99b09b1a668b16b3bbd5db7c01dd0e44.ppt
- Количество слайдов: 44

เทคโนโลยและการตรวจทางหอง ปฏบตการ โรคตดตออบตใหม การประชมเชงปฏบตการ เรองโรคตดตออบตใหม : ความทาทายสำหรบบคลากรทางการแพทยและสาธา รณสข วนท 10 -11 มกราคม 2556 โรงแรม อมาร แอรพอรต กรงเทพมหานคร มาลน จตตกานตพชย สถาบนวจยวทยาศาสตรสา ธารณสข กรมวทยาศาสตรการแพทย
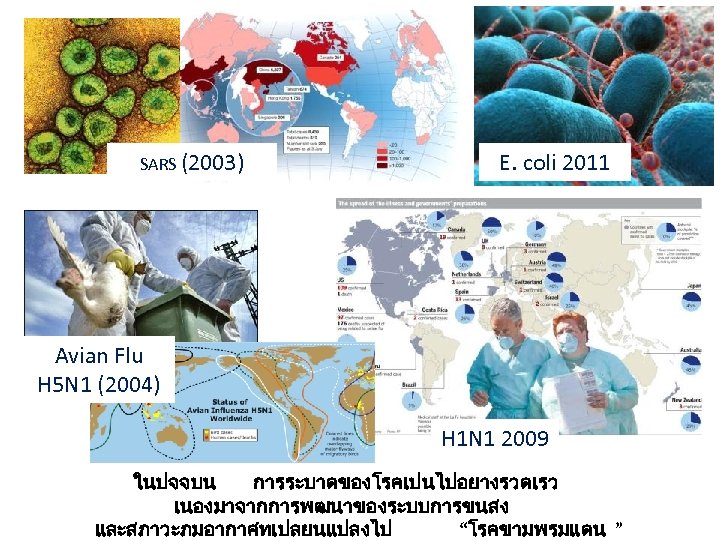
SARS (2003) E. coli 2011 Avian Flu H 5 N 1 (2004) H 1 N 1 2009 ในปจจบน การระบาดของโรคเปนไปอยางรวดเรว เนองมาจากการพฒนาของระบบการขนสง และสภาวะภมอากาศทเปลยนแปลงไป “โรคขามพรมแดน ”
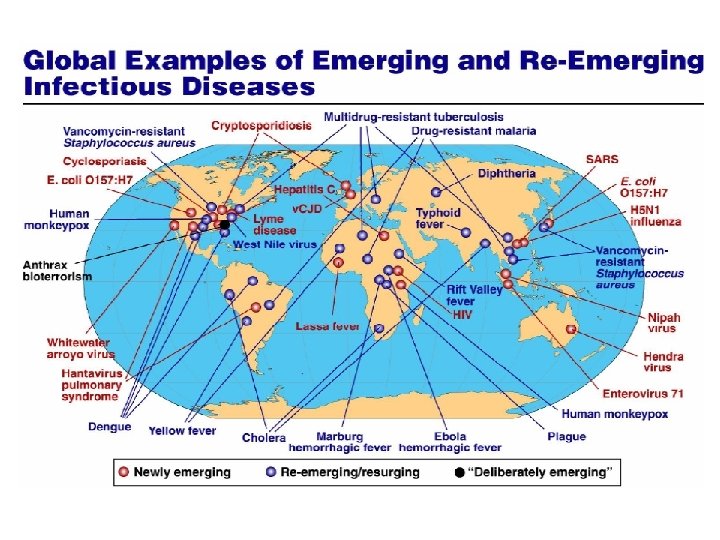
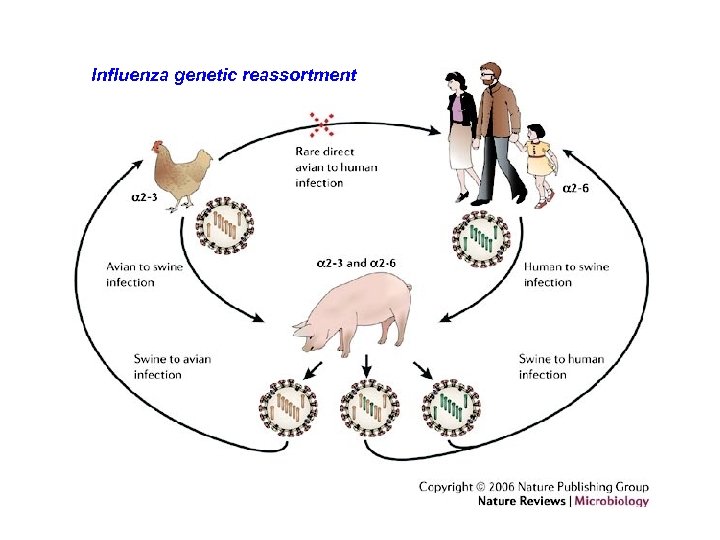
Influenza genetic reassortment
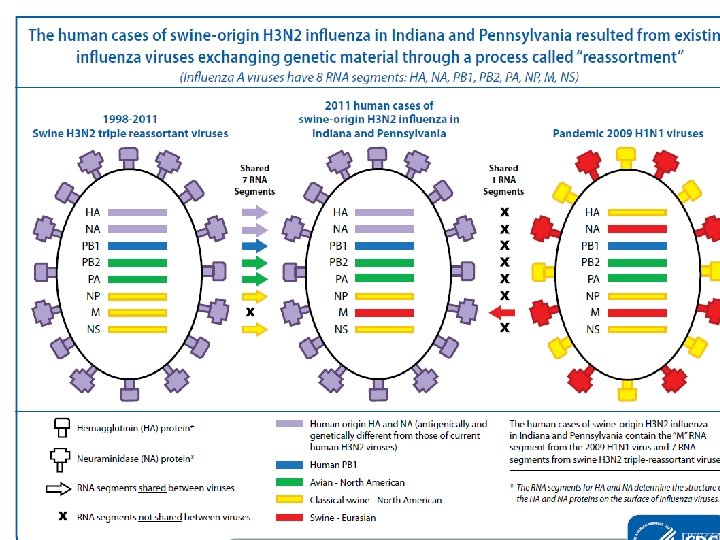

Diagnostic test need to………. . ØIdentify new variants of pathogens ØRapid detection & Highly throughput Ø Highly sensitivity & specificity

OIE ad hoc Group on Diagnostic Tests in Relation to New and Emerging Technologies The following new molecular diagnostic methodologies were identified: • Direct diagnostic assays • PCR-based assays • • o Real time; o Rapid detection in a disease outbreak; o Multiplex; o PCR robotics. • Isothermal amplification assays; • Microarray technologies; • Rapid sequencing technologies, phylogenic analysis/bioinformatics; • Genomic technologies to determine virulence; • Complete full length genome sequencing technologies; • Pen-side test technologies (lateral flow devices); • Portable PCR technologies for field use; • Nanotechnology; • Proximity ligation technologies; • In-situ hybridisation; • Proteomics (detection of proteins). Source: http: //www. oie. int/downld/SC/2008/A_BSC_sept 2008. pdf
