63a5b4b684161026b05cbc96b2475e88.ppt
- Количество слайдов: 41

A Gentle Introduction to UCSC Genome Browser 陳任志, 游岳齊

Options l l l l l I. Genome Browser II. ENCODE III. Blat IV. Table Browser V. Gene Sorter VI. In Silico PCR VII. Proteome Browser VIII. Utilities IX. Downloads

I. Genome Browser Human (Homo sapiens) Genome Browser Gateway l Provides any section of entire human genome l Non-Standard Join Certificates l – some sequence joins between adjacent clones in this assembly could not be computationally validated l the sequencing center responsible for the particular chromosome provides an electronic certificate – should state why the submitter thinks the join is valid
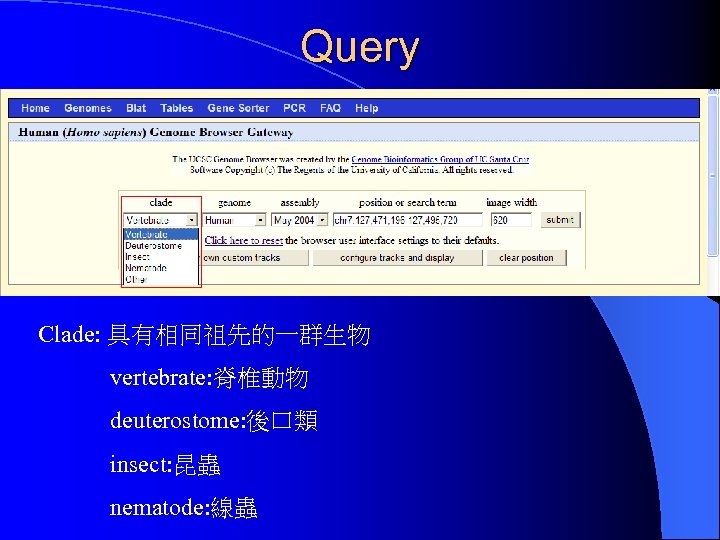
Query Clade: 具有相同祖先的一群生物 vertebrate: 脊椎動物 deuterostome: 後口類 insect: 昆蟲 nematode: 線蟲
