 25 e-05 40 Sos_MAPK* -> MAPK* +Sos* : : 10 MAPK-tyr + MKP-1 <> MAPK-tyr_MKP-1 : : 0. 000125 4 MAPK-tyr_MKP-1 -> MKP-1 +MAPK : : 1 MAPK* + MKP-1 <> MAPK*_MKP-1 : : 0. 000125 4 MAPK*_MKP-1 -> MKP-1 +MAPK-tyr : : 1 craf-1* + PPhosphatase 2 A <> craf-1*_PPhosphatase 2 A : : 3. 3 e-06 25 craf-1*_PPhosphatase 2 A -> PPhosphatase 2 A +craf-1 : : 6 MAPKK* + PPhosphatase 2 A <> MAPKK*_PPhosphatase 2 A : : 3. 3 e-06 25 MAPKK*_PPhosphatase 2 A -> PPhosphatase 2 A +MAPKK-ser : : 6 MAPKK-ser + PPhosphatase 2 A <> MAPKK-ser_PPhosphatase 2 A : : 3. 3 e-06 25 MAPKK-ser_PPhosphatase 2 A -> PPhosphatase 2 A +MAPKK : : 6 craf-1** + PPhosphatase 2 A <> craf-1**_PPhosphatase 2 A : : 3. 3 e-06 25 craf-1**_PPhosphatase 2 A -> PPhosphatase 2 A +craf-1* : : 6 inact-GEF + PKA-active <> inact-GEF_PKA-active : : 1 e-05 36 inact-GEF_PKA-active -> PKA-active +inact-GEF* : : 9 GDP-Ras + Shc*. Sos. Grb 2 <> GDP-Ras_Shc*. Sos. Grb 2 : : 3. 3 e-07 0. 08 GDP-Ras_Shc*. Sos. Grb 2 -> Shc*. Sos. Grb 2 +GTP-Ras : : 0. 02 PIP 2 + PLC-Ca <> PIP 2_PLC-Ca : : 4. 2 e-06 40 PIP 2_PLC-Ca -> PLC-Ca +DAG + IP 3 : : 10 PIP 2 + PLC-Ca-Gq <> PIP 2_PLC-Ca-Gq : : 8 e-05 192 PIP 2_PLC-Ca-Gq -> PLC-Ca-Gq +DAG + IP 3 : : 48 PIP 2 + Ca. PLC_g <> PIP 2_Ca. PLC_g : : 1. 2 e-06 56 PIP 2_Ca. PLC_g -> Ca. PLC_g +DAG + IP 3 : : 14 PIP 2 + Ca. PLC_g* <> PIP 2_Ca. PLC_g* : : 2. 4 e-05 228 PIP 2_Ca. PLC_g* -> Ca. PLC_g* +DAG + IP 3 : : 57 GDP-Ras + GEF-Gprot-bg <> GDP-Ras_GEF-Gprot-bg : : 3. 3 e-07 0. 08 GDP-Ras_GEF-Gprot-bg -> GEF-Gprot-bg +GTP-Ras : : 0. 02 GDP-Ras + GEF* <> GDP-Ras_GEF* : : 3. 3 e-07 0. 08 GDP-Ras_GEF* -> GEF* +GTP-Ras : : 0. 02 GTP-Ras + GAP <> GTP-Ras_GAP : : 0. 001666 1000 GTP-Ras_GAP -> GAP +GDP-Ras : : 10 GDP-Ras + Ca. M-GEF <> GDP-Ras_Ca. M-GEF : : 3. 3 e-07 0. 08 GDP-Ras_Ca. M-GEF -> Ca. M-GEF +GTP-Ras : : 0. 02 Ca. Transp-2 Ca -> Ca. Transp + Ca-sequester : : 25 Special topics on Electrical and Systems Engineering 2007 Systems Biology Ca +Ca. Transp +Ca <> Ca. Transp-2 Ca : : 1 e-08 144 IP 3 R +IP 3 <> IP 3 R* : : 1 e-20 1 Ca-sequester + capacitive_Ca_entry* <> inact_cap_entry : : 1. 2 e-11 EGFR +EGF <> L. EGFR : : 7 e-06 0. 25 SHC* -> SHC : : 0. 0016667 L. EGFR <> Internal_L. EGFR : : 0. 002 0. 00033 m. Glu. R +Glu <> Rec-Glu : : 2. 8 e-05 10 G-GDP -> G*GTP + Beta. Gamma : : 0. 0001 G*GDP +Beta. Gamma -> G-GDP : : 1 e-05 G*GTP -> G*GDP : : 0. 0133 G-GDP +Rec-Glu <> Rec-Glu-Gq : : 1 e-08 0. 0001 Glu +Rec-Gq <> Rec-Glu-Gq : : 2. 8 e-05 0. 1 Rec-Glu-Gq -> G*GTP + Beta. Gamma + Rec-Glu : : 0. 01 G-GDP +m. Glu. R <> Rec-Gq : : 1 e-06 1 Rec-Gq +m. Glu. RAntag <> Blocked-rec-Gq : : 0. 0001 0. 01 craf-1* +GTP-Ras <> Raf-GTP-Ras* : : 4 e-05 0. 5 PKC-cytosolic +Ca <> PKC-Ca : : 1 e-06 0. 5 DAG +PKC-Ca <> PKC-Ca-DAG : : 1. 3333 e-08 8. 6348 PKC-Ca <> PKC-Ca-memb* : : 1. 2705 3. 5026 PKC-Ca-DAG <> PKC-DAG-memb* : : 1 0. 1 PKC-Ca +AA <> PKC-Ca-AA* : : 2 e-09 0. 1 PKC-DAG-AA <> PKC-DAG-AA* : : 2 0. 2 PKC-cytosolic <> PKC-basal* : : 1 50 AA +PKC-cytosolic <> PKC-AA* : : 2 e-10 0. 1 PKC-cytosolic +DAG <> PKC-DAG : : 1 e-09 0. 1 PKC-DAG +AA <> PKC-DAG-AA : : 3 e-08 2 PLA 2 -cytosolic +Ca <> PLA 2 -Ca* : : 1. 6667 e-06 0. 1 temp-PIP 2 +PLA 2 -cytosolic <> PIP 2 -PLA 2* : : 2 e-09 0. 5 temp-PIP 2 +PLA 2 -Ca* <> PIP 2 -Ca-PLA 2* : : 2 e-08 0. 1 DAG +PLA 2 -Ca* <> DAG-Ca-PLA 2* : : 5 e-09 4 AA -> APC : : 0. 4 PLA 2* +Ca <> PLA 2*-Ca : : 1 e-05 0. 1 PLA 2* -> PLA 2 -cytosolic : : 0. 17 Ca +PLC <> PLC-Ca : : 5 e-06 1 IP 3 -> Inositol : : 2. 5 DAG -> PC : : 0. 15 G*GTP +PLC-Ca <> PLC-Ca-Gq : : 4. 2 e-05 1 PLC-Ca-Gq -> G*GDP + PLC-Ca : : 0. 0133 PLC +G*GTP <> PLC-Gq : : 4. 2 e-06 1 Ca +PLC-Gq <> PLC-Ca-Gq : : 5 e-05 1 PLC_g +Ca <> Ca. PLC_g : : 0. 0003 10 Ca +PLC_G* <> Ca. PLC_g* : : 2 e-05 10 Ca. PLC_g* -> Ca. PLC_g : : 0. 05 Beta. Gamma +inact-GEF <> GEF-Gprot-bg : : 1 e-05 1 GEF* -> inact-GEF : : 1 GTP-Ras -> GDP-Ras : : 0. 0001 GAP* -> GAP : : 0. 1 inact-GEF +Ca. M-Ca 4 <> Ca. M-GEF : : 0. 0001 1 inact-GEF* -> inact-GEF : : 1 Sos. Grb 2 +SHC* <> Shc*. Sos. Grb 2 : : 8. 333 e-07 0. 1 Sos* +Grb 2 <> Sos*. Grb 2 : : 4. 1667 e-08 0. 0168 Sos* -> Sos : : 0. 001 Grb 2 +Sos <> Sos. Grb 2 : : 4. 1667 e-08 0. 0168 1
![Leads to. . d[capacitive_ca_entry*]/dt = 1[inact_cap_entry] - 1. 2 e-11[ca-sequester][capacitive_ca_entry*] d[ca]/dt = 288[ca_caepump] + Leads to. . d[capacitive_ca_entry*]/dt = 1[inact_cap_entry] - 1. 2 e-11[ca-sequester][capacitive_ca_entry*] d[ca]/dt = 288[ca_caepump] +](https://present5.com/presentation/0dd6036afd1d8bfe6892be420651c564/image-11.jpg)
Leads to. . d[capacitive_ca_entry*]/dt = 1[inact_cap_entry] - 1. 2 e-11[ca-sequester][capacitive_ca_entry*] d[ca]/dt = 288[ca_caepump] + 144[catransp-2 ca] + 0. 5[pkc-ca] + 0. 1[pla 2 -ca*] + 0. 1[pla 2*-ca] + 1[plc-ca-gq] + 10[ca. plc_g*] - 0. 003[ca][caepump] - 1 e 08[ca][catransp][ca] - 1 e-06[pkc-cytosolic][ca] - 1. 6667 e-06[pla 2 -cytosolic][ca] - 1 e-05[pla 2*][ca] - 5 e-06[ca][plc] - 5 e-05[ca][plc-gq] - 0. 0003[plc_g][ca] - 2 e-05[ca][plc_g*] + 0. 005 * [capacitive_ca_entry*] * ( [ca-ext] - [ca] ) + 0. 004 * [ ca-leak-from-extracell] * ( [ca-ext] - [ca] ) + 8 * [ca-leak-to-cytoplasm] * ( [ca-sequester] - [ca] ) + 19. 2 * [ip 3 r*] * ( [ca-sequester] - [ca] ) d[ca-leak-from-extracell]/dt = d[ca-leak-to-cytoplasm]/dt = d[ip 3 r*]/dt = 1 e-20[ip 3 r][ip 3][ip 3] - 1[ip 3 r*] d[caepump]/dt = 288[ca_caepump] + 72[ca_caepump] - 0. 003[ca][caepump] d[ca_caepump]/dt = 0. 003[ca][caepump] - 288[ca_caepump] - 72[ca_caepump] d[ca. plc_g]/dt = 0. 8[ca. plc_g_l. egfr] + 56[pip 2_ca. plc_g] + 14[pip 2_ca. plc_g] + 0. 0003[plc_g][ca] + 0. 05[ca. plc_g*] - 5 e-06[ca. plc_g][l. egfr] - 1. 2 e-06[pip 2][ca. plc_g] - 10[ca. plc_g] d[l. egfr]/dt = 0. 8[ca. plc_g_l. egfr] + 0. 2[ca. plc_g_l. egfr] + 0. 8[shc_l. egfr] + 0. 2[shc_l. egfr] + 7 e-06[egfr][egf] + 0. 00033[internal_l. egfr] - 5 e-06[ca. plc_g][l. egfr] - 2 e-06[shc][l. egfr] - 0. 25[l. egfr] - 0. 002[l. egfr] d[ca. plc_g_l. egfr]/dt = 5 e-06[ca. plc_g][l. egfr] - 0. 8[ca. plc_g_l. egfr] - 0. 2[ca. plc_g_l. egfr] d[ca. plc_g*]/dt = 0. 2[ca. plc_g_l. egfr] + 228[pip 2_ca. plc_g*] + 57[pip 2_ca. plc_g*] + 2 e-05[ca][plc_g*] - 2. 4 e-05[pip 2][ca. plc_g*] - 10[ca. plc_g*] - 0. 05[ca. plc_g*] d[shc]/dt = 0. 8[shc_l. egfr] + 0. 