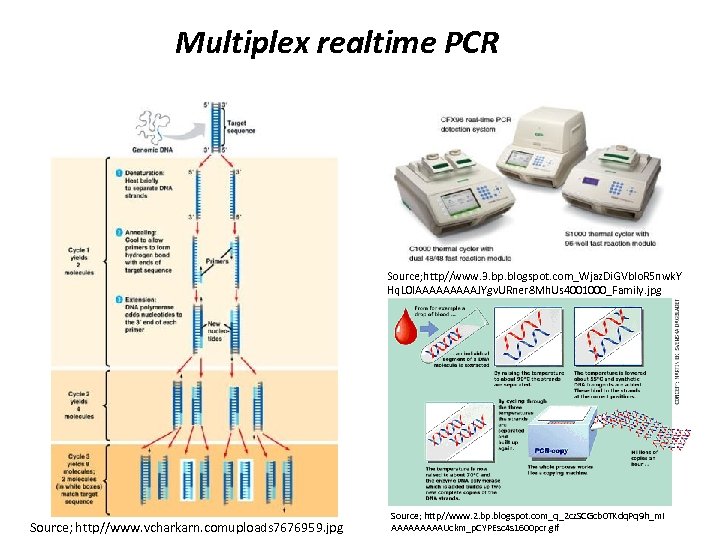
Multiplex realtime PCR Source; http//www. 3. bp. blogspot. com_Wjaz. Di. GVblo. R 5 nwk. Y Hq. L 0 IAAAAAJYgv. URner 8 Mh. Us 4001000_Family. jpg Source; http//www. vcharkarn. comuploads 7676959. jpg Source; http//www. 2. bp. blogspot. com_q_2 cz. SCGcb 0 TKdq. Pq 9 h_m. I AAAAAUckm_p. CYPEsc 4 s 1600 pcr. gif
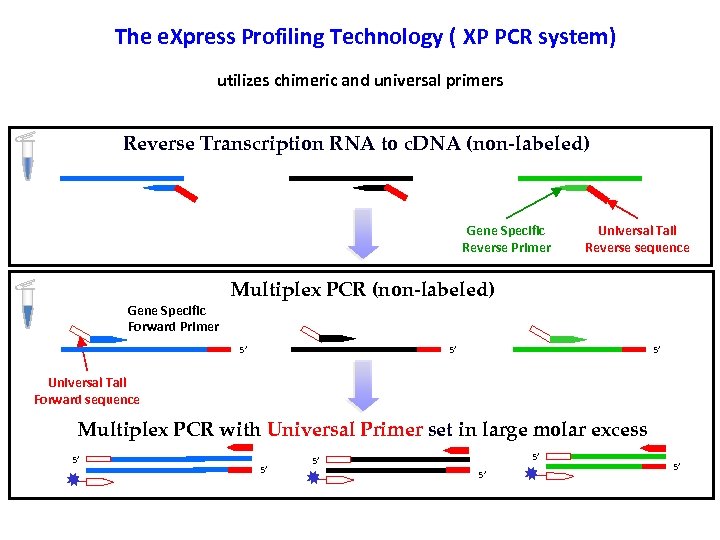
The e. Xpress Profiling Technology ( XP PCR system) utilizes chimeric and universal primers Reverse Transcription RNA to c. DNA (non-labeled) Gene Specific Reverse Primer Gene Specific Forward Primer Universal Tail Reverse sequence Multiplex PCR (non-labeled) 5’ 5’ 5’ Universal Tail Forward sequence Multiplex PCR with Universal Primer set in large molar excess 5’ 5’ 5’
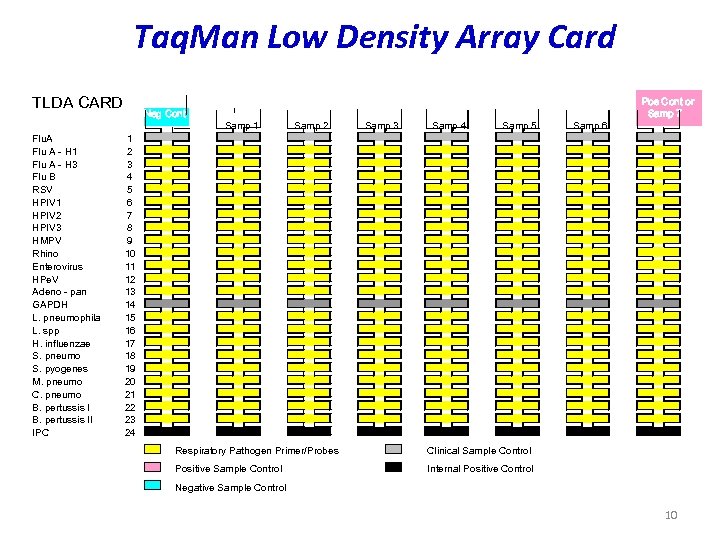
Taq. Man Low Density Array Card TLDA CARD Flu. A Flu A - H 1 Flu A - H 3 Flu B RSV HPIV 1 HPIV 2 HPIV 3 HMPV Rhino Enterovirus HPe. V Adeno - pan GAPDH L. pneumophila L. spp H. influenzae S. pneumo S. pyogenes M. pneumo C. pneumo B. pertussis II IPC Neg Cont 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Samp 1 Samp 2 Samp 3 Samp 4 Samp 5 Respiratory Pathogen Primer/Probes Clinical Sample Control Positive Sample Control Samp 6 Pos Cont or Samp 7 Internal Positive Control Negative Sample Control 10
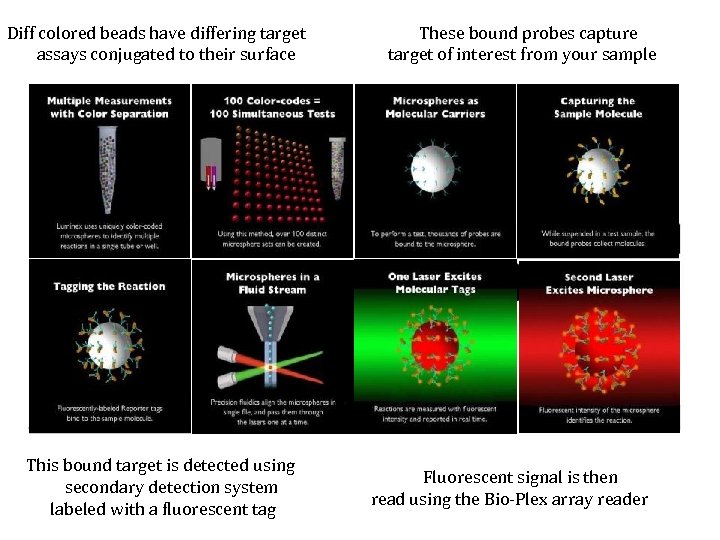