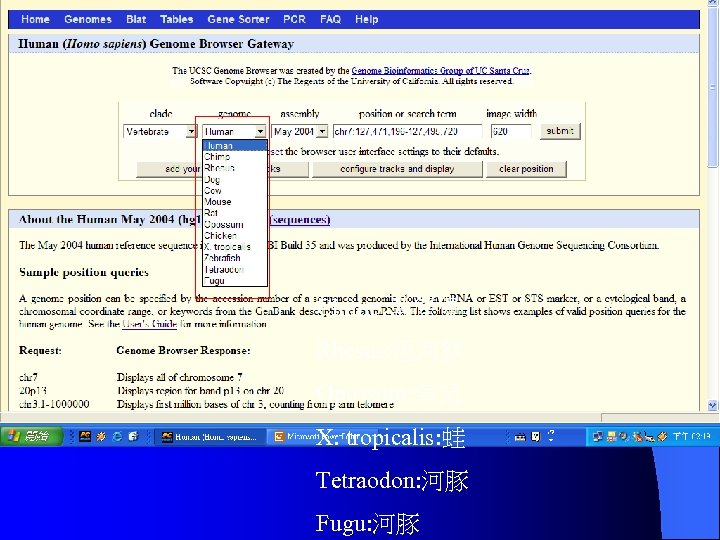
Chimp: 黑猩猩 Rhesus: 恆河猴 Opossum: 負鼠 X. tropicalis: 蛙 Tetraodon: 河豚 Fugu: 河豚
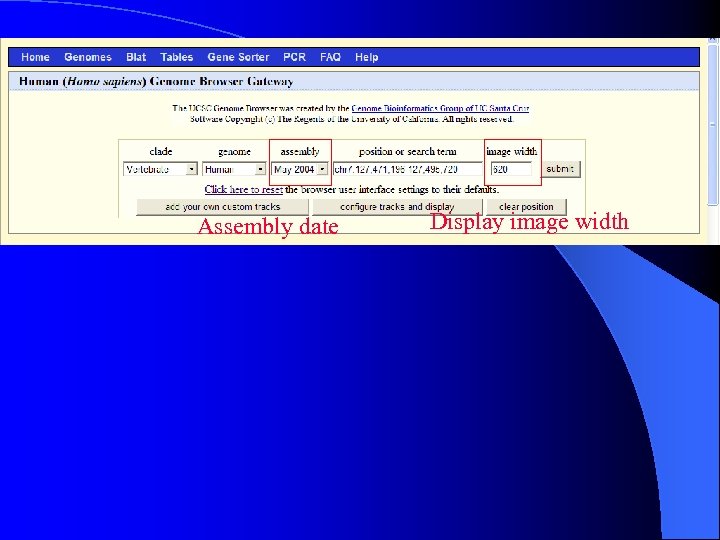
Assembly date Display image width
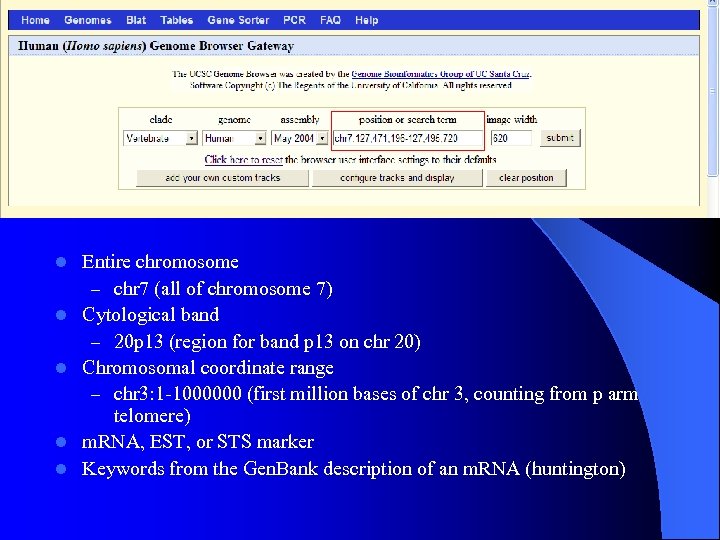
l l l Entire chromosome – chr 7 (all of chromosome 7) Cytological band – 20 p 13 (region for band p 13 on chr 20) Chromosomal coordinate range – chr 3: 1 -1000000 (first million bases of chr 3, counting from p arm telomere) m. RNA, EST, or STS marker Keywords from the Gen. Bank description of an m. RNA (huntington)
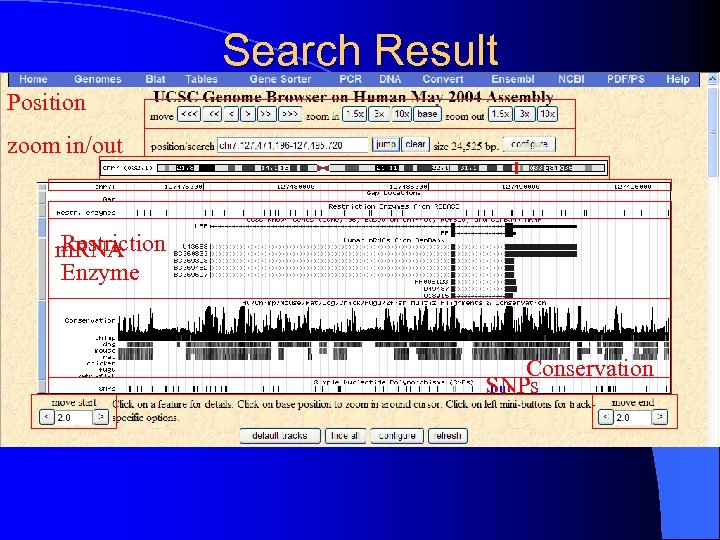
Search Result Position zoom in/out Restriction m. RNA Enzyme Conservation SNPs
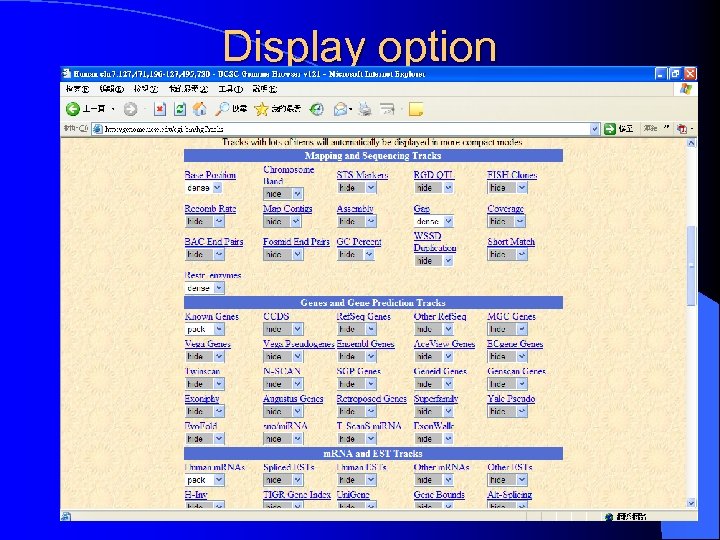
Display option

