cb293044db676bf80255a294452c7908.ppt
- Количество слайдов: 29
 A couple of approaches to modelling and analysis of biochemical networks ”Biomodelling” seminar, October 2006 more an inspiration for a discussion than a talk. . . Matúš Kalaš
A couple of approaches to modelling and analysis of biochemical networks ”Biomodelling” seminar, October 2006 more an inspiration for a discussion than a talk. . . Matúš Kalaš
 Contents 1. The variety of modelling paradigms 2. An example of systematic approach (M. Heiner & D. Gilbert) 3. Another example (GOALIE; B. Mishra, M. Antoniotti et al. ) 2
Contents 1. The variety of modelling paradigms 2. An example of systematic approach (M. Heiner & D. Gilbert) 3. Another example (GOALIE; B. Mishra, M. Antoniotti et al. ) 2
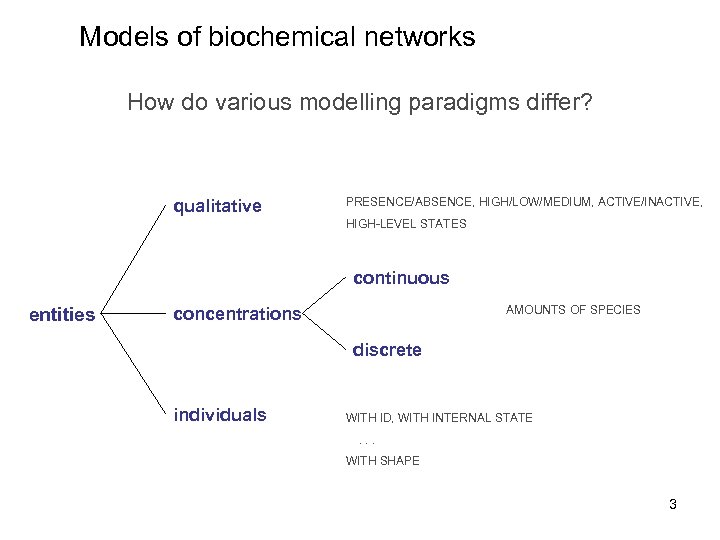 Models of biochemical networks How do various modelling paradigms differ? qualitative PRESENCE/ABSENCE, HIGH/LOW/MEDIUM, ACTIVE/INACTIVE, HIGH-LEVEL STATES continuous entities AMOUNTS OF SPECIES concentrations discrete individuals WITH ID, WITH INTERNAL STATE. . . WITH SHAPE 3
Models of biochemical networks How do various modelling paradigms differ? qualitative PRESENCE/ABSENCE, HIGH/LOW/MEDIUM, ACTIVE/INACTIVE, HIGH-LEVEL STATES continuous entities AMOUNTS OF SPECIES concentrations discrete individuals WITH ID, WITH INTERNAL STATE. . . WITH SHAPE 3
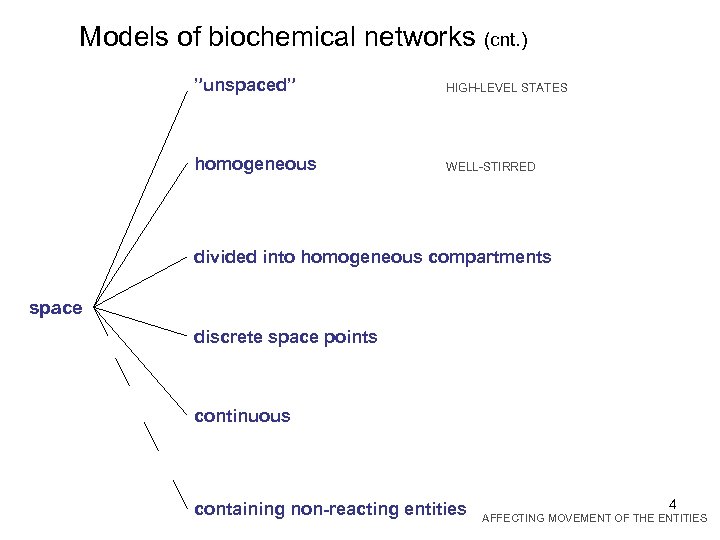 Models of biochemical networks (cnt. ) ”unspaced” HIGH-LEVEL STATES homogeneous WELL-STIRRED divided into homogeneous compartments space discrete space points continuous containing non-reacting entities 4 AFFECTING MOVEMENT OF THE ENTITIES
Models of biochemical networks (cnt. ) ”unspaced” HIGH-LEVEL STATES homogeneous WELL-STIRRED divided into homogeneous compartments space discrete space points continuous containing non-reacting entities 4 AFFECTING MOVEMENT OF THE ENTITIES
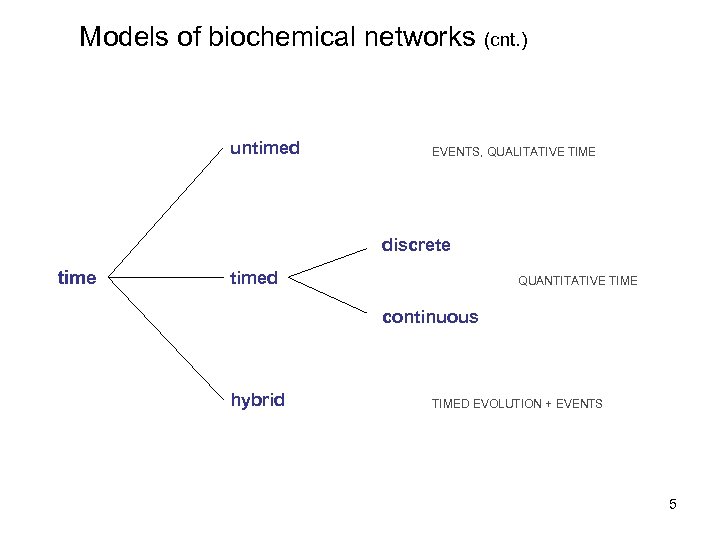 Models of biochemical networks (cnt. ) untimed EVENTS, QUALITATIVE TIME discrete timed QUANTITATIVE TIME continuous hybrid TIMED EVOLUTION + EVENTS 5
Models of biochemical networks (cnt. ) untimed EVENTS, QUALITATIVE TIME discrete timed QUANTITATIVE TIME continuous hybrid TIMED EVOLUTION + EVENTS 5