0016667[shc*] - 2 e-06[shc][l. egfr] d[shc_l. egfr]/dt = 2 e-06[shc][l. egfr] - 0. 8[shc_l. egfr] - 0. 2[shc_l. egfr] d[shc*]/dt = 0. 2[shc_l. egfr] + 0. 1[shc*. sos. grb 2] - 0. 0016667[shc*] - 8. 333 e-07[sos. grb 2][shc*] d[mapk]/dt = 0. 6[mapk_mapkk*] + 1[mapk-tyr_mkp-1] - 2. 7 e-05[mapk][mapkk*] d[mapkk*]/dt = 0. 6[mapk_mapkk*] + 0. 15[mapk_mapkk*] + 0. 6[mapk-tyr_mapkk*] + 0. 15[mapk-tyr_mapkk*] + 0. 105[mapkk-ser_raf-gtp-ras*] + 25[mapkk*_pphosphatase 2 a] - 2. 7 e-05[mapk][mapkk*] - 2. 7 e -05[mapk-tyr][mapkk*] - 3. 3 e-06[mapkk*][pphosphatase 2 a] d[mapk_mapkk*]/dt = 2. 7 e-05[mapk][mapkk*] - 0. 6[mapk_mapkk*] - 0. 15[mapk_mapkk*] d[mapk-tyr]/dt = 0. 15[mapk_mapkk*] + 0. 6[mapk-tyr_mapkk*] + 4[mapk-tyr_mkp-1] + 1[mapk*_mkp-1] - 2. 7 e-05[mapk-tyr][mapkk*] - 0. 000125[mapk-tyr][mkp-1] d[mapk-tyr_mapkk*]/dt = 2. 7 e-05[mapk-tyr][mapkk*] - 0. 6[mapk-tyr_mapkk*] - 0. 15[mapk-tyr_mapkk*] d[mapk*]/dt = 0. 15[mapk-tyr_mapkk*] + 80[pla 2 -cytosolic_mapk*] + 20[pla 2 -cytosolic_mapk*] + 40[craf-1*_ mapk*] + 10[craf-1*_mapk*] + 40[sos_mapk*] + 10[sos_mapk*] + 4[mapk*_mkp-1] - 6. 5 e-06[pla 2 -cytosolic][mapk*] - 3. 25 e-06[craf-1*][mapk*] - 3. 25 e-05[sos][mapk*] - 0. 000125[mapk*][mkp-1] d[mapkk]/dt = 0. 42[mapkk_raf-gtp-ras*] + 6[mapkk-ser_pphosphatase 2 a] - 5. 5 e-06[mapkk][raf-gtp-ras*] d[raf-gtp-ras*]/dt = 0. 42[mapkk_raf-gtp-ras*] + 0. 105[mapkk_raf-gtp-ras*] + 0. 42[mapkk-ser_raf-gtp-ras*] + 0. 105[mapkk-ser_raf-gtp-ras*] + 4 e-05[craf-1*][ tp-ras] - 5. 5 e-06[mapkk][raf-gtp-ras*] - 5. 5 eg 06[mapkk-ser][raf-gtp-ras*] - 0. 5[raf-gtp-ras*] d[mapkk_raf-gtp-ras*]/dt = 5. 5 e-06[mapkk][raf-gtp-ras*] - 0. 42[mapkk_raf-gtp-ras*] - 0. 105[mapkk_raf-gtp-ras*] d[mapkk-ser]/dt = 0. 105[mapkk_raf-gtp-ras*] + 0. 42[mapkk-ser_raf-gtp-ras*] + 6[mapkk*_pphosphatase 2 a] + 25[mapkk-ser_pphosphatase 2 a] - 5. 5 e-06[mapkk-ser][raf-gtp-ras*] - 3. 3 e-06[mapkkser][pphosphatase 2 a] d[mapkk-ser_raf-gtp-ras*]/dt = 5. 5 e-06[mapkk-ser][raf-gtp-ras*] - 0. 42[mapkk-ser_raf-gtp-ras*] - 0. 105[mapkk-ser_raf-gtp-ras*] d[craf-1]/dt = 16[craf-1_pkc-dag-aa*] + 16[craf-1_pkc-ca-memb*] + 16[craf-1_pkc-ca-aa*] + 16[craf-1_pkc-dag-memb*] + 16[craf-1_pkc-basal*] + 16[craf-1_pkc-aa*] + 6[craf-1*_pphosphatase 2 a] - 5 e 07[craf-1][pkc-dag-aa*] - 5 e-07[craf-1][pkc-ca-memb*] - 5 e-07[craf-1][pkc-ca-aa*] - 5 e-07[craf-1][pkc-dag-memb*] - 5 e-07[craf-1][pkc-basal*] - 5 e-07[craf-1][pkc-aa*] d[pkc-dag-aa*]/dt = 16[craf-1_pkc-dag-aa*] + 4[craf-1_pkc-dag-aa*] + 16[gap_pkc-dag-aa*] + 4[gap_pkc-dag-aa*] + 16[inact-gef_pkc-dag-aa*] + 4[inact-gef_pkc-dag-aa*] + 2[pkc-dag-aa] - 5 e-07[craf 1][pkc-dag-aa*] - 1 e-05[gap][pkc-dag-aa*] - 1 e-05[inact-gef][pkc-dag-aa*] - 0. 2[pkc-dag-aa*] d[craf-1_pkc-dag-aa*]/dt = 5 e-07[craf-1][pkc-dag-aa*] - 16[craf-1_pkc-dag-aa*] - 4[craf-1_pkc-dag-aa*] d[craf-1*]/dt = 4[craf-1_pkc-dag-aa*] + 4[craf-1_pkc-ca-memb*] + 4[craf-1_pkc-ca-aa*] + 4[craf-1_pkc-dag-memb*] + 4[craf-1_pkc-basal*] + 4[craf-1_pkc-aa*] + 40[craf-1*_ mapk*] + 25[craf 1*_pphosphatase 2 a] + 6[craf-1**_pphosphatase 2 a] + 0. 5[raf-gtp-ras*] - 3. 25 e-06[craf-1*][ apk*] - 3. 3 e-06[craf-1*][pphosphatase 2 a] - 4 e-05[craf-1*][gtp-ras] m d[gap]/dt = 16[gap_pkc-dag-aa*] + 16[gap_pkc-ca-memb*] + 16[gap_pkc-ca-aa*] + 16[gap_pkc-dag-memb*] + 16[gap_pkc-basal*] + 16[gap_pkc-aa*] + 1000[gtp-ras_gap] + 10[gtp-ras_gap] + 0. 1[gap*] 1 e-05[gap][pkc-dag-aa*] - 1 e-05[gap][pkc-ca-memb*] - 1 e-05[gap][pkc-ca-aa*] - 1 e-05[gap][pkc-dag-memb*] - 1 e-05[gap][pkc-basal*] - 1 e-05[gap][pkc-aa*] - 0. 001666[gtp-ras][gap] d[gap_pkc-dag-aa*]/dt = 1 e-05[gap][pkc-dag-aa*] - 16[gap_pkc-dag-aa*] - 4[gap_pkc-dag-aa*] d[gap*]/dt = 4[gap_pkc-dag-aa*] + 4[gap_pkc-ca-memb*] + 4[gap_pkc-ca-aa*] + 4[gap_pkc-dag-memb*] + 4[gap_pkc-basal*] + 4[gap_pkc-aa*] - 0. 1[gap*] d[inact-gef]/dt = 16[inact-gef_pkc-dag-aa*] + 16[inact-gef_pkc-ca-memb*] + 16[inact-gef_pkc-ca-aa*] + 16[inact-gef_pkc-dag-memb*] + 16[inact-gef_pkc-basal*] + 16[inact-gef_pkc-aa*] + 36[inact-gef_pkaactive] + 1[gef-gprot-bg] + 1[gef*] + 1[cam-gef] + 1[inact-gef*] - 1 e-05[inact-gef][pkc-dag-aa*] - 1 e-05[inact-gef][pkc-ca-memb*] - 1 e-05[inact-gef][pkc-ca-aa*] - 1 e-05[inact-gef][pkc-dag-memb*] - 1 e 05[inact-gef][pkc-basal*] - 1 e-05[inact-gef][pkc-aa*] - 1 e-05[inact-gef][pka-active] - 1 e-05[betagamma][inact-gef] - 0. 0001[inact-gef][cam-ca 4] d[inact-gef_pkc-dag-aa*]/dt = 1 e-05[inact-gef][pkc-dag-aa*] - 16[inact-gef_pkc-dag-aa*] - 4[inact-gef_pkc-dag-aa*] d[gef*]/dt = 4[inact-gef_pkc-dag-aa*] + 4[inact-gef_pkc-ca-memb*] + 4[inact-gef_pkc-ca-aa*] + 4[inact-gef_pkc-dag-memb*] + 4[inact-gef_pkc-basal*] + 4[inact-gef_pkc-aa*] + 0. 08[gdp-ras_gef*] + 0. 02[gdp-ras_gef*] - 3. 3 e-07[gdp-ras][gef*] - 1[gef*] d[pkc-ca-memb*]/dt = 16[craf-1_pkc-ca-memb*] + 4[craf-1_pkc-ca-memb*] + 16[gap_pkc-ca-memb*] + 4[gap_pkc-ca-memb*] + 16[inact-gef_pkc-ca-memb*] + 4[inact-gef_pkc-ca-memb*] + 1. 2705[pkc-ca] - 5 e-07[craf-1][pkc-ca-memb*] - 1 e-05[gap][pkc-ca-memb*] - 1 e-05[inact-gef][pkc-ca-memb*] - 3. 5026[pkc-ca-memb*] d[craf-1_pkc-ca-memb*]/dt = 5 e-07[craf-1][pkc-ca-memb*] - 16[craf-1_pkc-ca-memb*] - 4[craf-1_pkc-ca-memb*] d[gap_pkc-ca-memb*]/dt = 1 e-05[gap][pkc-ca-memb*] - 16[gap_pkc-ca-memb*] - 4[gap_pkc-ca-memb*] d[inact-gef_pkc-ca-memb*]/dt = 1 e-05[inact-gef][pkc-ca-memb*] - 16[inact-gef_pkc-ca-memb*] - 4[inact-gef_pkc-ca-memb*] d[pkc-ca-aa*]/dt = 16[craf-1_pkc-ca-aa*] + 4[craf-1_pkc-ca-aa*] + 16[gap_pkc-ca-aa*] + 4[gap_pkc-ca-aa*] + 16[inact-gef_pkc-ca-aa*] + 4[inact-gef_pkc-ca-aa*] + 2 e-09[pkc-ca][aa] - 5 e-07[craf-1][pkc-caaa*] - 1 e-05[gap][pkc-ca-aa*] - 1 e-05[inact-gef][pkc-ca-aa*] - 0. 1[pkc-ca-aa*] d[craf-1_pkc-ca-aa*]/dt = 5 e-07[craf-1][pkc-ca-aa*] - 16[craf-1_pkc-ca-aa*] - 4[craf-1_pkc-ca-aa*] d[gap_pkc-ca-aa*]/dt = 1 e-05[gap][pkc-ca-aa*] - 16[gap_pkc-ca-aa*] - 4[gap_pkc-ca-aa*] d[glu]/dt = 10[rec-glu] + 0. 1[rec-glu-gq] - 2. 8 e-05[mglur][glu] - 2. 8 e-05[glu][rec-gq] d[rec-glu]/dt = 2. 8 e-05[mglur][glu] + 0. 0001[rec-glu-gq] + 0. 01[rec-glu-gq] - 10[rec-glu] - 1 e-08[g-gdp][rec-glu] d[g-gdp]/dt = 1 e-05[g*gdp][betagamma] + 0. 0001[rec-glu-gq] + 1[rec-gq] - 0. 0001[g-gdp] - 1 e-08[g-gdp][rec-glu] - 1 e-06[g-gdp][mglur] d[g*gtp]/dt = 0. 0001[g-gdp] + 0. 01[rec-glu-gq] + 1[plc-ca-gq] + 1[plc-gq] - 0. 