II. ENCODE Stands for “Encyclopedia Of DNA Elements” l Public research consortium to carry out a project to identify all functional elements in the human genome sequence l Launched by The National Human Genome Research Institute (NHGRI) l Conducted in three phases: l – pilot project phase (survey existing methods) – technology development phase (develop new methods) – planned production phase (…)

ENCODE Formats l Browser Extensible Data Format (BED) – for efficient access to genomic annotations l General Feature Format (GFF) – for data where there a set of linked features l Gene Transfer Format (GTF) – a refinement of GFF that tightens the specification l Multiple Alignment Format (MAF) – a series of multiple alignments in one format l Wiggle Format (WIG) – for continuous-valued data in track format

ENCODE Options l Regions (hg 16) – old database (+m. RNA, EST, & STS markers) l Regions (hg 17) – new database (+m. RNA, EST, & STS markers) l Data Status – the current status of ENCODE datasets l Downloads – sequence and annotation data downloads l Submission – for the submission of ENCODE-related data
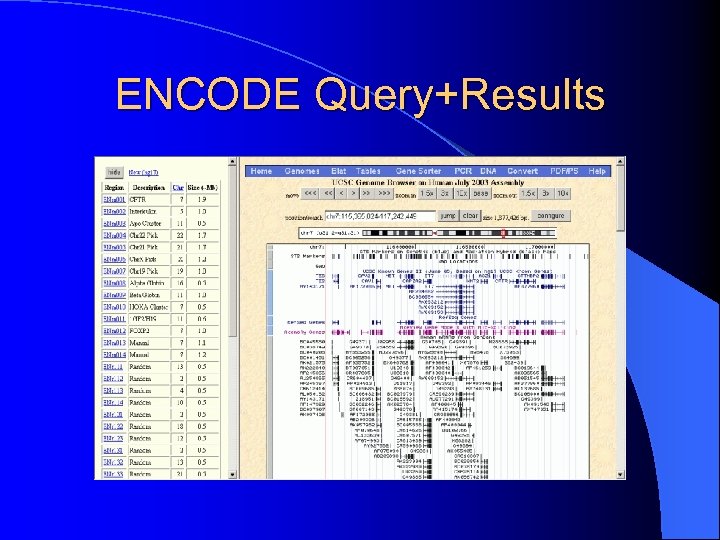
ENCODE Query+Results
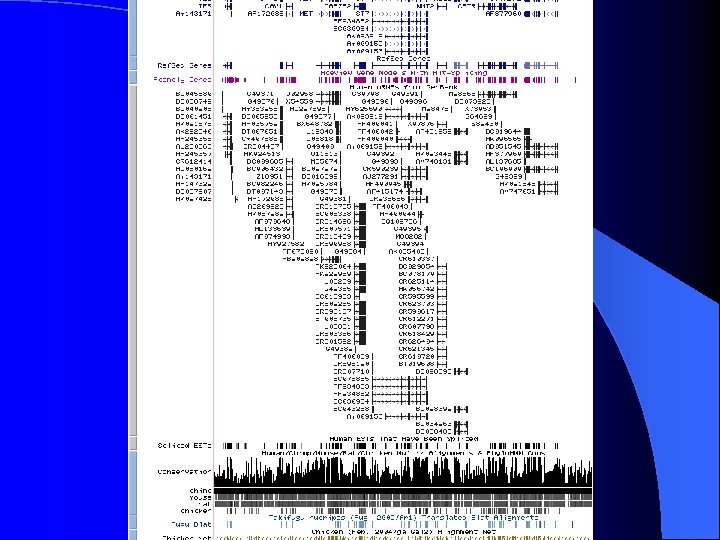
ENCODE Details hg 16
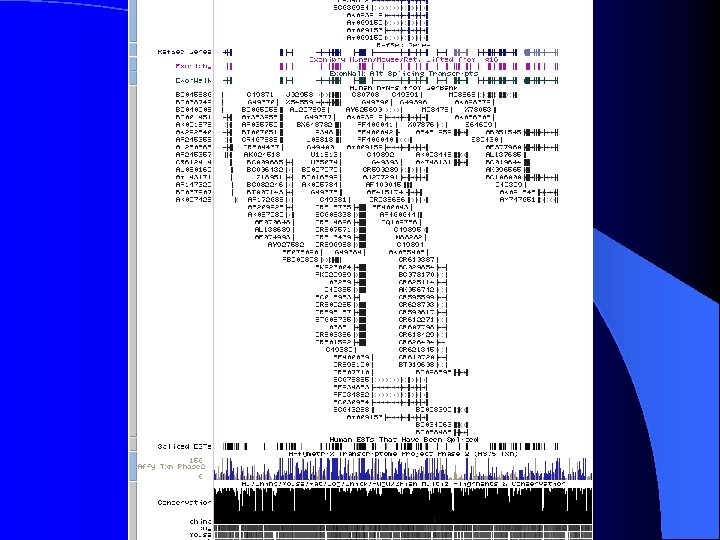
ENCODE Details hg 17