Diff colored beads have differing target assays conjugated to their surface This bound target is detected using secondary detection system labeled with a fluorescent tag These bound probes capture target of interest from your sample Fluorescent signal is then read using the Bio-Plex array reader
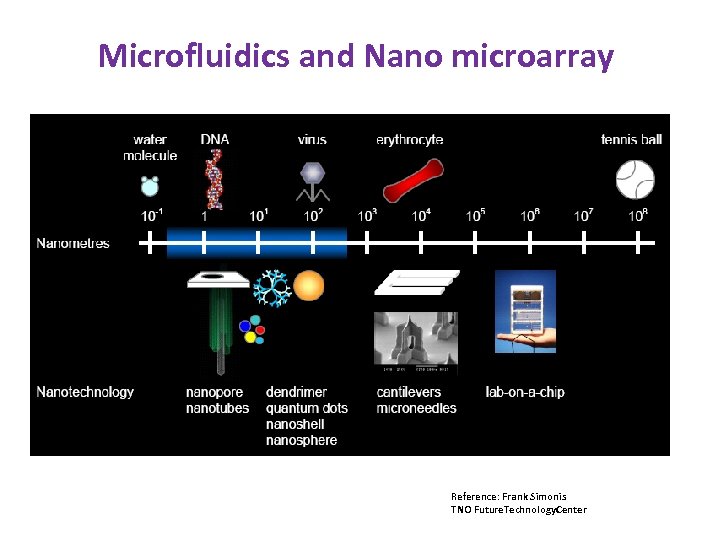
Microfluidics and Nano microarray Reference: Frank Simonis TNO Future. Technology. Center
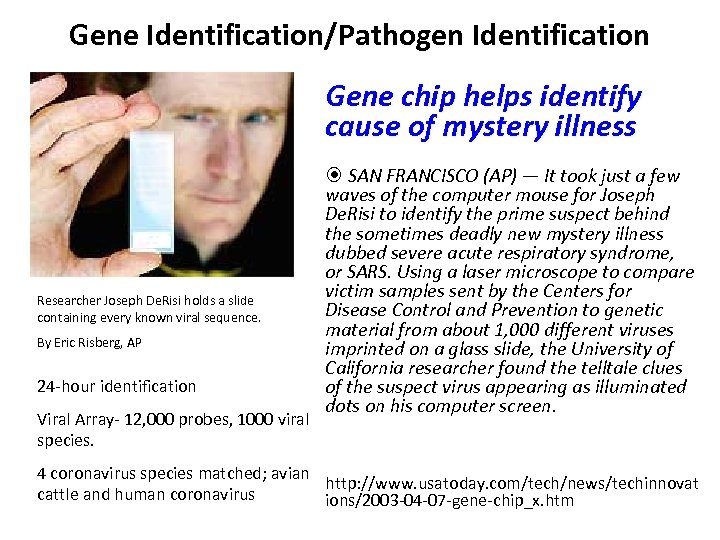

Gene Identification/Pathogen Identification Gene chip helps identify cause of mystery illness Researcher Joseph De. Risi holds a slide containing every known viral sequence. By Eric Risberg, AP 24 -hour identification Viral Array- 12, 000 probes, 1000 viral species. SAN FRANCISCO (AP) — It took just a few waves of the computer mouse for Joseph De. Risi to identify the prime suspect behind the sometimes deadly new mystery illness dubbed severe acute respiratory syndrome, or SARS. Using a laser microscope to compare victim samples sent by the Centers for Disease Control and Prevention to genetic material from about 1, 000 different viruses imprinted on a glass slide, the University of California researcher found the telltale clues of the suspect virus appearing as illuminated dots on his computer screen. 4 coronavirus species matched; avian http: //www. usatoday. com/tech/news/techinnovat cattle and human coronavirus ions/2003 -04 -07 -gene-chip_x. htm
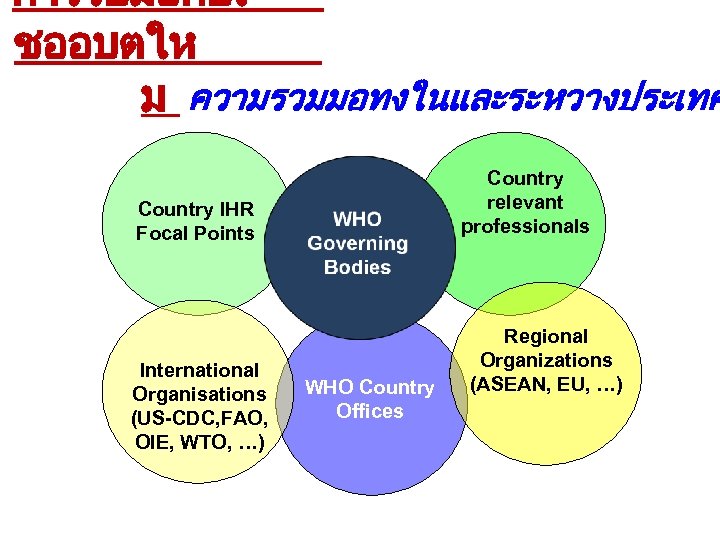
การรบมอกบเ ชออบตให ม ความรวมมอทงในและระหวางประเทศ Country relevant professionals Country IHR Focal Points International Organisations (US-CDC, FAO, OIE, WTO, …) WHO Country Offices Regional Organizations (ASEAN, EU, …)

