c870716b85b31635d48fa916ddb9c49b.ppt
- Количество слайдов: 83

0 GBCS ecosystem for NGS Data Management with em. BASE Data analysis with Galaxy
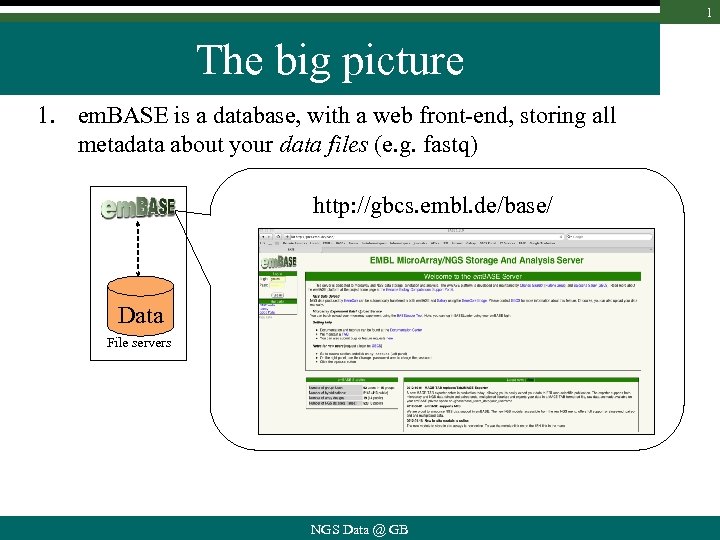
1 The big picture 1. em. BASE is a database, with a web front-end, storing all metadata about your data files (e. g. fastq) http: //gbcs. embl. de/base/ Data File servers NGS Data @ GB
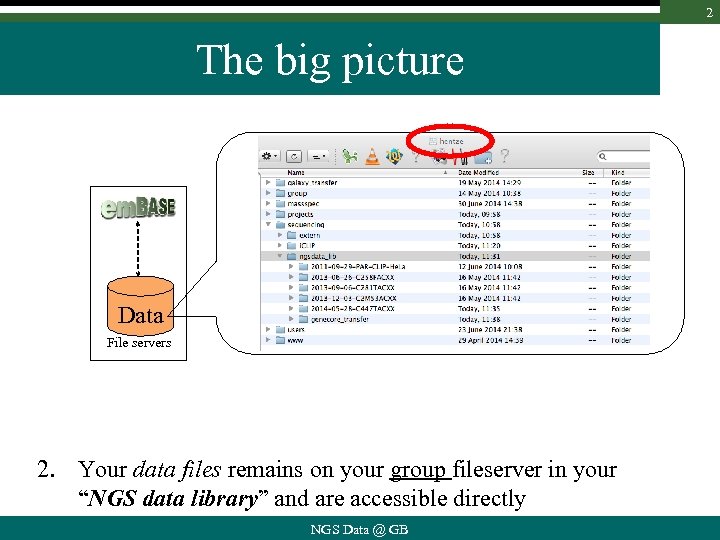
2 The big picture Data File servers 2. Your data files remains on your group fileserver in your “NGS data library” and are accessible directly NGS Data @ GB
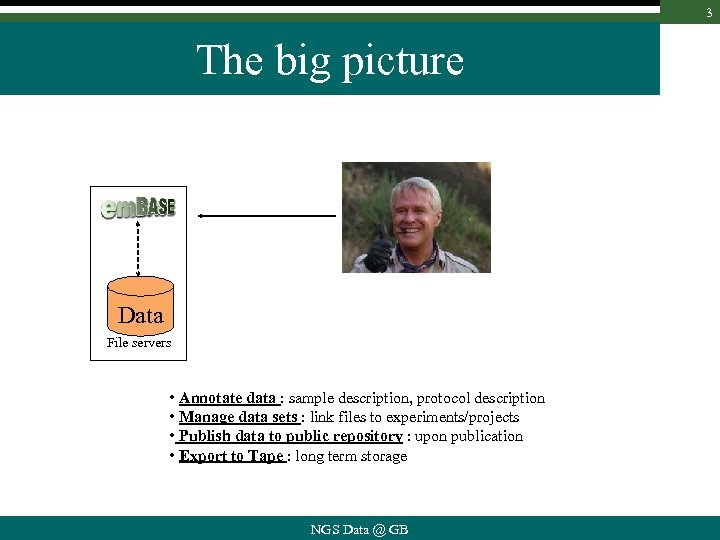

3 The big picture Data File servers • Annotate data : sample description, protocol description • Manage data sets : link files to experiments/projects • Publish data to public repository : upon publication • Export to Tape : long term storage NGS Data @ GB
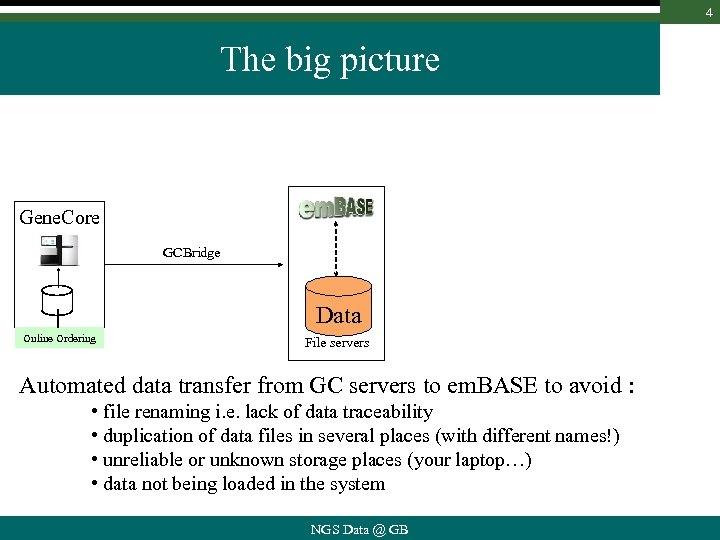
4 The big picture Gene. Core GCBridge Data Online Ordering File servers Automated data transfer from GC servers to em. BASE to avoid : • file renaming i. e. lack of data traceability • duplication of data files in several places (with different names!) • unreliable or unknown storage places (your laptop…) • data not being loaded in the system NGS Data @ GB
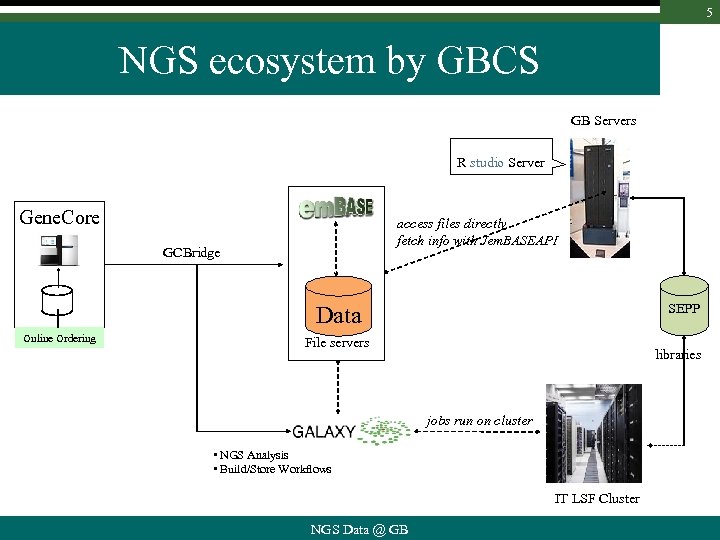
5 NGS ecosystem by GBCS GB Servers R studio Server Gene. Core access files directly fetch info with Jem. BASEAPI GCBridge SEPP Data Online Ordering File servers libraries jobs run on cluster • NGS Analysis • Build/Store Workflows IT LSF Cluster NGS Data @ GB