III. Blat l To quickly find sequences of 95% and greater similarity of length 40 bases or more l BLAST-Like Alignment Tool, not BLAST l Use: Paste in a query sequence to find its location in the genome l takes up just under 1 GB of RAM
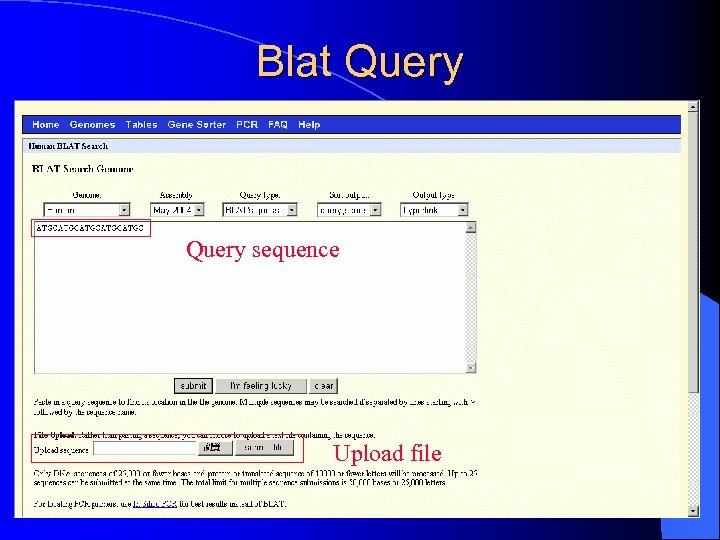
Blat Query sequence Upload file
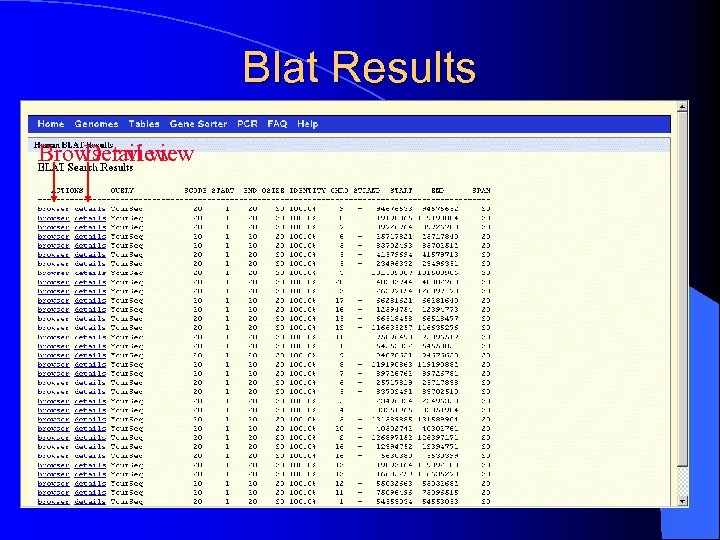
Blat Results Browser view Detail view

