MSU & SkolTech Replication in bacteria Replication Chemistry


MSU & SkolTech Replication in bacteria
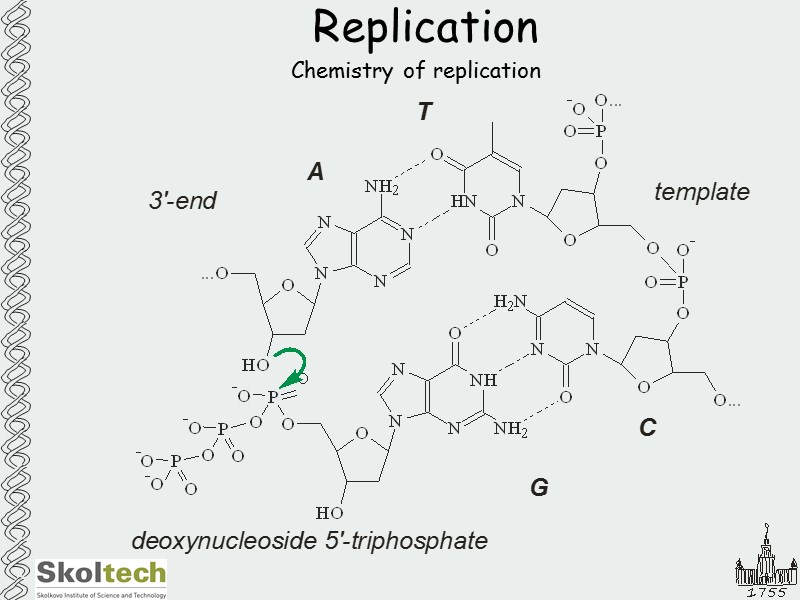
Replication Chemistry of replication
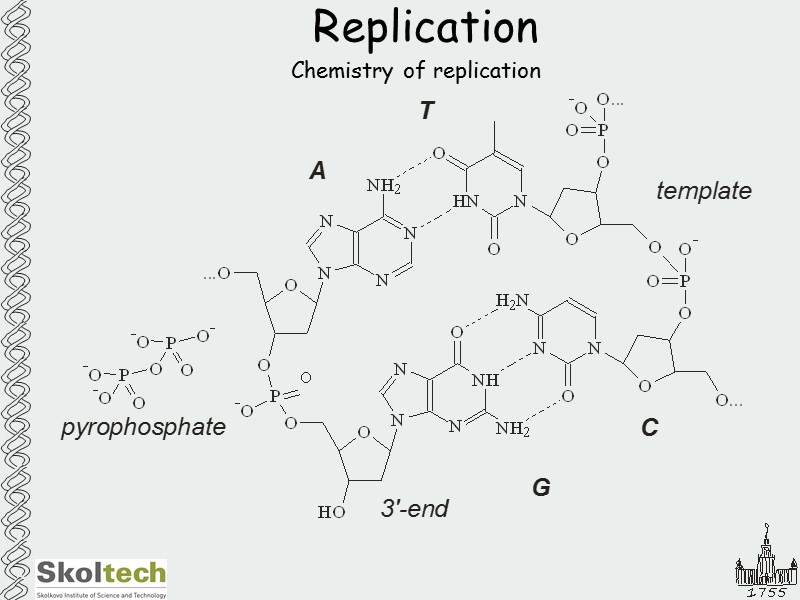
Replication Chemistry of replication
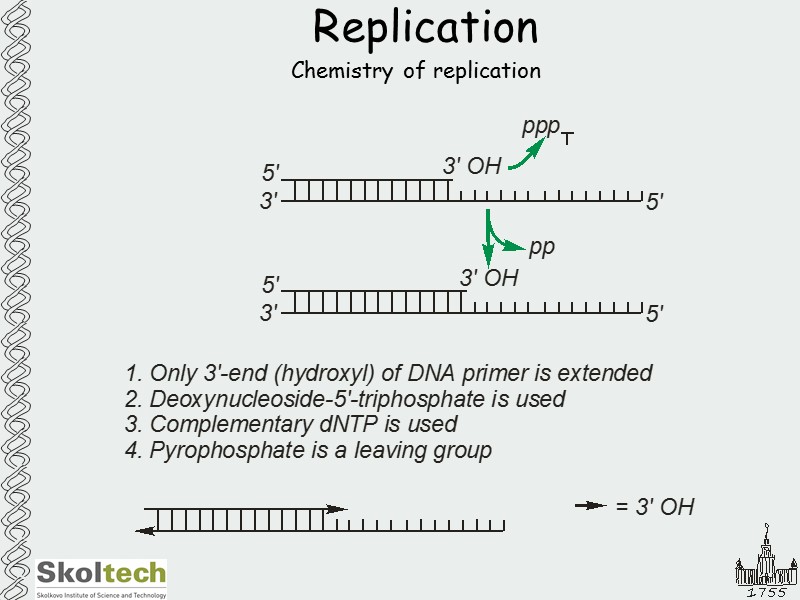
Replication Chemistry of replication
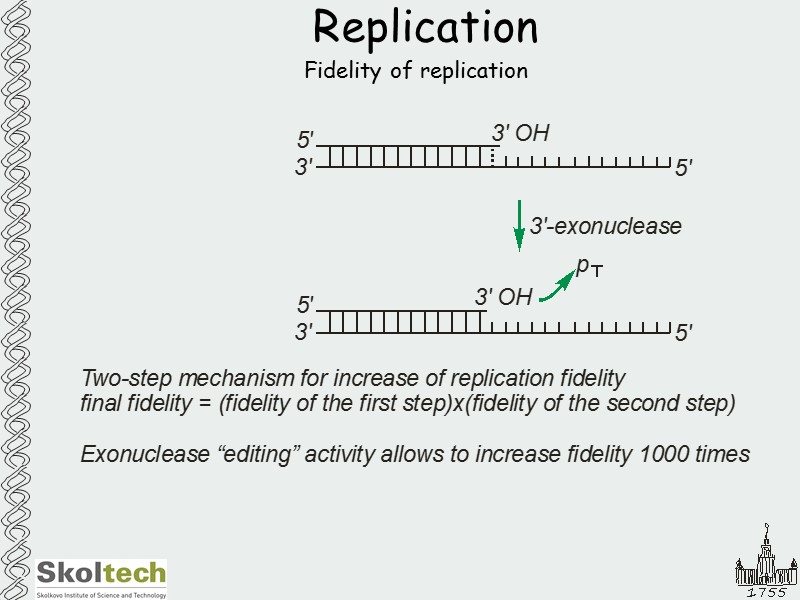
Replication Fidelity of replication
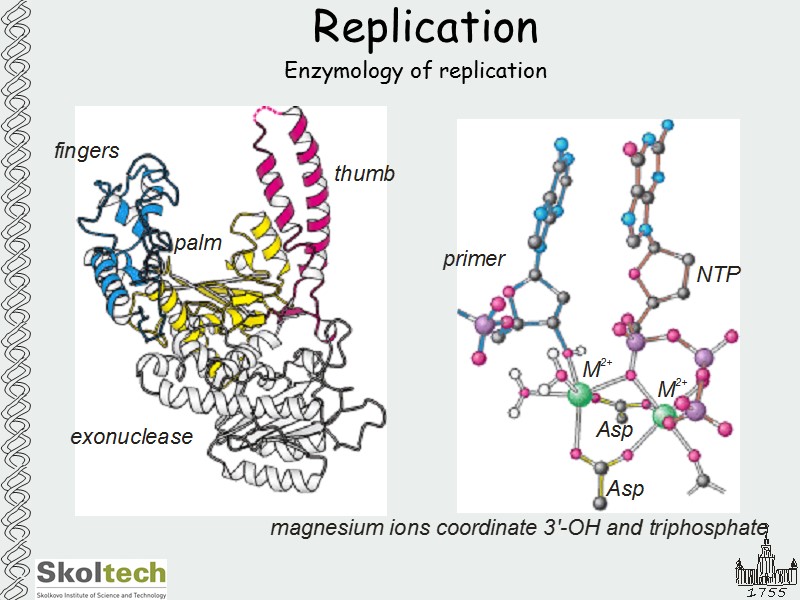
Replication Enzymology of replication

Replication Enzymology of replication Replicative DNA polymerases DNA polymerase I + 3' exo + 5' exo 103 kD DNA polymerase III a 130 kD polymerase e 27 kD 3'-exo q 9 kD d 39 kD c 17 kD y 15 kD g 48 kD b 41 kD processivity factor t 71 kD