6 0. What is the “data”
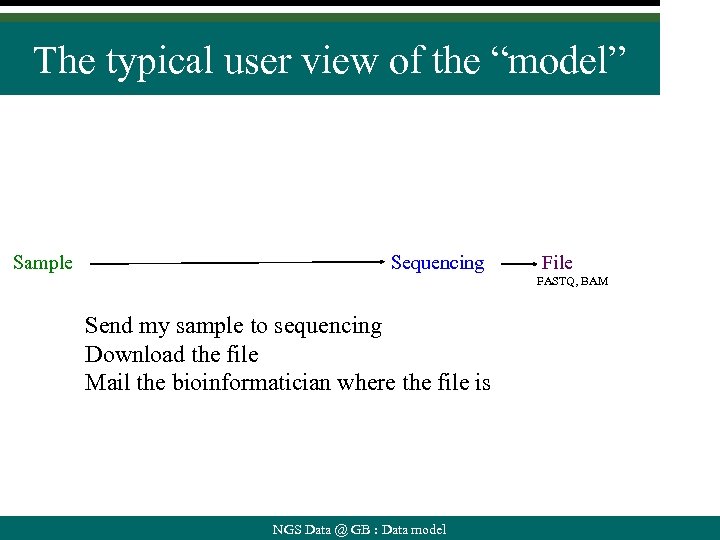

The typical user view of the “model” Sample Sequencing File FASTQ, BAM Send my sample to sequencing Download the file Mail the bioinformatician where the file is NGS Data @ GB : Data model
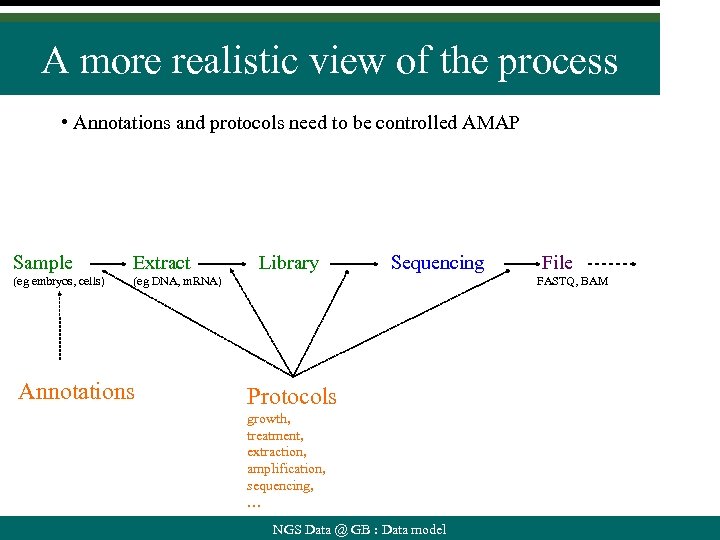
A more realistic view of the process • Annotations and protocols need to be controlled AMAP Sample Extract (eg embryos, cells) (eg DNA, m. RNA) Annotations Library Sequencing File FASTQ, BAM Protocols growth, treatment, extraction, amplification, sequencing, … NGS Data @ GB : Data model
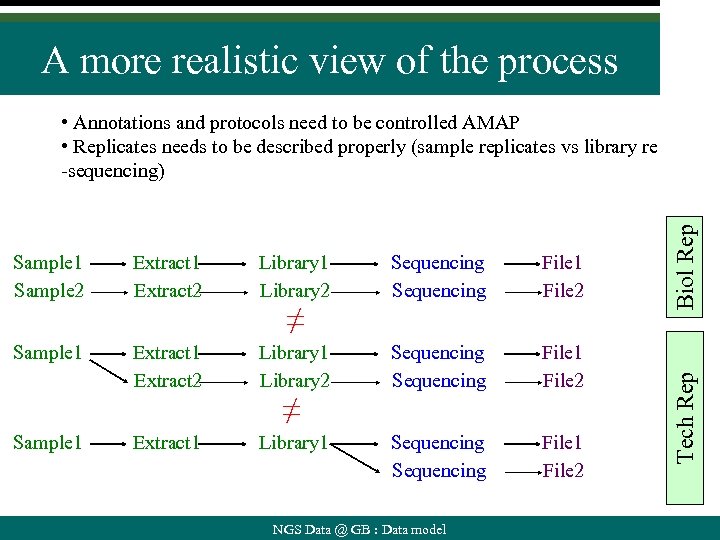
A more realistic view of the process Extract 1 Extract 2 Library 1 Library 2 Sequencing File 1 File 2 Sample 1 Extract 2 Library 1 Library 2 Sequencing File 1 File 2 Extract 1 Library 1 Sequencing File 1 File 2 Sample 1 ≠ ≠ NGS Data @ GB : Data model Tech Rep Sample 1 Sample 2 Biol Rep • Annotations and protocols need to be controlled AMAP • Replicates needs to be described properly (sample replicates vs library re -sequencing)
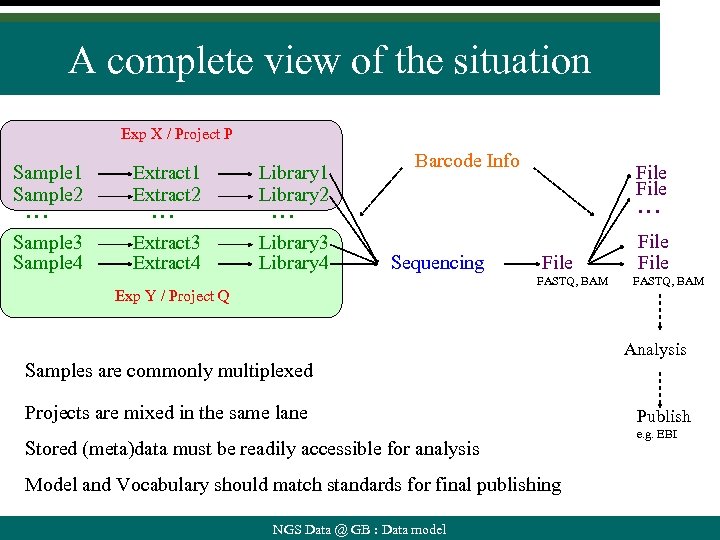
A complete view of the situation Exp X / Project P Sample 1 Sample 2 Extract 1 Extract 2 Library 1 Library 2 Sample 3 Sample 4 Extract 3 Extract 4 Library 3 Library 4 … … Barcode Info File … … Sequencing File FASTQ, BAM Exp Y / Project Q File FASTQ, BAM Analysis Samples are commonly multiplexed Projects are mixed in the same lane Stored (meta)data must be readily accessible for analysis Model and Vocabulary should match standards for final publishing NGS Data @ GB : Data model Publish e. g. EBI

11 1. em. BASE “Data management, organization, annotations and publication”
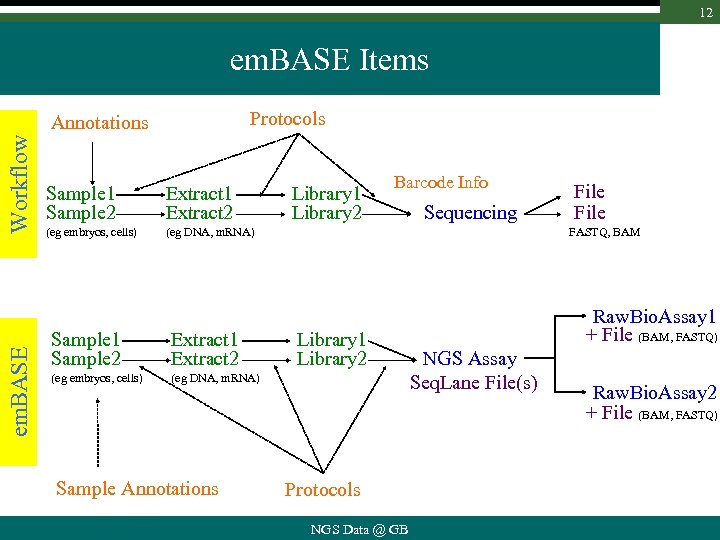
12 em. BASE Items Protocols em. BASE Workflow Annotations Sample 1 Sample 2 Extract 1 Extract 2 (eg embryos, cells) Barcode Info (eg DNA, m. RNA) Sample 1 Sample 2 Extract 1 Extract 2 (eg embryos, cells) Library 1 Library 2 FASTQ, BAM (eg DNA, m. RNA) Sample Annotations Sequencing File Library 1 Library 2 Protocols NGS Data @ GB Raw. Bio. Assay 1 + File (BAM, FASTQ) NGS Assay Seq. Lane File(s) Raw. Bio. Assay 2 + File (BAM, FASTQ)
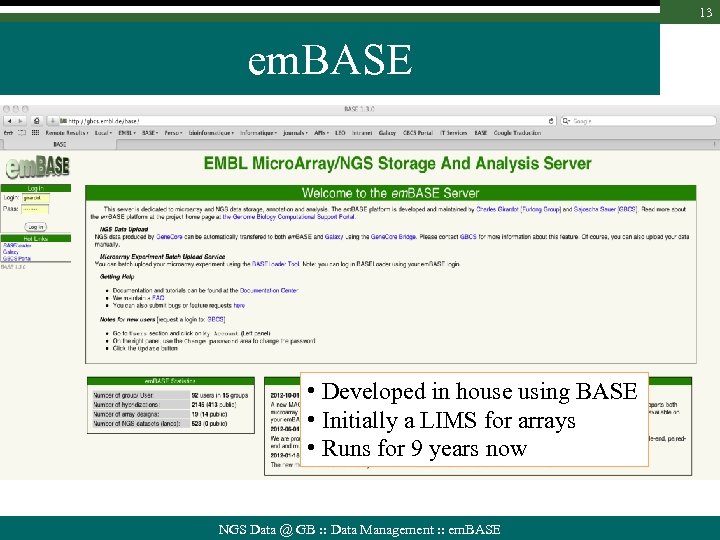
13 em. BASE • Developed in house using BASE • Initially a LIMS for arrays • Runs for 9 years now NGS Data @ GB : : Data Management : : em. BASE
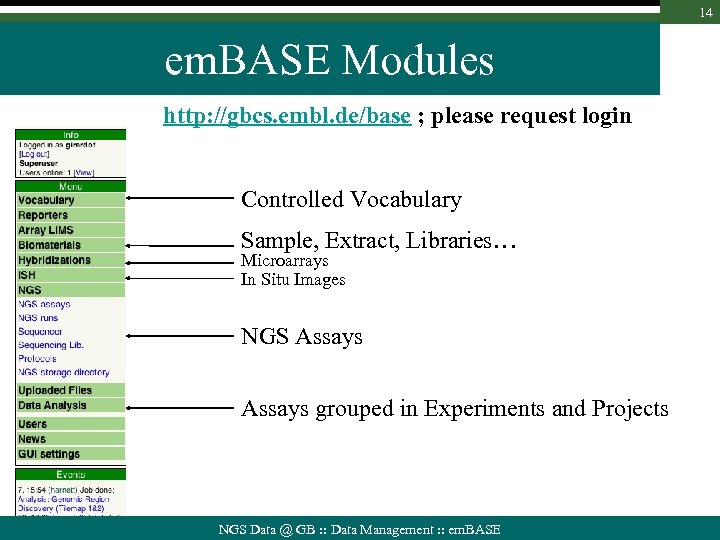

14 em. BASE Modules http: //gbcs. embl. de/base ; please request login Controlled Vocabulary Sample, Extract, Libraries… Microarrays In Situ Images NGS Assays grouped in Experiments and Projects NGS Data @ GB : : Data Management : : em. BASE
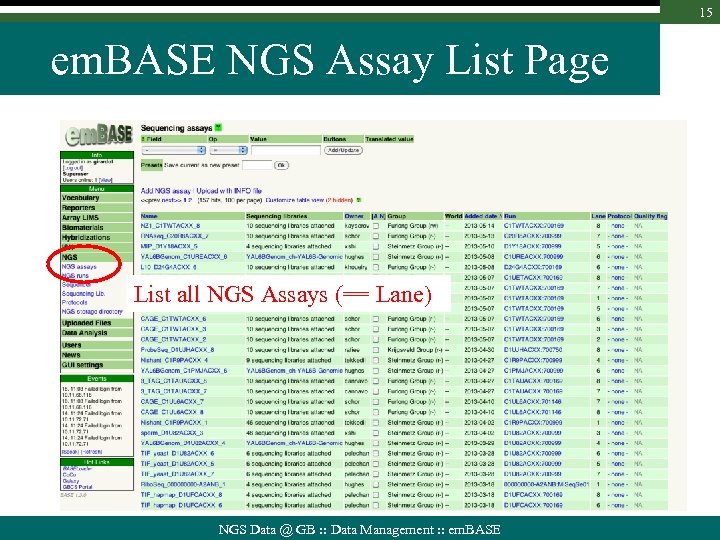
15 em. BASE NGS Assay List Page List all NGS Assays (== Lane) NGS Data @ GB : : Data Management : : em. BASE
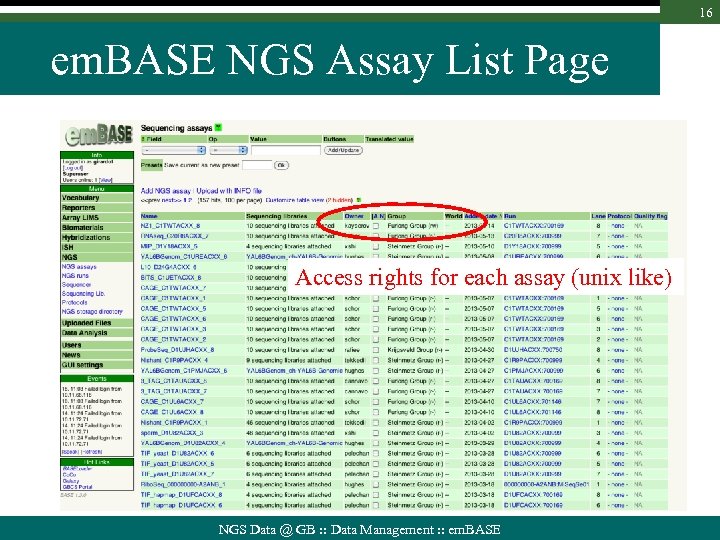
16 em. BASE NGS Assay List Page Access rights for each assay (unix like) NGS Data @ GB : : Data Management : : em. BASE
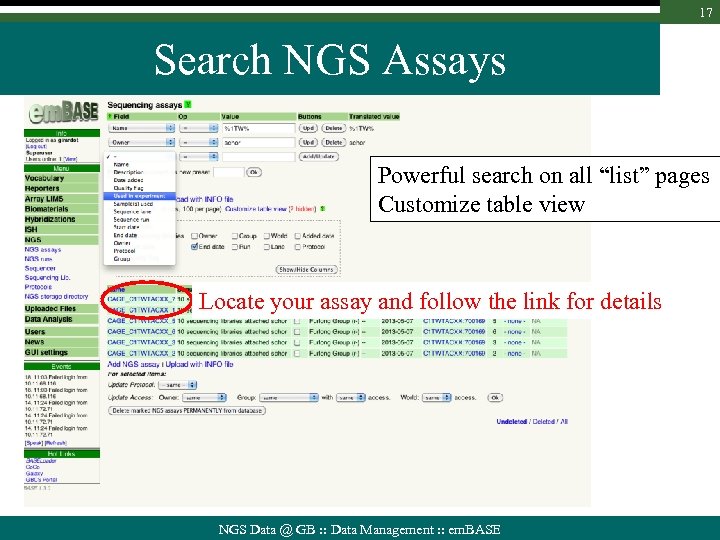
17 Search NGS Assays Powerful search on all “list” pages Customize table view Locate your assay and follow the link for details NGS Data @ GB : : Data Management : : em. BASE
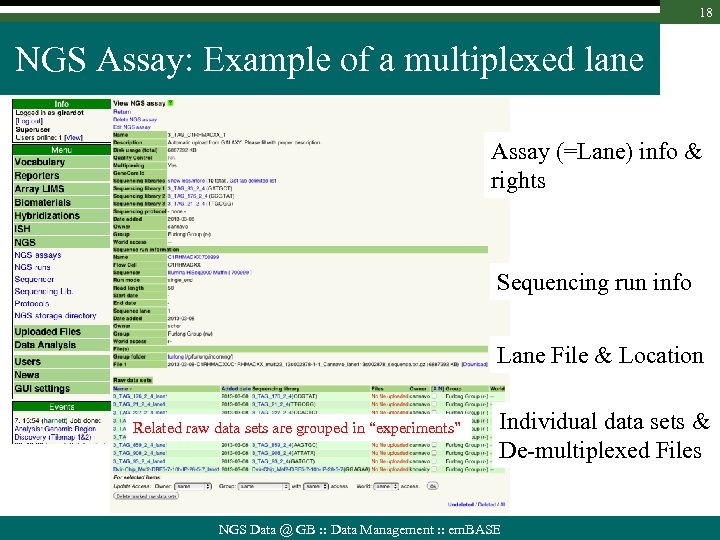