0133[g*gtp] - 4. 2 e-05[g*gtp][plc-ca] - 4. 2 e-06[plc][g*gtp] d[betagamma]/dt = 0. 0001[g-gdp] + 0. 01[rec-glu-gq] + 1[gef-gprot-bg] - 1 e-05[g*gdp][betagamma] - 1 e-05[betagamma][inact-gef] d[g*gdp]/dt = 0. 0133[g*gtp] + 0. 0133[plc-ca-gq] - 1 e-05[g*gdp][betagamma] d[rec-glu-gq]/dt = 1 e-08[g-gdp][rec-glu] + 2. 8 e-05[glu][rec-gq] - 0. 0001[rec-glu-gq] - 0. 01[rec-glu-gq] d[rec-gq]/dt = 0. 1[rec-glu-gq] + 1 e-06[g-gdp][mglur] + 0. 01[blocked-rec-gq] - 2. 8 e-05[glu][rec-gq] - 1[rec-gq] - 0. 0001[rec-gq][mglurantag] d[mglurantag]/dt = 0. 01[blocked-rec-gq] - 0. 0001[rec-gq][mglurantag] d[blocked-rec-gq]/dt = 0. 0001[rec-gq][mglurantag] - 0. 01[blocked-rec-gq] d[pkc-cytosolic]/dt = 0. 5[pkc-ca] + 50[pkc-basal*] + 0. 1[pkc-aa*] + 0. 1[pkc-dag] - 1 e-06[pkc-cytosolic][ca] - 1[pkc-cytosolic] - 2 e-10[aa][pkc-cytosolic] - 1 e-09[pkc-cytosolic][dag] d[pkc-ca]/dt = 1 e-06[pkc-cytosolic][ca] + 8. 6348[pkc-ca-dag] + 3. 5026[pkc-ca-memb*] + 0. 1[pkc-ca-aa*] - 0. 5[pkc-ca] - 1. 3333 e-08[dag][pkc-ca] - 1. 2705[pkc-ca] - 2 e-09[pkc-ca][aa] d[pkc-ca-dag]/dt = 1. 3333 e-08[dag][pkc-ca] + 0. 1[pkc-dag-memb*] - 8. 6348[pkc-ca-dag] - 1[pkc-ca-dag] d[pkc-dag-aa]/dt = 0. 2[pkc-dag-aa*] + 3 e-08[pkc-dag][aa] - 2[pkc-dag-aa] d[pkc-dag]/dt = 1 e-09[pkc-cytosolic][dag] + 2[pkc-dag-aa] - 0. 1[pkc-dag] - 3 e-08[pkc-dag][aa] d[temp-pip 2]/dt = 0. 5[pip 2 -pla 2*] + 0. 1[pip 2 -ca-pla 2*] - 2 e-09[temp-pip 2][pla 2 -cytosolic] - 2 e-08[temp-pip 2][pla 2 -ca*] d[plc]/dt = 1[plc-ca] + 1[plc-gq] - 5 e-06[ca][plc] - 4. 2 e-06[plc][g*gtp] d[inositol]/dt = 2. 5[ip 3] d[pc]/dt = 0. 15[dag] d[plc-gq]/dt = 4. 2 e-06[plc][g*gtp] + 1[plc-ca-gq] - 1[plc-gq] - 5 e-05[ca][plc-gq] d[plc_g]/dt = 10[ca. plc_g] - 0. 0003[plc_g][ca] d[plc_g*]/dt = 10[ca. plc_g*] - 2 e-05[ca][plc_g*] d[cam-ca 4]/dt = 1[cam-gef] - 0. 0001[inact-gef][cam-ca 4] d[sos. grb 2]/dt = 0. 1[shc*. sos. grb 2] + 4. 1667 e-08[grb 2][sos] - 8. 333 e-07[sos. grb 2][shc*] - 0. 0168[sos. grb 2] d[grb 2]/dt = 0. 0168[sos*. grb 2] + 0. 0168[sos. grb 2] - 4. 1667 e-08[sos*][grb 2] - 4. 1667 e-08[grb 2][sos] d[sos*. grb 2]/dt = 4. 1667 e-08[sos*][grb 2] - 0. 0168[sos*. grb 2] d[ca-ext]/dt = 72[ca_caepump] - 0. 005 * [capacitive_ca_entry*] * ( [ca-ext] - [ca] ) - 0. 004 * [ca-leak-from- extracell] * ( [ca-ext] - [ca] ) d[ca-sequester]/dt = 25[catransp-2 ca] + 1[inact_cap_entry] - 1. 2 e-11[ca-sequester][capacitive_ca_entry*] - 1. 2 e-11[ca-sequester][ca- d[inact-gef_pkc-ca-aa *]/dt = 1 e-05[inact-gef][pkc-ca-aa*] - 16[inact-gef_pkc-ca-aa*] - 4[inact-gef_pkc-ca-aa*] d[pkc-dag-memb*]/dt = 16[craf-1_pkc-dag-memb*] + 4[craf-1_pkc-dag-memb*] + 16[gap_pkc-dag-memb*] + 4[gap_pkc-dag-memb*] + 16[inact-gef_pkc-dag-memb*] + 4[inact-gef_pkc-dag-memb*] + 1[pkc-ca-dag] - 5 e-07[craf-1][pkc-dagmemb*] - 1 e-05[gap][pkc-dag-memb*] - 1 e-05[inact-gef][pkc-dag-memb*] - 0. 1[pkc-dag-memb*] d[craf-1_pkc-dag-memb*]/dt = 5 e-07[craf-1][pkc-dag-memb*] - 16[craf-1_pkc-dag-memb*] - 4[craf-1_pkc-dag-memb*] d[gap_pkc-dag-memb*]/dt = 1 e-05[gap][pkc-dag-memb*] - 16[gap_pkc-dag-memb*] - 4[gap_pkc-dag-memb*] d[inact-gef_pkc-dag-memb*]/dt = 1 e-05[inact-gef][pkc-dag-memb*] - 16[inact-gef_pkc-dag-memb*] - 4[inact-gef_pkc-dag-memb*] d[pkc-basal*]/dt = 16[craf-1_pkc-basal*] + 4[craf-1_pkc-basal*] + 16[gap_pkc-basal*] + 4[gap_pkc-basal*] + 16[inact-gef_pkc-basal*] + 4[inact-gef_pkc-basal*] + 1[pkc-cytosolic] - 5 e-07[craf-1][pkc-basal*] - 1 e-05[gap][pkc-basal*] - 1 e-05[inact -gef][pkc-basal*] - 50[pkc-basal*] d[craf-1_pkc-basal*]/dt = 5 e-07[craf-1][pkc-basal*] - 16[craf-1_pkc-basal*] - 4[craf-1_pkc-basal*] d[gap_pkc-basal*]/dt = 1 e-05[gap][pkc-basal*] - 16[gap_pkc-basal*] - 4[gap_pkc-basal*] d[inact-gef_pkc-basal*]/dt = 1 e-05[inact-gef][pkc-basal*] - 16[inact-gef_pkc-basal*] - 4[inact-gef_pkc-basal*] d[pkc-aa*]/dt = 16[craf-1_pkc-aa*] + 4[craf-1_pkc-aa*] + 16[gap_pkc-aa*] + 4[gap_pkc-aa*] + 16[inact-gef_pkc-aa*] + 4[inact-gef_pkc-aa*] + 2 e-10[aa][pkc-cytosolic] - 5 e-07[craf-1][pkc-aa*] - 1 e-05[gap][pkc-aa*] - 1 e-05[inact-gef][pkc-aa*] 0. 1[pkc-aa*] d[craf-1_pkc-aa*]/dt = 5 e-07[craf-1][pkc-aa*] - 16[craf-1_pkc-aa*] - 4[craf-1_pkc-aa*] d[gap_pkc-aa*]/dt = 1 e-05[gap][pkc-aa*] - 16[gap_pkc-aa*] - 4[gap_pkc-aa*] d[inact-gef_pkc-aa*]/dt = 1 e-05[inact-gef][pkc-aa*] - 16[inact-gef_pkc-aa*] - 4[inact-gef_pkc-aa*] d[apc]/dt = 21. 6[apc_pla 2 -ca*] + 44. 16[apc_pip 2 -pla 2*] + 144[apc_pip 2 -ca-pla 2*] + 240[apc_dag-ca-pla 2*] + 480[apc_pla 2*-ca] + 0. 4[aa] - 2. 25 e-06[apc][pla 2 -ca*] - 4. 6 e-06[apc][pip 2 -pla 2*] - 1. 5 e-05[apc][pip 2 -ca-pla 2*] - 2. 5 e-05[apc][dag-capla 2*] - 5 e-05[apc][pla 2*-ca] d[pla 2 -ca*]/dt = 21. 6[apc_pla 2 -ca*] + 5. 4[apc_pla 2 -ca*] + 1. 6667 e-06[pla 2 -cytosolic][ca] + 0. 1[pip 2 -ca-pla 2*] + 4[dag-ca-pla 2*] - 2. 25 e-06[apc][pla 2 -ca*] - 0. 1[pla 2 -ca*] - 2 e-08[temp-pip 2][pla 2 -ca*] - 5 e-09[dag][pla 2 -ca*] d[apc_pla 2 -ca*]/dt = 2. 25 e-06[apc][pla 2 -ca*] - 21. 6[apc_pla 2 -ca*] - 5. 4[apc_pla 2 -ca*] d[aa]/dt = 5. 4[apc_pla 2 -ca*] + 11. 04[apc_pip 2 -pla 2*] + 36[apc_pip 2 -ca-pla 2*] + 60[apc_dag-ca-pla 2*] + 120[apc_pla 2*-ca] + 0. 1[pkc-ca-aa*] + 0. 1[pkc-aa*] + 2[pkc-dag-aa] - 2 e-09[pkc-ca][aa] - 2 e-10[aa][pkc-cytosolic] - 3 e-08[pkc-dag][aa] 0. 4[aa] d[pip 2 -pla 2*]/dt = 44. 16[apc_pip 2 -pla 2*] + 11. 04[apc_pip 2 -pla 2*] + 2 e-09[temp-pip 2][pla 2 -cytosolic] - 4. 6 e-06[apc][pip 2 -pla 2*] - 0. 5[pip 2 -pla 2*] d[apc_pip 2 -pla 2*]/dt = 4. 6 e-06[apc][pip 2 -pla 2*] - 44. 16[apc_pip 2 -pla 2*] - 11. 04[apc_pip 2 -pla 2*] d[pip 2 -ca-pla 2*]/dt = 144[apc_pip 2 -ca-pla 2*] + 36[apc_pip 2 -ca-pla 2*] + 2 e-08[temp-pip 2][pla 2 -ca*] - 1. 5 e-05[apc][pip 2 -ca-pla 2*] - 0. 1[pip 2 -ca-pla 2*] d[apc_pip 2 -ca-pla 2*]/dt = 1. 5 e-05[apc][pip 2 -ca-pla 2*] - 144[apc_pip 2 -ca-pla 2*] - 36[apc_pip 2 -ca-pla 2*] d[dag-ca-pla 2*]/dt = 240[apc_dag-ca-pla 2*] + 60[apc_dag-ca-pla 2*] + 5 e-09[dag][pla 2 -ca*] - 2. 5 e-05[apc][dag-ca-pla 2*] - 4[dag-ca-pla 2*] d[apc_dag-ca-pla 2*]/dt = 2. 5 e-05[apc][dag-ca-pla 2*] - 240[apc_dag-ca-pla 2*] - 60[apc_dag-ca-pla 2*] d[pla 2*-ca]/dt = 480[apc_pla 2*-ca] + 120[apc_pla 2*-ca] + 1 e-05[pla 2*][ca] - 5 e-05[apc][pla 2*-ca] - 0. 