Replication Enzymology of replication Reparative DNA polymerases DNA polymerase II + 3' exo 90 kD DNA polymerase IV 40 kD DNA polymerase V umuD’ 15 kD umuC’ 48 kD
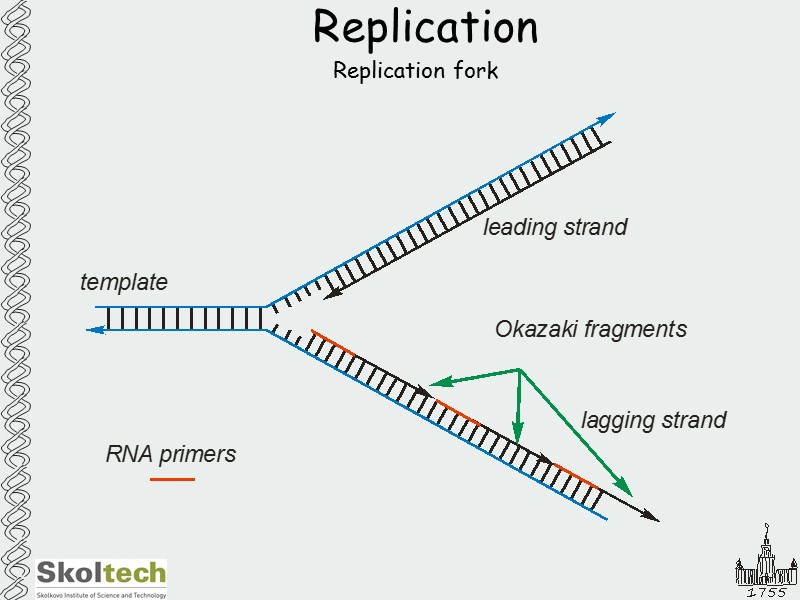
Replication Replication fork
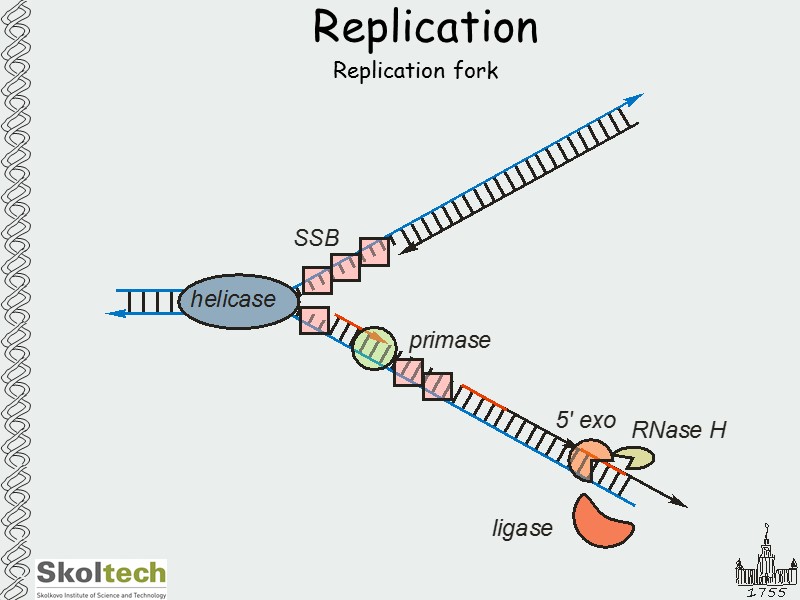
Replication Replication fork
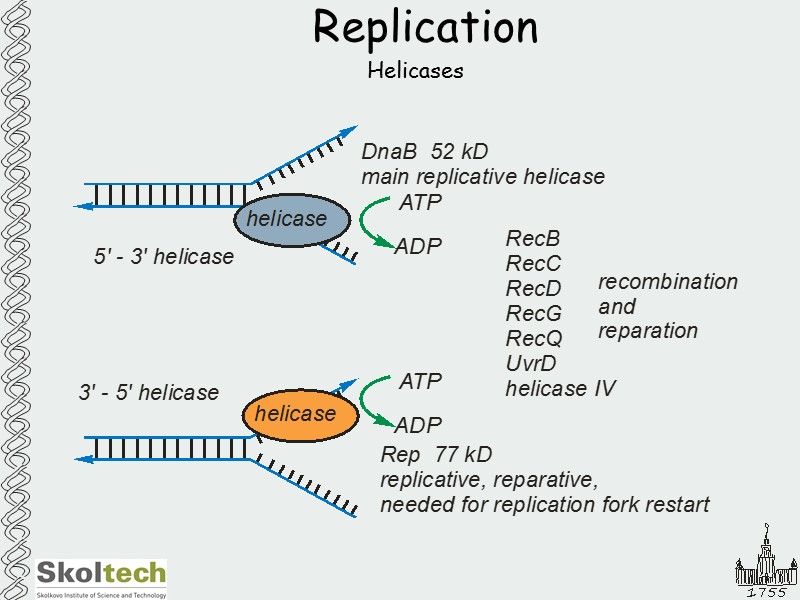
Replication Helicases
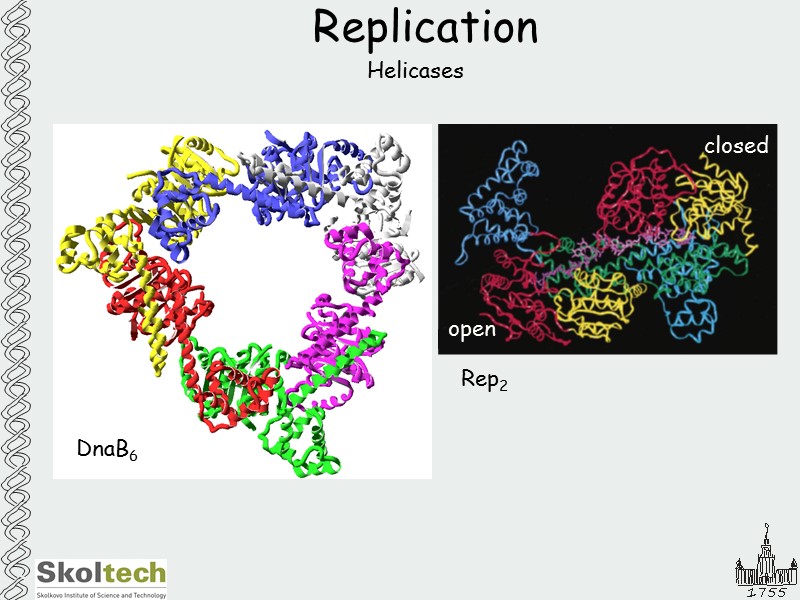
Replication Helicases DnaB6 Rep2 open closed
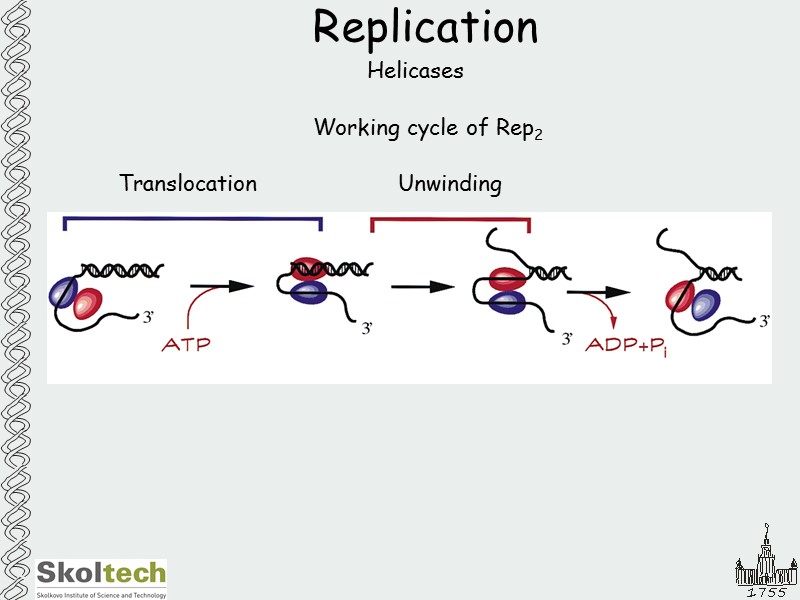
Replication Helicases Working cycle of Rep2 Translocation Unwinding
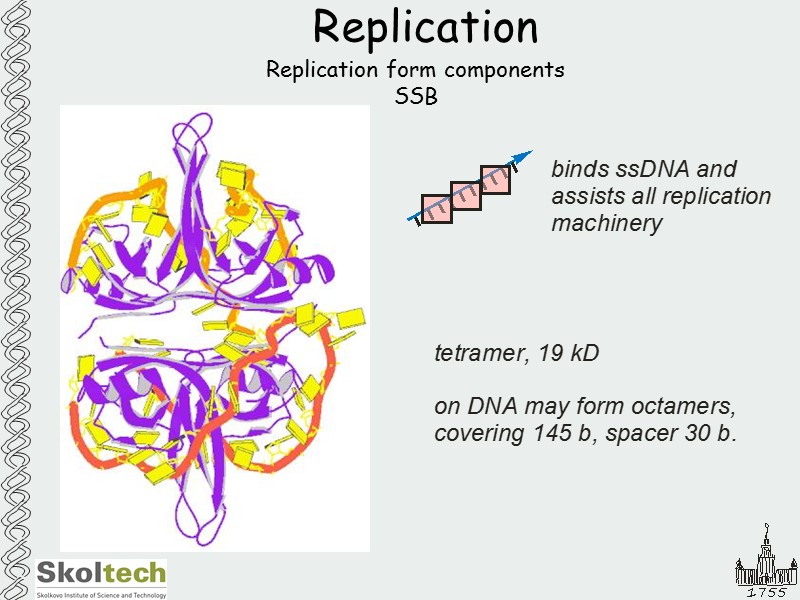