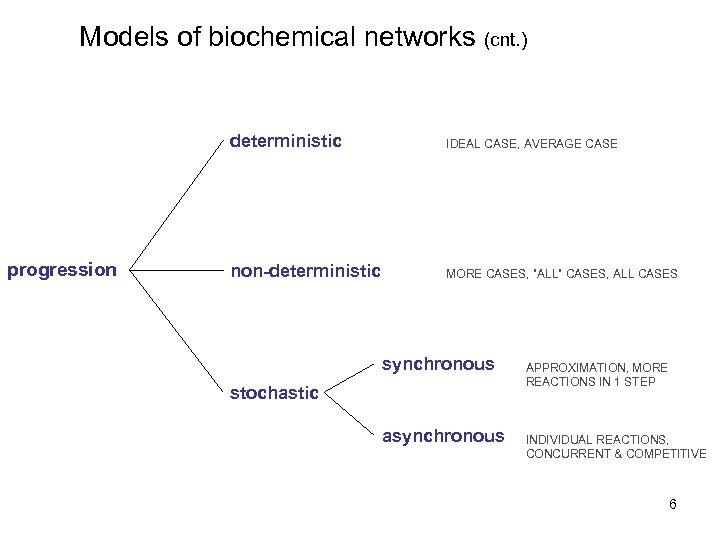 Models of biochemical networks (cnt. ) deterministic progression IDEAL CASE, AVERAGE CASE non-deterministic MORE CASES, ”ALL” CASES, ALL CASES synchronous stochastic asynchronous APPROXIMATION, MORE REACTIONS IN 1 STEP INDIVIDUAL REACTIONS, CONCURRENT & COMPETITIVE 6
Models of biochemical networks (cnt. ) deterministic progression IDEAL CASE, AVERAGE CASE non-deterministic MORE CASES, ”ALL” CASES, ALL CASES synchronous stochastic asynchronous APPROXIMATION, MORE REACTIONS IN 1 STEP INDIVIDUAL REACTIONS, CONCURRENT & COMPETITIVE 6
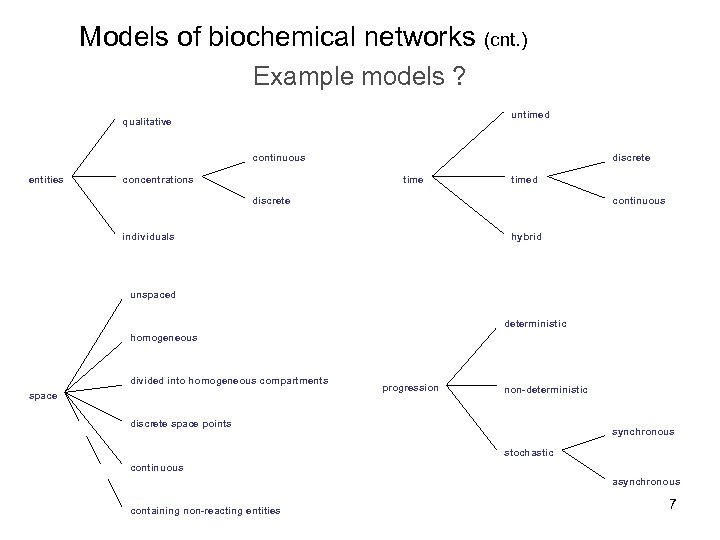 Models of biochemical networks (cnt. ) Example models ? untimed qualitative continuous entities concentrations discrete timed continuous discrete hybrid individuals unspaced deterministic homogeneous divided into homogeneous compartments space progression non-deterministic discrete space points synchronous stochastic continuous asynchronous containing non-reacting entities 7
Models of biochemical networks (cnt. ) Example models ? untimed qualitative continuous entities concentrations discrete timed continuous discrete hybrid individuals unspaced deterministic homogeneous divided into homogeneous compartments space progression non-deterministic discrete space points synchronous stochastic continuous asynchronous containing non-reacting entities 7
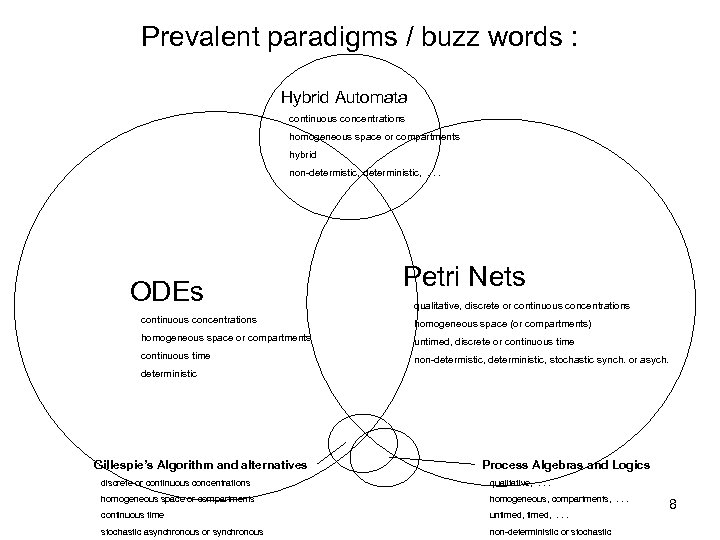 Prevalent paradigms / buzz words : Hybrid Automata continuous concentrations homogeneous space or compartments hybrid non-determistic, deterministic, . . . ODEs Petri Nets qualitative, discrete or continuous concentrations homogeneous space (or compartments) homogeneous space or compartments untimed, discrete or continuous time non-determistic, deterministic, stochastic synch. or asych. deterministic Gillespie’s Algorithm and alternatives Process Algebras and Logics discrete or continuous concentrations qualitative, . . . homogeneous space or compartments homogeneous, compartments, . . . continuous time untimed, . . . stochastic asynchronous or synchronous non-deterministic or stochastic 8
Prevalent paradigms / buzz words : Hybrid Automata continuous concentrations homogeneous space or compartments hybrid non-determistic, deterministic, . . . ODEs Petri Nets qualitative, discrete or continuous concentrations homogeneous space (or compartments) homogeneous space or compartments untimed, discrete or continuous time non-determistic, deterministic, stochastic synch. or asych. deterministic Gillespie’s Algorithm and alternatives Process Algebras and Logics discrete or continuous concentrations qualitative, . . . homogeneous space or compartments homogeneous, compartments, . . . continuous time untimed, . . . stochastic asynchronous or synchronous non-deterministic or stochastic 8