การเตรยมพรอมของหองปฏบ ตการตอโรคอบตใหม q. Strengthen/upgrade : Genetic identification , BSL-3 Laboratory Network Variety of cell culture isolation : Good collaboration and Rapid response q National Bio-Medical Resources Center q. Information Technology Support
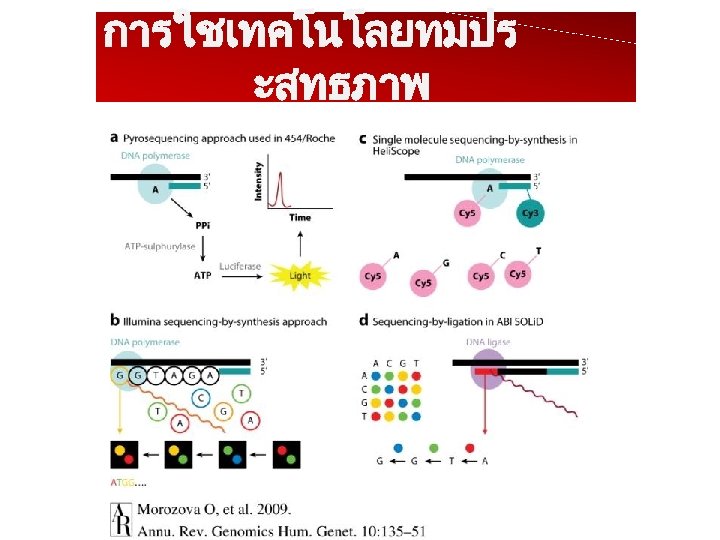
การใชเทคโนโลยทมปร ะสทธภาพ
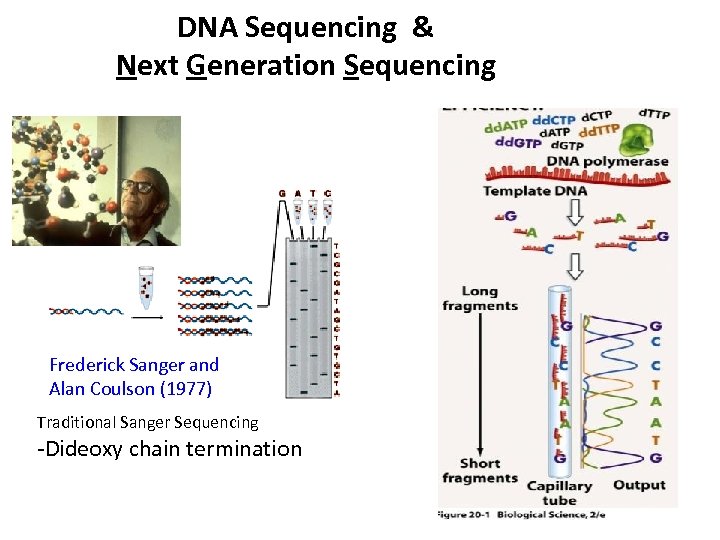
DNA Sequencing & Next Generation Sequencing Frederick Sanger and Alan Coulson (1977) Traditional Sanger Sequencing -Dideoxy chain termination
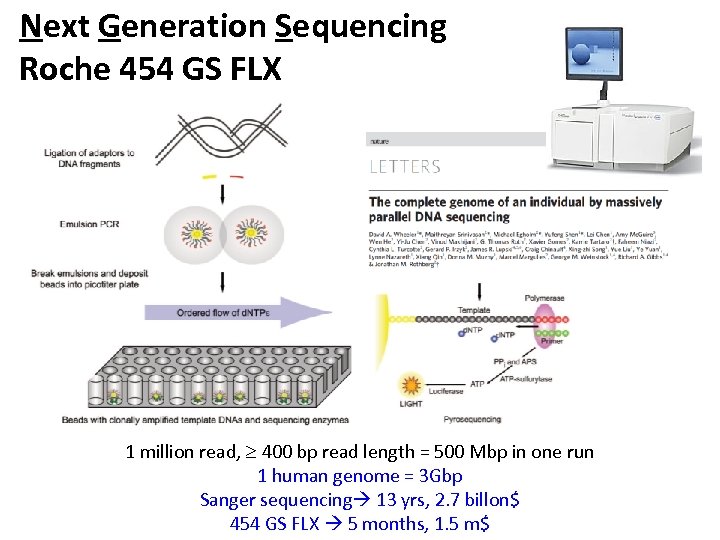
Next Generation Sequencing Roche 454 GS FLX 1 million read, 400 bp read length = 500 Mbp in one run 1 human genome = 3 Gbp Sanger sequencing 13 yrs, 2. 7 billon$ 454 GS FLX 5 months, 1. 5 m$
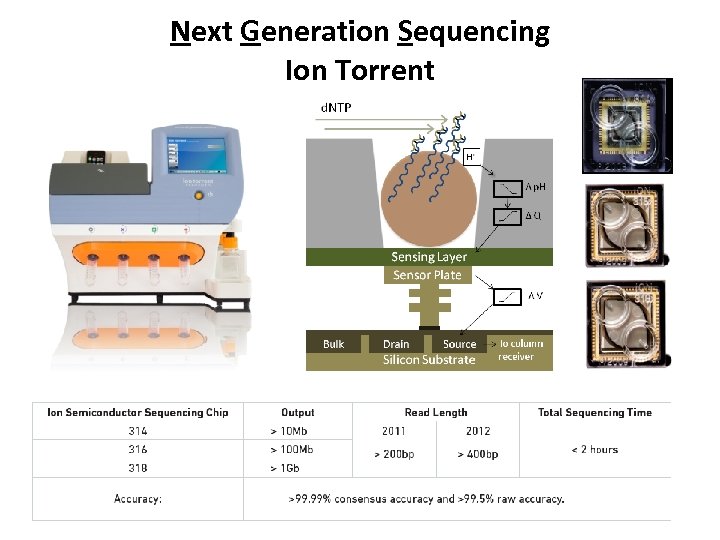
Next Generation Sequencing Ion Torrent
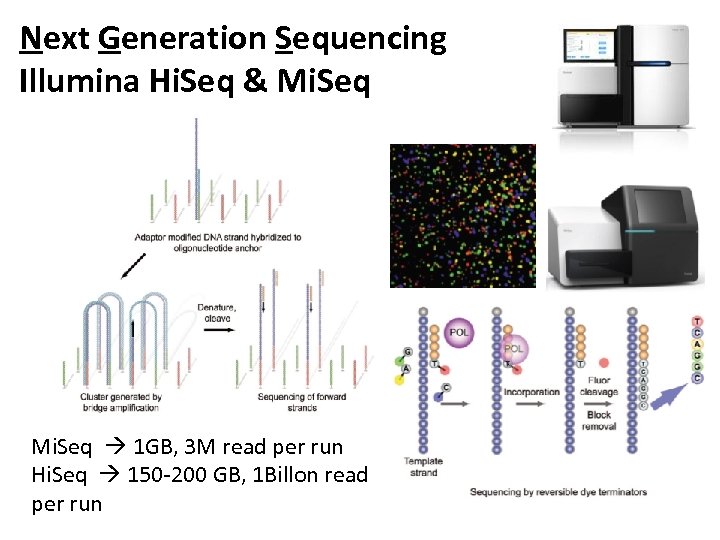
Next Generation Sequencing Illumina Hi. Seq & Mi. Seq 1 GB, 3 M read per run Hi. Seq 150 -200 GB, 1 Billon read per run
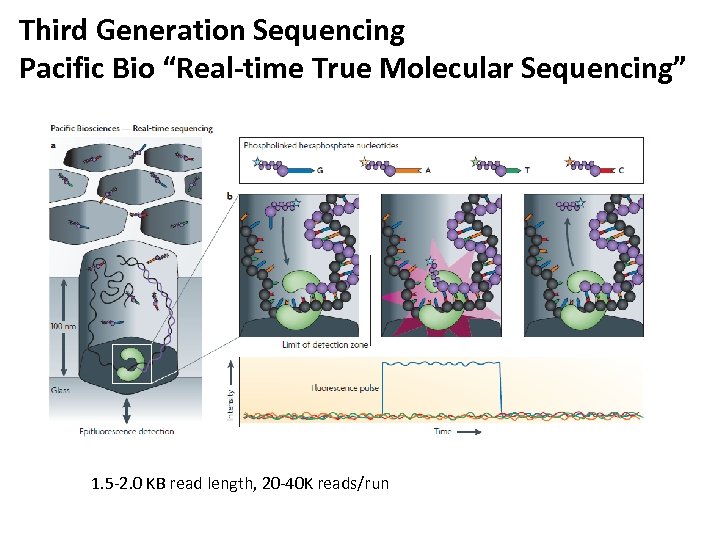
Third Generation Sequencing Pacific Bio “Real-time True Molecular Sequencing” 1. 5 -2. 0 KB read length, 20 -40 K reads/run
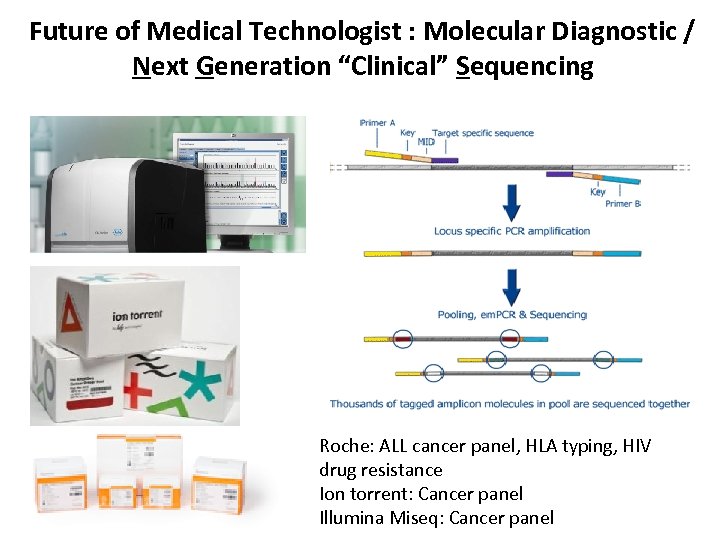
Future of Medical Technologist : Molecular Diagnostic / Next Generation “Clinical” Sequencing Roche: ALL cancer panel, HLA typing, HIV drug resistance Ion torrent: Cancer panel Illumina Miseq: Cancer panel