1[pla 2*-ca] d[apc_pla 2*-ca]/dt = 5 e-05[apc][pla 2*-ca] - 480[apc_pla 2*-ca] - 120[apc_pla 2*-ca] d[pla 2 -cytosolic]/dt = 80[pla 2 -cytosolic_mapk*] + 0. 1[pla 2 -ca*] + 0. 5[pip 2 -pla 2*] + 0. 17[pla 2*] - 6. 5 e-06[pla 2 -cytosolic][mapk*] - 1. 6667 e-06[pla 2 -cytosolic][ca] - 2 e-09[temp-pip 2][pla 2 -cytosolic] d[pla 2 -cytosolic_mapk*]/dt = 6. 5 e-06[pla 2 -cytosolic][mapk*] - 80[pla 2 -cytosolic_mapk*] - 20[pla 2 -cytosolic_mapk*] d[pla 2*]/dt = 20[pla 2 -cytosolic_mapk*] + 0. 1[pla 2*-ca] - 1 e-05[pla 2*][ca] - 0. 17[pla 2*] d[craf-1*_mapk*]/dt = 3. 25 e-06[craf-1*][mapk*] - 40[craf-1*_mapk*] - 10[craf-1*_mapk*] d[craf-1**]/dt = 10[craf-1*_mapk*] + 25[craf-1**_pphosphatase 2 a] - 3. 3 e-06[craf-1**][pphosphatase 2 a] d[sos]/dt = 40[sos_mapk*] + 0. 001[sos*] + 0. 0168[sos. grb 2] - 3. 25 e-05[sos][mapk*] - 4. 1667 e-08[grb 2][sos] d[sos_mapk*]/dt = 3. 25 e-05[sos][mapk*] - 40[sos_mapk*] - 10[sos_mapk*] d[sos*]/dt = 10[sos_mapk*] + 0. 0168[sos*. grb 2] - 4. 1667 e-08[sos*][grb 2] - 0. 001[sos*] d[mkp-1]/dt = 4[mapk-tyr_mkp-1] + 1[mapk-tyr_mkp-1] + 4[mapk*_mkp-1] + 1[mapk*_mkp-1] - 0. 000125[mapk-tyr][mkp-1] - 0. 000125[mapk*][mkp-1] d[mapk-tyr_mkp-1]/dt = 0. 000125[mapk-tyr][mkp-1] - 4[mapk-tyr_mkp-1] - 1[mapk-tyr_mkp-1] d[mapk*_mkp-1]/dt = 0. 000125[mapk*][mkp-1] - 4[mapk*_mkp-1] - 1[mapk*_mkp-1] d[pphosphatase 2 a]/dt = 25[craf-1*_pphosphatase 2 a] + 6[craf-1*_pphosphatase 2 a] + 25[mapkk*_pphosphatase 2 a] + 6[mapkk*_pphosphatase 2 a] + 25[mapkk-ser_pphosphatase 2 a] + 6[mapkk-ser_pphosphatase 2 a] + 25[craf 1**_pphosphatase 2 a] + 6[craf-1**_pphosphatase 2 a] - 3. 3 e-06[craf-1*][pphosphatase 2 a] - 3. 3 e-06[mapkk-ser][pphosphatase 2 a] - 3. 3 e-06[craf-1**][pphosphatase 2 a] d[craf-1*_pphosphatase 2 a]/dt = 3. 3 e-06[craf-1*][pphosphatase 2 a] - 25[craf-1*_pphosphatase 2 a] - 6[craf-1*_pphosphatase 2 a] d[mapkk*_pphosphatase 2 a]/dt = 3. 3 e-06[mapkk*][pphosphatase 2 a] - 25[mapkk*_pphosphatase 2 a] - 6[mapkk*_pphosphatase 2 a] d[mapkk-ser_pphosphatase 2 a]/dt = 3. 3 e-06[mapkk-ser][pphosphatase 2 a] - 25[mapkk-ser_pphosphatase 2 a] - 6[mapkk-ser_pphosphatase 2 a] d[craf-1**_pphosphatase 2 a]/dt = 3. 3 e-06[craf-1**][pphosphatase 2 a] - 25[craf-1**_pphosphatase 2 a] - 6[craf-1**_pphosphatase 2 a] d[pka-active]/dt = 36[inact-gef_pka-active] + 9[inact-gef_pka-active] - 1 e-05[inact-gef][pka-active] d[inact-gef_pka-active]/dt = 1 e-05[inact-gef][pka-active] - 36[inact-gef_pka-active] - 9[inact-gef_pka-active] d[inact-gef*]/dt = 9[inact-gef_pka-active] - 1[inact-gef*] d[gdp-ras]/dt = 0. 08[gdp-ras_shc*. sos. grb 2] + 0. 08[gdp-ras_gef-gprot-bg] + 0. 08[gdp-ras_gef*] + 10[gtp-ras_gap] + 0. 08[gdp-ras_cam-gef] + 0. 0001[gtp-ras] - 3. 3 e-07[gdp-ras][shc*. sos. grb 2] - 3. 3 e-07[gdp-ras][gef-gprot-bg] - 3. 3 e-07[gdpras][gef*] - 3. 3 e-07[gdp-ras][cam-gef] d[shc*. sos. grb 2]/dt = 0. 08[gdp-ras_shc*. sos. grb 2] + 0. 02[gdp-ras_shc*. sos. grb 2] + 8. 333 e-07[sos. grb 2][shc*] - 3. 3 e-07[gdp-ras][shc*. sos. grb 2] - 0. 1[shc*. sos. grb 2] d[gdp-ras_shc*. sos. grb 2]/dt = 3. 3 e-07[gdp-ras][shc*. sos. grb 2] - 0. 08[gdp-ras_shc*. sos. grb 2] - 0. 02[gdp-ras_shc*. sos. grb 2] d[gtp-ras]/dt = 0. 02[gdp-ras_shc*. sos. grb 2] + 0. 02[gdp-ras_gef-gprot-bg] + 0. 02[gdp-ras_gef*] + 1000[gtp-ras_gap] + 0. 02[gdp-ras_cam-gef] + 0. 5[raf-gtp-ras*] - 0. 001666[gtp-ras][gap] - 4 e-05[craf-1*][ gtp-ras] - 0. 0001[gtp-ras] d[pip 2]/dt = 40[pip 2_plc-ca] + 192[pip 2_plc-ca-gq] + 56[pip 2_ca. plc_g] + 228[pip 2_ca. plc_g*] - 4. 2 e-06[pip 2][plc-ca] - 8 e-05[pip 2][plc-ca-gq] - 1. 2 e-06[pip 2][ca. plc_g] - 2. 4 e-05[pip 2][ca. plc_g*] d[plc-ca]/dt = 40[pip 2_plc-ca] + 10[pip 2_plc-ca] + 5 e-06[ca][plc] + 1[plc-ca-gq] + 0. 0133[plc-ca-gq] - 4. 2 e-06[pip 2][plc-ca] - 1[plc-ca] - 4. 2 e-05[g* tp][plc-ca] g d[pip 2_plc-ca]/dt = 4. 2 e-06[pip 2][plc-ca] - 40[pip 2_plc-ca] - 10[pip 2_plc-ca] d[dag]/dt = 10[pip 2_plc-ca] + 48[pip 2_plc-ca-gq] + 14[pip 2_ca. plc_g] + 57[pip 2_ca. plc_g*] + 8. 6348[pkc-ca-dag] + 0. 1[pkc-dag] + 4[dag-ca-pla 2*] - 1. 3333 e-08[dag][pkc-ca] - 1 e-09[pkc-cytosolic][dag] - 5 e-09[dag][pla 2 -ca*] - 0. 15[dag] d[ip 3]/dt = 10[pip 2_plc-ca] + 48[pip 2_plc-ca-gq] + 14[pip 2_ca. plc_g] + 57[pip 2_ca. plc_g*] + 1[ip 3 r*] - 1 e-20[ip 3 r][ip 3][ip 3] - 1 e-20[ip 3 r][ip 3][ip 3] - 2. 5[ip 3] d[plc-ca-gq]/dt = 192[pip 2_plc-ca-gq] + 48[pip 2_plc-ca-gq] + 4. 2 e-05[g* gtp][plc-ca] + 5 e-05[ca][plc-gq] - 8 e-05[pip 2][plc-ca-gq] - 1[plc-ca-gq] - 0. 0133[plc-ca-gq] - 1[plc-ca-gq] d[pip 2_plc-ca-gq]/dt = 8 e-05[pip 2][plc-ca-gq] - 192[pip 2_plc-ca-gq] - 48[pip 2_plc-ca-gq] d[pip 2_ca. plc_g]/dt = 1. 2 e-06[pip 2][ca. plc_g] - 56[pip 2_ca. plc_g] - 14[pip 2_ca. plc_g] d[pip 2_ca. plc_g*]/dt = 2. 4 e-05[pip 2][ca. plc_g*] - 228[pip 2_ca. plc_g*] - 57[pip 2_ca. plc_g*] d[gef-gprot-bg]/dt = 0. 08[gdp-ras_gef-gprot-bg] + 0. 02[gdp-ras_gef-gprot-bg] + 1 e-05[betagamma][inact-gef] - 3. 3 e-07[gdp-ras][gef-gprot-bg] - 1[gef-gprot-bg] d[gdp-ras_gef-gprot-bg]/dt = 3. 3 e-07[gdp-ras][gef-gprot-bg] - 0. 08[gdp-ras_gef-gprot-bg] - 0. 02[gdp-ras_gef-gprot-bg] d[gdp-ras_gef*]/dt = 3. 3 e-07[gdp-ras][gef*] - 0. 08[gdp-ras_gef*] - 0. 02[gdp-ras_gef*] d[gtp-ras_gap]/dt = 0. 001666[gtp-ras][gap] - 1000[gtp-ras_gap] - 10[gtp-ras_gap] d[cam-gef]/dt = 0. 08[gdp-ras_cam-gef] + 0. 02[gdp-ras_cam-gef] + 0. 0001[inact-gef][cam-ca 4] - 3. 3 e-07[gdp-ras][cam-gef] - 1[cam-gef] d[gdp-ras_cam-gef]/dt = 3. 3 e-07[gdp-ras][cam-gef] - 0. 08[gdp-ras_cam-gef] - 0. 02[gdp-ras_cam-gef] d[catransp-2 ca]/dt = 1 e-08[ca][catransp][ca] - 25[catransp-2 ca] - 144[catransp-2 ca] d[catransp]/dt = 25[catransp-2 ca] + 144[catransp-2 ca] - 1 e-08[ca][catransp][ca] d[ip 3 r]/dt = 1[ip 3 r*] - 1 e-20[ip 3 r][ip 3][ip 3] d[inact_cap_entry]/dt = 1. 2 e-11[ca-sequester][capacitive_ca_entry*] - 1[inact_cap_entry] d[egfr]/dt = 0. 25[l. egfr] - 7 e-06[egfr][egf] d[egf]/dt = 0. 25[l. egfr] - 7 e-06[egfr][egf] d[internal_l. egfr]/dt = 0. 002[l. egfr] - 0. 00033[internal_l. egfr] d[mglur]/dt = 10[rec-glu] + 1[rec-gq] - 2. 8 e-05[mglur][glu] - 1 e-06[g-gdp][mglur] sequester][capacitive_ca_entry*] - 8 * [ca-leak-to-cytoplasm] * ( [ca-sequester] - [ca] ) - 19. 2 * [ip 3 r*] * ( [ca-sequester] - [ca] ) Special topics on Electrical and Systems Engineering 2007 Systems Biology