 Now an instant introduction to Petri Nets. . . 9
Now an instant introduction to Petri Nets. . . 9
 An example of systematic modelling: Step-wise modelling David Gilbert, Monika Heiner: From Petri Nets to Differential Equations – An Integrative Approach for Biochemical Network Analysis ICATPN 2006, TR 2005 . . . a tutorial example of • different useful features of different modelling paradigms • step-wise modelling 10
An example of systematic modelling: Step-wise modelling David Gilbert, Monika Heiner: From Petri Nets to Differential Equations – An Integrative Approach for Biochemical Network Analysis ICATPN 2006, TR 2005 . . . a tutorial example of • different useful features of different modelling paradigms • step-wise modelling 10
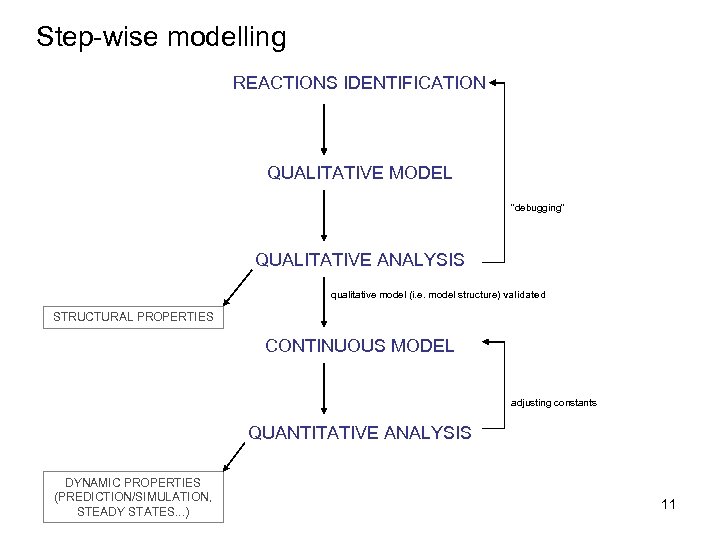 Step-wise modelling REACTIONS IDENTIFICATION QUALITATIVE MODEL ”debugging” QUALITATIVE ANALYSIS qualitative model (i. e. model structure) validated STRUCTURAL PROPERTIES CONTINUOUS MODEL adjusting constants QUANTITATIVE ANALYSIS DYNAMIC PROPERTIES (PREDICTION/SIMULATION, STEADY STATES. . . ) 11
Step-wise modelling REACTIONS IDENTIFICATION QUALITATIVE MODEL ”debugging” QUALITATIVE ANALYSIS qualitative model (i. e. model structure) validated STRUCTURAL PROPERTIES CONTINUOUS MODEL adjusting constants QUANTITATIVE ANALYSIS DYNAMIC PROPERTIES (PREDICTION/SIMULATION, STEADY STATES. . . ) 11
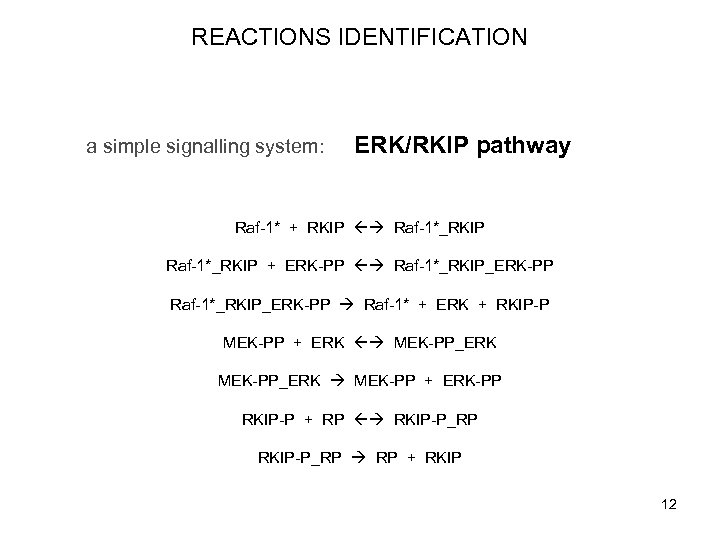 REACTIONS IDENTIFICATION a simple signalling system: ERK/RKIP pathway Raf-1* + RKIP Raf-1*_RKIP + ERK-PP Raf-1*_RKIP_ERK-PP Raf-1* + ERK + RKIP-P MEK-PP + ERK MEK-PP_ERK MEK-PP + ERK-PP RKIP-P + RP RKIP-P_RP RP + RKIP 12
REACTIONS IDENTIFICATION a simple signalling system: ERK/RKIP pathway Raf-1* + RKIP Raf-1*_RKIP + ERK-PP Raf-1*_RKIP_ERK-PP Raf-1* + ERK + RKIP-P MEK-PP + ERK MEK-PP_ERK MEK-PP + ERK-PP RKIP-P + RP RKIP-P_RP RP + RKIP 12
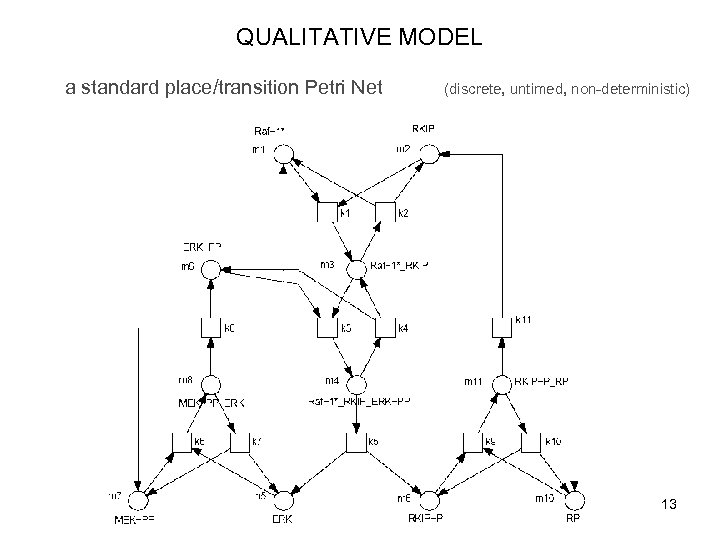 QUALITATIVE MODEL a standard place/transition Petri Net (discrete, untimed, non-deterministic) 13
QUALITATIVE MODEL a standard place/transition Petri Net (discrete, untimed, non-deterministic) 13