18 NGS Assay: Example of a multiplexed lane Assay (=Lane) info & rights Sequencing run info Lane File & Location Related raw data sets are grouped in “experiments” Individual data sets & De-multiplexed Files NGS Data @ GB : : Data Management : : em. BASE
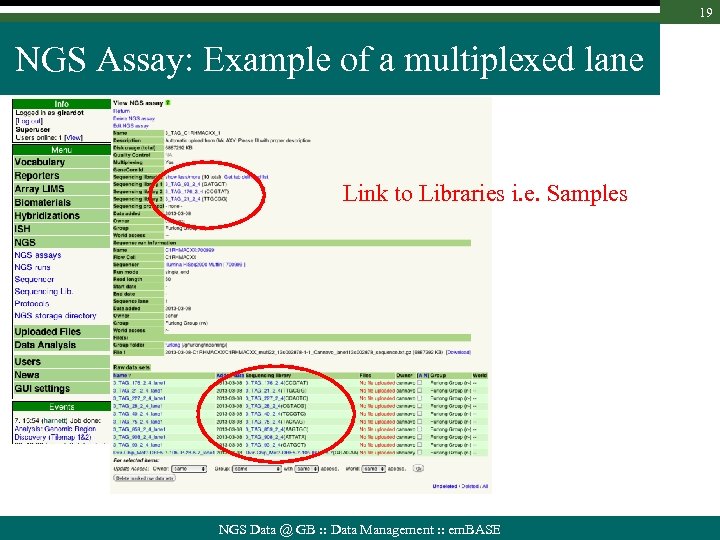
19 NGS Assay: Example of a multiplexed lane Link to Libraries i. e. Samples NGS Data @ GB : : Data Management : : em. BASE
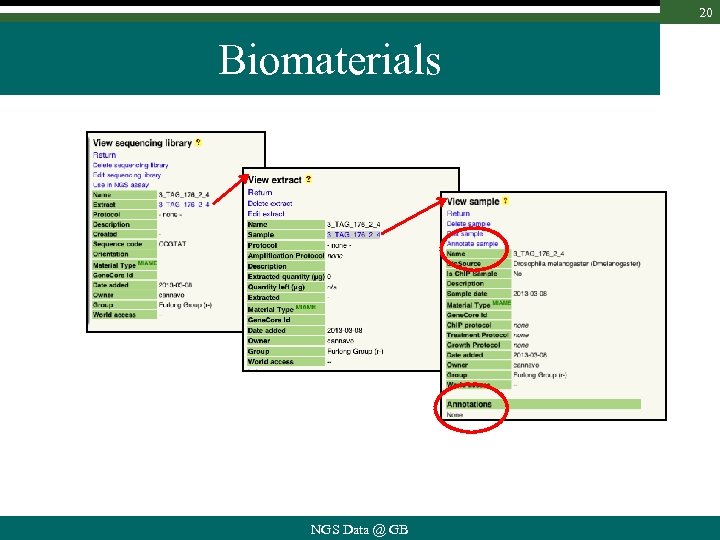
20 Biomaterials NGS Data @ GB
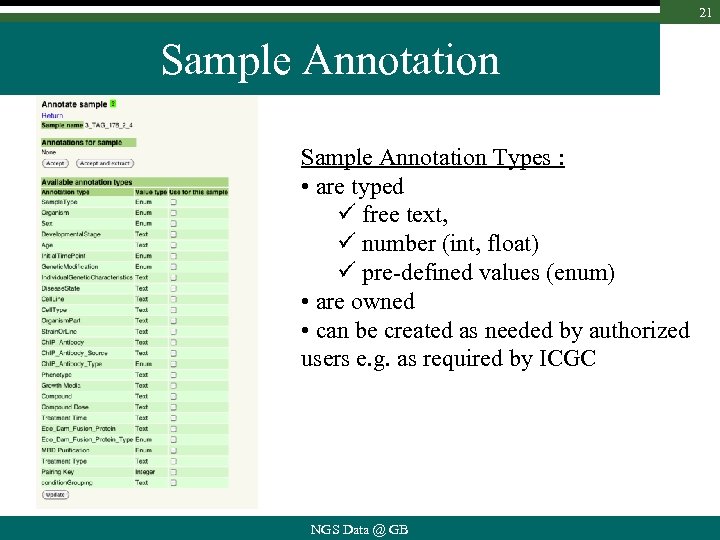

21 Sample Annotation Types : • are typed ü free text, ü number (int, float) ü pre-defined values (enum) • are owned • can be created as needed by authorized users e. g. as required by ICGC NGS Data @ GB
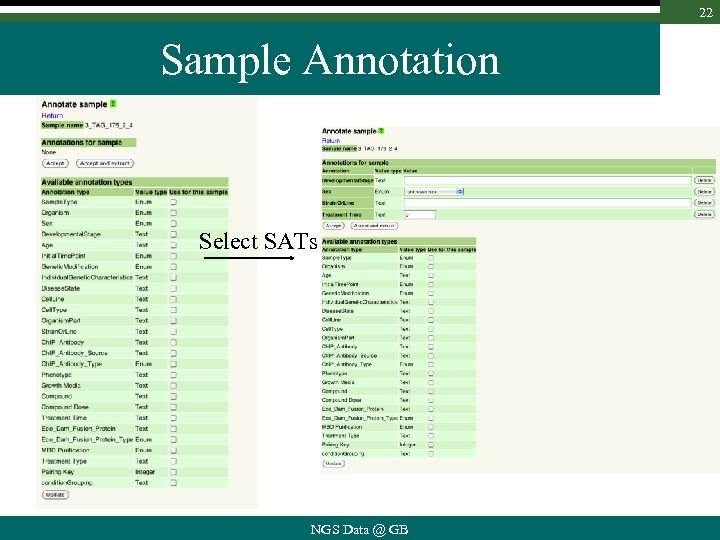
22 Sample Annotation Select SATs NGS Data @ GB
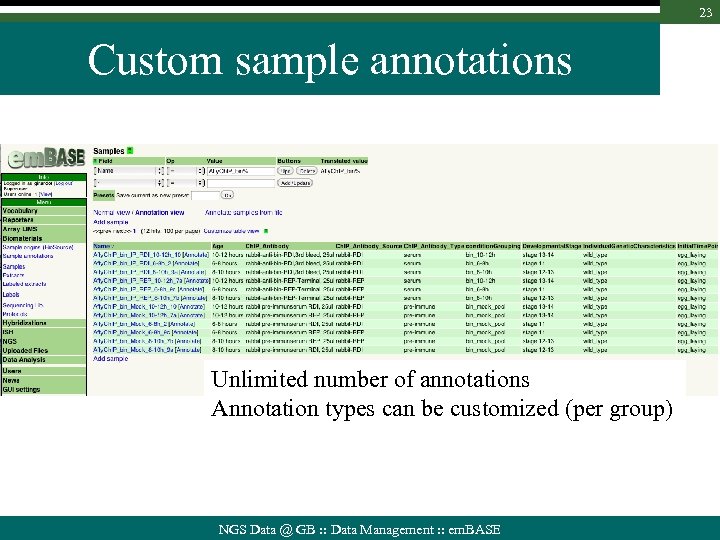

23 Custom sample annotations Unlimited number of annotations Annotation types can be customized (per group) NGS Data @ GB : : Data Management : : em. BASE
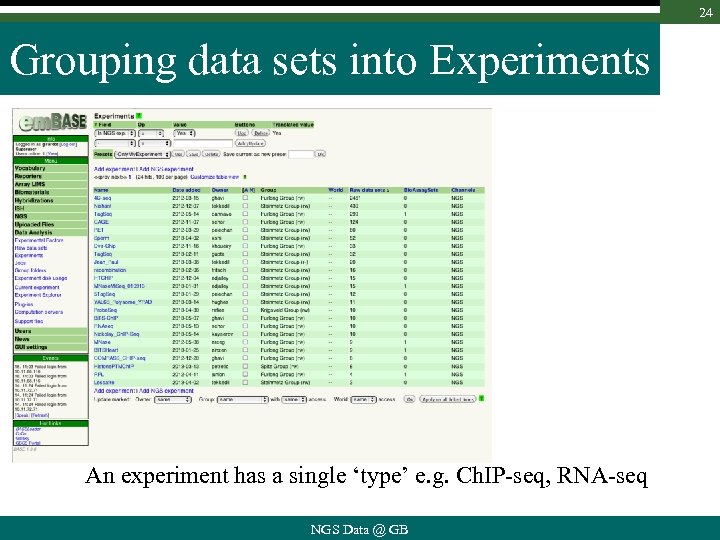
24 Grouping data sets into Experiments An experiment has a single ‘type’ e. g. Ch. IP-seq, RNA-seq NGS Data @ GB
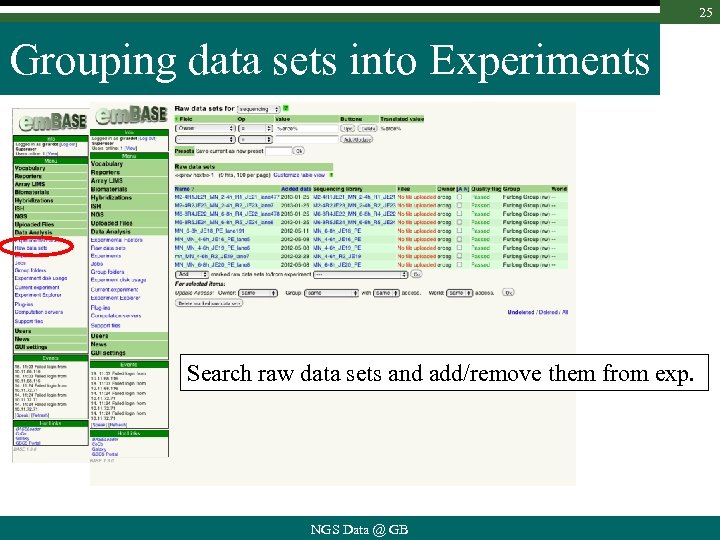
25 Grouping data sets into Experiments Search raw data sets and add/remove them from exp. NGS Data @ GB
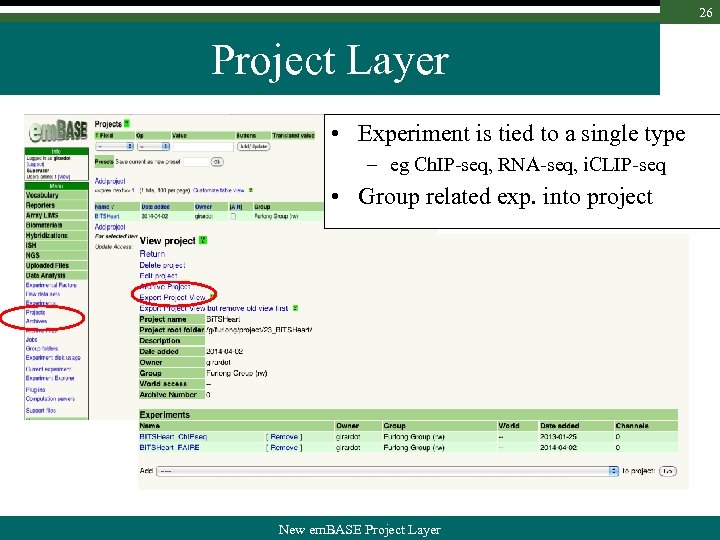
26 Project Layer • Experiment is tied to a single type – eg Ch. IP-seq, RNA-seq, i. CLIP-seq • Group related exp. into project New em. BASE Project Layer