Novel Coronavirus Recently Detected Coronaviruses are common around the world. They usually cause colds. However, this year, a novel coronavirus was identified as the cause of severe respiratory illness in adults from Middle Eastern countries. Learn more about this novel coronavirus
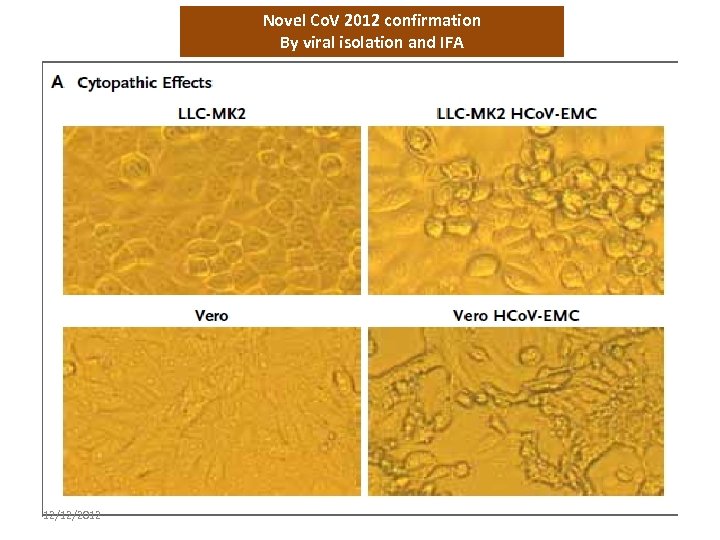
Novel Co. V 2012 confirmation By viral isolation and IFA 12/12/2012
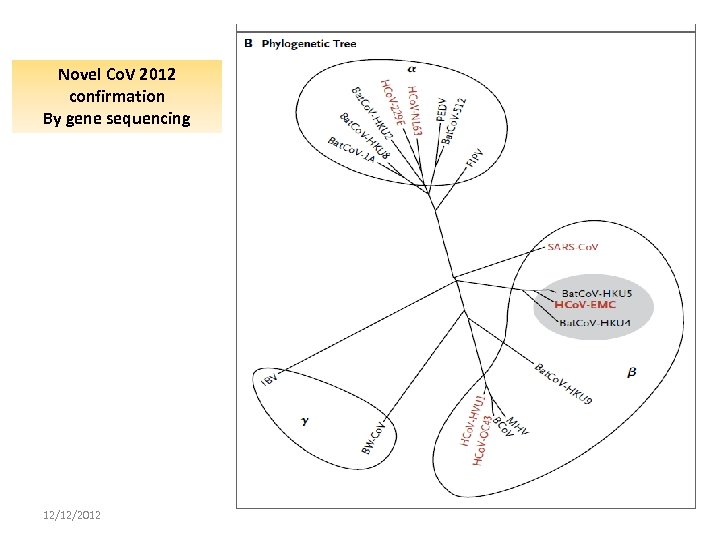
Novel Co. V 2012 confirmation By gene sequencing 12/12/2012

ศกยภาพหองปฏบตการในป จจบน และการพฒนาในอนาคต
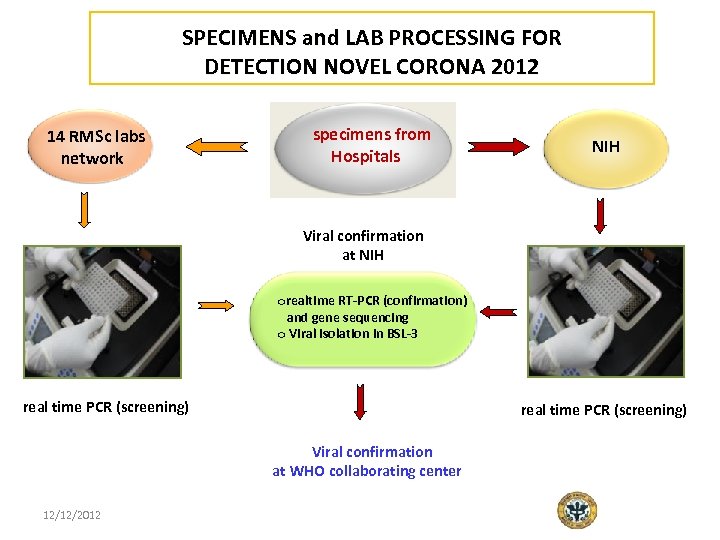
SPECIMENS and LAB PROCESSING FOR DETECTION NOVEL CORONA 2012 14 RMSc labs network specimens from Hospitals NIH Viral confirmation at NIH orealtime RT-PCR (confirmation) and gene sequencing o Viral isolation in BSL-3 real time PCR (screening) Viral confirmation at WHO collaborating center 12/12/2012
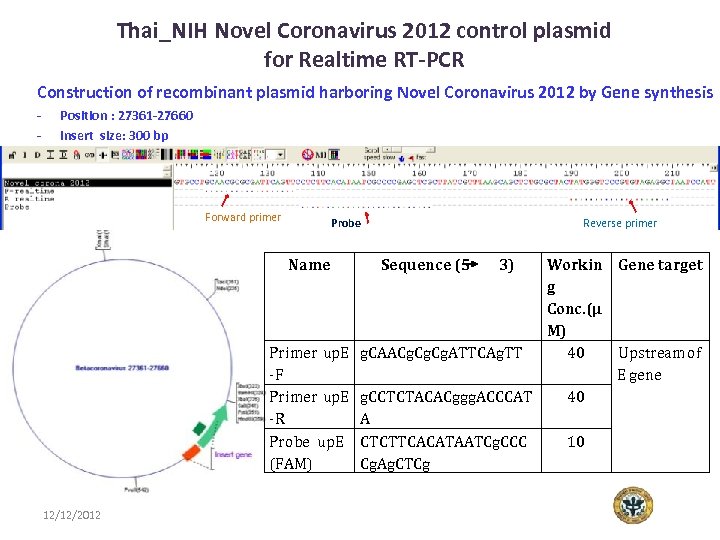
Thai_NIH Novel Coronavirus 2012 control plasmid for Realtime RT-PCR Construction of recombinant plasmid harboring Novel Coronavirus 2012 by Gene synthesis - Position : 27361 -27660 Insert size: 300 bp Forward primer Probe Name Reverse primer Workin Gene target g Conc. (µ M) Primer up. E g. CAACg. Cg. ATTCAg. TT 40 Upstream of -F E gene Primer up. E g. CCTCTACACggg. ACCCAT 40 -R A Probe up. E CTCTTCACATAATCg. CCC 10 (FAM) Cg. Ag. CTCg 12/12/2012 Sequence (5 3)
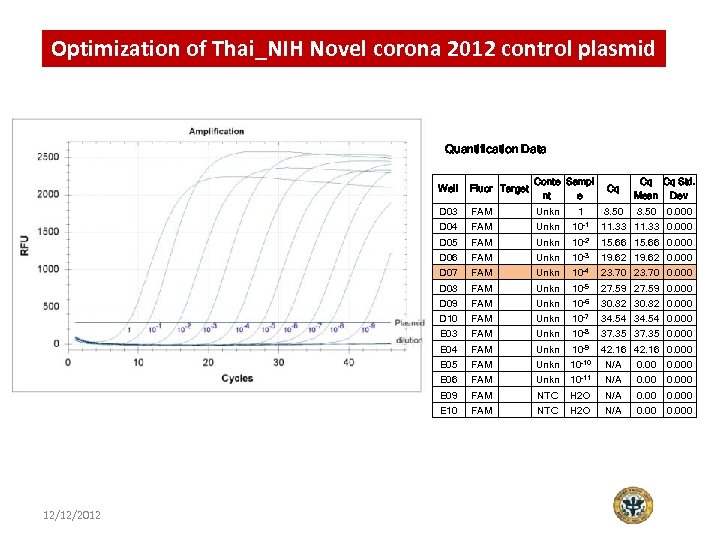
Optimization of Thai_NIH Novel corona 2012 control plasmid Quantification Data Well Fluor Target Conte Sampl nt e D 03 FAM Unkn 10 -1 11. 33 0. 000 Unkn 10 -2 15. 66 0. 000 Unkn 10 -3 19. 62 0. 000 23. 70 0. 000 D 04 D 05 D 06 FAM FAM Cq 8. 