Replication Replication form components SSB
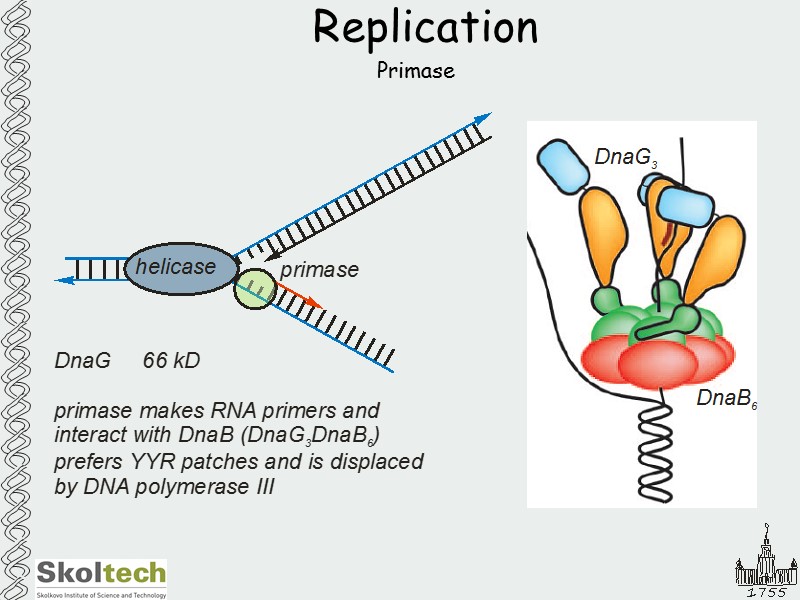
Replication Primase

Replication Primase
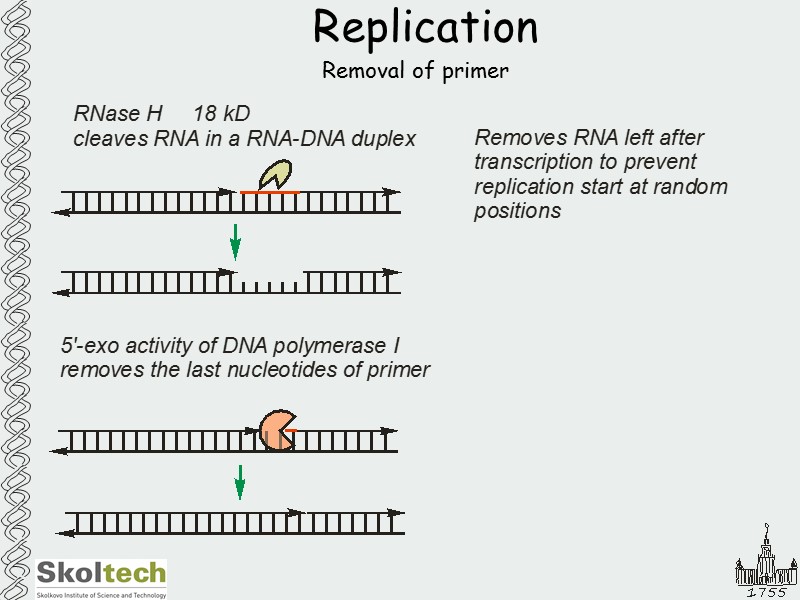
Replication Removal of primer
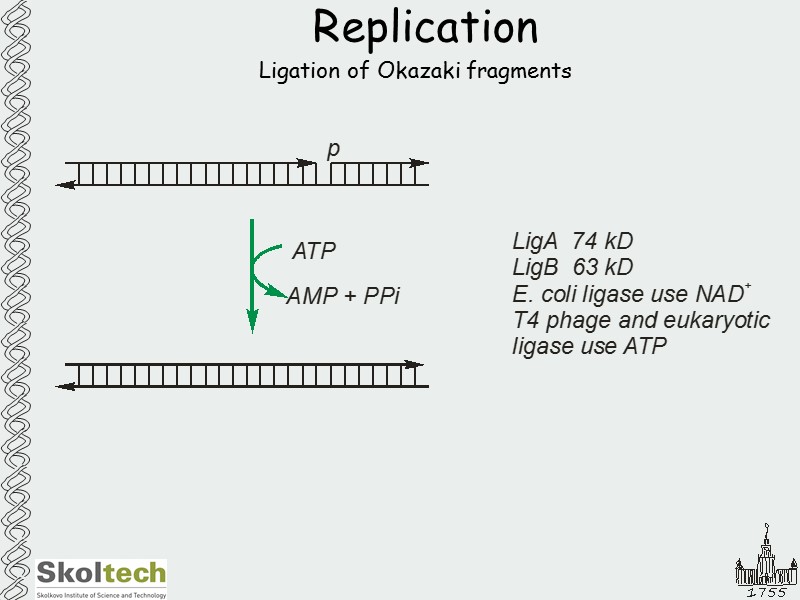
Replication Ligation of Okazaki fragments
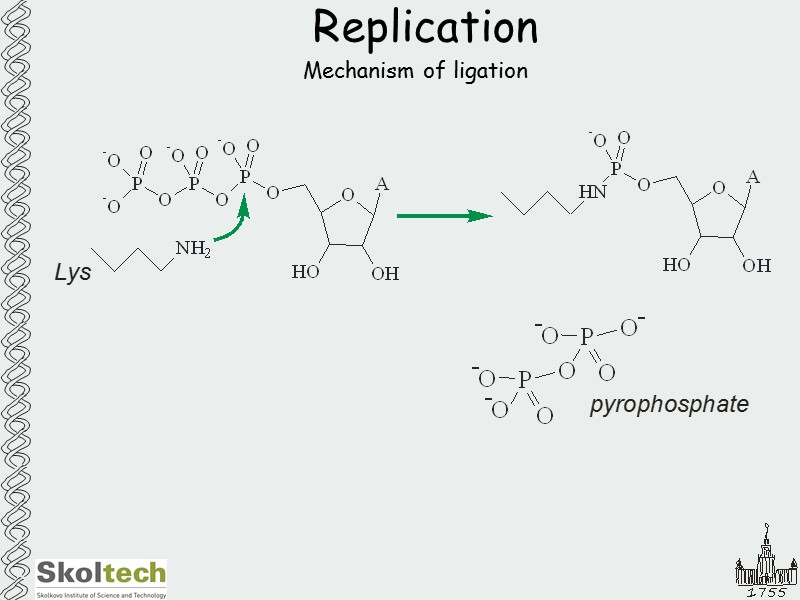
Replication Mechanism of ligation
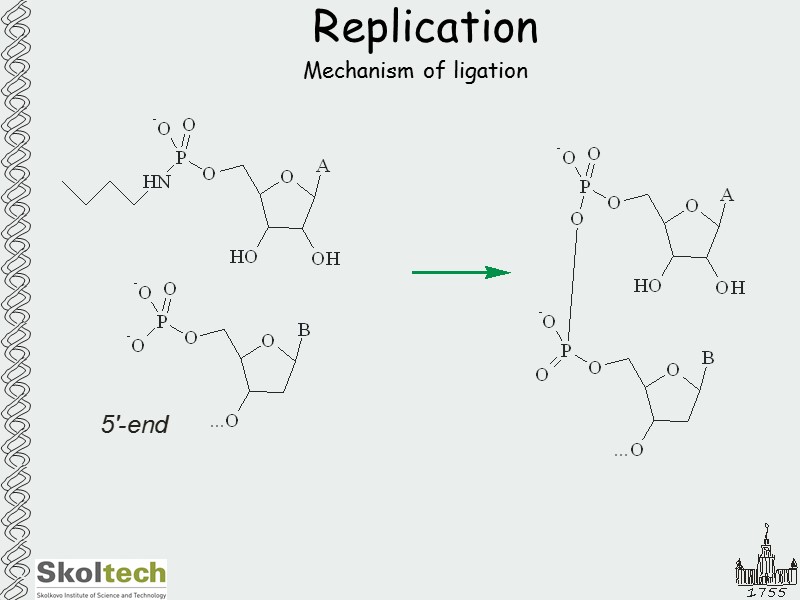
Replication Mechanism of ligation
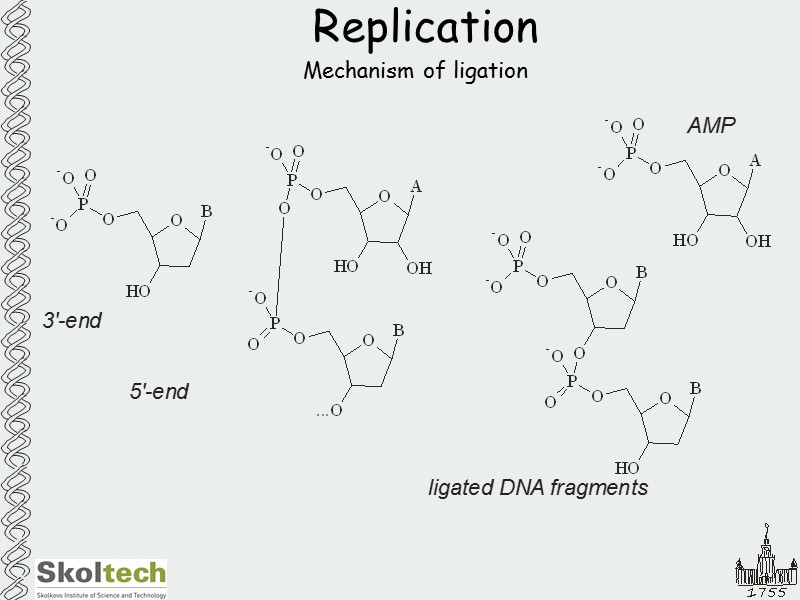
Replication Mechanism of ligation
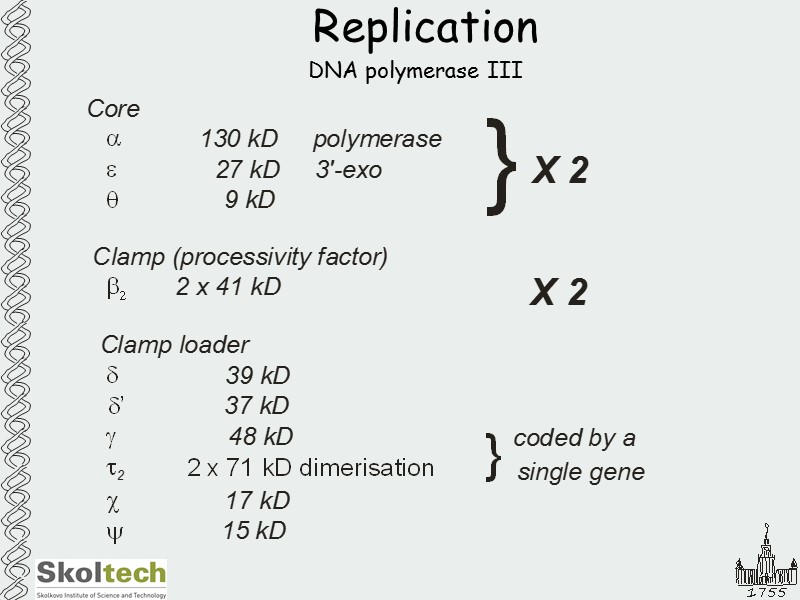

Replication DNA polymerase III
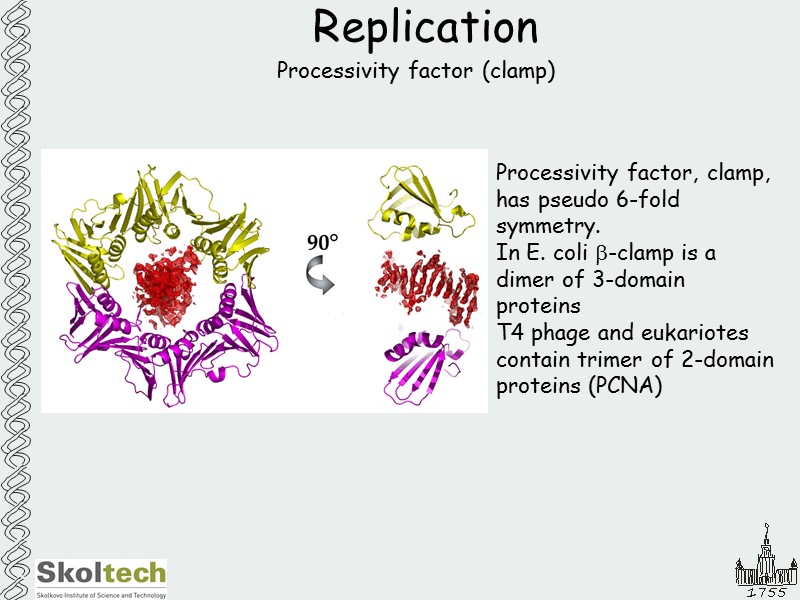