Parameters l 1: Science. 1999 Jan 15; 283(5400): 381 -7. Emergent properties of networks of biological signaling pathways. Bhalla US, Iyengar R. Special topics on Electrical and Systems Engineering 2007 Systems Biology
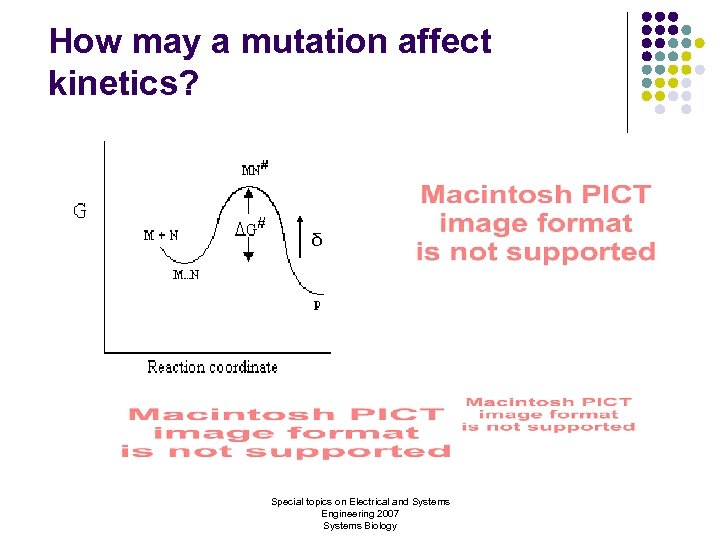