27 Wait a sec. . . Do we really have to fill all these web forms ? !? ! NO ! 1. GCBridge: all “items” are pre-created for you 2. Protocols and sample annotations remain to be done NGS Data @ GB

28 2. Decentralized NGS File Data Lib “Your data lives on your file server and is readily accessible”
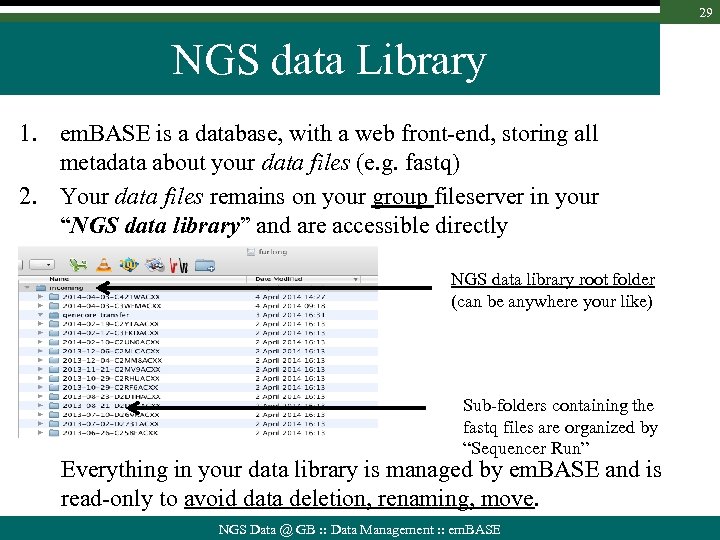
29 NGS data Library 1. em. BASE is a database, with a web front-end, storing all metadata about your data files (e. g. fastq) 2. Your data files remains on your group fileserver in your “NGS data library” and are accessible directly NGS data library root folder (can be anywhere your like) Sub-folders containing the fastq files are organized by “Sequencer Run” Everything in your data library is managed by em. BASE and is read-only to avoid data deletion, renaming, move. NGS Data @ GB : : Data Management : : em. BASE
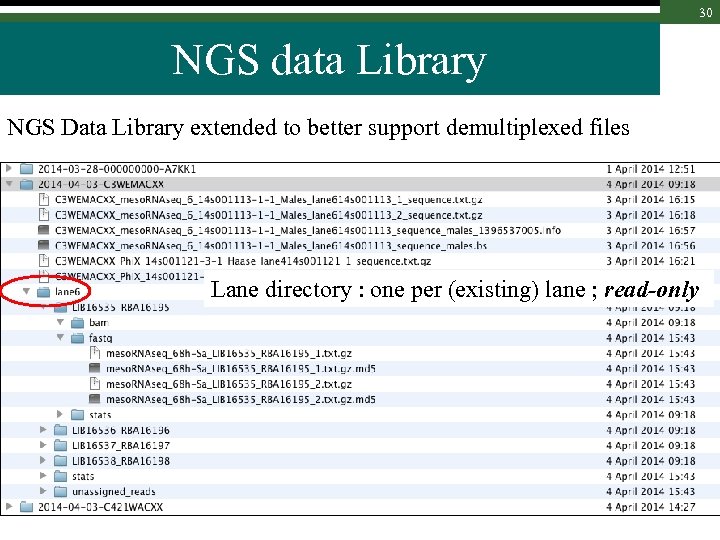
30 NGS data Library NGS Data Library extended to better support demultiplexed files Lane directory : one per (existing) lane ; read-only
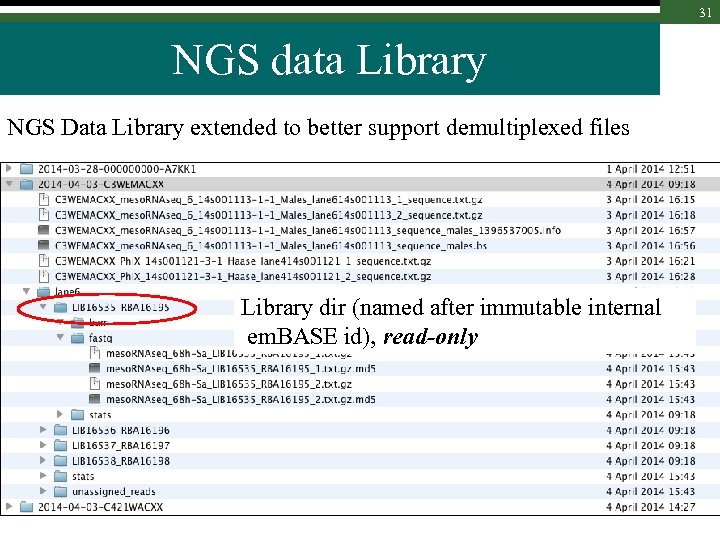

31 NGS data Library NGS Data Library extended to better support demultiplexed files Library dir (named after immutable internal em. BASE id), read-only
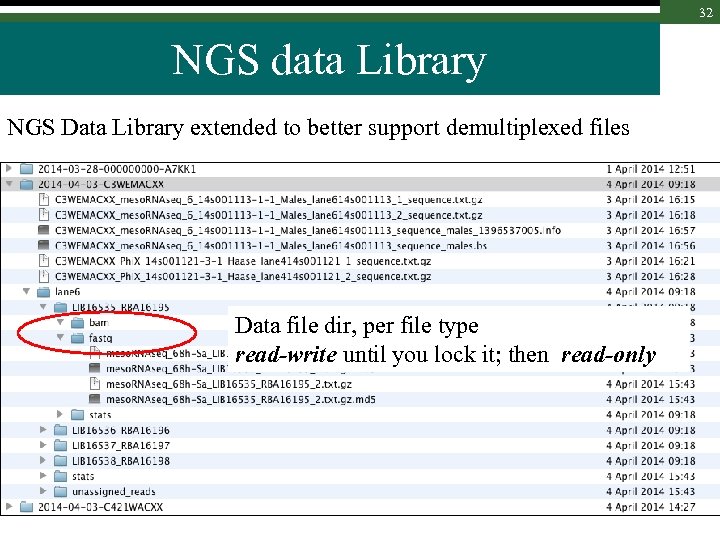
32 NGS data Library NGS Data Library extended to better support demultiplexed files Data file dir, per file type read-write until you lock it; then read-only

33 Locking / Unlocking concept 1. Library file sub-directories are unlocked (writable for group) – you can work and replace files as you wish 2. At some point, files are ready and directories can be locked (only readable): 1. em. BASE starts, at this point, to track these files 2. em. BASE will allow lane file deletion when all its multiplexed libraries are locked. 3. Locking is operated via the web interface, on the whole lane or per library (case of shared lanes)

34 3. GC Bridge “Ensuring smooth data transfer between Gene. Core to em. BASE”
35 GCBridge : Making your life as easy as possible Gene. Core 1. Transfer file NGS Lib Online Ordering NGS Data @ GB : : Automated Data Transfer
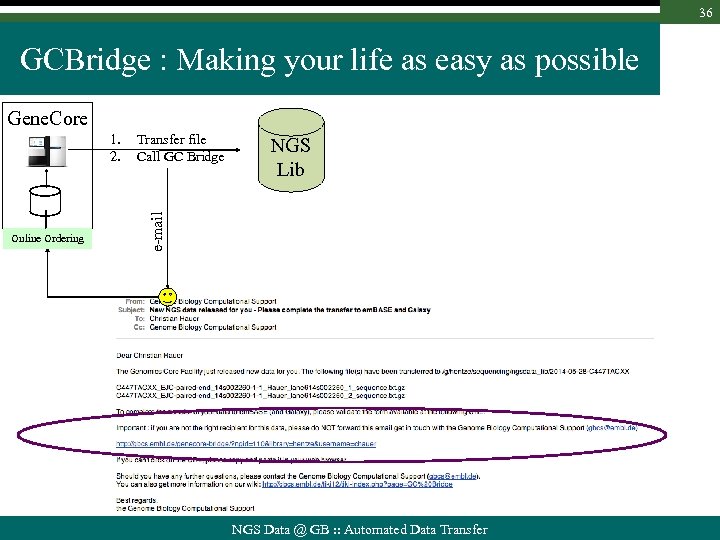

36 GCBridge : Making your life as easy as possible Gene. Core Online Ordering Transfer file Call GC Bridge NGS Lib e-mail 1. 2. NGS Data @ GB : : Automated Data Transfer

37 GCBridge : Making your life as easy as possible Gene. Core Online Ordering Transfer file Call GC Bridge NGS Lib e-mail 1. 2. NGS Data @ GB : : Automated Data Transfer
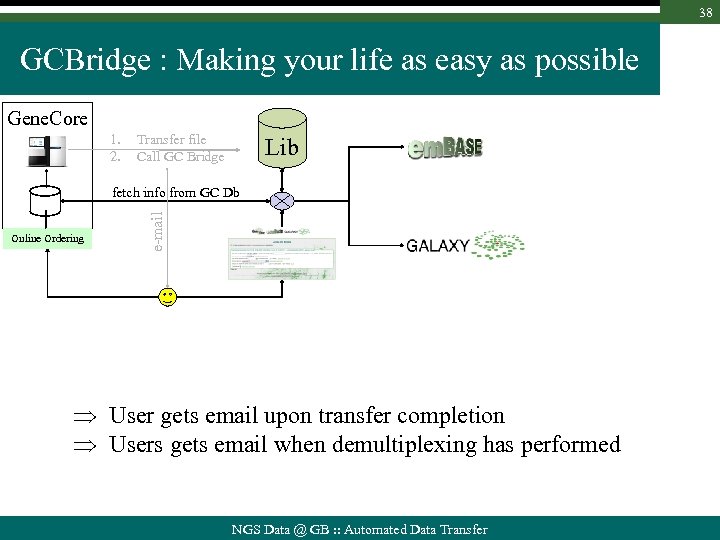
38 GCBridge : Making your life as easy as possible Gene. Core 1. 2. Transfer file Call GC Bridge Lib Online Ordering e-mail fetch info from GC Db User gets email upon transfer completion Users gets email when demultiplexing has performed NGS Data @ GB : : Automated Data Transfer