IV. Table Browser l To get the data associated with a track in text format, to calculate intersections between tracks, and to retrieve DNA sequence covered by a track
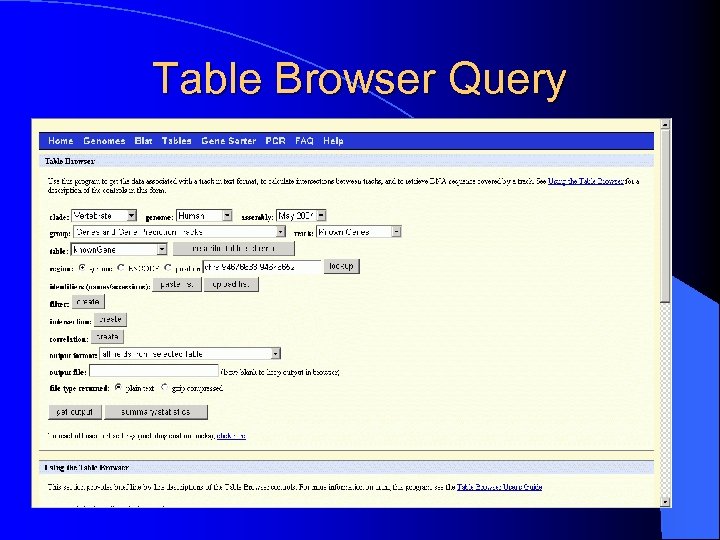
Table Browser Query
Table Browser Results

Table Browser Options l Describe Table Schema – schema for SQL table format l Filter – regular expression filter – range control l Intersection? ? l Correlation? ? l Summary Statistics
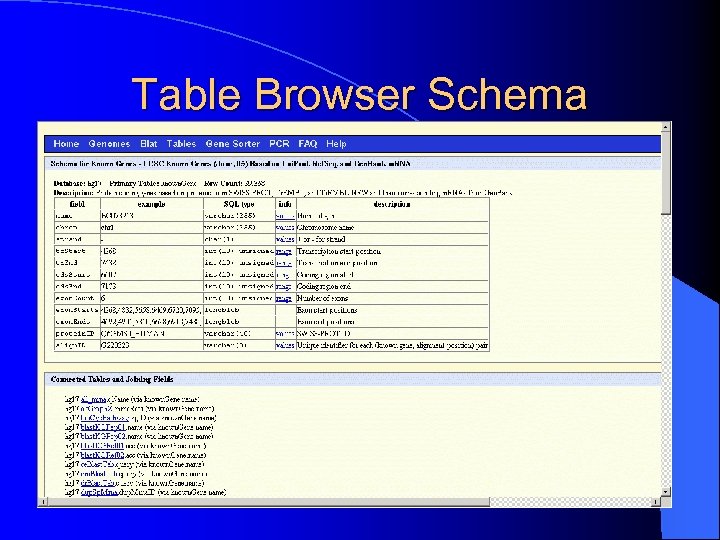
Table Browser Schema
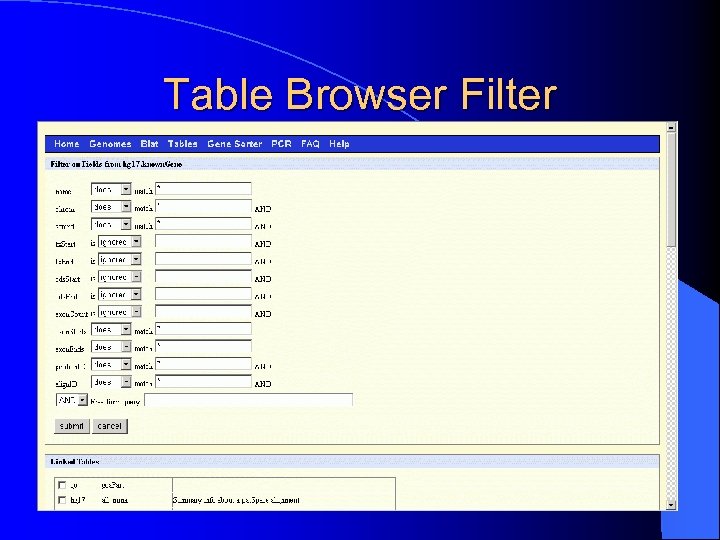
Table Browser Filter

Table Browser Summary Statistics

V. Gene Sorter l Displays a sorted table of genes that are related to one another l Correlation is color-coded – a highly expressed gene is colored – a less expressed gene is shown in green
Gene Sorter Query