How may a mutation affect kinetics? δ Special topics on Electrical and Systems Engineering 2007 Systems Biology

Effects on signaling A mutation between two unique species Affects Reactant State Affects Product State Affects Transition State Special topics on Electrical and Systems Engineering 2007 Systems Biology

Cancer as a cumulative process l l Multiple Mutations at different sites contribute to tumor transformation Tumor may exist as “families” of differentially mutated cells l Overrides resistance of tumor to treatment Special topics on Electrical and Systems Engineering 2007 Systems Biology
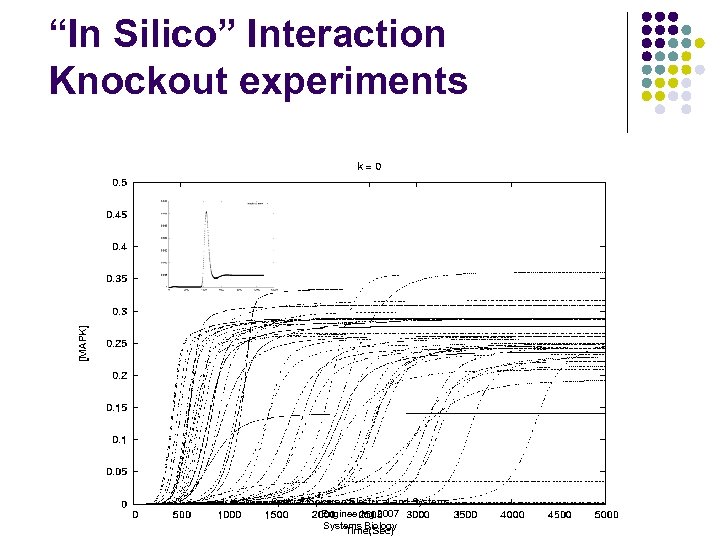
“In Silico” Interaction Knockout experiments Special topics on Electrical and Systems Engineering 2007 Systems Biology
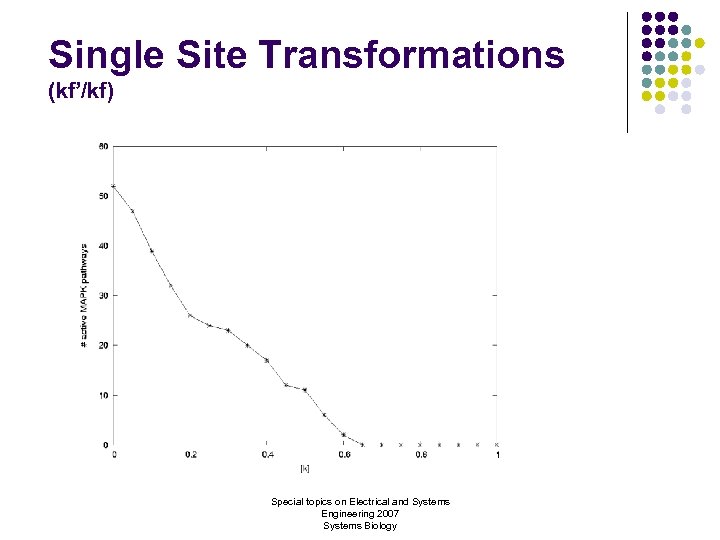
Single Site Transformations (kf’/kf) Special topics on Electrical and Systems Engineering 2007 Systems Biology
Top Ranked Mutations X* + pp 2 a X*. pp 2 a 0. 75 X*. pp 2 a X + pp 2 a 0. 75 raf* + gtp-ras raf*. gtp. ras 0. 60 Gtp-ras. gap + gdp-ras 0. 55 raf* + pp 2 a raf*. pp 2 a 0. 55 gtp-ras + gap gtp. ras_gap 0. 55 AA apc 0. 55 Ca. Pump capump + ca-ext 0. 55 X + Raf. GTP. Ras X. Raf. GTP. Ras 0. 50 PKC. DAG + AA PKC. DAG. AA* 0. 50 Special topics on Electrical and Systems Engineering 2007 Systems Biology
Top Ranked expression level mutations Species 0. 4 MKP-1 1. 9 GEF 0. 5 GAP 2. 8 MEK 0. 7 PP 2 -A 3. 0 PLC 1. 4 PKC-I 3. 4 PLC-g 1. 6 Raf 3. 6 ERK 1. 7 Ras Special topics on Electrical and Systems Engineering 2007 Systems Biology