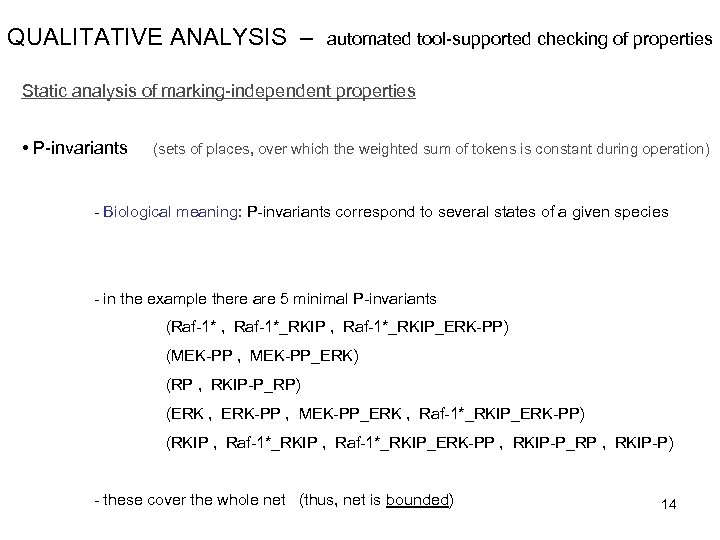 QUALITATIVE ANALYSIS – automated tool-supported checking of properties Static analysis of marking-independent properties • P-invariants (sets of places, over which the weighted sum of tokens is constant during operation) - Biological meaning: P-invariants correspond to several states of a given species - in the example there are 5 minimal P-invariants (Raf-1* , Raf-1*_RKIP_ERK-PP) (MEK-PP , MEK-PP_ERK) (RP , RKIP-P_RP) (ERK , ERK-PP , MEK-PP_ERK , Raf-1*_RKIP_ERK-PP) (RKIP , Raf-1*_RKIP_ERK-PP , RKIP-P_RP , RKIP-P) - these cover the whole net (thus, net is bounded) 14
QUALITATIVE ANALYSIS – automated tool-supported checking of properties Static analysis of marking-independent properties • P-invariants (sets of places, over which the weighted sum of tokens is constant during operation) - Biological meaning: P-invariants correspond to several states of a given species - in the example there are 5 minimal P-invariants (Raf-1* , Raf-1*_RKIP_ERK-PP) (MEK-PP , MEK-PP_ERK) (RP , RKIP-P_RP) (ERK , ERK-PP , MEK-PP_ERK , Raf-1*_RKIP_ERK-PP) (RKIP , Raf-1*_RKIP_ERK-PP , RKIP-P_RP , RKIP-P) - these cover the whole net (thus, net is bounded) 14
 QUALITATIVE ANALYSIS Static analysis of marking-independent properties (cnt. ) • T-invariants - can be also read as the relative firing rates of transitions (reactions/phases in sysbio) (this corresponds to the steady-state behaviour) - minimal T-invariants characterise minimal self-contained subnetworks with an enclosed biological meaning - useful to comprehend the network if it is very complex {not in this tutorial example} - example net is covered by T-invariants - only 1 non-trivial minimal T-invariant: (k 1; k 3; k 5; (k 6; k 8), (k 9; k 11)) 15
QUALITATIVE ANALYSIS Static analysis of marking-independent properties (cnt. ) • T-invariants - can be also read as the relative firing rates of transitions (reactions/phases in sysbio) (this corresponds to the steady-state behaviour) - minimal T-invariants characterise minimal self-contained subnetworks with an enclosed biological meaning - useful to comprehend the network if it is very complex {not in this tutorial example} - example net is covered by T-invariants - only 1 non-trivial minimal T-invariant: (k 1; k 3; k 5; (k 6; k 8), (k 9; k 11)) 15