Gene Sorter Results
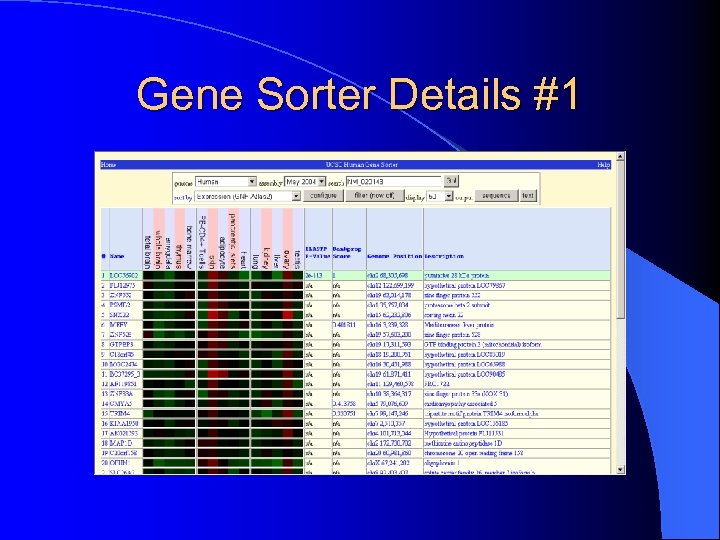
Gene Sorter Details #1
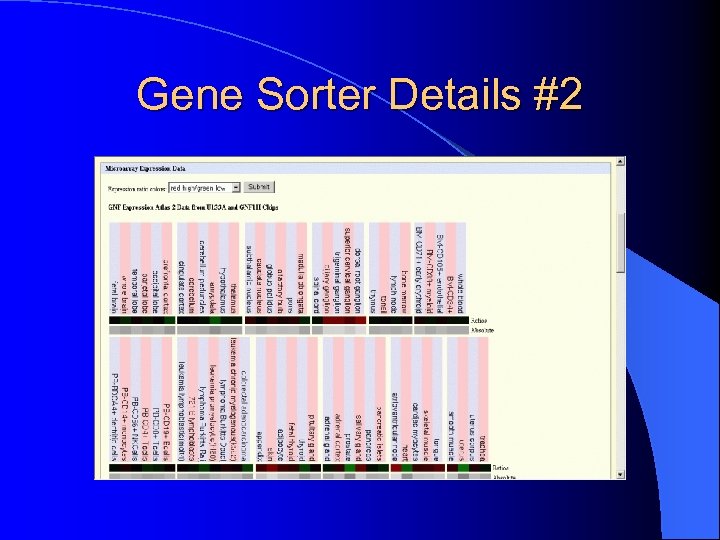

Gene Sorter Details #2

VI. In Silico PCR l In-Silico PCR searches a sequence database with a pair of PCR primers l Returns: a sequence output file in fasta format containing all sequence in the database that lie between and include the primer pair
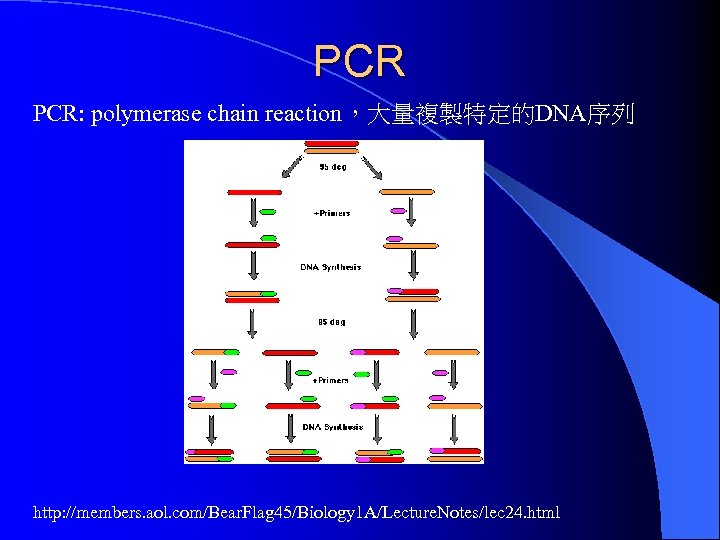
PCR PCR: polymerase chain reaction,大量複製特定的DNA序列 http: //members. aol. com/Bear. Flag 45/Biology 1 A/Lecture. Notes/lec 24. html
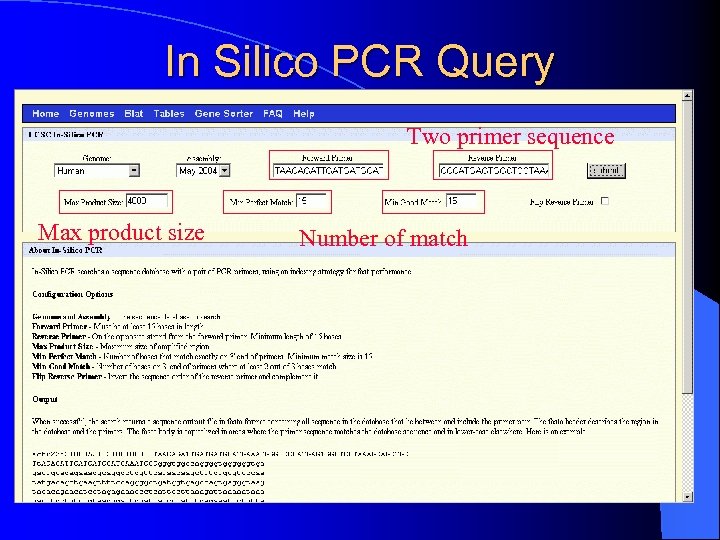