50 Cq Cq Std. Mean Dev 8. 50 0. 000 D 07 FAM Unkn 10 -4 D 08 FAM Unkn 10 -5 27. 59 0. 000 Unkn 10 -6 30. 82 0. 000 Unkn 10 -7 34. 54 0. 000 Unkn 10 -8 37. 35 0. 000 Unkn 10 -9 42. 16 0. 000 D 09 D 10 E 03 E 04 FAM FAM E 05 Unkn N/A 0. 000 E 06 FAM Unkn 10 -11 N/A 0. 000 E 09 FAM NTC H 2 O N/A 0. 000 E 10 12/12/2012 FAM 10 -10 FAM NTC H 2 O N/A 0. 000
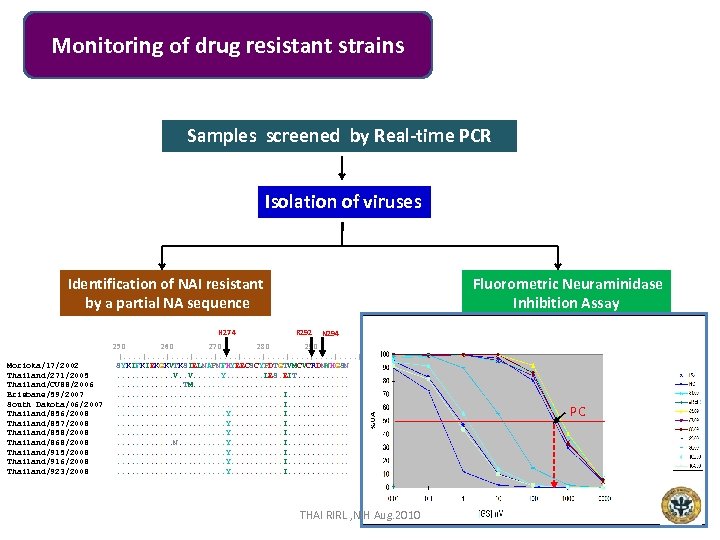
Monitoring of drug resistant strains Samples screened by Real-time PCR Isolation of viruses Weekly random submission Identification of NAI resistant by a partial NA sequence H 274 Morioka/17/2002 Thailand/271/2005 Thailand/CU 88/2006 Brisbane/59/2007 South Dakota/06/2007 Thailand/856/2008 Thailand/857/2008 Thailand/858/2008 Thailand/868/2008 Thailand/915/2008 Thailand/916/2008 Thailand/923/2008 Fluorometric Neuraminidase Inhibition Assay R 292 N 294 250 260 270 280 290 |. . . . | SYKIFKIEKGKVTKSIELNAPNFHYEECSCYPDTGTVMCVCRDNWHGSN. . . V. . . . Y. . . . LES. EIT. . . TM. . . . . I. . . . . . . . Y. . . . . I. . . . . Y. . . I. . . N. . . . . Y. . . . . I. . . . Y. . . I. . . . . Y. . . I. . . THAI RIRL , NIH Aug. 2010 PC 30
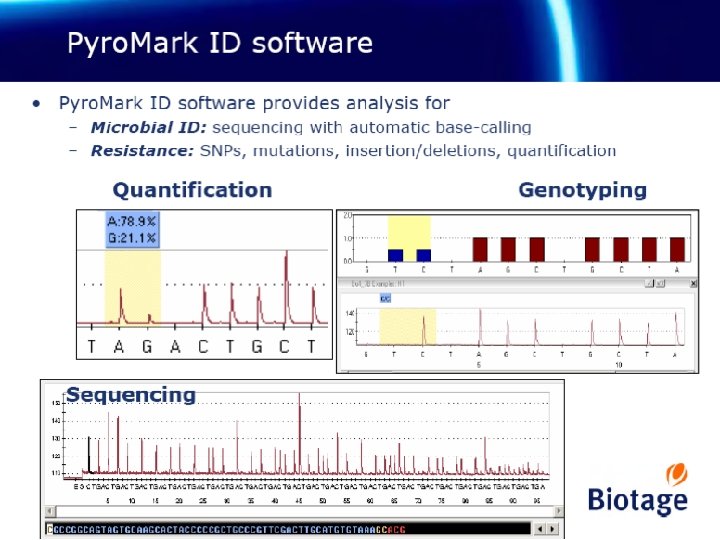
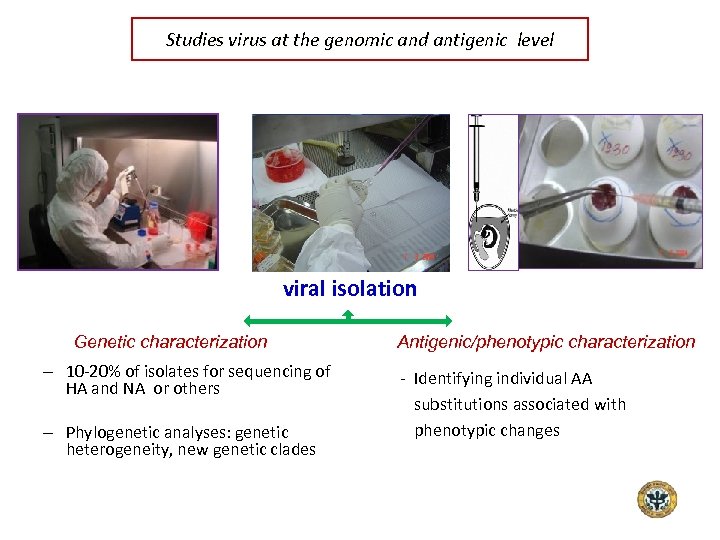
Studies virus at the genomic and antigenic level viral isolation Genetic characterization – 10 -20% of isolates for sequencing of HA and NA or others – Phylogenetic analyses: genetic heterogeneity, new genetic clades Antigenic/phenotypic characterization - Identifying individual AA substitutions associated with phenotypic changes