Replication Processivity factor (clamp) Processivity factor, clamp, has pseudo 6-fold symmetry. In E. coli b-clamp is a dimer of 3-domain proteins Т4 phage and eukariotes contain trimer of 2-domain proteins (PCNA)
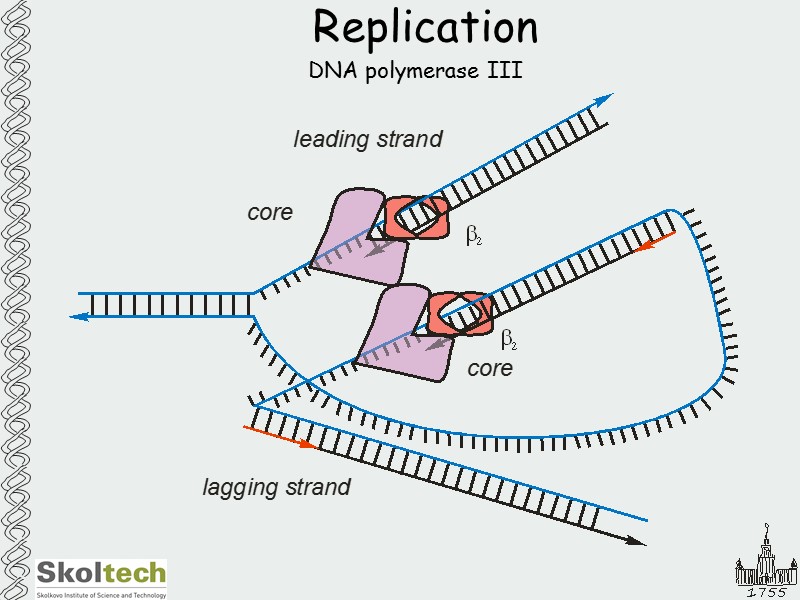
Replication DNA polymerase III
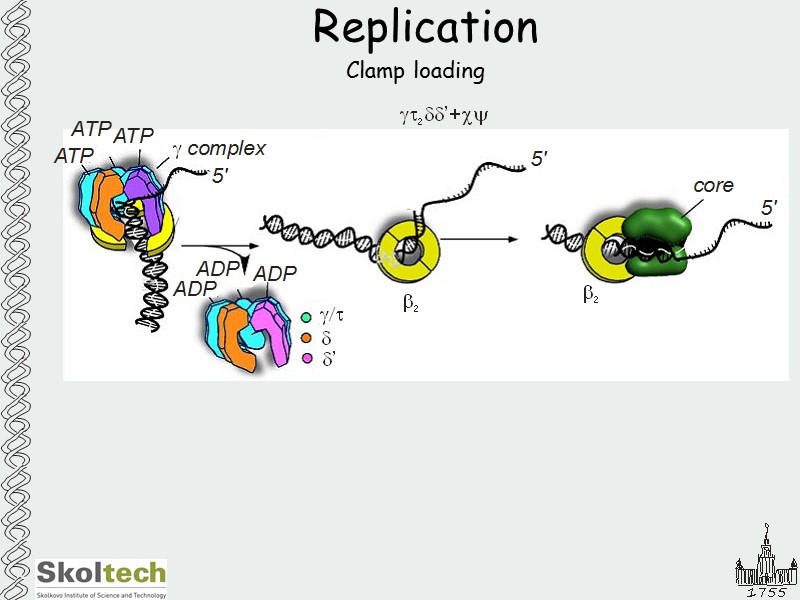
Replication Clamp loading
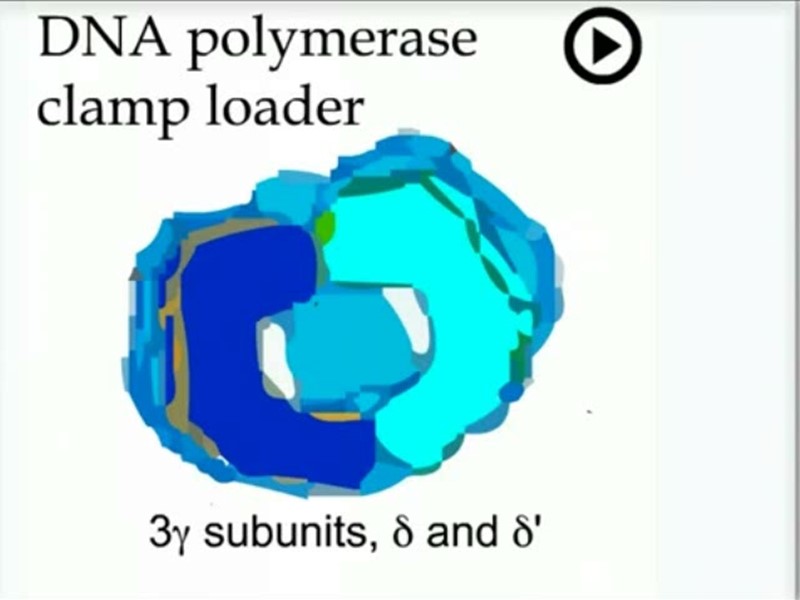

Replication Clamp loading
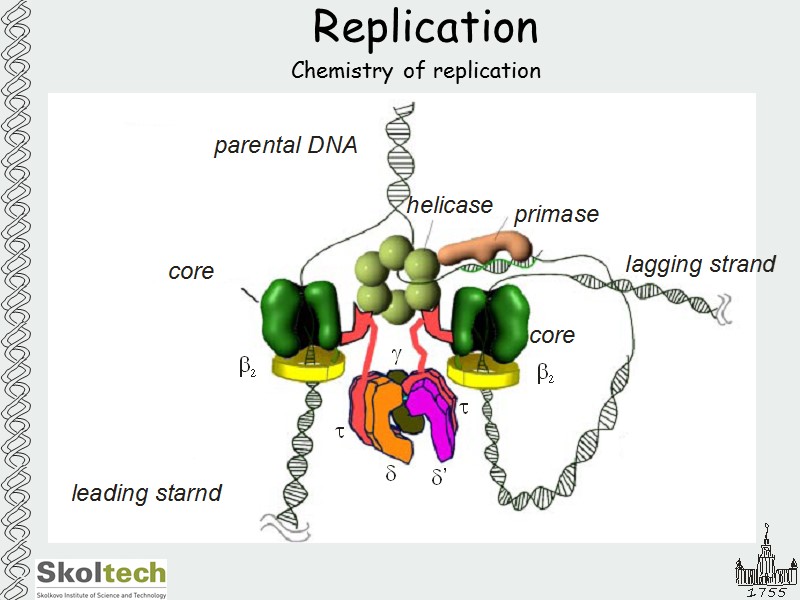
Replication Chemistry of replication
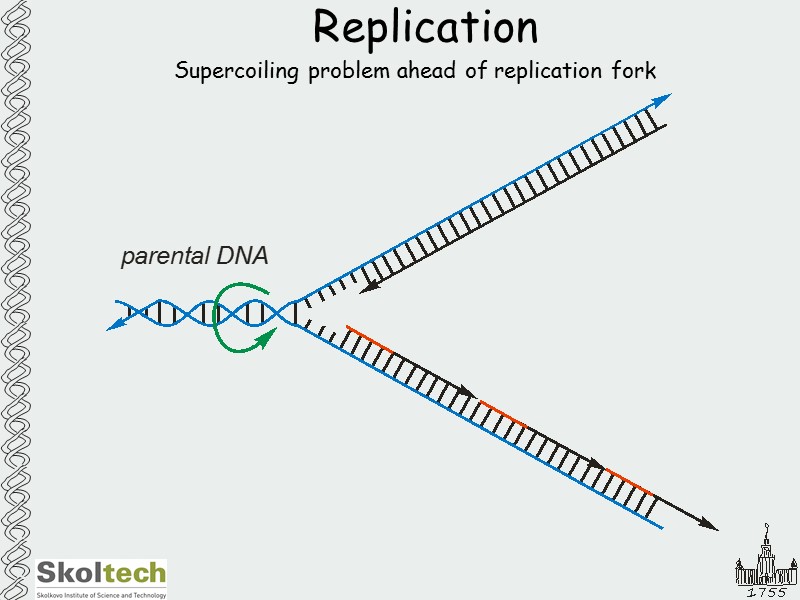
Replication Supercoiling problem ahead of replication fork
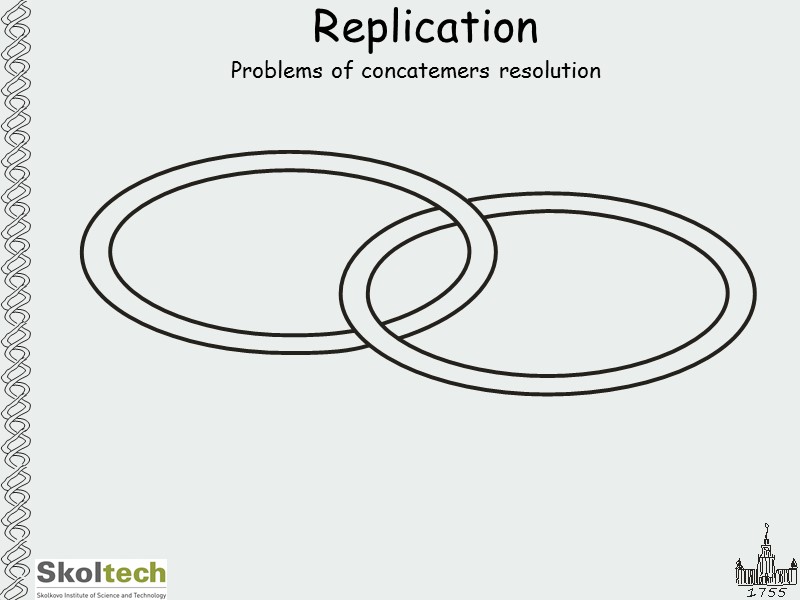
Replication Problems of concatemers resolution