In Silico PCR Query Two primer sequence Max product size Number of match
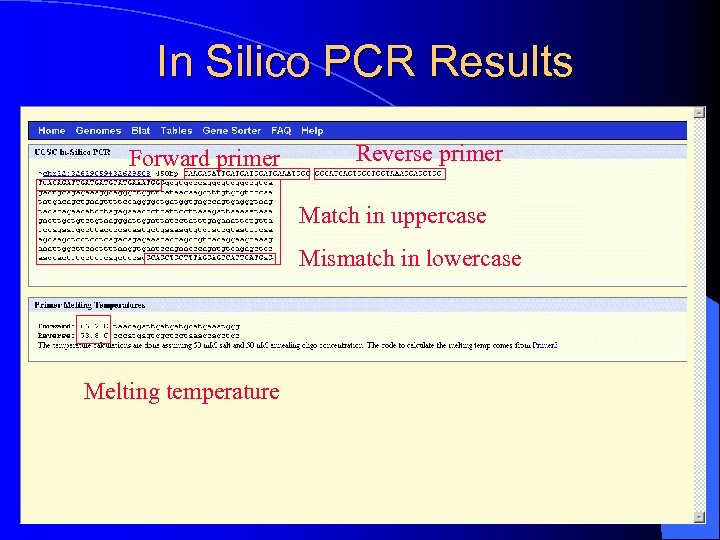

In Silico PCR Results Forward primer Reverse primer Match in uppercase Mismatch in lowercase Melting temperature

VII. Protein Browser UCSC Proteome Browser Gateway l provides a wealth of protein information presented in the form of graphical images and links to external internet sites l – – – Swiss. Prot information Proteome browser tracks Protein property histograms UCSC links / Domain information Comparative 3 D structures Pathways / Fasta format
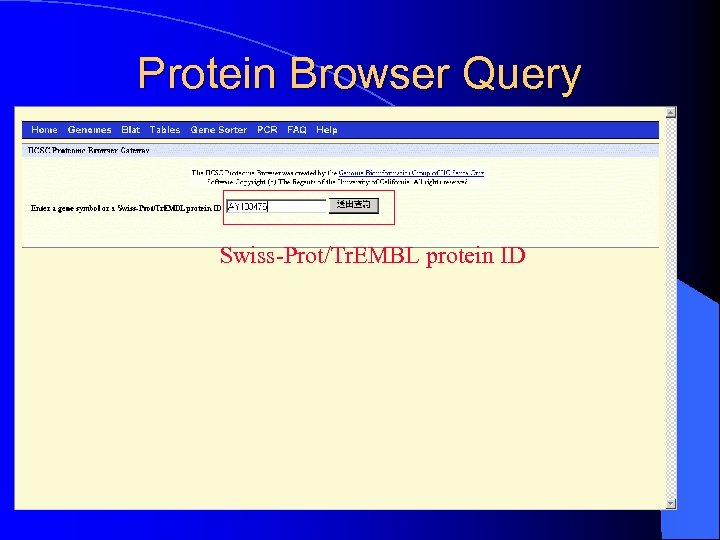
Protein Browser Query Swiss-Prot/Tr. EMBL protein ID
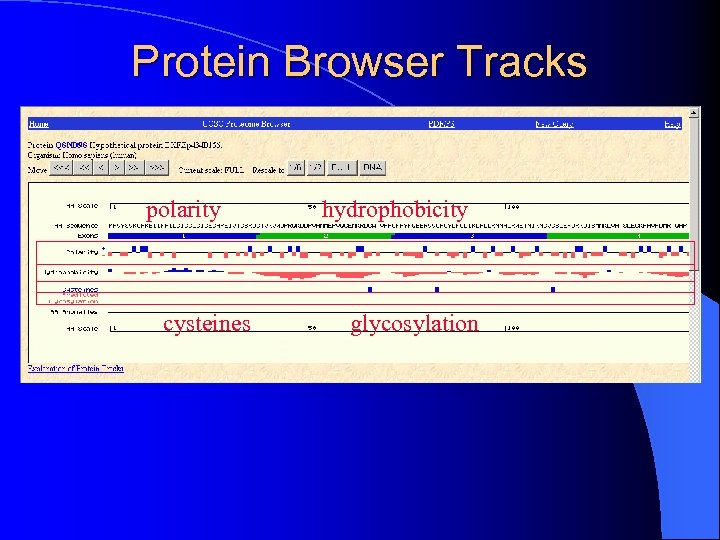