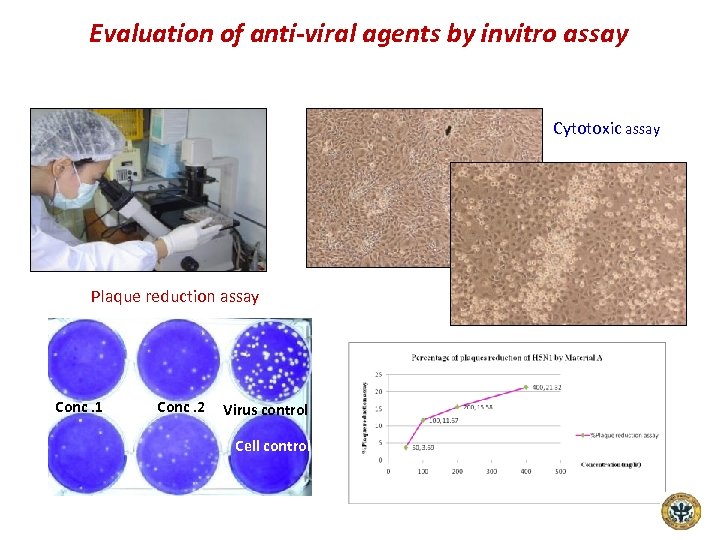
Evaluation of anti-viral agents by invitro assay Cytotoxic assay Plaque reduction assay Conc. 1 Conc. 2 Virus control Cell control
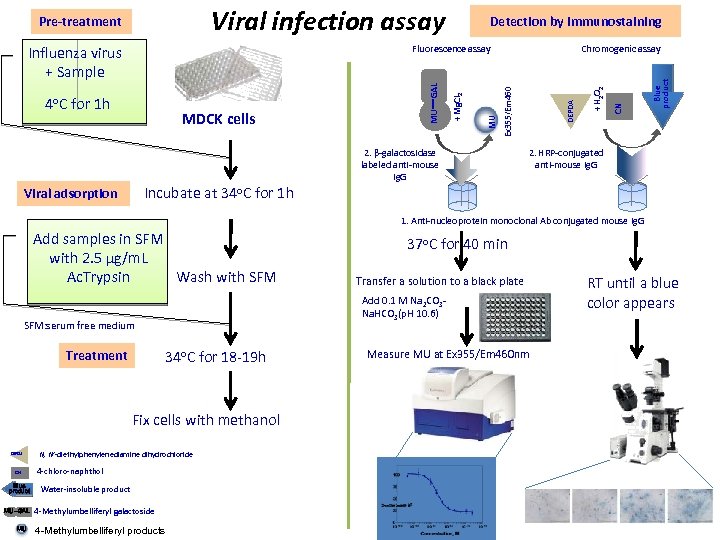
Viral infection assay 2. β-galactosidase labeled anti-mouse Ig. G DEPDA Blue product Chromogenic assay Ex 355/Em 460 MU MDCK cells + Mg. Cl 2 4 o. C for 1 h MUーGAL Fluorescence assay CN Influenza virus + Sample Detection by immunostaining + H 2 O 2 Pre-treatment 2. HRP-conjugated anti-mouse Ig. G Incubate at 34 o. C for 1 h Viral adsorption 1. Anti-nucleoprotein monoclonal Ab conjugated mouse Ig. G Add samples in SFM with 2. 5 µg/m. L Wash with SFM Ac. Trypsin 34 o. C for 18 -19 h Fix cells with methanol DEPDA CN Blue product MUーGAL MU Transfer a solution to a black plate Add 0. 1 M Na 2 CO 3 Na. HCO 3(p. H 10. 6) SFM: serum free medium Treatment 37 o. C for 40 min N, N’-diethylphenylenediamine dihydrochloride 4 -chloro-naphthol Water-insoluble product 4 -Methylumbelliferyl galactoside 4 -Methylumbelliferyl products Measure MU at Ex 355/Em 460 nm RT until a blue color appears
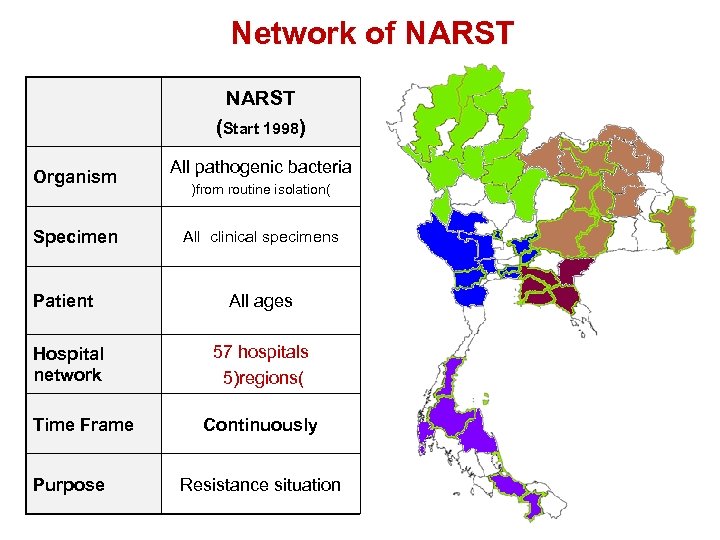
Network of NARST (Start 1998) Organism Specimen Patient Hospital network Time Frame Purpose All pathogenic bacteria )from routine isolation( All clinical specimens All ages 57 hospitals 5)regions( Continuously Resistance situation
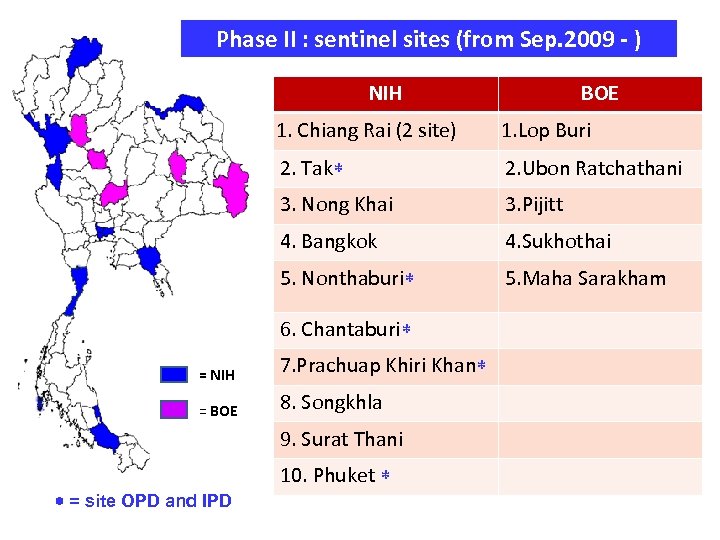