39 3. Practical steps Validate GCBridge Transfer Form Annotate Samples, link protocols
40 Data released email Click the link to get to the GCBridge Transfer Form
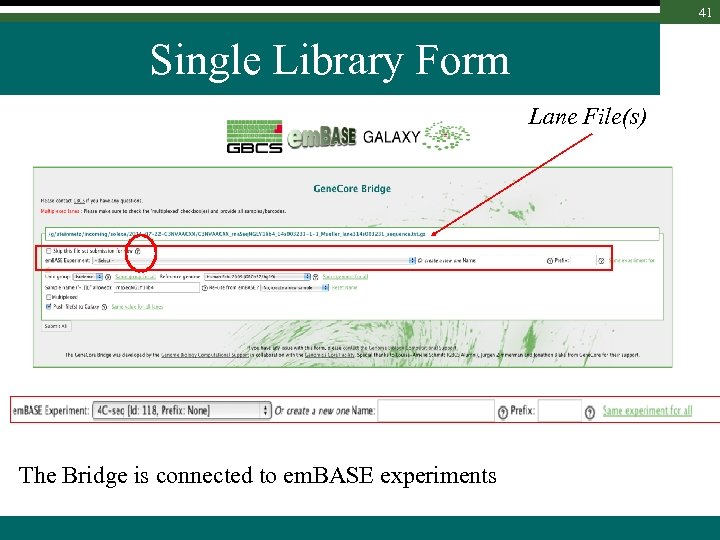

41 Single Library Form Lane File(s) The Bridge is connected to em. BASE experiments
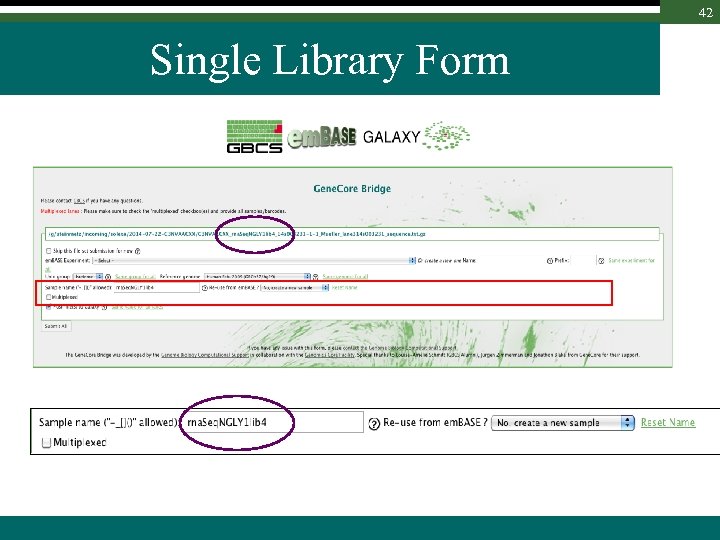
42 Single Library Form
43 Single Library Form
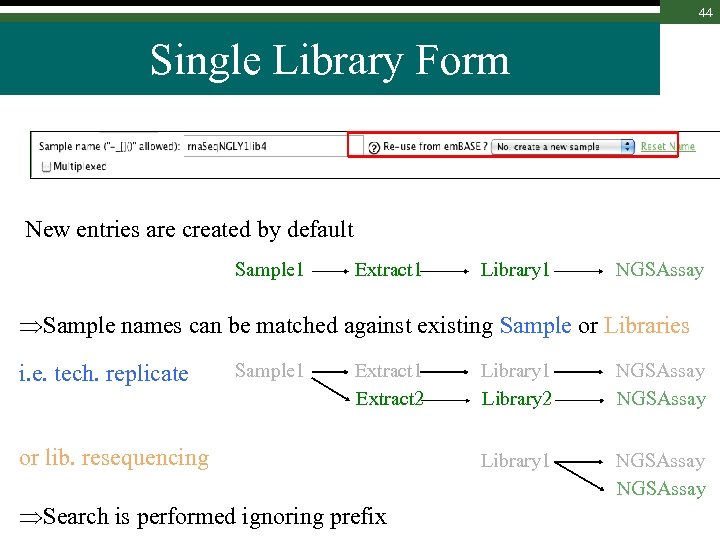

44 Single Library Form New entries are created by default Sample 1 Extract 1 Library 1 NGSAssay Sample names can be matched against existing Sample or Libraries i. e. tech. replicate Sample 1 Extract 2 or lib. resequencing Search is performed ignoring prefix Library 1 Library 2 NGSAssay Library 1 NGSAssay
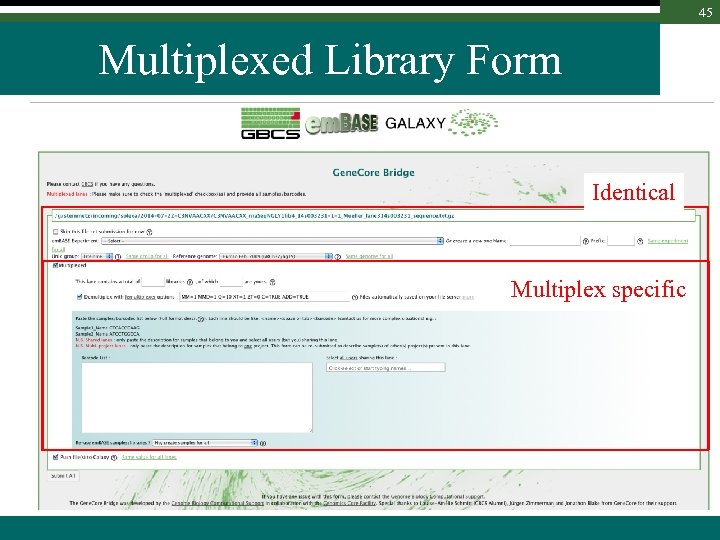
45 Multiplexed Library Form Identical Multiplex specific
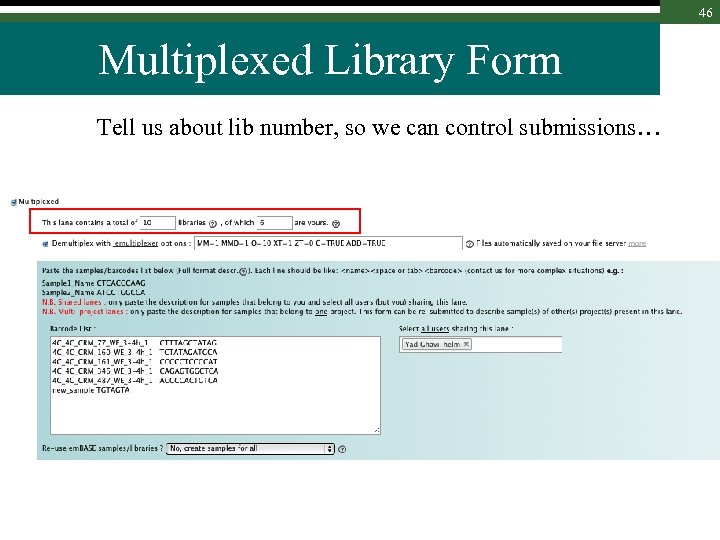
46 Multiplexed Library Form Tell us about lib number, so we can control submissions…
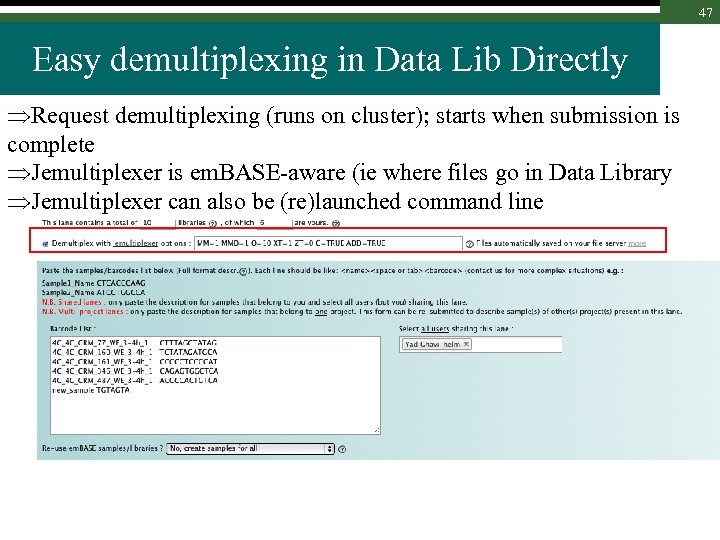

47 Easy demultiplexing in Data Lib Directly Request demultiplexing (runs on cluster); starts when submission is complete Jemultiplexer is em. BASE-aware (ie where files go in Data Library Jemultiplexer can also be (re)launched command line
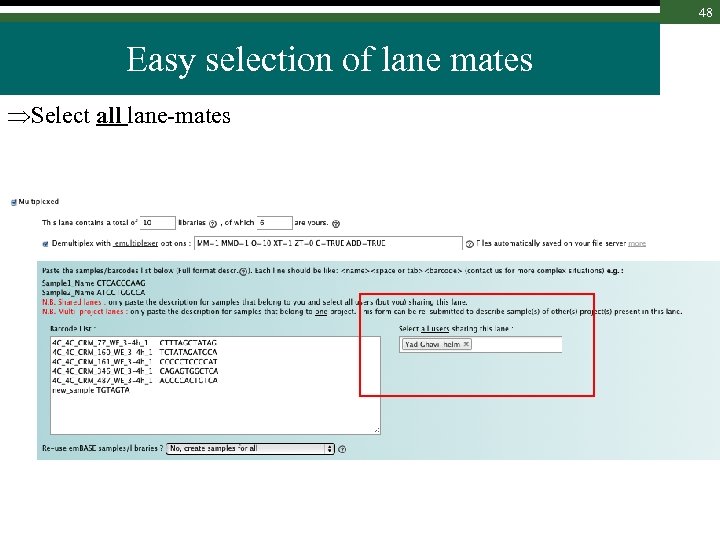
48 Easy selection of lane mates Select all lane-mates
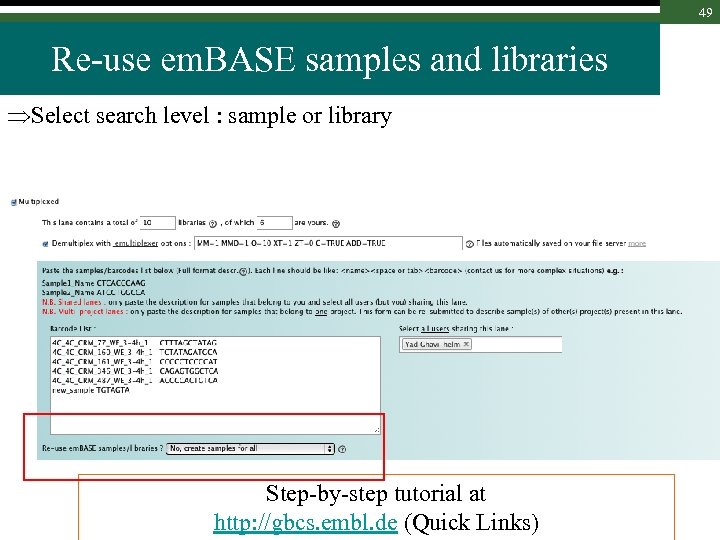
49 Re-use em. BASE samples and libraries Select search level : sample or library Step-by-step tutorial at http: //gbcs. embl. de (Quick Links)
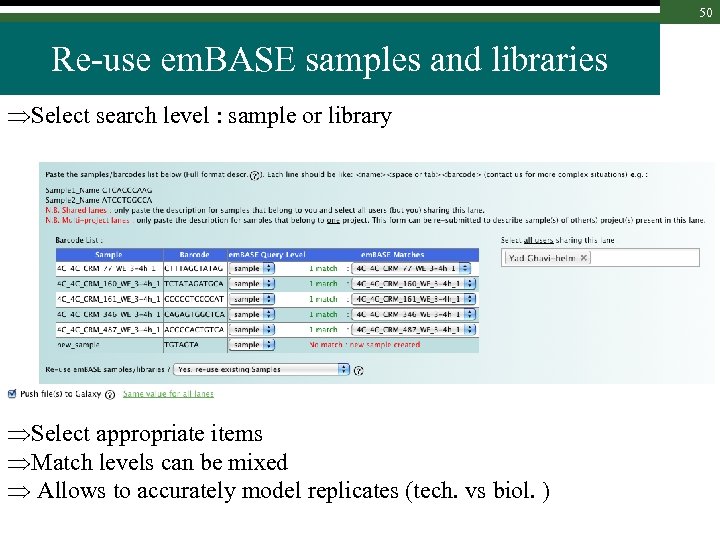