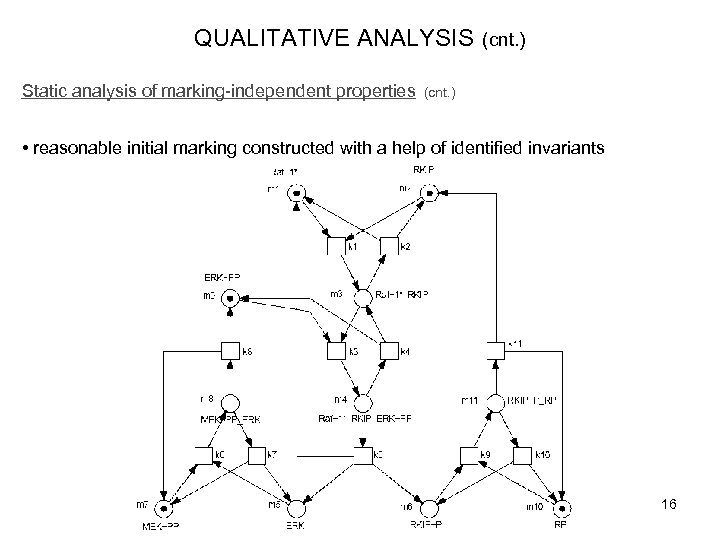 QUALITATIVE ANALYSIS Static analysis of marking-independent properties (cnt. ) • reasonable initial marking constructed with a help of identified invariants 16
QUALITATIVE ANALYSIS Static analysis of marking-independent properties (cnt. ) • reasonable initial marking constructed with a help of identified invariants 16
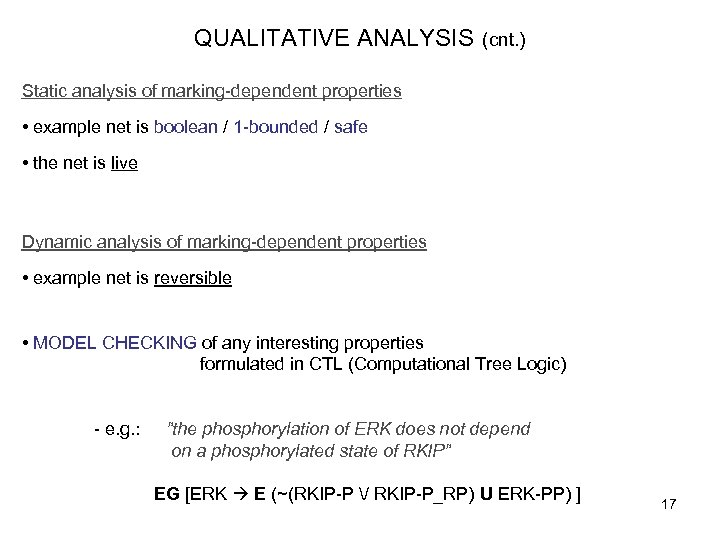 QUALITATIVE ANALYSIS (cnt. ) Static analysis of marking-dependent properties • example net is boolean / 1 -bounded / safe • the net is live Dynamic analysis of marking-dependent properties • example net is reversible • MODEL CHECKING of any interesting properties formulated in CTL (Computational Tree Logic) - e. g. : ”the phosphorylation of ERK does not depend on a phosphorylated state of RKIP” EG [ERK E (~(RKIP-P / RKIP-P_RP) U ERK-PP) ] 17
QUALITATIVE ANALYSIS (cnt. ) Static analysis of marking-dependent properties • example net is boolean / 1 -bounded / safe • the net is live Dynamic analysis of marking-dependent properties • example net is reversible • MODEL CHECKING of any interesting properties formulated in CTL (Computational Tree Logic) - e. g. : ”the phosphorylation of ERK does not depend on a phosphorylated state of RKIP” EG [ERK E (~(RKIP-P / RKIP-P_RP) U ERK-PP) ] 17
 QUALITATIVE ANALYSIS (cnt. ) VALIDATION OF THE QUALITATIVE MODEL (i. e. structure of the system) ü all expected structural and general behavioural properties hold ü covered by P-invariants ü no minimal P-invariant without biological interpretation ü covered by T-invariants ü no minimal T-invariant without biological interpretation ü no known biological behaviour without corresponding T-invariant ü all expected logic-formulated properties hold a break? 18
QUALITATIVE ANALYSIS (cnt. ) VALIDATION OF THE QUALITATIVE MODEL (i. e. structure of the system) ü all expected structural and general behavioural properties hold ü covered by P-invariants ü no minimal P-invariant without biological interpretation ü covered by T-invariants ü no minimal T-invariant without biological interpretation ü no known biological behaviour without corresponding T-invariant ü all expected logic-formulated properties hold a break? 18