Phase II : sentinel sites (from Sep. 2009 - ) NIH BOE 1. Chiang Rai (2 site) 1. Lop Buri 2. Tak 2. Ubon Ratchathani 3. Nong Khai 3. Pijitt 4. Bangkok 4. Sukhothai 5. Nonthaburi 5. Maha Sarakham 6. Chantaburi = NIH 7. Prachuap Khiri Khan = BOE 8. Songkhla 9. Surat Thani 10. Phuket = site OPD and IPD
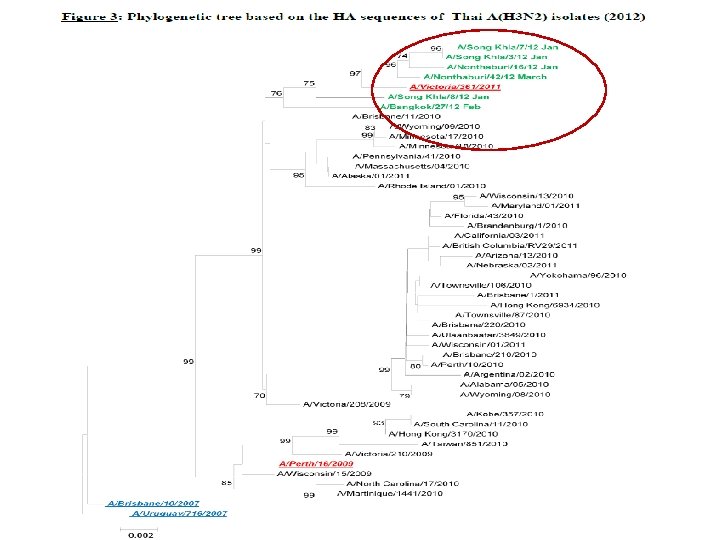
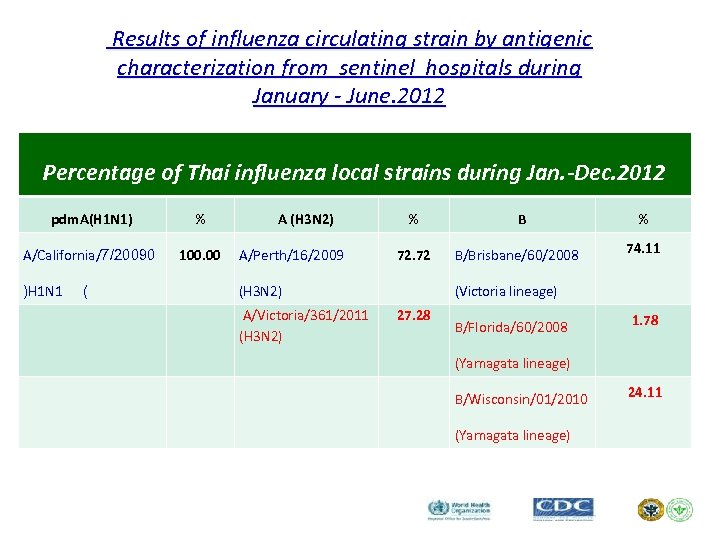
Results of influenza circulating strain by antigenic characterization from sentinel hospitals during January - June. 2012 Percentage of Thai influenza local strains during Jan. -Dec. 2012 pdm. A(H 1 N 1) % A/California/7/20090 100. 00 )H 1 N 1 ( A (H 3 N 2) A/Perth/16/2009 % 72. 72 (H 3 N 2) A/Victoria/361/2011 (H 3 N 2) B B/Brisbane/60/2008 % 74. 11 (Victoria lineage) 27. 28 B/Florida/60/2008 1. 78 (Yamagata lineage) B/Wisconsin/01/2010 (Yamagata lineage) 24. 11

Mae Sot Hospital Thai NIH_13 Oct. 2011 Nong Khai Hospital

Phra Pok Klao hospital Thai NIH_13 Oct. 2011 Hat Yai hospital

Chiang Saen Hospital Mae Chan Hospital Koh Chang Hospital Koh Samui Hospital Bangkok Hospital Samui Thai NIH_13 Oct. 2011
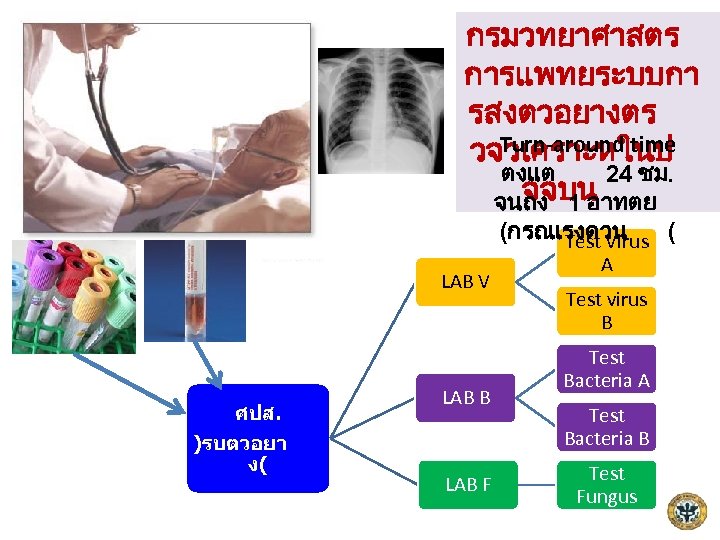
กรมวทยาศาสตร การแพทยระบบกา รสงตวอยางตร Turn-around time วจวเคราะหในป ตงแต 24 ชม. จจบน จนถง 1 อาทตย (กรณเรงดวน Test Virus ( LAB V ศปส. )รบตวอยา ง( LAB B LAB F A Test virus B Test Bacteria A Test Bacteria B Test Fungus
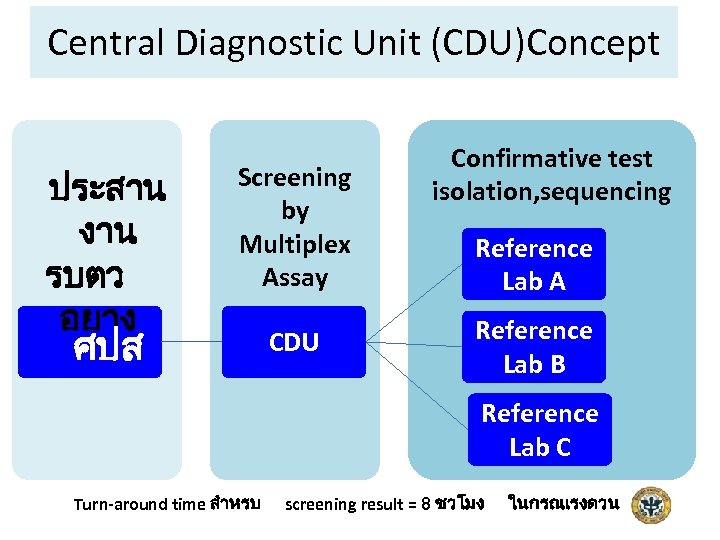
Central Diagnostic Unit (CDU)Concept ประสาน งาน รบตว อยาง Screening by Multiplex Assay ศปส CDU Confirmative test isolation, sequencing Reference Lab A Reference Lab B Reference Lab C Turn-around time สำหรบ screening result = 8 ชวโมง ในกรณเรงดวน

ขอบคณคะ
99b09b1a668b16b3bbd5db7c01dd0e44.ppt