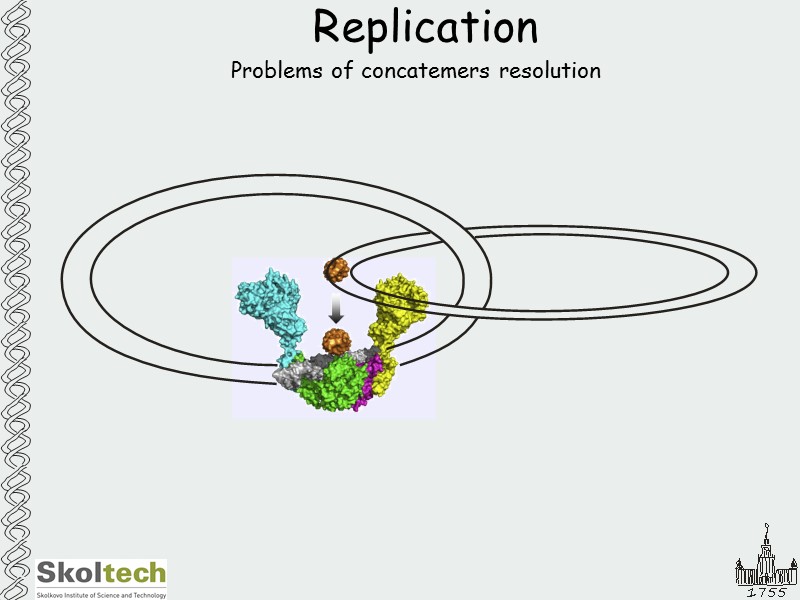
Replication Problems of concatemers resolution
Replication Problems of concatemers resolution
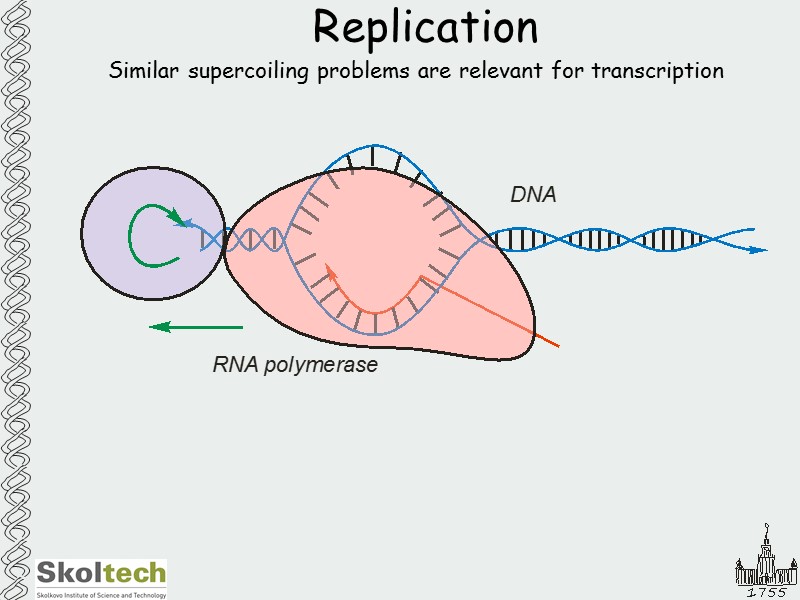
Replication Similar supercoiling problems are relevant for transcription
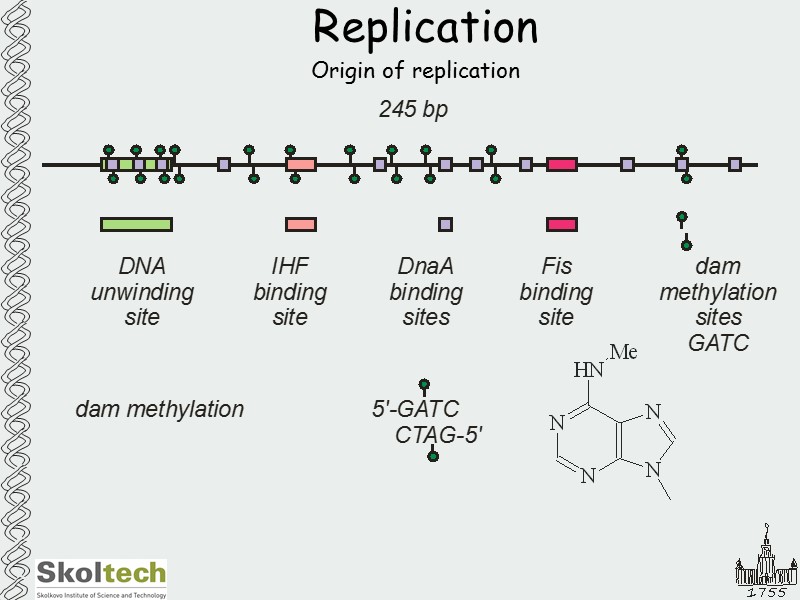
Replication Origin of replication
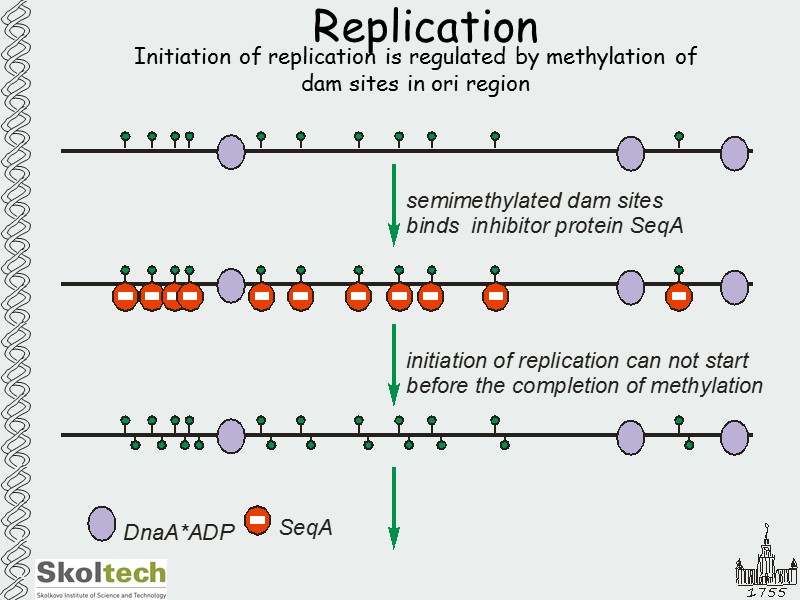
Replication Initiation of replication is regulated by methylation of dam sites in ori region
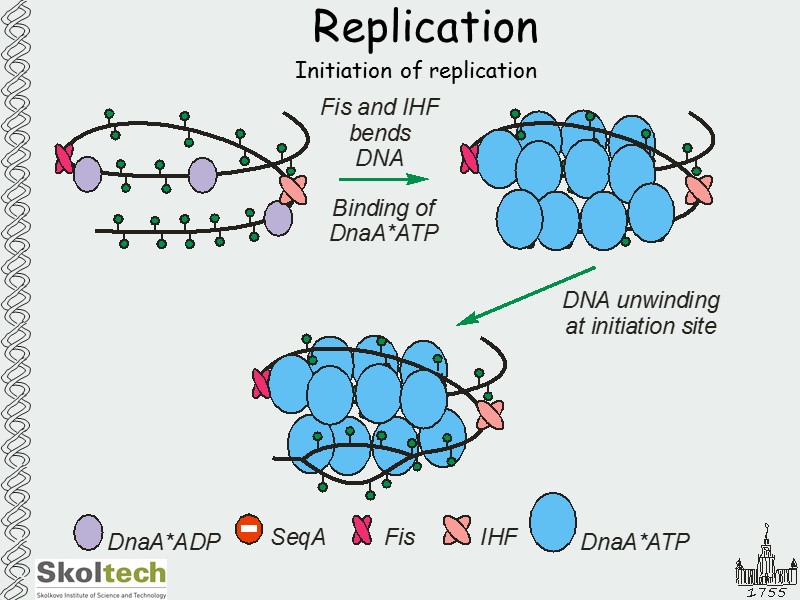
Replication Initiation of replication
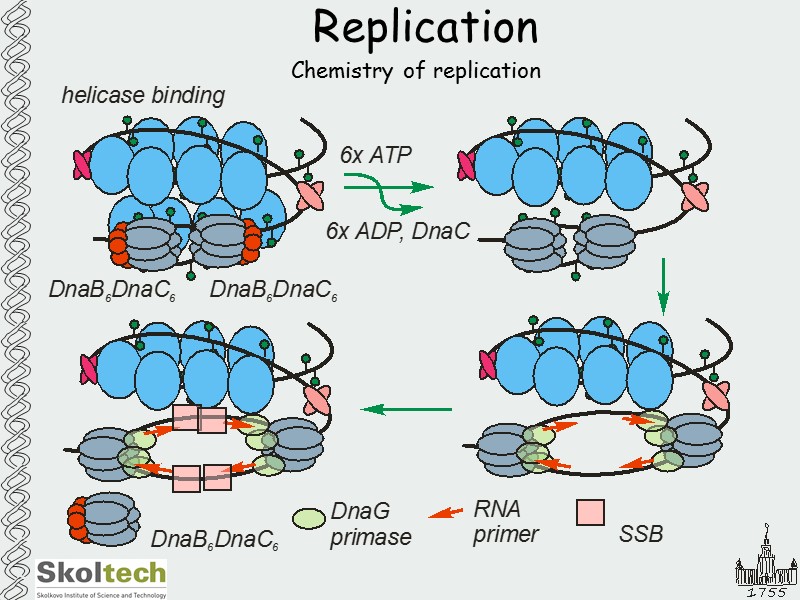
Replication Chemistry of replication
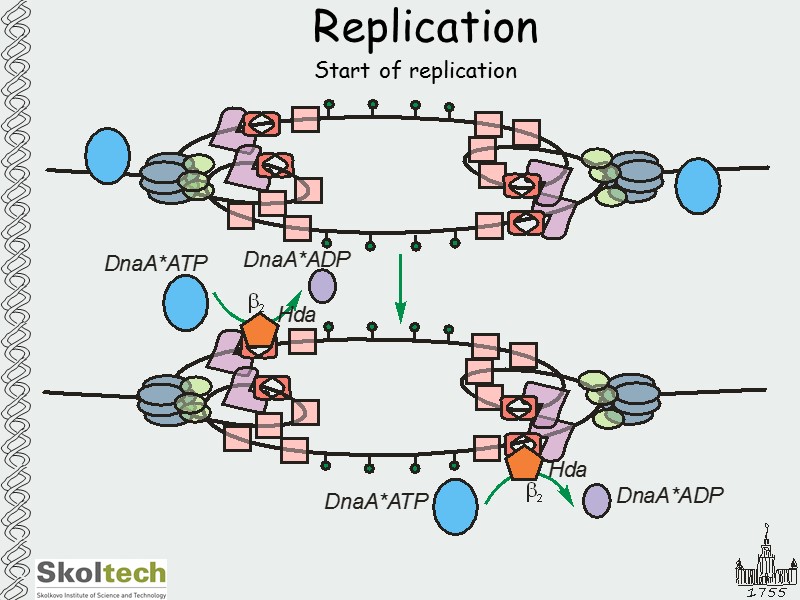