 CONTINUOUS QUANTITATIVE MODEL Continuous Petri Net - tokens: real numbers - transitions associated with a rate - semantics: a set of ODEs (e. g. reaction-rate equation) - thus a continuous, timed (continuously) and deterministic model - basically a set of ODEs enhanced with a graphical representation - within this step, all we need is to find suitable rate constants (e. g. to fit in-vivo or in-vitro quantitative experiments) 19
CONTINUOUS QUANTITATIVE MODEL Continuous Petri Net - tokens: real numbers - transitions associated with a rate - semantics: a set of ODEs (e. g. reaction-rate equation) - thus a continuous, timed (continuously) and deterministic model - basically a set of ODEs enhanced with a graphical representation - within this step, all we need is to find suitable rate constants (e. g. to fit in-vivo or in-vitro quantitative experiments) 19
 QUANTITATIVE ANALYSIS • Prediction (easy) - both qualitative and quantitative • Steady-state properties, oscillations, sensibility, . . . (hard) (. . . you know better. . . ) 20
QUANTITATIVE ANALYSIS • Prediction (easy) - both qualitative and quantitative • Steady-state properties, oscillations, sensibility, . . . (hard) (. . . you know better. . . ) 20
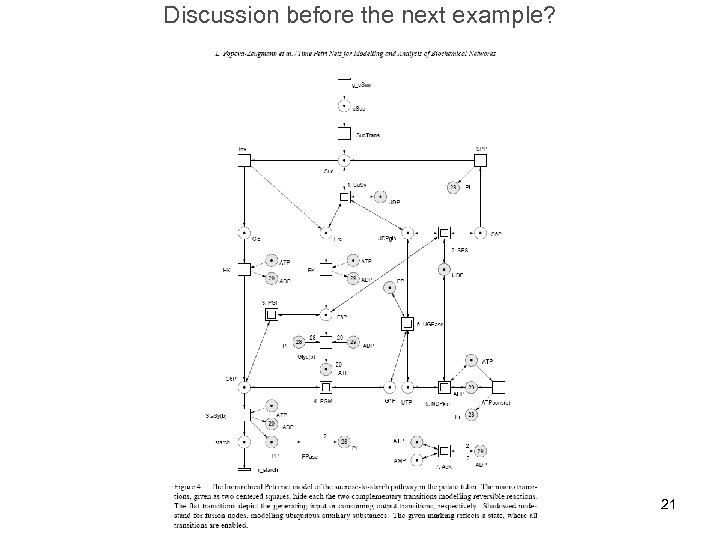 Discussion before the next example? 21
Discussion before the next example? 21
 Another example: Automated modelling. . . building a model in order to understand very complex processes. . . Samantha Kleinberg, Marco Antoniotti, Satish Tadepalli, Naren Ramakrishnan, Bud Mishra: Remembrance of Experiments Past: A redescription based tool for discovery in complex systems ICCS 2006 Marco Antoniotti, Naren Ramakrishnan, Bud Mishra: GOALIE, A Common Lisp Application to Discover Kripke Models: Redescribing Biological Processes from Time-Course Data ILC 2005 22
Another example: Automated modelling. . . building a model in order to understand very complex processes. . . Samantha Kleinberg, Marco Antoniotti, Satish Tadepalli, Naren Ramakrishnan, Bud Mishra: Remembrance of Experiments Past: A redescription based tool for discovery in complex systems ICCS 2006 Marco Antoniotti, Naren Ramakrishnan, Bud Mishra: GOALIE, A Common Lisp Application to Discover Kripke Models: Redescribing Biological Processes from Time-Course Data ILC 2005 22
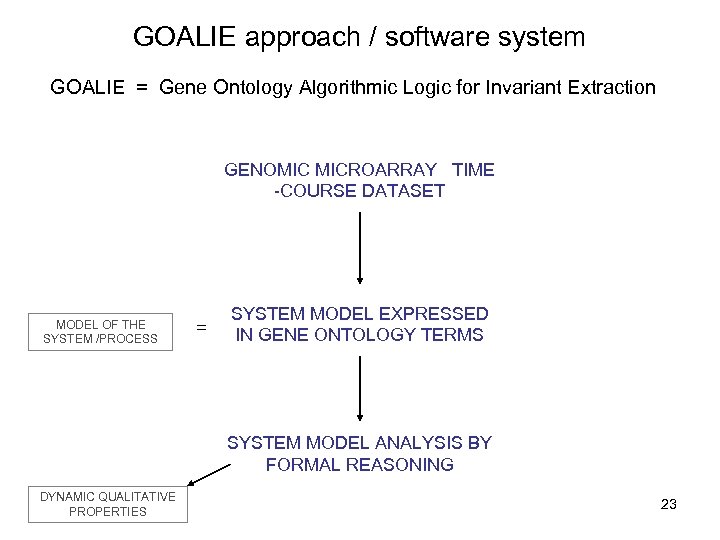 GOALIE approach / software system GOALIE = Gene Ontology Algorithmic Logic for Invariant Extraction GENOMIC MICROARRAY TIME -COURSE DATASET MODEL OF THE SYSTEM /PROCESS = SYSTEM MODEL EXPRESSED IN GENE ONTOLOGY TERMS SYSTEM MODEL ANALYSIS BY FORMAL REASONING DYNAMIC QUALITATIVE PROPERTIES 23