50 Re-use em. BASE samples and libraries Select search level : sample or library Select appropriate items Match levels can be mixed Allows to accurately model replicates (tech. vs biol. )
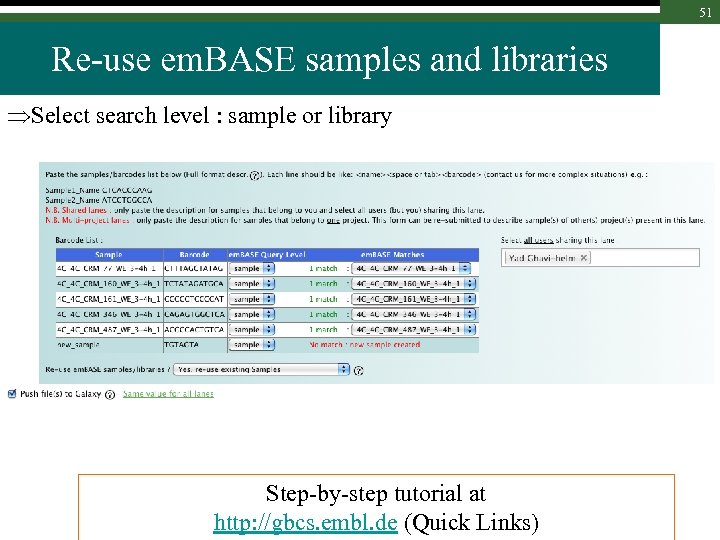
51 Re-use em. BASE samples and libraries Select search level : sample or library Step-by-step tutorial at http: //gbcs. embl. de (Quick Links)
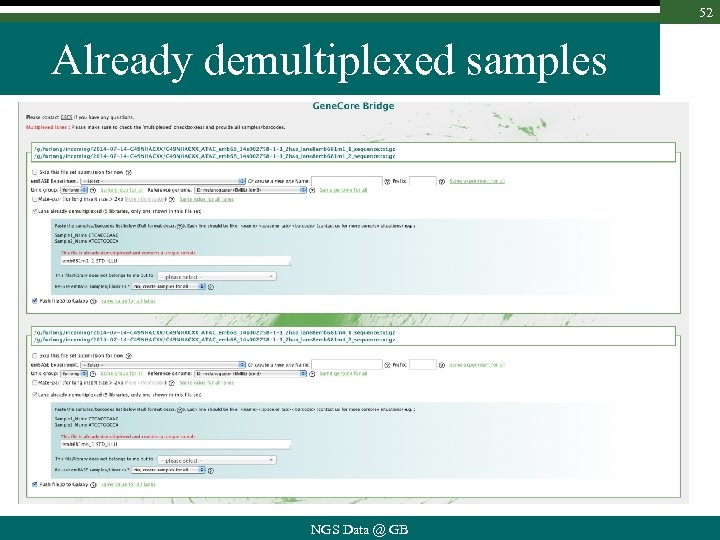
52 Already demultiplexed samples NGS Data @ GB
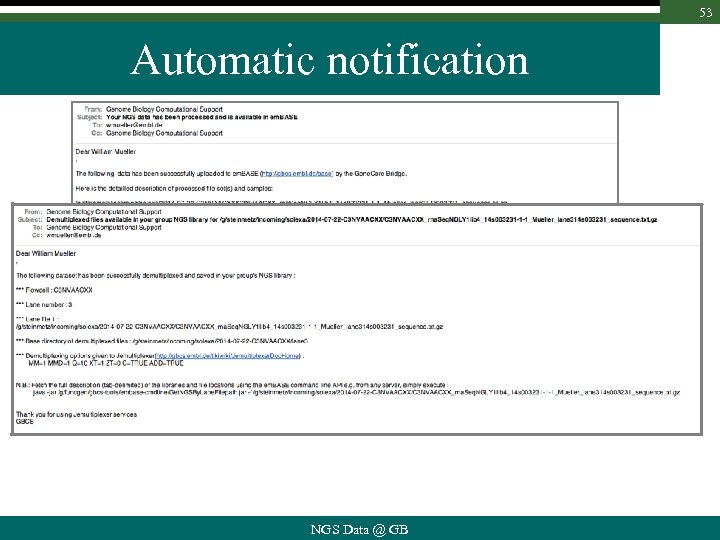
53 Automatic notification NGS Data @ GB

54 Now what ? 1. GCBridge: all “items” are pre-created for you 2. Protocols and sample annotations remain to be done in em. BASE NGS Data @ GB
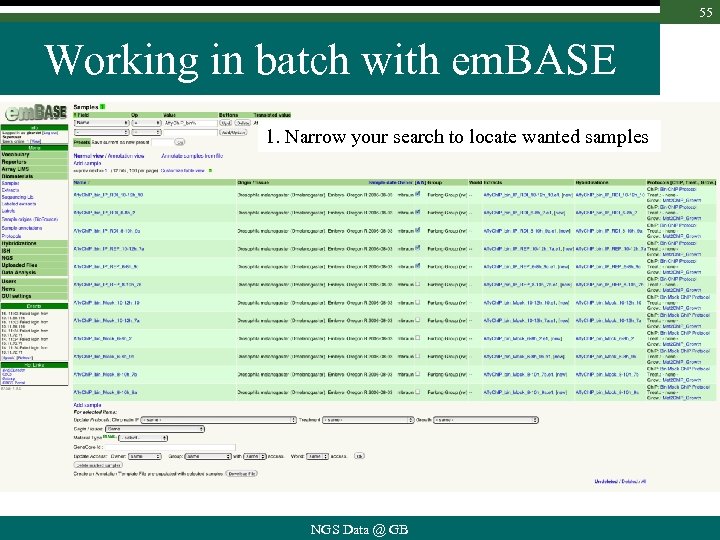
55 Working in batch with em. BASE 1. Narrow your search to locate wanted samples NGS Data @ GB
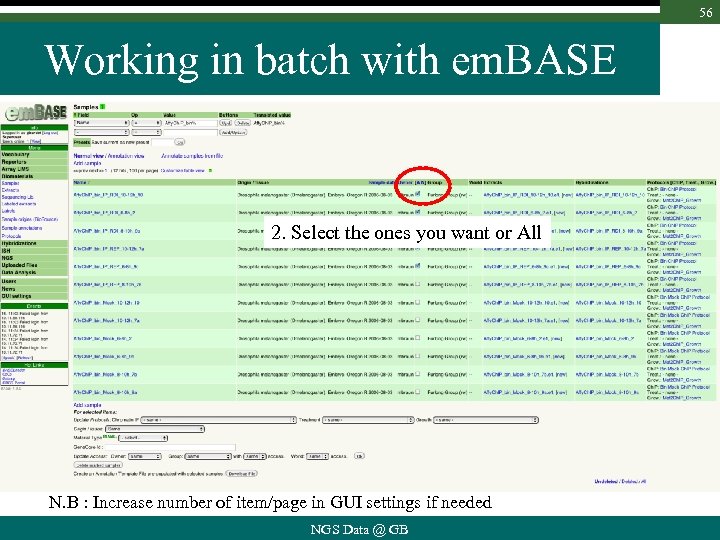
56 Working in batch with em. BASE 2. Select the ones you want or All N. B : Increase number of item/page in GUI settings if needed NGS Data @ GB
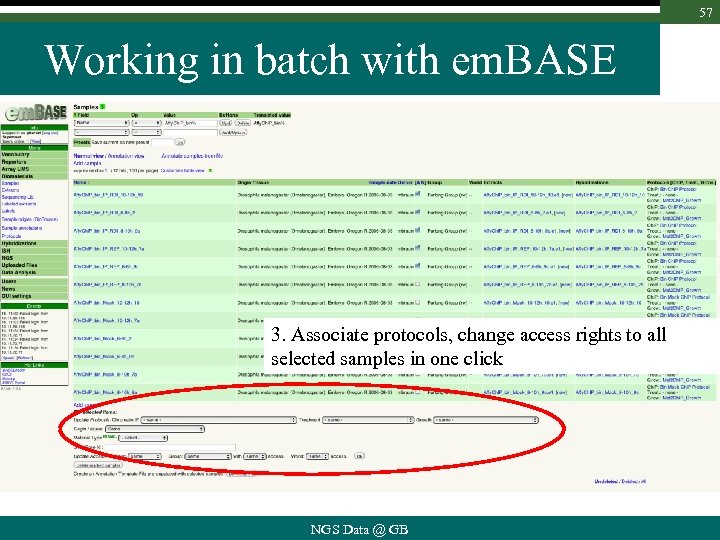
57 Working in batch with em. BASE 3. Associate protocols, change access rights to all selected samples in one click NGS Data @ GB
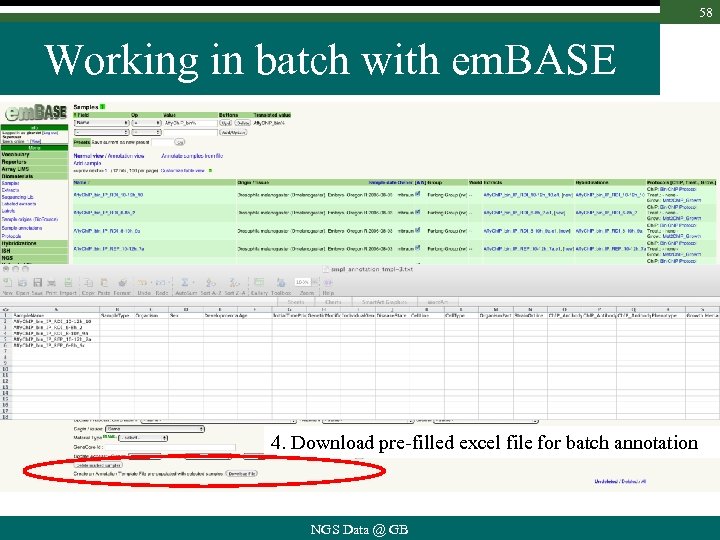
58 Working in batch with em. BASE 4. Download pre-filled excel file for batch annotation NGS Data @ GB
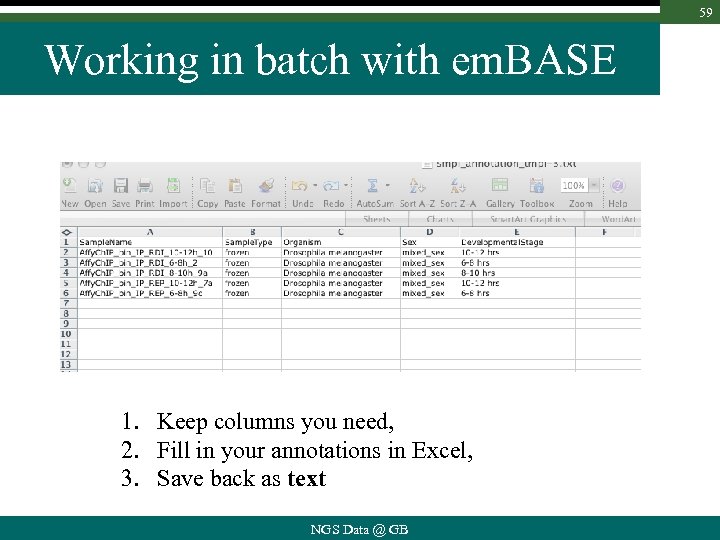