Advantages of In Silico Pathway Analysis l Using experimentally validated models of a working signal pathway and a simple kinetic view of mutations l One can: l Quantify (and rank) l Single Interaction Mutations l Multiple Interaction Mutations l Initial State Mutations l Many results are validated in the literature l Several new and non-obvious predictions Special topics on Electrical and Systems Engineering 2007 Systems Biology

In Silico Cancer Cells l We have developed a family of “in silico” cancer cells as models of pathways What is this good for? Special topics on Electrical and Systems Engineering 2007 Systems Biology

The effect of Drugs on Signaling in mutated cells Competitive Binding Noncompetitive binding Special topics on Electrical and Systems Engineering 2007 Systems Biology

ERK Target Special topics on Electrical and Systems Engineering 2007 Systems Biology

MEK Target Special topics on Electrical and Systems Engineering 2007 Systems Biology
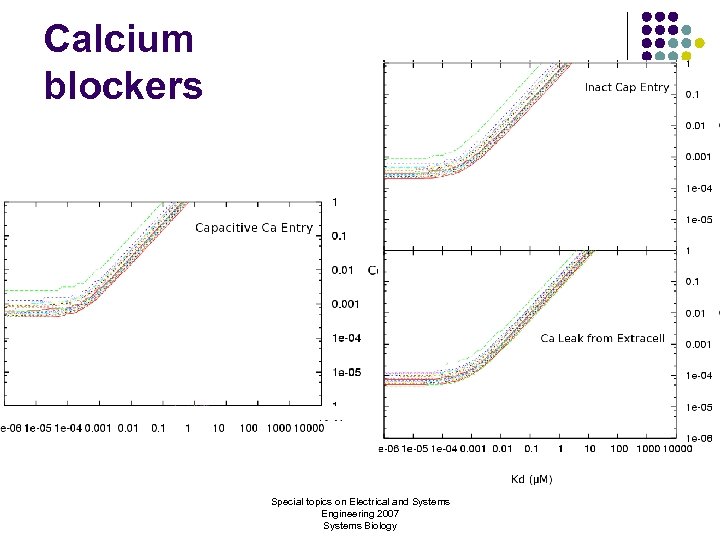
Calcium blockers Special topics on Electrical and Systems Engineering 2007 Systems Biology