GOALIE approach / software system GOALIE = Gene Ontology Algorithmic Logic for Invariant Extraction GENOMIC MICROARRAY TIME -COURSE DATASET MODEL OF THE SYSTEM /PROCESS = SYSTEM MODEL EXPRESSED IN GENE ONTOLOGY TERMS SYSTEM MODEL ANALYSIS BY FORMAL REASONING DYNAMIC QUALITATIVE PROPERTIES 23
 GOALIE approach / software system (cnt. ) Model: Kripke Structure - called also ”Hidden Kripke Model” in GOALIE - annotated by Gene Ontology terms (propositional logic) - qualitative, high-level, untimed and non-deterministic model with clear biological meaning 24
GOALIE approach / software system (cnt. ) Model: Kripke Structure - called also ”Hidden Kripke Model” in GOALIE - annotated by Gene Ontology terms (propositional logic) - qualitative, high-level, untimed and non-deterministic model with clear biological meaning 24
 GOALIE approach / software system (cnt. ) Controlled vocabulary: Gene Ontology 8517 possible GO process ontology terms 25
GOALIE approach / software system (cnt. ) Controlled vocabulary: Gene Ontology 8517 possible GO process ontology terms 25
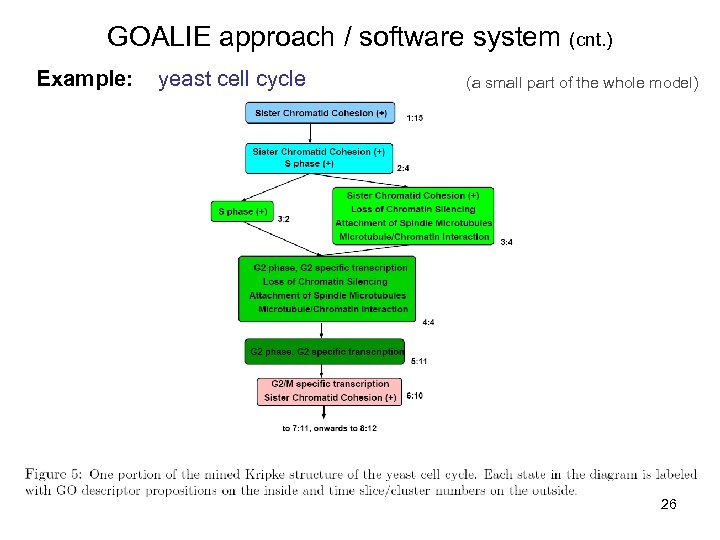 GOALIE approach / software system (cnt. ) Example: yeast cell cycle (a small part of the whole model) 26
GOALIE approach / software system (cnt. ) Example: yeast cell cycle (a small part of the whole model) 26
 GOALIE approach / software system (cnt. ) Techniques used to automatically build a model: - time-windowed clustering (k-means) data-to-GO association done by Go. Miner software Fisher exact test (p-values) empirical Bayes approach (Benjamini-Hochberg test) information bottleneck principle (generalised Shannon-Kolmogorov’s rate-distortion theory) - connecting annotated clusters (Jaccard’s coefficient) Analysis: - propositional temporal-logic reasoning (model checking of temporal invariants (CTL)) - graph rewriting rules for projection and collapsing, preserving ”bisimulation-like” relations getting higher-level clusters - process / dataset alignment (similarity of cellular processes) 27