59 Working in batch with em. BASE 1. Keep columns you need, 2. Fill in your annotations in Excel, 3. Save back as text NGS Data @ GB
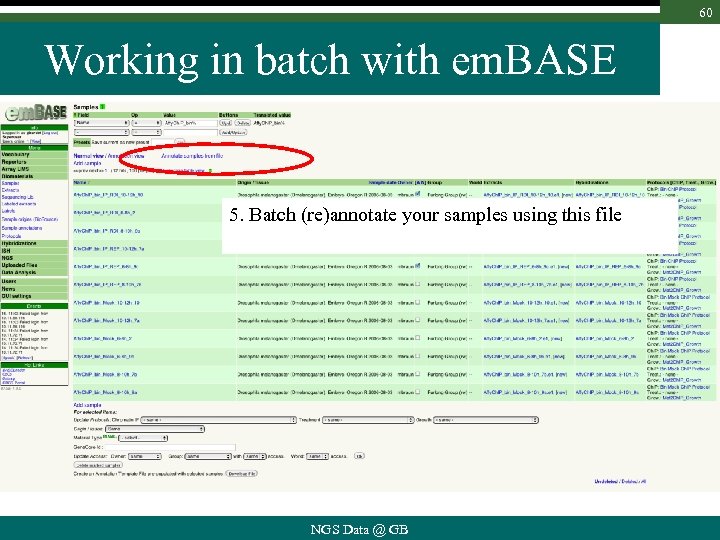
60 Working in batch with em. BASE 5. Batch (re)annotate your samples using this file NGS Data @ GB

61 em. BASE Advanced Features (for the command line user)

62 Working with em. BASE 1. Export experiment or project views using the web interface 2. Use the new command line em. BASE API to learn where files are or should be placed – These commands extracts all info from em. BASE for a lane, an experiment or a project Documentation at : http: //gbcs. embl. de/
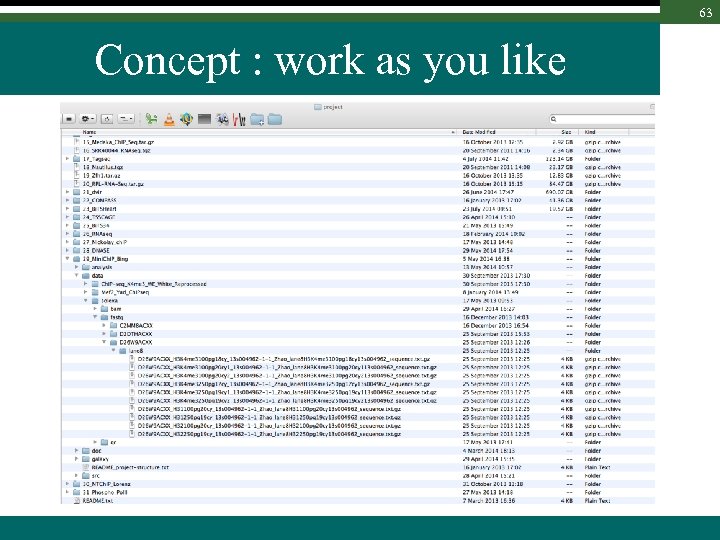
63 Concept : work as you like
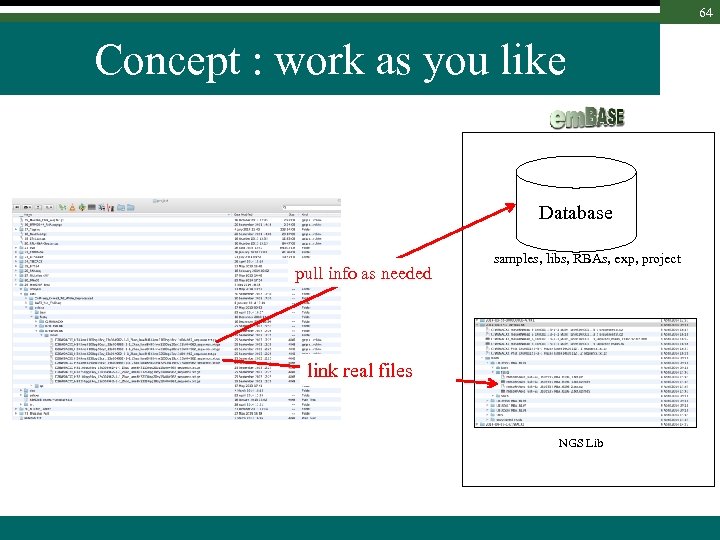
64 Concept : work as you like Database pull info as needed samples, libs, RBAs, exp, project link real files NGS Lib
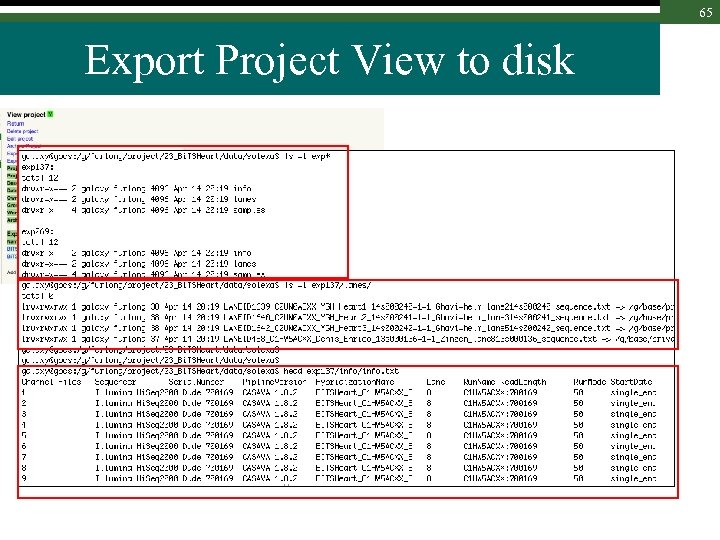
65 Export Project View to disk
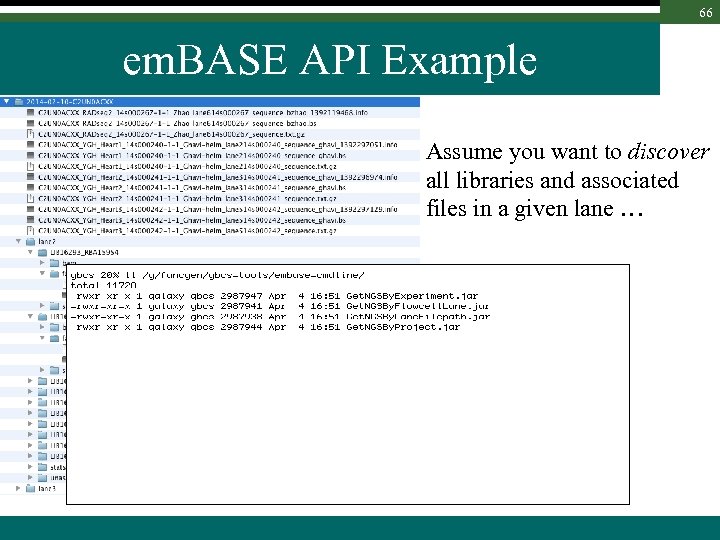
66 em. BASE API Example Assume you want to discover all libraries and associated files in a given lane …

67 em. BASE API Example • Available from anywhere • Logged in user used to authenticate in em. BASE • Rights apply the same way as in em. BASE
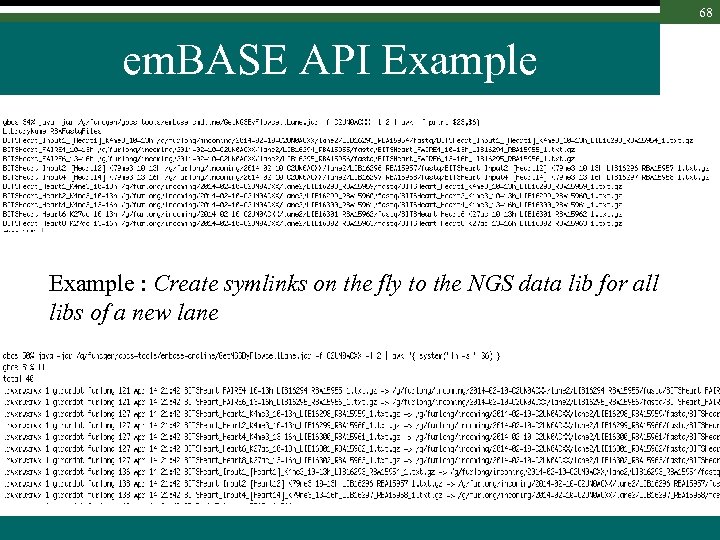

68 em. BASE API Example : Create symlinks on the fly to the NGS data lib for all libs of a new lane
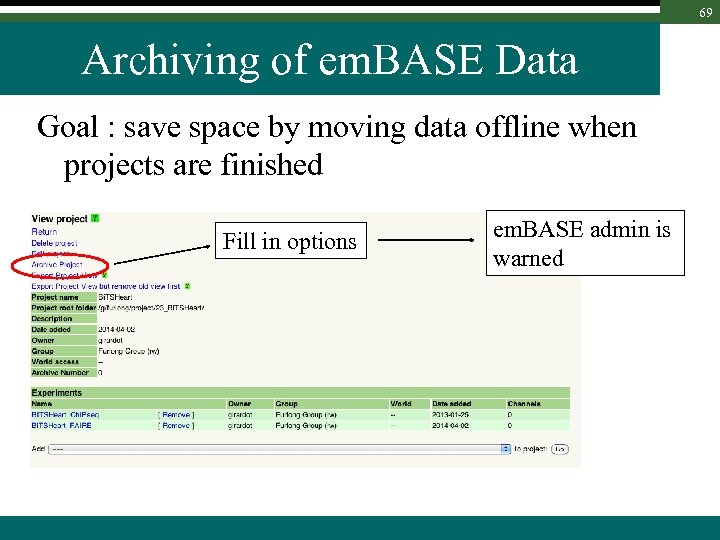

69 Archiving of em. BASE Data Goal : save space by moving data offline when projects are finished Fill in options em. BASE admin is warned

70 Archiving of em. BASE Data What happens next ? • All data files connected to the experiments are exported • IT performs back up on tape • We delete ‘deletable’ files (concept of active experiment): – em. BASE knows which files can be deleted, which ones have been deleted and how to get them back, if needed – delete files are locally replaced with the a small file containing back up information • You can follow the archiving status in em. BASE This is a couple of clicks on your side but remember that you still pay the bill !