Threshold “Ranking” Special topics on Electrical and Systems Engineering 2007 Systems Biology

Experimental Assays l l l l l PLC- - peptide based inhibitors PLC - U 73122 PKC - Tamoxifen , Go 6976, SPC-100221 PLA-2 - Quinacrine (anti-inflammatory) ERK MEK - AZD 6244/ARRY-142886, CI-1040 , PD 98059 Ras - R 115577, BMS-214462 SCH-66336 Raf - Bay 43 -9006 ISIS-5132 GEF Calcium Blockers Verapamil , Nicardipine Special topics on Electrical and Systems Engineering 2007 Systems Biology

Overview l l l Ras/Gap mutations predicted PP 2 A mutations predicted Ras/Raf interaction “importance” predicted Much of the overexpression/underexpression mutations predicted Drug targets are for the most part have experimental counter parts. Special topics on Electrical and Systems Engineering 2007 Systems Biology
Transport? Special topics on Electrical and Systems Engineering 2007 Systems Biology
How to treat real cells? Special topics on Electrical and Systems Engineering 2007 Systems Biology
Confocal Imaging and Quantum Dots (Andres Kriete) QDots Tunable Photostable Narrow Emission Spectra Broad Absorption Spectrum Confocal Imaging Live Cells and fixed cells sub micron imaging Special topics on Electrical and Systems Engineering 2007 Systems Biology

Quantitative Signaling Analysis Data from Confocal Imaging - derivation of “in vivo” rate constants through global analysis Given time sliced images and model Derived rate constants will “fit” experimental images best (global optimization problem) with confidence intervals - wealth of data helps prevent over fitting C(x, y, z, t) - “Automated” Sensitivity Analysis will be used for fitting Special topics on Electrical and Systems Engineering 2007 Systems Biology
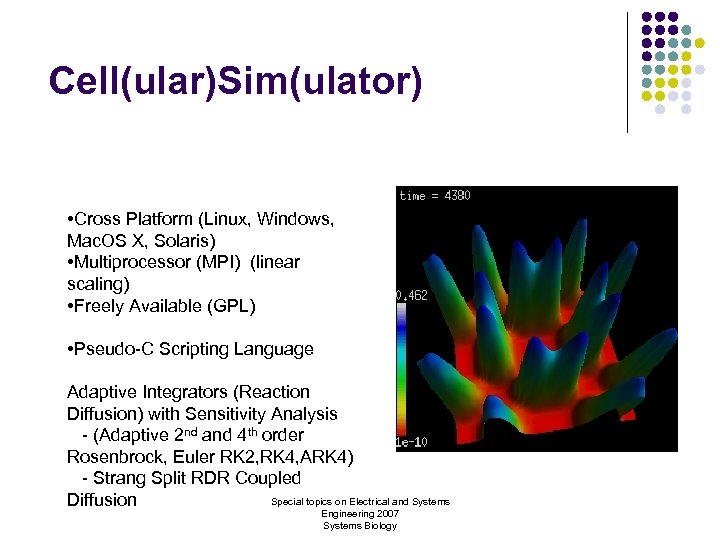

Cell(ular)Sim(ulator) • Cross Platform (Linux, Windows, Mac. OS X, Solaris) • Multiprocessor (MPI) (linear scaling) • Freely Available (GPL) • Pseudo-C Scripting Language Adaptive Integrators (Reaction Diffusion) with Sensitivity Analysis - (Adaptive 2 nd and 4 th order Rosenbrock, Euler RK 2, RK 4, ARK 4) - Strang Split RDR Coupled Special topics on Electrical and Systems Diffusion Engineering 2007 Systems Biology

Analysis Optimization Tools l l l l Sensitivity Analysis Module (Automatic generation of Jacobian, Hessian) MCA Module (Conservation Relationships) Global Optimization Tools - Use tools derived by the Protein Folding Community (and elsewhere) - Simulated Annealing - Monte Carlo + Minimization - GA / Evolutionary Programming Special topics on Electrical and Systems Engineering 2007 Systems Biology

Numerical Methods Equations are stiff, Jacobian is Sparse Reaction - Diffusion Reaction - Advection RADAR Must ||’ize well and must be accurate Special topics on Electrical and Systems Engineering 2007 Systems Biology

Reaction Operator : : Rosenbrock We have this for solving Inherently || for our distribution scheme Special topics on Electrical and Systems Engineering 2007 Systems Biology

Diffusion Operator Simple || algorithm based on explicit (FTCS) method Simple single processor method based on Crank. Nicolson || Crank-Nicolson method based on the fact that you are solving a slew of independent 1 D tri-diagonal solves Special topics on Electrical and Systems Engineering 2007 Systems Biology

Advection Operator Based on Lax (explicit, inherently || ) Nonlinear part of advection operator moved to reaction Operator Special topics on Electrical and Systems Engineering 2007 Systems Biology
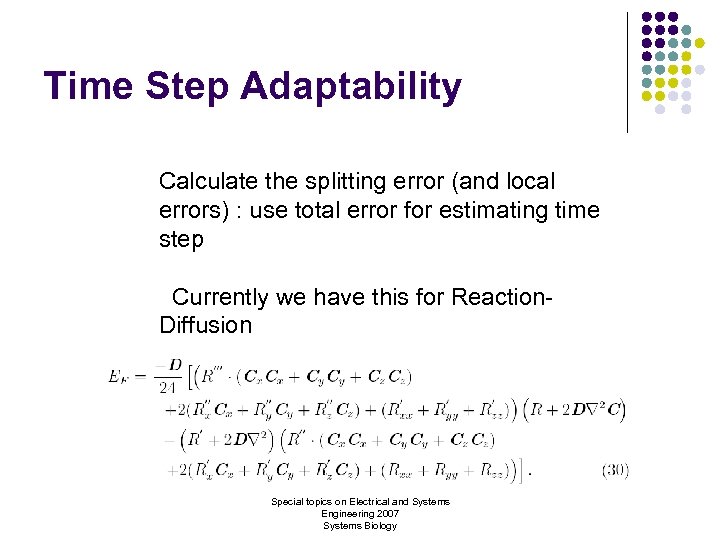

Time Step Adaptability Calculate the splitting error (and local errors) : use total error for estimating time step Currently we have this for Reaction. Diffusion Special topics on Electrical and Systems Engineering 2007 Systems Biology
Visualization Cross Platform QT/Open. GL Code Base Client / Server (Socket Based) GUI “Live” Interaction with Numerical Code 1 D/2 D/3 D time lapse contour plots Mouse-based interactive manipulation Flexible Modular Design To Be Implemented Point and Click graphical Entry of Biological Pathways, Grid Geometry, Initial Parameters and Control Special topics on Electrical and Systems Engineering 2007 Systems Biology

Who did the real work? l Dhruv Pant Ph. D. l Dave Miller Ray Zou Yihua Wang Hanbing Lin Tom Shortell Travis Hoppe Aparna Kumar l l l http: //bio. physics. drexel. edu/systems. Biology Email : avijit@physics. drexel. edu Special topics on Electrical and Systems Engineering 2007 Systems Biology
0dd6036afd1d8bfe6892be420651c564.ppt