GOALIE approach / software system (cnt. ) Techniques used to automatically build a model: - time-windowed clustering (k-means) data-to-GO association done by Go. Miner software Fisher exact test (p-values) empirical Bayes approach (Benjamini-Hochberg test) information bottleneck principle (generalised Shannon-Kolmogorov’s rate-distortion theory) - connecting annotated clusters (Jaccard’s coefficient) Analysis: - propositional temporal-logic reasoning (model checking of temporal invariants (CTL)) - graph rewriting rules for projection and collapsing, preserving ”bisimulation-like” relations getting higher-level clusters - process / dataset alignment (similarity of cellular processes) 27
 A couple of diverse systematic approaches: • C. Wiggins, I. Nemenman: Process Pathway Inference via Time Series Analysis, 2006 • M. Calder, S. Gilmore, J. Hillston: Automatically deriving ODEs from process algebra models of signalling pathways, CMSB 2005 • N. Chabrier-Rivier, M. Chiaverini, V. Danos, F. Fages, V. Schächter: Modeling and Querying Biomolecular Interaction Networks, TCS 2004 • A. Arkin, P. Shen, J. Ross: A Test Case of Correlation Metric Construction of a Reaction Pathway from Measurements, Science 1997 • M. Chen, R. Hofestädt: A medical bioinformatics approach for metabolic disorders: Biomedical data prediction, modeling, and systematic analysis, JBMI 2006 28
A couple of diverse systematic approaches: • C. Wiggins, I. Nemenman: Process Pathway Inference via Time Series Analysis, 2006 • M. Calder, S. Gilmore, J. Hillston: Automatically deriving ODEs from process algebra models of signalling pathways, CMSB 2005 • N. Chabrier-Rivier, M. Chiaverini, V. Danos, F. Fages, V. Schächter: Modeling and Querying Biomolecular Interaction Networks, TCS 2004 • A. Arkin, P. Shen, J. Ross: A Test Case of Correlation Metric Construction of a Reaction Pathway from Measurements, Science 1997 • M. Chen, R. Hofestädt: A medical bioinformatics approach for metabolic disorders: Biomedical data prediction, modeling, and systematic analysis, JBMI 2006 28
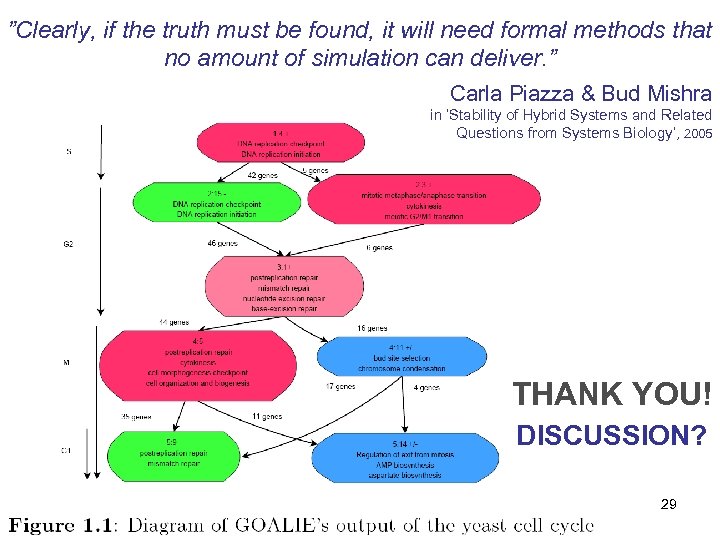 ”Clearly, if the truth must be found, it will need formal methods that no amount of simulation can deliver. ” Carla Piazza & Bud Mishra in ’Stability of Hybrid Systems and Related Questions from Systems Biology’, 2005 THANK YOU! DISCUSSION? 29
”Clearly, if the truth must be found, it will need formal methods that no amount of simulation can deliver. ” Carla Piazza & Bud Mishra in ’Stability of Hybrid Systems and Related Questions from Systems Biology’, 2005 THANK YOU! DISCUSSION? 29