Protein Browser Tracks polarity cysteines hydrophobicity glycosylation
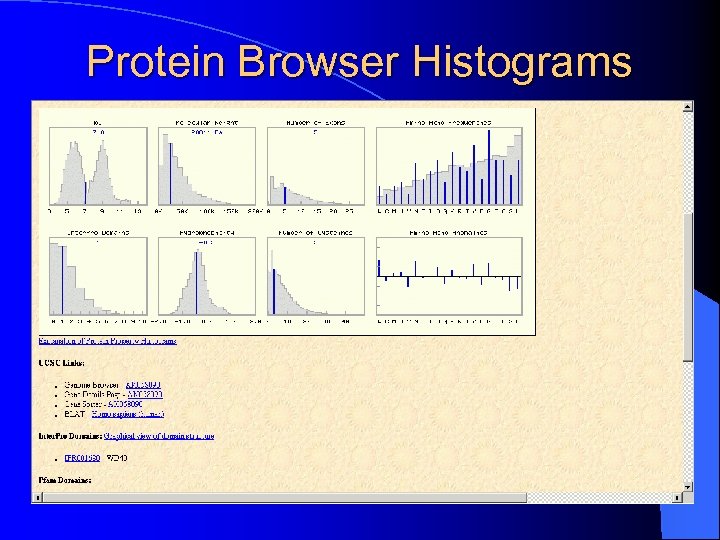
Protein Browser Histograms
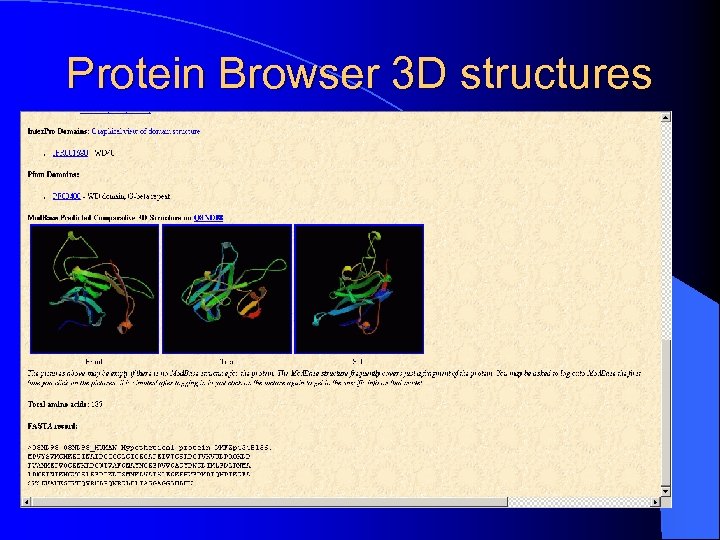
Protein Browser 3 D structures

VIII. Utilities l Some tools (for preparing input) – Batch Coordinate Conversion (lift. Over) l l converts genome coordinates and genome annotation files between assemblies WHY? – occasionally, a chunk of sequence may be moved to an entirely different chromosome as the map is refined – DNA Duster l formatting tool – Protein Duster l formatting tool

IX. Downloads l Offers downloads to complete genomes – – – – – Human Chimpanzee Rhesus Dog Cow Mouse Rat Opossum Chicken
63a5b4b684161026b05cbc96b2475e88.ppt