Replication Start of replication
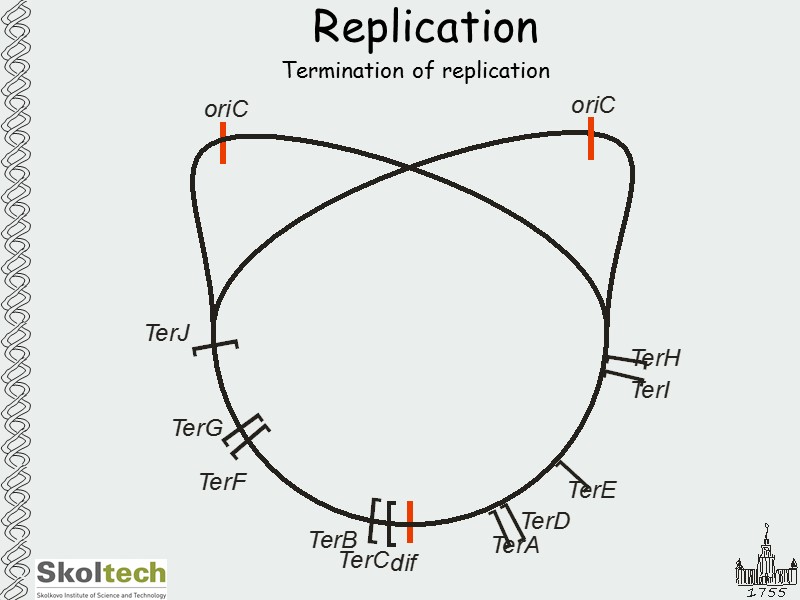
Replication Termination of replication
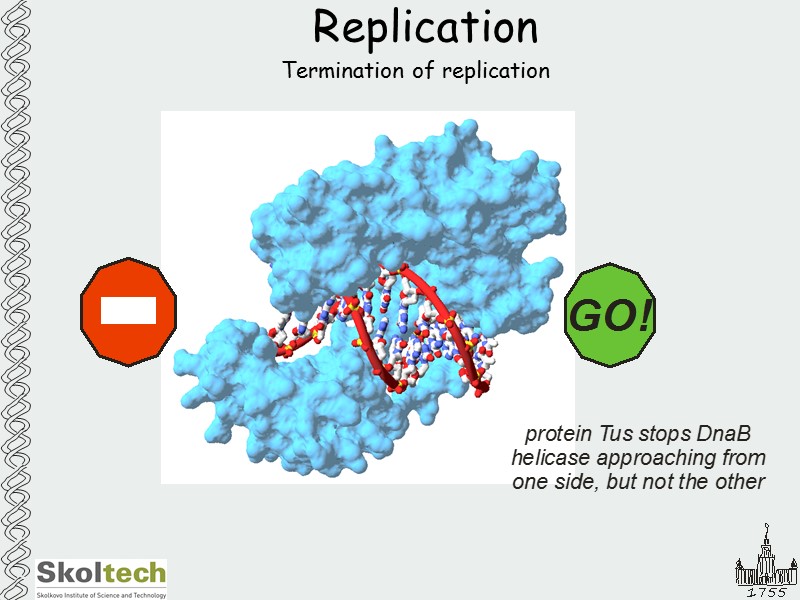
Replication Termination of replication
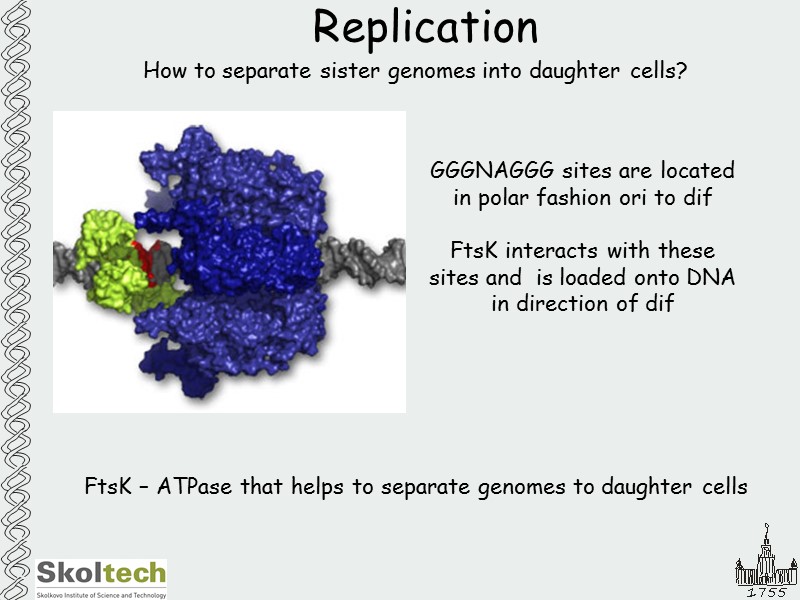

Replication How to separate sister genomes into daughter cells? FtsK – ATPase that helps to separate genomes to daughter cells GGGNAGGG sites are located in polar fashion ori to dif FtsK interacts with these sites and is loaded onto DNA in direction of dif
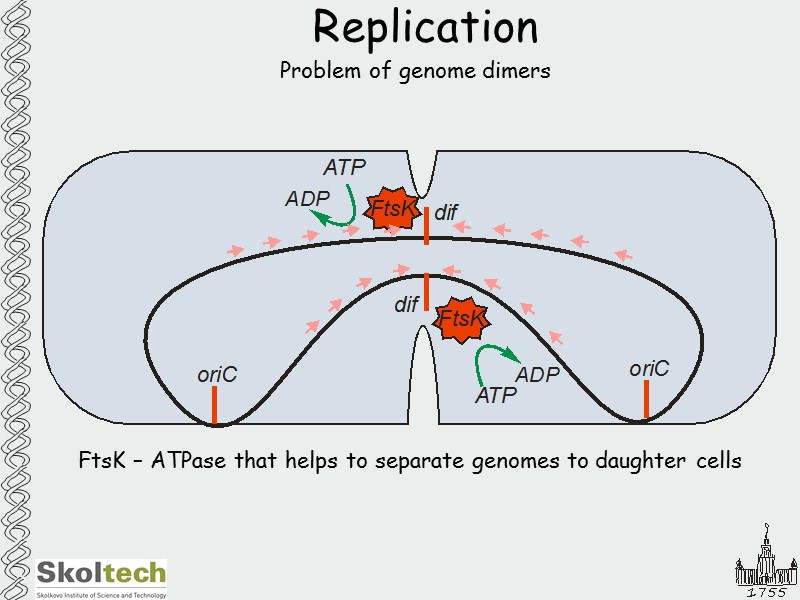
Replication Problem of genome dimers FtsK – ATPase that helps to separate genomes to daughter cells
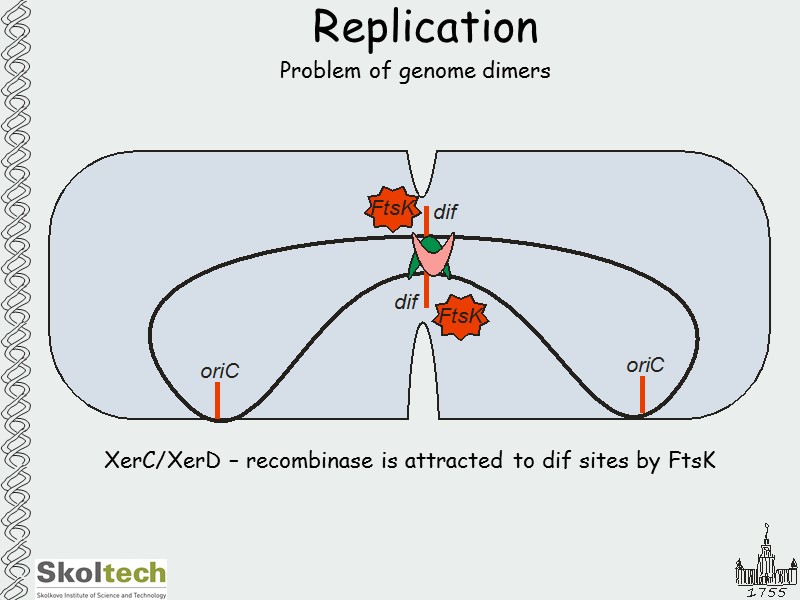