71 Galaxy (First Steps) “Powerful data analysis made easy and reproducible ”
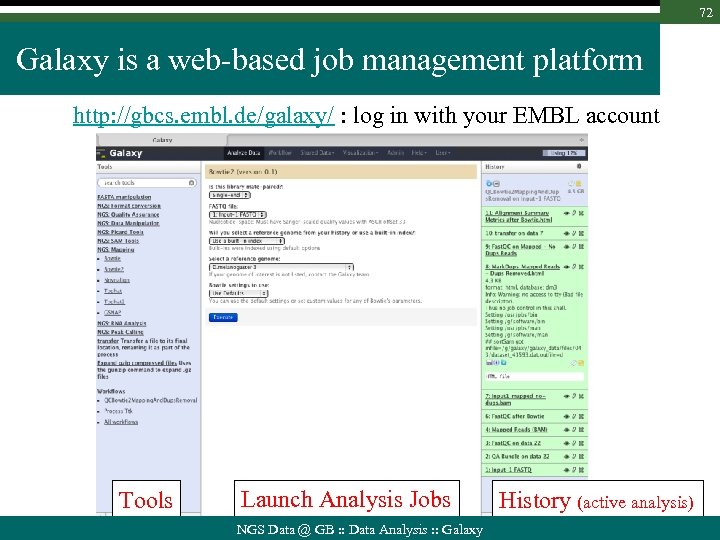
72 Galaxy is a web-based job management platform http: //gbcs. embl. de/galaxy/ : log in with your EMBL account Tools Launch Analysis Jobs NGS Data @ GB : : Data Analysis : : Galaxy History (active analysis)
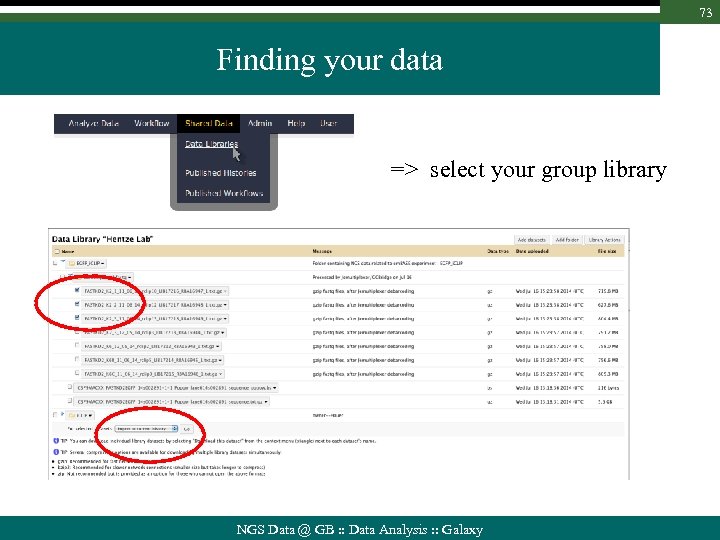
73 Finding your data => select your group library NGS Data @ GB : : Data Analysis : : Galaxy
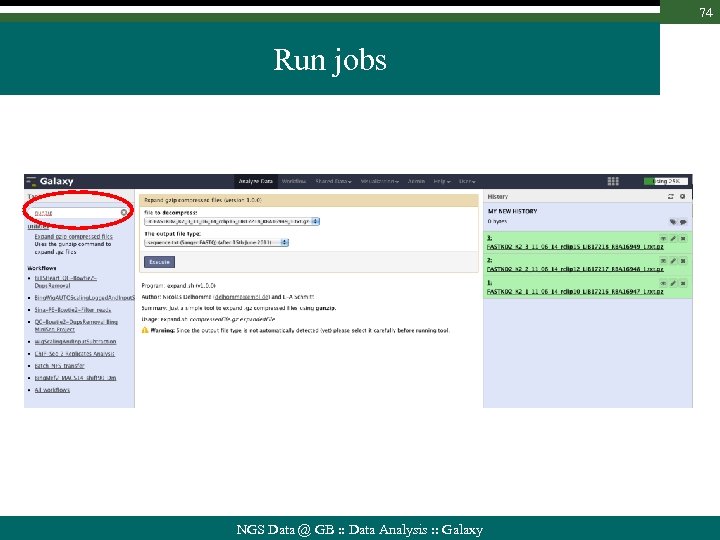
74 Run jobs NGS Data @ GB : : Data Analysis : : Galaxy
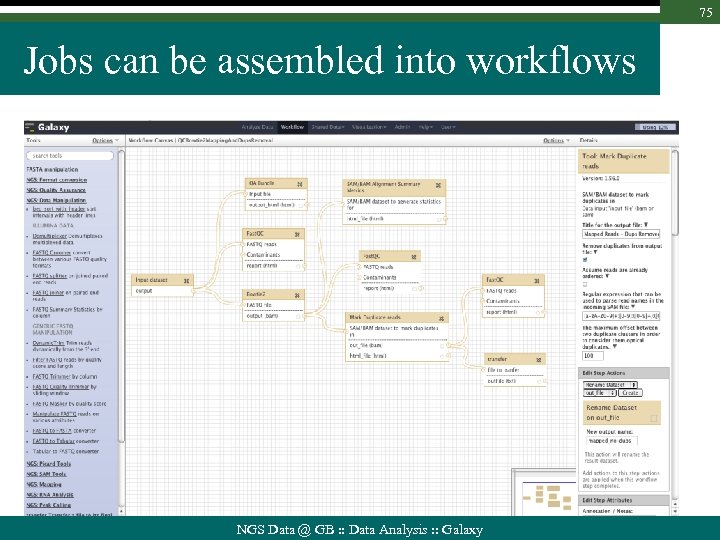
75 Jobs can be assembled into workflows NGS Data @ GB : : Data Analysis : : Galaxy
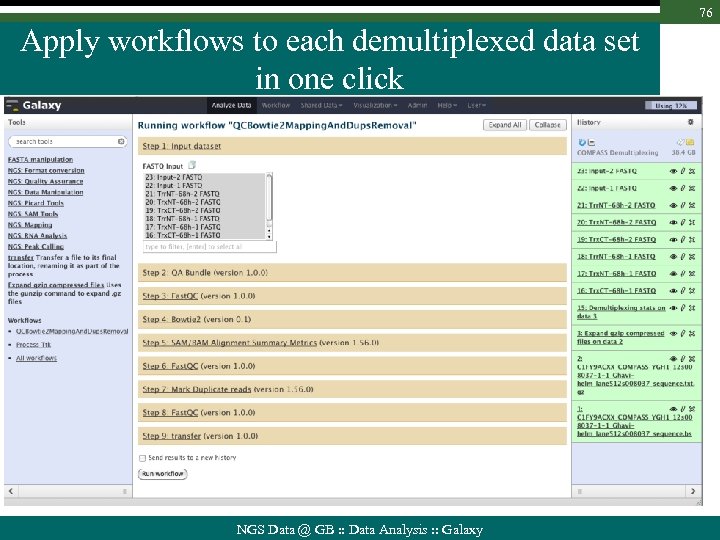
76 Apply workflows to each demultiplexed data set in one click NGS Data @ GB : : Data Analysis : : Galaxy
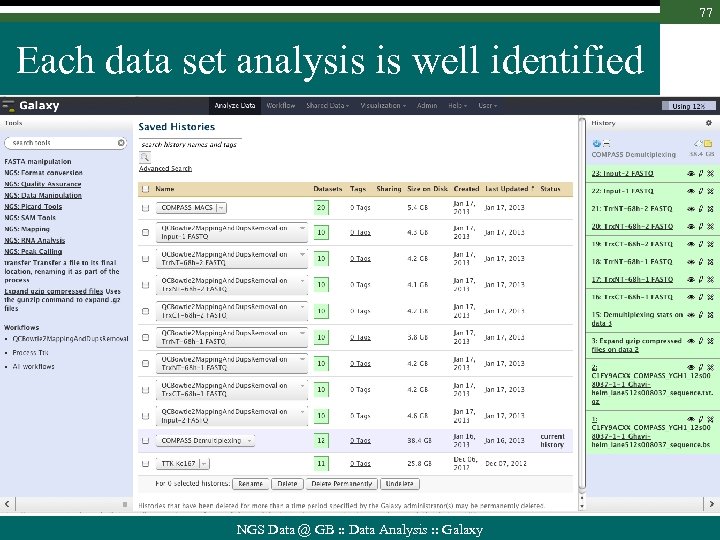
77 Each data set analysis is well identified NGS Data @ GB : : Data Analysis : : Galaxy

78 Galaxy Summary 1. Galaxy is a job management / analysis platform • Run standard analysis (trimming, QC, mapping, peak calling, …) • Assemble workflows and perform parallel processing 2. Jobs are sent to the new LSF EMBL cluster • We implement cluster good practices (copy to local /tmp, …) • Tools are available under BCR/SEPP 3. Continuous update/addition of tools & indices 4. Open source and very active project NGS Data @ GB : : Data Analysis

79 Galaxy Summary 1. Galaxy is a job management / analysis platform • Run standard analysis (trimming, QC, mapping, peak calling, …) • Assemble workflows and perform parallel processing 2. Jobs are sent to the new LSF EMBL cluster • We implement cluster good practices (copy to local /tmp, …) • Tools are available under BCR/SEPP 3. Continuous update/addition of tools & indices 4. Open source and very active project NGS Data @ GB : : Data Analysis

80 Galaxy Summary 1. Galaxy is a job management / analysis platform • Run standard analysis (trimming, QC, mapping, peak calling, …) • Assemble workflows and perform parallel processing 2. Jobs are sent to the new LSF EMBL cluster • We implement cluster good practices (copy to local /tmp, …) • Tools are available under BCR/SEPP 3. Continuous update/addition of tools & indices 4. Galaxy uses the data from your NGS Data library directly 5. Easy transfer of results from Galaxy to your own disks NGS Data @ GB : : Data Analysis

81 Conclusion There absolutely no drawbacks in using our system, only benefits ! NGS Data @ GB : : Data Analysis

82 Thank you Joscha Sauer Shu-yi Su Laura O’Donovan Matthias Monfort Alumni Aziz Moussa M. Chaturvedi L-A Schmitt Nicolas Delhomme Leila Tlili Arnaud Huaulme IT Services Michael Wahlers Andres Lindau All GB members Chenchen Zhu Simon Anders Tobias Rausch Frank Thommen (CBB) Eileen Furlong Gene. Core Jonathon Blake Juergen Zimmermann Markus Fritz Vladimir Benes
c870716b85b31635d48fa916ddb9c49b.ppt