Replication Problem of genome dimers XerC/XerD – recombinase is attracted to dif sites by FtsK
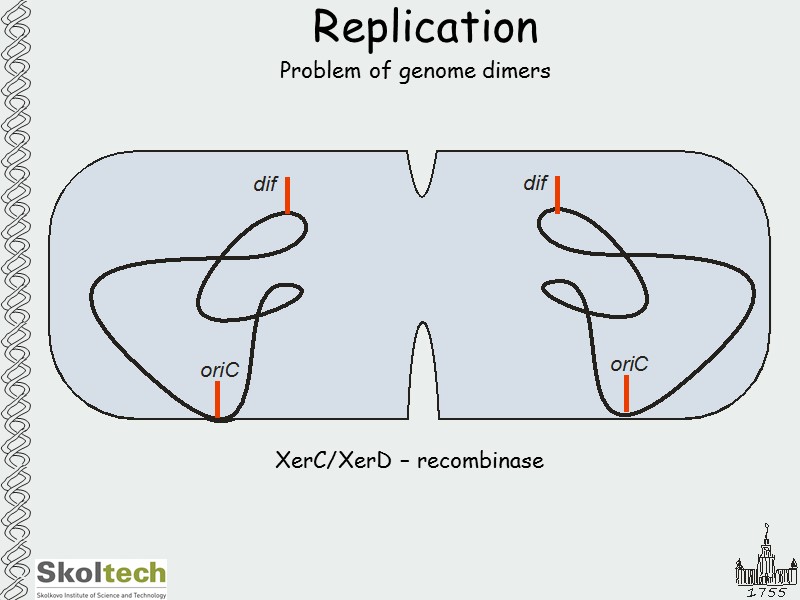
Replication Problem of genome dimers XerC/XerD – recombinase
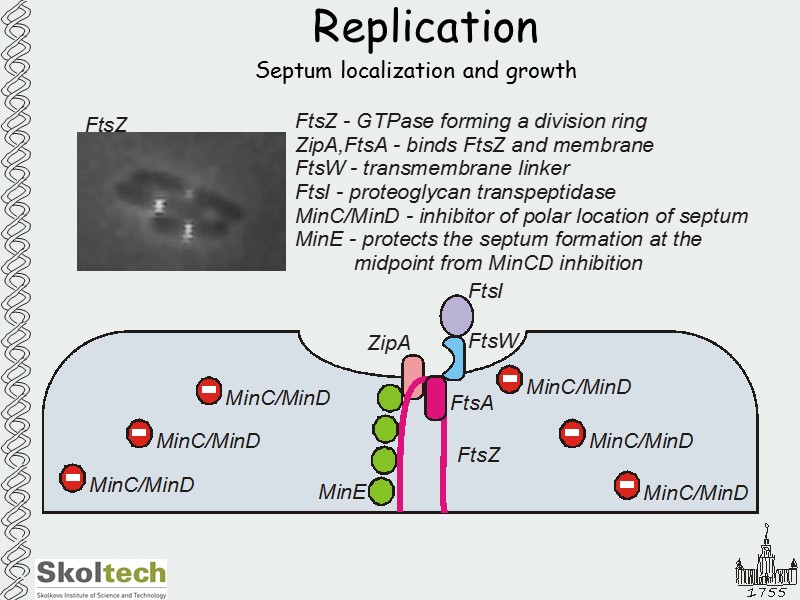
Replication Septum localization and growth
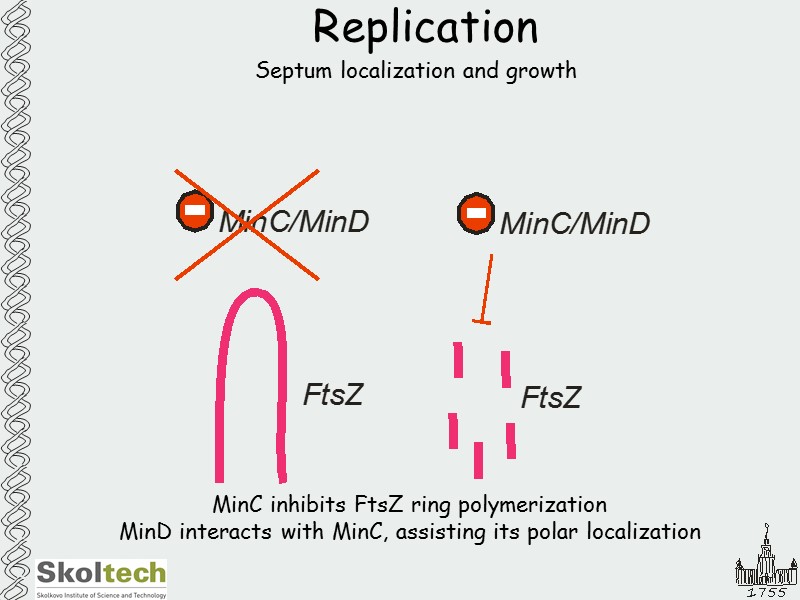
Replication Septum localization and growth MinC inhibits FtsZ ring polymerization MinD interacts with MinC, assisting its polar localization
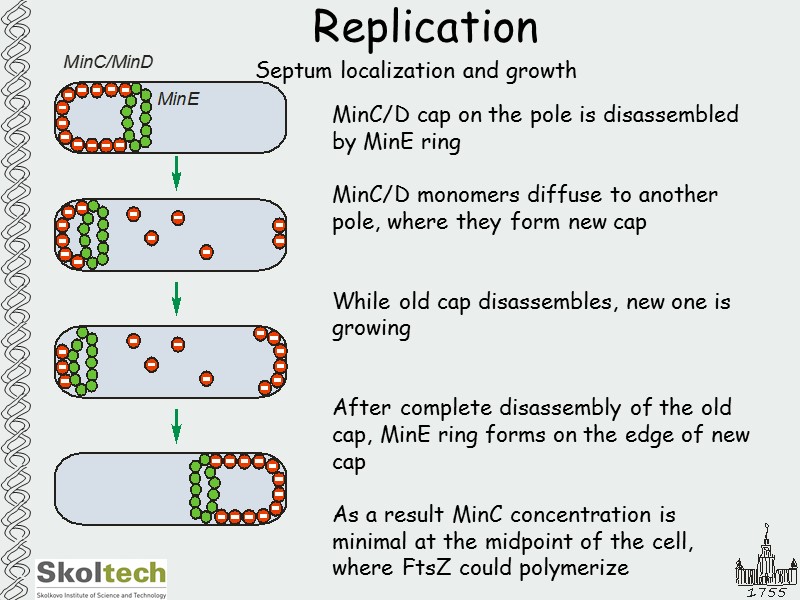
Replication Septum localization and growth MinC/D cap on the pole is disassembled by MinE ring MinC/D monomers diffuse to another pole, where they form new cap While old cap disassembles, new one is growing After complete disassembly of the old cap, MinE ring forms on the edge of new cap As a result MinC concentration is minimal at the midpoint of the cell, where FtsZ could polymerize
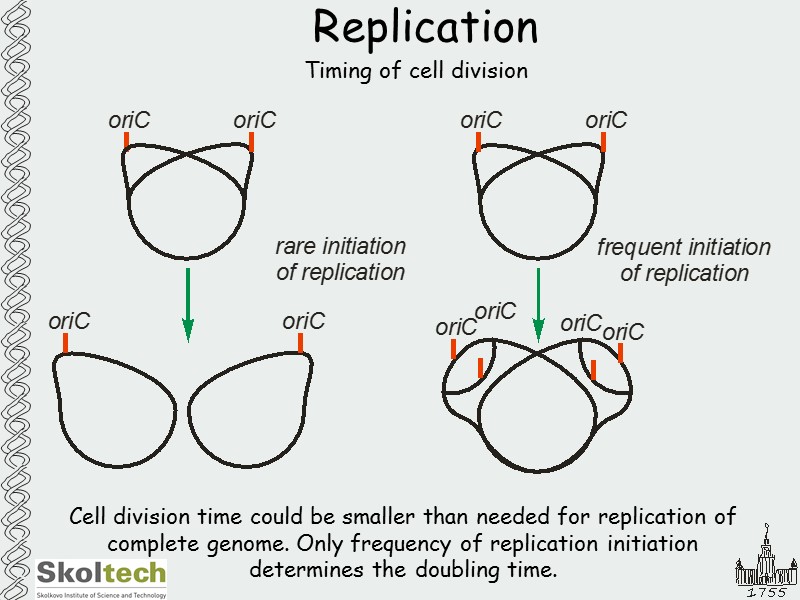
Replication Timing of cell division Cell division time could be smaller than needed for replication of complete genome. Only frequency of replication initiation determines the doubling time.
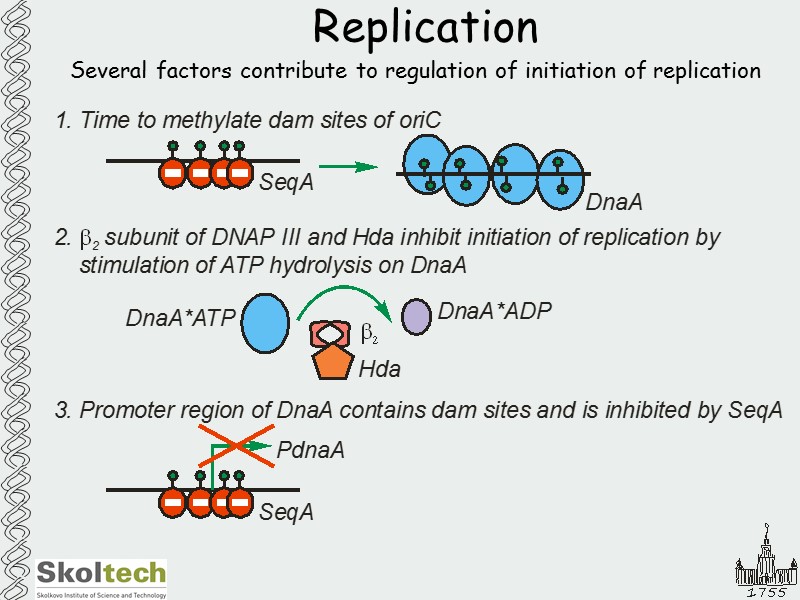
Replication Several factors contribute to regulation of initiation of replication
31628-7._replication_in_bacteria.ppt
- Количество слайдов: 48

